Abstract
Lysine (K) methyltransferase 2a (Kmt2a) and other regulators of H3 lysine 4 methylation, a histone modification enriched at promoters and enhancers, are widely expressed throughout the brain, but molecular and cellular phenotypes in subcortical areas remain poorly explored. We report that Kmt2a conditional deletion in postnatal forebrain is associated with excessive nocturnal activity and with absent or blunted responses to stimulant and dopaminergic agonist drugs, in conjunction with near-complete loss of spike-timing-dependent long-term potentiation in medium spiny neurons (MSNs). Selective ablation of Kmt2a, but not the ortholog Kmt2b, in adult ventral striatum/nucleus accumbens neurons markedly increased anxiety scores in multiple behavioral paradigms. Striatal transcriptome sequencing in adult mutants identified 262 Kmt2a-sensitive genes, mostly downregulated in Kmt2a-deficient mice. Transcriptional repression includes the 5-Htr2a serotonin receptor, strongly associated with anxiety- and depression-related disorders in human and animal models. Consistent with the role of Kmt2a in promoting gene expression, the transcriptional regulators Bahcc1, Isl1, and Sp9 were downregulated and affected by H3K4 promoter hypomethylation. Therefore, Kmt2a regulates synaptic plasticity in striatal neurons and provides an epigenetic drug target for anxiety and dopamine-mediated behaviors.
Similar content being viewed by others
INTRODUCTION
Mood and anxiety spectrum disorders cause significant morbidity and mortality with an estimated 30–40% of subjects only partially responding to conventional antidepressant and anxiolytic treatments including monoamine reuptake inhibitors (Krishnan and Nestler, 2010). Therefore, it will be necessary to explore, in the preclinical model, novel therapeutic avenues, including ‘epigenetic’ therapies targeting chromatin structure and function (Lattal et al, 2007; Mahgoub and Monteggia, 2013; Vialou et al, 2013). Of particular interest are regulators of mono-, di-, and trimethyl-histone H3-lysine 4 (H3K4me1/2/3)—chromatin marks associated with promoters and active enhancers and gene expression (Zhou et al, 2011)—which show a surprisingly strong link to the genetic risk architecture of mood and psychosis spectrum disorders and other common psychiatric disease (Psychiatric Genomics Consortium, 2015). Similarly, animal models identified multiple H3K4-specific histone methyltransferase and demethylase enzymes as critical regulators of cognition and memory, fear and anxiety (Gupta et al, 2010; Kerimoglu et al, 2013; Jakovcevski et al, 2015), and substance abuse and dependence (Aguilar-Valles et al, 2014; Shen et al, 2014). For example, conditional deletion of lysine (K) methyltransferase 2a (Kmt2a)/mixed-lineage leukemia 1 (Mll1) in mouse prefrontal cortex neurons is associated with increased anxiety and impaired working memory (Jakovcevski et al, 2015), a cognitive defect often encountered in subjects diagnosed with mood or psychosis spectrum disorder (Hall et al, 2014; Okon-Singer et al, 2015). Defective cognition in Kmt2a-deficient mice was associated with dysregulated H3K4(me3) methylation at a subset of gene promoters important for cognition and emotion (Jakovcevski et al, 2015). However, little is known about functional and behavioral phenotypes resulting from Kmt2a deletion in subcortical structures implicated in anxiety and depression. These include the ventral (including nucleus accumbens) and dorsal (caudate, putamen) striatum, both important for reward and stress processing (Sesack and Grace, 2010; Admon et al, 2015). Interestingly, stimulant drug-associated reward memory is sensitive to small RNA-mediated Kmt2a knockdown in the nucleus accumbens (Aguilar-Valles et al, 2014), a finding that could point to a broader role of Kmt2a in the regulation of motivational and emotional behaviors. Here, we further explore this hypothesis with a highly integrative approach focused on anxiety- and depression-related phenotypes. Our study includes behavioral assays in conditional mutant mice, slice recordings to measure striatal synaptic plasticity, and profiling of striatal transcriptomes and epigenomes.
MATERIALS AND METHODS
Animals
All animal experiments were approved by the Animal Use and Care committee of the participating institutions. Mice were kept under specific pathogen-free constant conditions (21±1 °C; 60% humidity) and mice of both sexes were used for the experiments, with each mutant mouse matched to a control mouse of the same gender. Food and water was supplied ad libitum in an animal facility with a regular 12 h light–dark cycle (light on at 0700 hours). All experiments were in accordance with the guidelines of the Institutional Animal Care and Use Committee (IACUC) of the participating institutions. The present study included male and female mice, all from the same colony in a predominant C57BL6/J background, back-crossed for at least eight generations. For all test conditions, the male:female ratio was ~1:1, and each mutant animal was analyzed in parallel to one or two sex-matched wild-type (WT) control of the same age ±2 weeks.
Experiments After Conditional Mll1 Ablation in Postnatal Forebrain Neurons
Conditional deletion of Mll1 were obtained by breeding mice carrying a previously described Mll1flox/floxallele (Jude et al, 2007; Jakovcevski et al, 2015) with a CamKIIα-Cre (CamK-Cre) transgenic line that recombines in forebrain neurons starting at the time of birth, resulting in widespread Cre-mediated deletion in forebrain before postnatal day 18 (Akbarian et al, 2002). These animals and their controls (Cre-negative litter mate controls) were used for diurnal and stimulant effects of amphetamine and D1-agonist-induced locomotor activity as well as for striatal slice recordings. A summary and timeline for the various experiments is provided in Figure 1a.
Striatum-related phenotypes after neuron-specific Mll1 deletion in the forebrain. (a) Experimental outline, including age (range) for slice recordings and behavioral assays. (b) Increased locomotion during active/dark phase in Camk2a-Cre+, Mll1flox/flox (orange) mice in comparison with Camk2a-Cre−, Mll1flox/flox (blue) littermates. Effect of ‘genotype’ F(1,14)=57.39, p<0.001, ‘hour’ F(23,322)=8.23 p<0.001, and interaction between ‘genotype × hour’ F(23,322)=1.8, p<0.05. (c) Camk2a-Cre+, Mll1flox/flox mice show a blunted response to amphetamine (3 mg/kg) during the dark phase. Effect of ‘genotype’ F(1,10)=29.17, p<0.001, ‘interval/drug’ F(11,110)=9.92, p<0.001, and interaction between ‘genotype × interval/drug’ F(11,110)=11.96, p<0.001 and (d) during the light phase (amphetamine 6 mg/kg). Effect of ‘genotype’ F(1,10)=30.21, p<0.001, ‘interval/drug’ F(11,110)=1.94, p<0.05, and interaction between ‘genotype × interval/drug’ F(11,110)=3.99, p<0.001, (e) Decreased response to the D1/D5 agonist SKF81297 (0.03 mg/kg) in Camk2a-Cre+, Mll1flox/flox mice. Effect of ‘genotype’ F(1,12)=5.4, p<0.05, effect of interval/drug F(23,276)=6.2, p<0.001, interaction ‘genotype × interval/drug’, F(23,276)=2.13, p<0.01; n=6–9 animals per genotype. *p<0.05, **p<0.01, ***p<0.001 post hoc after analysis of variance (ANOVA), difference between wild type (WT) and knockout. (f–h) Spike-timing-dependent long-term potentiation (tLTP) in medium spiny neurons (MSNs) in the ventral striatum. (f) Excitatory postsynaptic potential (EPSP) amplitude, from a representative core MSN, showing tLTP. (g) (from up to down) EPSPs in mV, resting membrane potential (RMP) in mV, and input resistance (Rin) in mΩ from neuron shown in (f) were stable over entire recording period. Note the lack of change in membrane resting potential and input resistance 20 min after induction. Solid horizontal bars indicate average of 30 consecutive EPSP recorded before (red bar) and 20 min after (red bar) AP-EPSP pairing. (h) EPSP amplitudes (% of control, preinduction)/tLTPs in Camk2a-Cre+, Mll1flox/flox (scatter plot, orange symbols) and Camk2a-Cre−, Mll1flox/flox (blue symbols) MSNs; n=11 and 12 recorded cells per genotype. *p<0.05, χ2. Solid black circle with S.E.M. indicates averaged tLTD amplitude in Mll1 mutant mice.
Diurnal, Amphetamine and SKF 81297-Induced Locomotor Activity
Mice were individually housed in standard macrolon cages. On the testing day, they were placed into the frame of a photobeam activity system for home cage (San Diego Instruments). For accessing the diurnal home-cage activity, mice were left undisturbed for the entire period of 24 h. For measurements of stimulant and D1 agonist responses, the baseline activity was recorded for 30 min immediately after mice were injected with 0.9% saline (vehicle). Then, the recording was put on hold and mice were injected either with amphetamine (3 mg/kg during the dark phase or 6 mg/kg during the light phase) or the D1/D5 receptor agonist SKF81297 (0.03 mg/kg during the dark phase), and the locomotor activity was recorded either for 30 min (amphetamine) or 90 min (SKF81297). The SKF81297 dose administered here was similar to the dose administered systemically in a previous study reporting altered cognition and behavior in rats exposed to low-dose SKF81297 (Hotte et al, 2005).
Slice Recordings: Animals and Slice Preparation
We used 21–30-day-old conditional Mll1 CamK-Cre mutant and control male mice to prepare slices from fresh brain tissue. A full description of the electrophysiological recordings is available in the Supplementary Methods.
RNA-seq
Striatal Ribo-Zero depleted RNA from N=4 animals was used to prepare sequencing libraries, using a standard Illumina protocol. Libraries were sequenced on a HiSeq 2000 (Illumina). Reads were mapped to the mouse genome (build 38, mm10) using the TopHat2 package (Trapnell et al, 2009). Differential gene expression between n=2 control and n=2 Camk2aCre+, Mll1flox/flox conditional mutant mice was determined using the Voom-Limma package (Law et al, 2014) at cutoffs of p-value <0.01 and fold changes >2.
RT-qPCR
Total RNA was extracted from Mll1 WT and KO striatum and transcribed into cDNA using SuperScript III (Invitrogen) and random hexamer primers. cDNA was thereafter amplified on a Roche 480 LightCycler using SYBR Green PCR Master Mix Kits (Qiagen/Roche). Primer pairs were designed using Primer 3. The primers used for Isl1 (GenBank: NM_021459.4), Bahcc1 (GenBank:NM_198423.3), and Sp9 (GenBank:NM_001005343.2) transcripts are listed in Supplementary Table S1. Gapdh (GenBank:NM_001289726.1) transcripts were used for normalization.
ChIP-qPCR
ChIP was performed using either 4 μl of anti-H3K4me1 antibody (Diagenode, C15410037) or 10 μl of anti-H3K4me3 antibody (Cell Signaling Technology; no. 97635), respectively (experimental details are provided in Supplementary Methods). For quantification, 1 μl out of 40 μl of ChIP-DNA or input DNA was used for quantitative PCR (qPCR) quantification using QuantiFast SYBR Green PCR Master Mix on a Roche 480 LightCycler 2.0 (Roche Diagnostics). PCRs were run in triplicates. Primer pairs design for Isl1, Bahcc1, and Sp9 promoter regions were listed in Supplementary Table S1. The enrichment for each mark was normalized to its own input before being compared between Mll1 WT and KO samples.
Experiments After Conditional Kmt2a/Mll1 and Kmt2b/Mll2 Ablation in Adult Ventral Striatum
A summary and timeline of the experiments is provided in Figure 2a. Stereotactic injection of AAV-Cre: For all experiments, the male:female ratio was ~1:1 for each genotype tested, with each mutant mouse matched to an age- and gender-matched control. Adult Mll1flox/flox mice and previously described Mll2flox/flox animals (Glaser et al, 2009; Kerimoglu et al, 2013) were subjected to Cre-mediated deletion in the ventral striatum, as described in the following paragraph. Stereotactic delivery of adeno-associated virus, serotype 8 (AAV) for the expression of a CreGFP (Cre-green fluorescent protein) fusion protein under the control of the neuron-specific SYNAPSIN1 promoter (Nakajima et al, 2012) was performed as follows: mice were anesthetized with a ketamine/xylazine cocktail (intraperitoneally: 100 mg/kg, 15 mg/kg; Sigma-Aldrich) and 1 μl of virus for each hemisphere (~4.7 × 109 genomic copies) was injected at a rate of 0.10 μl/min using a Hamilton syringe (Reno, NV), a micropump and sterotactic frame (Stoelting). Coordinates for injection were: +1.6 mm anterior/posterior; ±1.5 mm medial/lateral; −4.4 mm dorsal/ventral. All (AAV-Cre) experiments were performed at least 2 weeks after surgery.
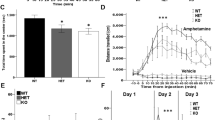
Neuronal Mll1 deletion in the adult ventral striatum affects anxiety-related behaviors. (a) Experimental timeline for AAV8hSYN1-CreGFP-injected adult (>10 weeks of age) Mll1flox/flox, Mll2flox/flox, and control mice. (b, left) NeuN neuronal marker (red) immunostained, and DAPI (4',6-diamidino-2-phenylindole) (blue) counterstained coronal section from AAV8hSYN1-CreGF-injected animal, showing robust CreGFP (Cre-green fluorescent protein) expression (green) in the ventral striatum. (Right) Ventral striatal CreGFP+ nuclei (green, two examples marked by arrowheads) express NeuN immunoreactivity (red) and DAPI counterstain (blue), confirming neuronal transduction with CreGFP fusion protein. Scale bars, (left) 1 mm and (right) 20 μm. (c) Performance in the elevated plus maze (EPM), including percentage of entries into open arms and time spent on the open arms. (d) Anxiety-like behavior in the light/dark test (L/D), comprising latency to enter the light compartment and time spent in each compartment. (e) Performance in the open field test (OF), left to right: distance moved, 5 min intervals (effect of genotype, F(2,138)=14.9, p<0.001; interval, F(2,138)=7.5, p<0.001), total distance moved in 15 min, and time in the center. (f and g) Behavioral despair paradigms, (f) forced swim test and (g) tail suspension test; n=13–19 animals per genotype, *p<0.05 and **p<0.01 Newman–Keuls post hoc. (h, left) Flow cytometry to count GFP+ nuclei from AAV8hSYN1-CreGFP-injected ventral striatum; (middle) representative sorting plot, nuclei were NeuN+ (red) immunostained for 4-way sort (NeuN+/GFP+, NeuN+/GFP−, NeuN−/GFP+, NeuN−/GFP−). Note the near-complete absence of NeuN−/GFP+ nuclei, further confirming overwhelming neuron-specific expression of CreGFP (right). Total number of NeuN+/GFP+ and NeuN−/GFP+ nuclei sorted from AAV8hSYN1-CreGFP-injected Mll1flox/flox and Mll2flox/flox striata (n=3 per group), note similar numbers of GFP+ nuclei in both genotypes, with (y axis log scale) ~100-fold higher proportion of NeuN+/GFP+ compared with NeuN−/GFP+. ST, striatum.
Anxiety Assays
Anxiety testing began 2 weeks after stereotactic injection: (1) Open-field locomotion test: This test was monitored in test chambers using a photocell-beam-based computer detecting system (OmniTech Electronics). The apparatus consisted of an arena (40 × 40 cm2) surrounded by 40 cm high walls made from clear plastic. Animals were introduced into one corner of the test chamber and allowed free exploration for 20 min individually under standard illumination conditions. The duration of time the animals spent in the central and peripheral areas were calculated. Additionally, locomotor activity was recorded and plotted in 5 min intervals to evaluate the spontaneous locomotor activity. (2) The light/dark box test: This test was performed in the open-field arena with a black box insert (20 cm L × 20 cm W × 40 cm H), dividing the arena into dark and light components. Animals were able to roam between the two components via a small portal in the divider. Animals were introduced into the dark chamber and allowed free exploration in the arena for 10 min. Latency to enter the lit compartment and duration of time spent in the dark and light chambers were calculated. (3) Elevated plus maze test: The elevated plus maze (Med Associates) consists of a center square (6 × 6 cm2), two open arms, and two closed arms (measuring 35 × 6 cm2 each). The closed arms were enclosed by black polypropylene walls measuring 20 cm in height. Mice were placed in the center square facing one of the closed arms. Time spent in each arm was recorded and scored by the EthoVision video tracking system (Noldus, Wageningen, The Netherlands). A decrease in time spent in the open arms reflects a state of increased anxiety. Animals that fell/jumped off the maze were removed from the study.
Behavioral Despair/Depression Assays
The following despair-related behaviors were assessed in the third week after injection: (1) Tail suspension test: Animals were suspended with duct tape by the tail onto a suspension bar for the duration of 5 min. Latency to freeze and the time spent immobile was evaluated by EthoVision software. Animals showing tail climbing behaviors were removed from the further analysis. (2) Forced swim test: Animals were placed into a 4 L Pyrex beaker (13 cm diameter, 24 cm height) filled with 22 °C water to a height of 17 cm. The test was 5 min in duration. Latency to freeze and time spent immobile was evaluated by EthoVision software.
Amphetamine-Induced Locomotor Activity
After completion of the experiments described above, an independent cohort of adult age- and gender-matched Mll1flox/flox and Mll1+/+ mice was injected with AAV-Cre (~4.5 × 109 genomic copies per μl, 2 μl of virus for each hemisphere) in the ventral striatum. These mice were tested for amphetamine-induced locomotor activity at least 3 weeks after AAV expression. All mice were tested in an open-field design after treatment with sterile saline followed by amphetamine (intraperitoneally; Sigma; A5880). Mice were given saline (intraperitoneally) and recorded for 30 min, followed by administration of amphetamine intraperitoneally; 6 mg/kg) and additional recording for 30 min.
Histology and Imaging
Mouse brains were processed as described in Supplementary Methods.
Flow Cytometry
Nuclei were extracted and purified from striatal tissue punches as described (Jakovcevski et al, 2015), and immunostained with anti-NeuN antibody using Alexa 544 as conjugate. Nuclei counts based on NeuN and GFP were carried out with a FACSVantage SE flow cytometer.
Quantification of Intranuclear RNA
Reverse-transcriptase qPCR (RT-qPCR) was conducted on intranuclear RNA. To confirm the efficiency of conditional gene deletions, striatal neuronal nuclei were separated and sorted based on Cre expression, as described above, with ~5000–9000 nuclei per sample. RNA was quantified with primers sensitive for the conditional Mll1 and Mll2 gene deletions as described previously (Jakovcevski et al, 2015), with 18S RNA for normalization, using Mll1 deletion primer (forward, Fwd) 5'-TAATCCTAGCCGTTAGGCCG-3' and (reverse, Rev) 5′-TTGGGGCAGGTTTGGGTTAG-3′, and Mll2 deletion primer Fwd, 5′-CATCTTCCCTGACCCACCAC-3′ and Rev, 5′-CTCCCCTGAGGTAGGTGTGA-3′, and additional primers for Bahcc1, Isl1, and Sp9 transcripts as described above.
Statistical Analysis
All behavioral, molecular (except for the RNA-seq data, see paragraph RNA-seq), and electrophysiological data were expressed as mean±SEM and the significance of group differences were evaluated by two-tailed t-test or, when indicated (repeated measurements or >2 groups) by ANOVA, followed by post hoc tests.
RESULTS
Neuronal Loss of Mll1 in the Forebrain is Associated with Behavioral Phenotypes, Indicative of Altered Dopaminergic Signaling, and Loss of tLTPs in the Striatum/Nucleus Accumbens
We recently described a conditional Mll1 mutant line (Jakovcevski et al, 2015), obtained by breeding a Camk2a-Cre transgenic line (for Cre-mediated deletions in postnatal (P) forebrain neurons at or before P18; Akbarian et al, 2002), with mice carrying a conditional Mll1 allele, with loxP sites flanking exons 3 and 4 (Jude et al, 2007). Conditional Camk2a-Cre-mediated deletion resulted in strongly reduced MLL1 immunoreactivity and Mll1 RNA levels in telencephalic areas, in conjunction with mildly (<8–10%) reduced brain weight but normal brain morphology and cytoarchitecture (Jakovcevski et al, 2015). During the course of our initial investigations, which largely were focused on molecular and cellular phenotypes in the cerebral cortex (Jakovcevski et al, 2015), we noticed that Mll1 mutant mice are strikingly hyperactive in the ‘active’ or dark phase of the light–dark cycle in comparison with littermate controls with WT MLL1 levels (Figure 1b), which could be consistent with altered dopaminergic signaling (Zhu et al, 2012). Therefore, we hypothesized that Mll1-deficient mice will show an abnormal response to dopaminergic drugs. Indeed, independent batches of Camk2aCre+, Mll1flox/flox conditional mutants, in comparison with Camk2aCre−, Mll1flox/flox controls, robustly exhibited blunted activity after treatment with the stimulant drug amphetamine (6 mg/kg intraperitoneally) during the light phase (Figure 1d) or during the dark phase (3 mg/kg intraperitoneally) (Figure 1c) and showed consistently a blunted locomotor response for the first hour after acute administration of the dopamine D1 receptor agonist SKF81927 (0.03 mg/kg intraperitoneally) (Figure 1e).
Importantly, some of these behavioral phenotypes, including the failure to show a hyperlocomotor response after amphetamine administration, have been previously linked to a disruption of striatal long-term potentiation (LTP) in mice with defective dopamine D1 signaling (Napolitano et al, 2010). Therefore, we speculated that synaptic plasticity in the striatum, including its ventral portions (nucleus accumbens), which is broadly implicated in psychiatric diseases including depression and addiction, is altered in our Mll1 mutant mice. However, conventional LTP involving high-frequency stimulation (ie, 100 Hz) is not ideal for the study of synaptic plasticity given that MSNs in the ventral striatum typically fire between 1 and 10 Hz in freely moving animals (Carelli and Ijames, 2000; Hollander et al, 2002; Krause et al, 2010). Unsurprisingly, <20% of ventral striatal MSN show induced plasticity in conventional LTP paradigms (Pennartz et al, 1991; Kombian and Malenka, 1994). To overcome these limitations, we recently introduced spike-timing-dependent plasticity as a stimulation paradigm that reflects more closely the in vivo firing patterns of mouse nucleus accumbens core MSN and their afferents (Ji and Martin, 2012; Figures 1f and g), compared with paradigms that are based on the high-frequency LTP model of Bliss and Lomo (1973). Indeed, we observed a preferential abolishment of timing-dependent LTP (tLTP) (Figure 1h) in Mll1-deficient MSN from the nucleus accumbens. Thus, while in MSN of the ventral striatum of Camk2aCre−, Mll1flox/flox mice, tLTP was observed in approximately one-half of recorded cells (6/11, 148±15.5% of baseline control, n=11), only 1 out of 12 recorded Camk2aCre+, Mll1flox/flox MSN mutant neurons showed tLTP (Figure 1h). This difference was significant (χ2 (d.f. 1)=2.84, p<0.05). These effects were highly specific, because the magnitude of tLTD was completely preserved in Camk2aCre+, Mll1flox/flox MSN (81.7±8.4% of baseline control) and thus very comparable to that of Camk2aCre−, Mll1flox/flox MSN with normal MLL1 levels (79.6±7.5% of baseline control) (Figure 1h).
Deletion of Mll1, but not Mll2, in the Striatum Leads to Increased Anxiety Behaviors
Having shown that stimulant drug- and dopamine agonist-mediated locomotor behaviors, and tLTP in the ventral striatum are affected in Camk2aCre+, Mll1flox/flox conditional mutant mice, we then asked whether other types of complex behaviors could be dependent on striatal Mll1. We focused on behavioral phenotypes related to anxiety and depression, in part, because previous work had shown that Mll1 ablation in mature prefrontal cortex is anxiogenic (Jakovcevski et al, 2015). Given that both prefrontal cortex and ventral striatum are key nodes in the neural circuitry underlying mood and anxiety spectrum disorders, we reasoned that Mll1 ablation in mature MSN of the ventral striatum, too, could affect depression- and anxiety-related behaviors. To selectively ablate Mll1 from ventral striatal neurons, we bilaterally injected adeno-associated virus expressing CreGFP fusion protein (AAV8hSYN1-CreGFP) into adult (>10 weeks of age) nucleus accumbens. Furthermore, because mammalian genomes harbor at least 11 H3K4-specific methyltransferase genes (Black et al, 2012), we wanted to know whether any of the behavioral phenotypes after gene deletion in ventral striatal neurons are specific for Mll1 or whether such phenotypes would reflect a nonspecific response to methyltransferase gene deletions. Therefore, we conducted parallel studies in mice homozygous for the Mll1 ortholog, Mll2/Kmt2bflox/flox (Kerimoglu et al, 2013; Ladopoulos et al, 2013).
Quantification of Mll1 and Mll2 transcripts with deletion-sensitive primer pairs in the RT-qPCR, conducted separately on striatal neuronal NeuN+, GFP+, and NeuN+, GFP− nuclei sorted and collected from the same tissues (see Materials and Methods), showed >10-fold decrease in transcript levels after Cre-mediated gene ablation (Supplementary Figure S1), confirming the efficacy of the conditional Mll1 and Mll2 mutagenesis.
Anxiety- (open-field, light-dark box and elevated plus maze tests) and depression-related behaviors (forced swim and tail suspension tests) were assessed 15–25 days after the AAV-Cre injections (Figure 2a), followed by brain harvest. The bilateral AAV-Cre injections were tolerated well by all mice and genotypes, and body weight at the time of brain harvest was indistinguishable between groups (Supplementary Figure S2). Interestingly, AAV8hSYN1-CreGFP-injected Mll1flox/flox mice, but not AAV8hSYN1-CreGFP-injected Mll2flox/flox mice or Mll1+/+ control mice, showed significantly increased anxiety-related behaviors across all three tests, as evidenced by the significantly less open arm entries in the elevated plus maze (F(2,41)=4.07, p<0.05) (Figure 2c), and increased latency to enter and decreased time spent in the (anxiogenic) brighter compartment of the light dark box (F(2,45)=3.39, p<0.05) while more time spent in the dark compartment (F(2,45)=3.39, p<0.05) (Figure 2d). In the open field test, there was overall decreased locomotion (F(2,46)=6.064, p<0.01) and time spent in the (anxiogenic) center of the open field (F(2,46)=9.4, p<0.001) (Figure 2e). Post hoc tests confirmed that these changes, detected by ANOVA, were solely due to changes in the AAV8hSYN1-CreGFP-injected Mll1flox/flox mice (Figures 2c–e). In contrast to these robust (~30–50% from controls) increases in anxiety-related behaviors across all three anxiety tests (Figures 2c–e), AAV8hSYN1-CreGFP-injected Mll1flox/flox mice showed, in comparison with AAV8hSYN1-CreGFP-injected Mll2flox/flox and Mll1+/+ control mice, minimal (~5–7%) differences in immobility scores in 2/2 depression- and despair-related tests (Figures 2f and g). Importantly, the number of CreGFP+ striatal neurons, measured by quantitative flow cytometry, was very similar between genotypes (n=3, mean±S.E.M., Mll1flox/flox 6.409±1.157; Mll2flox/flox 8.694±3.116; Figure 2h). We conclude that Mll1 ablation in <7000 ventral striatal neurons suffices to elicit a robust, prodepressant and anxiogenic behavioral phenotype, whereas no alterations were observed when similar numbers were targeted for Mll2 ablation.
Furthermore, to confirm that the aforementioned severely attenuated and blunted locomotor response after stimulant and dopaminergic agonist treatment in Camk2aCre+, Mll1flox/flox conditional mutant mice is due to Mll1/Kmt2a loss in adult striatum, we studied another cohort of Mll1flox/flox and control mice (independent of the animals that underwent testing for anxiety and depression), doubling the amount of AAV8hSYN1-CreGFP injected into adult striatum bilaterally (see Materials and Methods). Notably, the AAV8hSYN1-CreGFP-injected Mll1flox/flox mice, in comparison with injected Mll1+/+ controls, showed a severely attenuated response after 3 mg/kg amphetamine intraperitoneally (Supplementary Figure S3). Therefore, the severely blunted or total lack of amphetamine-induced locomotor response in Camk2aCre+, Mll1flox/flox conditional mutants (Figure 1c) is likely to result from loss of Kmt2a/Mll1 in striatal neurons.
Striatal Transcriptome and H3K4 Methylation After Mll1 Ablation
Gene expression changes in the cerebral cortex and hippocampus after conditional, neuron-specific ablation of either Mll1 or the related gene Mll2 has been reported to be surprisingly subtle overall (Kerimoglu et al, 2013; Jakovcevski et al, 2015). To explore transcriptome alterations of the Mll1-deficient striatum, we performed RNA-seq on the rostral striatum of Camk2aCre+, Mll1flox/flox conditional mutants, in comparison with Camk2aCre−, Mll1flox/flox controls (n=2 per group), generating 48–71 M mappable reads for each sample. When filtered for >1.5-fold difference, p<0.01, 186 transcripts were downgulated and 76 were upregulated in the Mll1-deficient striatum, marking a 2.3-fold excess of decreased (vs increased) striatal RNAs after Mll1 ablation. This finding resonates with the notion that Mll1 and Mll1-regulated H3K4 methylation are, on a genome-wide scale, associated with open chromatin and the RNA polymerase II transcriptional complex (Guenther et al, 2005). Of note, the list of Mll1-sensitive genes expressed at decreased levels in the mutant striatum included a number of signaling molecules with a critical role in the regulation of cognition and mood (Supplementary Table S2). For example, the 5-Htr2a serotonin receptor is implicated in depression and anxiety (Sakata and Duke, 2014), the nicotinic acetylcholine receptor subunit Chrna6 is important for dopaminergic signaling in the striatum (Kamens et al, 2012), and the Slc38a8 sodium-coupled amino-acid transporter is widely expressed in neurons (Hagglund et al, 2015). Furthermore, several genes regulating chromatin and transcription emerged from the Mll1 RNA-seq screen. These included the Lim homeodomain transcription factor Isl1 important for the development of striatonigral projection neurons (Lu et al, 2014), and the Sp9 transcription factor and the Bahcc1 bromo adjacent homology domain-containing protein (Figure 3a). Of note, H3K4-specific methyltransferases (other than Mll1/Kmt2a), including Setd1a/b and Mll2-4, or the H3K4-specific demethylases Kdm1a/2a/4a/5a-d showed minimal (and nonsignificant) changes in the mutant striatum (Supplementary Table S3). This finding suggests that compensatory changes in other H3K4 methylation regulators are unlikely to have a role in the Kmt2a-deficient striatum. Furthermore, a subset of Mll1-sensitive striatal transcripts (including Slc38a8, Isl1, Sp9, Bahcc11) show similar changes in Mll1-deficient cortex (Supplementary Table S2 and Jakovcevski et al, 2015), suggesting a subset of MLL1-dependent regulatory mechanisms are common to both brain regions.
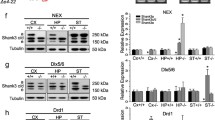
Altered gene expression and H3K4 methylation in Mll1-deficient striata. (a) Striatal RNA-seq tracks of (pink, ‘KO’) Camk2a-Cre+, Mll1flox/flox and (blue,‘WT’) Camk2a-Cre−, Mll1flox/flox, showing (top to bottom) 81 kb Kmt2a/Mll1 gene confirming (gray shaded area) conditional deletion of exons III and IV in mutant; 15 kb Isl1, 63 kb Bahcc1, and 7.8 kb Sp9 genes all with near-complete loss of exonic tags in mutant RNA-seq tracks. (b) In situ hybridization (sagittal sections) from the Allen Institute website www.alleninstitute.org, showing robust expression of Kmt2a/Mll1, Isl1, Bahcc1, and Sp9 genes in the area of ventral striatum (vSt). (c) cDNA qPCR and (d) ChIP PCR, from the striatum of (pink, ‘KO’) Camk2a-Cre+, Mll1flox/flox and (blue, ‘WT’) Camk2a-Cre−, Mll1flox/flox mice. Note significant deficits in striatal Isl1, Bahcc1, and Sp9 expression and H3K4me1 and H3K4me3 promoter methylation (c, d n=3 per group; two-tailed t-test, *p<0.05, **p<0.01). ChIP PCR, chromatin immunoprecipitation PCR; H3K4me, histone H3-lysine 4; KO, knockout; qPCR, quantitative PCR; WT, wild type.
Focusing on the three aforementioned transcriptional regulators, Isl1, Bahcc1, and Sp9 (all of which moderately or robustly expressed in adult striatum; Figure 3b), we first verified downregulated expression (which initially was observed in the striatal RNA-seq datasets) by processing additional striatal samples from Camk2aCre+, Mll1flox/flox mutant and Camk2aCre−, Mll1flox/flox control mice (n=3 per group), using RT-qPCR (Figure 3c).
Next, we tested whether the downregulation in expression of the three transcriptional regulators (Isl1, Bahcc1, and Sp9) in striatal tissue homogenate of adult Camk2aCre+, Mll1flox/flox mice could be replicated in sorted striatal neuronal nuclei that underwent Mll1 ablation by AAV8hSYN1-CreGFP injection into the adult striatum. Striatal NeuN+ neuron nuclei were sorted and separated into NeuN+, CreGFP+ and NeuN+, CreGFP− fractions, and intranuclear RNA was quantified by qRT-PCR. Indeed, consistent downregulation in expression in Mll1-deficient striatal neuronal nuclei was observed for Sp9 and Bahcc1 genes, in comparison with their surrounding Cre−, NeuN+ nuclei (Supplementary Figure S4). Importantly, the levels of Bahcc1 and Sp9 transcripts showed a nonsignificant increase, whereas Isl1 showed a nonsignificant decrease in NeuN+, CreGFP+ nuclei of AAV8hSYN1-CreGFP-injected striatum of adult Mll22lox/2lox (Supplementary Figure S5). We draw three conclusions: First, Mll1, in a cell autonomous manner, has a critical role for proper expression of a subset of transcriptional regulators in adult striatal neurons. Second, these effects are specific to Mll1, because Mll2-deficient striatal nuclei did not show significant changes in the expression of the Mll1-sensitive transcripts. Third, these findings draw a strong molecular connection to the above-described behavioral phenotypes of mice with striatal Mll1 deficiency.
Next, we then quantified striatal histone H3K4 methylation levels at Bahcc1, Isl1 and Sp9 promoters, using ChIP-PCR. Because MLL1 methyltransferase regulates the mono-, di- and trimethylated forms (Del Rizzo and Trievel, 2011), we measured two of the three H3K4 methylation forms, H3K4me1 and H3K4me3, that have been most closely linked to active transcription. Of note, Isl1, Bahcc1, and Sp9 promoters of Mll1-deficient striata consistently showed deficits in H3K4me1 and H3K4me3 occupation (37–65% decrease) in comparison with controls (Figure 3d). These results resonate with previous reports linking MLL1 to the regulation of multiple H3K4 methylation forms in the cerebral cortex (Jakovcevski et al, 2015) and peripheral tissues (Del Rizzo and Trievel, 2011).
DISCUSSION
The present study on mice with conditional Kmt2a/Mll1 gene deletions provides multiple lines of evidence for a critical role of this histone methyltransferase for striatal signaling and related behaviors. Neuronal Mll1 ablation in postnatal forebrain, including striatum, was associated with marked alterations in dopamine-mediated behaviors, including a marked hyperactivity in the active phase of the dark–light cycle, and a blunted locomotor response after systemic injection with dopamine D1 agonist and failure to show a hyperlocomotor response after acute exposure to the stimulant drug, amphetamine. Such types of behavioral alterations had been previously linked to defective synaptic long-term potentiation in the striatum (Napolitano et al, 2010). Indeed, spike-timing-dependent LTP in the ventral striatum, a type of neural plasticity mediated by medium spiny neurons as the striatum’s major neuronal constituency, was nearly completely abolished in Mll1 mutant mice. Consistent with these findings from Camk2a Cre+, Mll1flox/flox conditional mutant mice, region-specific ablation of neuronal Mll1 in the adult ventral striatum, a brain region that among other functions serves as an important node in the neural circuits of anxiety and depression (Vialou et al, 2010; Velasques et al, 2014), was highly anxiogenic in 3/3 anxiety-related behavioral paradigms (open field, light dark box, and elevated plus maze). These behavioral alterations were highly specific for Mll1. This is because striatal ablation of Kmt2b/Mll2, a gene closely related to Mll1 (Rao and Dou, 2015) and expressed at similar levels in the adult striatum (Supplementary Figure S2B), was not associated with altered anxiety (Figures 2c–e).
What types of molecular defects in Mll1 conditional mutant could underlie the defects in striatal LTP, and abnormal response to dopaminergic and stimulant drugs? In the present study, deep sequencing of the Mll1-deficient striatal transcriptome identified 262 genes with altered expression (186 downregulated and 76 upregulated), many of which implicated with a critical role in cognition and complex behaviors and depression and anxiety, including, for example, 5-Htr2a serotonin receptor (Sakata and Duke, 2014) and Chrna6 nicotinic acetylcholine receptor subunit (Kamens et al, 2012). These alterations could, in turn, be related to deficits in H3K4 methylation at gene proximal promoters and other sequences for the Isl1, Sp9, and Bahcc1 transcriptional regulators, and additional Mll1-sensitive genes, which could compromise neuronal function with additional, secondary changes in the neuronal transcriptome. Importantly, these limited alterations of the Mll1-deficient striatal transcriptome, with subtle changes in expression of altogether <262 genes, resonate with recent studies in peripheral tissues and brain reporting that only a small portion of the transcriptome and H3K4 methylated promoters become dysregulated in the context Mll1 or Mll2 gene deletions (Wang et al, 2009; Kerimoglu et al, 2013; Denissov et al, 2014; Jakovcevski et al, 2015). The specific set of transcripts altered in Mll1-deficient neurons may vary dependent on neuronal subtype and brain region. In Camk2aCre+, Mll1flox/flox conditional mutants, there is only limited overlap between striatal and cortical transcriptome alterations and <6% of Mll1-sensitive transcripts downregulated in Mll1 mutant striatum show a concurrent change in Mll1-deficient cortex (Supplementary Table S2).
Therefore, Mll1 could fine-tune, in a region-specific manner, a small set of transcripts with a critical role in neuronal signaling and behavior. Such type of region-specific transcriptome alterations resonates with the prevailing view that each of the four MLL1-4 histone methyltransferase family members regulates only a very small subset of genes in a tissue-specific manner, or in case of the brain, in a region-specific manner (Black et al, 2012; Shen et al, 2014), whereas another subgroup of SET-domain containing H3K4-specific methyltransferases, including the neuropsychiatric risk gene SETD1A (Singh et al, 2016; Takata et al, 2016), are attributed with the regulation of much larger portion of the genome (Black et al, 2012). The molecular mechanisms underlying these effects remain to be determined but differential binding of H3K4 methyltransferase complex-associated proteins, including the chromatin regulator, Menin, may have a role (Shen et al, 2014).
Remarkably, region-specific ablation of Mll1 in either the prefrontal cortex (Jakovcevski et al, 2015) or the ventral striatum (present study) results in an excessively anxious mouse. This finding could point to novel therapeutic avenues, given that treatment-resistant anxiety and depression, including insufficient response to serotonin reuptake inhibitors, cognitive behavioral, and other types of therapy, are estimated to afflict 40% of the patient population (Bystritsky, 2006; Ipser et al, 2006). Poor treatment outcomes significantly contribute to human suffering, reduced quality of life, and increased economic burden, considering a prevalence of 13% in the United States for anxiety disorders alone (Bystritsky et al, 2013). Therefore, the behavioral effects of small-molecule drugs targeted towards MLL1 and some its binding partners, including the MLL-associated WRAD complex (WDR5, RbBP5, ASH2L, DPY30) (Karatas et al, 2013; Rao and Dou, 2015; Cao et al, 2014), warrant further exploration in preclinical models for anxiety and depression. Importantly, a subset of extremely broad H3K4me3 peaks extending over several kilobases are highly conserved in human and mouse cortical neurons, including many genes associated with dopaminergic and glutamatergic signaling, two neurotransmitter pathways frequently implicated in the pathophysiology of mood, and psychosis spectrum disorders (Dincer et al, 2015).
Finally, the findings presented here draw a firm link between regulation of H3K4 methylation and maintenance of neuronal function in the striatum. The study presented here will have broader implications for neurological conditions associated with striatal dysfunction and mood and motor symptoms, such as Huntington’s disease for which alterations in H3K4 methylation and additional histone modifications had been reported (Valor et al, 2013; Vashishtha et al, 2013; Dong et al, 2015).
FUNDING AND DISCLOSURE
The authors declare no conflict of interest.
References
Admon R, Holsen LM, Aizley H, Remington A, Whitfield-Gabrieli S, Goldstein JM et al (2015). Striatal hypersensitivity during stress in remitted individuals with recurrent depression. Biol Psychiatry 78: 67–76.
Aguilar-Valles A, Vaissiere T, Griggs EM, Mikaelsson MA, Takacs IF, Young EJ et al (2014). Methamphetamine-associated memory is regulated by a writer and an eraser of permissive histone methylation. Biol Psychiatry 76: 57–65.
Akbarian S, Rios M, Liu RJ, Gold SJ, Fong HF, Zeiler S et al (2002). Brain-derived neurotrophic factor is essential for opiate-induced plasticity of noradrenergic neurons. J Neurosci 22: 4153–4162.
Black JC, Van Rechem C, Whetstine JR (2012). Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol Cell 48: 491–507.
Bliss TV, Lomo T (1973). Long-lasting potentiation of synaptic transmission in the dentate area of the anaesthetized rabbit following stimulation of the perforant path. J Physiol 232: 331–356.
Bystritsky A (2006). Treatment-resistant anxiety disorders. Mol Psychiatry 11: 805–814.
Bystritsky A, Khalsa SS, Cameron ME, Schiffman J (2013). Current diagnosis and treatment of anxiety disorders. PT 38: 30–57.
Cao F, Townsend EC, Karatas H, Xu J, Li L, Lee S et al (2014). Targeting MLL1 H3K4 methyltransferase activity in mixed-lineage leukemia. Mol Cell 53: 247–261.
Carelli RM, Ijames SG (2000). Nucleus accumbens cell firing during maintenance, extinction, and reinstatement of cocaine self-administration behavior in rats. Brain Res 866: 44–54.
Del Rizzo PA, Trievel RC (2011). Substrate and product specificities of SET domain methyltransferases. Epigenetics 6: 1059–1067.
Denissov S, Hofemeister H, Marks H, Kranz A, Ciotta G, Singh S et al (2014). Mll2 is required for H3K4 trimethylation on bivalent promoters in embryonic stem cells, whereas Mll1 is redundant. Development 141: 526–537.
Dincer A, Gavin DP, Xu K, Zhang B, Dudley JT, Schadt EE et al (2015). Deciphering H3K4me3 broad domains associated with gene-regulatory networks and conserved epigenomic landscapes in the human brain. Transl Psychiatry 5: e679.
Dong X, Tsuji J, Labadorf A, Roussos P, Chen JF, Myers RH et al (2015). The role of H3K4me3 in transcriptional regulation is altered in Huntington's disease. PLoS One 10: e0144398.
Glaser S, Lubitz S, Loveland KL, Ohbo K, Robb L, Schwenk F et al (2009). The histone 3 lysine 4 methyltransferase, Mll2, is only required briefly in development and spermatogenesis. Epigenet Chromatin 2: 5.
Guenther MG, Jenner RG, Chevalier B, Nakamura T, Croce CM, Canaani E et al (2005). Global and Hox-specific roles for the MLL1 methyltransferase. Proc Natl Acad Sci USA 102: 8603–8608.
Gupta S, Kim SY, Artis S, Molfese DL, Schumacher A, Sweatt JD et al (2010). Histone methylation regulates memory formation. J Neurosci 30: 3589–3599.
Hall MH, Levy DL, Salisbury DF, Haddad S, Gallagher P, Lohan M et al (2014). Neurophysiologic effect of GWAS derived schizophrenia and bipolar risk variants. Am J Med Genet B 165B: 9–18.
Hagglund MG, Hellsten SV, Bagchi S, Philippot G, Lofqvist E, Nilsson VC et al (2015). Transport of L-glutamine, L-alanine, L-arginine and L-histidine by the neuron-specific Slc38a8 (SNAT8) in CNS. J Mol Biol 427: 1495–1512.
Hollander JA, Ijames SG, Roop RG, Carelli RM (2002). An examination of nucleus accumbens cell firing during extinction and reinstatement of water reinforcement behavior in rats. Brain Res 929: 226–235.
Hotte M, Naudon L, Jay TM (2005). Modulation of recognition and temporal order memory retrieval by dopamine D1 receptor in rats. Neurobiol Learn Mem 84 (2): 85–92.
Ipser JC, Carey P, Dhansay Y, Fakier N, Seedat S, Stein DJ (2006). Pharmacotherapy augmentation strategies in treatment-resistant anxiety disorders. Cochrane Database Syst Rev CD005473.
Jakovcevski M, Ruan H, Shen EY, Dincer A, Javidfar B, Ma Q et al (2015). Neuronal Kmt2a/Mll1 histone methyltransferase is essential for prefrontal synaptic plasticity and working memory. J Neurosci 35: 5097–5108.
Ji X, Martin GE (2012). New rules governing synaptic plasticity in core nucleus accumbens medium spiny neurons. Eur J Neurosci 36: 3615–3627.
Jude CD, Climer L, Xu D, Artinger E, Fisher JK, Ernst P (2007). Unique and independent roles for MLL in adult hematopoietic stem cells and progenitors. Cell Stem Cell. 1: 324–337.
Kamens HM, Hoft NR, Cox RJ, Miyamoto JH, Ehringer MA (2012). The alpha6 nicotinic acetylcholine receptor subunit influences ethanol-induced sedation. Alcohol 46: 463–471.
Karatas H, Townsend EC, Cao F, Chen Y, Bernard D, Liu L et al (2013). High-affinity, small-molecule peptidomimetic inhibitors of MLL1/WDR5 protein-protein interaction. J Am Chem Soc. 135: 669–682.
Kerimoglu C, Agis-Balboa RC, Kranz A, Stilling R, Bahari-Javan S, Benito-Garagorri E et al (2013). Histone-methyltransferase MLL2 (KMT2B) is required for memory formation in mice. J Neurosci 33: 3452–3464.
Kombian SB, Malenka RC (1994). Simultaneous LTP of non-NMDA and LTD of NMDA-receptor-mediated responses in the nucleus accumbens. Nature 368: 242–246.
Krause M, German PW, Taha SA, Fields HL (2010). A pause in nucleus accumbens neuron firing is required to initiate and maintain feeding. J Neurosci 30: 4746–4756.
Krishnan V, Nestler EJ (2010). Linking molecules to mood: new insight into the biology of depression. Am J Psychiatry 167: 1305–1320.
Ladopoulos V, Hofemeister H, Hoogenkamp M, Riggs AD, Stewart AF, Bonifer C (2013). The histone methyltransferase KMT2B is required for RNA polymerase II association and protection from DNA methylation at the MagohB CpG island promoter. Mol Cell Biol 33: 1383–1393.
Lattal KM, Barrett RM, Wood MA (2007). Systemic or intra hippocampal delivery of histone deacetylase inhibitors facilitates fear extinction. Behav Neurosci 121: 1125–1131.
Law CW, Chen Y, Shi W, Smyth GK (2014). voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol 15: R29.
Lu KM, Evans SM, Hirano S, Liu FC (2014). Dual role for Islet-1 in promoting striatonigral and repressing striatopallidal genetic programs to specify striatonigral cell identity. Proc Natl Acad Sci USA 111: E168–E177.
Mahgoub M, Monteggia LM (2013). Epigenetics and psychiatry. Neurotherapeutics 10: 734–741.
Nakajima H, Kubo T, Semi Y, Itakura M, Kuwamura M, Izawa T et al (2012). A rapid, targeted, neuron-selective, in vivo knockdown following a single intracerebroventricular injection of a novel chemically modified siRNA in the adult rat brain. J Biotechnol 157: 326–333.
Napolitano F, Bonito-Oliva A, Federici M, Carta M, Errico F, Magara S et al (2010). Role of aberrant striatal dopamine D1 receptor/cAMP/protein kinase A/DARPP32 signaling in the paradoxical calming effect of amphetamine. J Neurosci 30: 11043–11056.
Okon-Singer H, Hendler T, Pessoa L, Shackman AJ (2015). The neurobiology of emotion–cognition interactions: fundamental questions and strategies for future research. Front Hum Neurosci 9: 58.
Pennartz CMA, Boejinga PH, Lopes da silva FH (1991). Contributions of NMDA receptors to postsynaptic potentials and paired-pluses facilitation in identified neurons of the rat nucleus accumbens in vitro. Exp Brain Res 86: 190–198.
Psychiatric Genomics Consortium, Network and Pathway Analysis Subgroup (2015). Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat Neurosci 18: 199–209.
Rao RC, Dou Y (2015). Hijacked in cancer: the KMT2 (MLL) family of methyltrans-ferases. Nat Rev Cancer 15: 334–346.
Sakata K, Duke SM (2014). Lack of BDNF expression through promoter IV disturbs expression of monoamine genes in the frontal cortex and hippocampus. Neuroscience 260: 265–275.
Sesack SR, Grace AA (2010). Cortico-basal ganglia reward network: microcircuitry. Neuropsychopharmacology 35: 27–47.
Shen E, Shulha H, Weng Z, Akbarian S (2014). Regulation of histone H3K4 methylation in brain development and disease. Philos Trans R Soc Lond Ser B 369: 20130514.
Singh T, Kurki MI, Curtis D, Purcell SM, Crooks L, McRae J et al (2016). Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat Neurosci 19: 571–577.
Takata A, Ionita-Laza I, Gogos JA, Xu B, Karayiorgou M (2016). De novo synonymous mutations in regulatory elements contribute to the genetic etiology of autism and schizophrenia. Neuron 89: 940–947.
Trapnell C, Pachter L, Salzberg SL (2009). TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25: 1105–1111.
Vashishtha M, Ng CW, Yildirim F, Gipson TA, Kratter IH, Bodai L et al (2013). Targeting H3K4 trimethylation in Huntington disease. Proc Natl Acad Sci USA 110: E3027–E3036.
Valor LM, Guiretti D, Lopez-Atalaya JP, Barco A (2013). Genomic landscape of transcriptional and epigenetic dysregulation in early onset polyglutamine disease. J Neurosci 33: 10471–10482.
Velasques B, Diniz C, Teixeira S, Cartier C, Peressutti C, Silva F et al (2014). Deep brain stimulation: a new treatment in mood and anxiety disorders. CNS Neurol Disord Drug Targets 13: 961–971.
Vialou V, Feng J, Robison AJ, Nestler EJ (2013). Epigenetic mechanisms of depression and antidepressant action. Annu Rev Pharmacol Toxicol 53: 59–87.
Vialou V, Maze I, Renthal W, LaPlant QC, Watts EL, Mouzon E et al (2010). Serum response factor promotes resilience to chronic social stress through the induction of DeltaFosB. J Neurosci 30: 14585–14592.
Wang P, Lin C, Smith ER, Guo H, Sanderson BW, Wu M et al (2009). Global analysis of H3K4 methylation defines MLL family member targets and points to a role for MLL1-mediated H3K4 methylation in the regulation of transcriptional initiation by RNA polymerase II. Mol Cell Biol 29: 6074–6085.
Zhou VW, Goren A, Bernstein BE (2011). Charting histone modifications and the functional organization of mammalian genomes. Nat Rev Genet 12: 7–18.
Zhu J, Zhang X, Xu Y, Spencer TJ, Biederman J, Bhide PG (2012). Prenatal nicotine exposure mouse model showing hyperactivity, reduced cingulate cortex volume, reduced dopamine turnover, and responsiveness to oral methylphenidate treatment. J Neurosci 32: 9410–9418.
Acknowledgements
Sequencing of the RNAseq libraries was performed at the New York Genome Center. This work was supported by NIH Grants P50MH096890, R01MH086509 (to SA), and OD011103 and R01DA032283 (to WDY), and by a Marie Curie Intra European Fellowship within the 7th European Community Framework Program and a NARSAD Young investigator grant from the Brain and Behavior Research Foundation (to MJ).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on the Neuropsychopharmacology website
Supplementary information
Rights and permissions
About this article
Cite this article
Shen, E., Jiang, Y., Javidfar, B. et al. Neuronal Deletion of Kmt2a/Mll1 Histone Methyltransferase in Ventral Striatum is Associated with Defective Spike-Timing-Dependent Striatal Synaptic Plasticity, Altered Response to Dopaminergic Drugs, and Increased Anxiety. Neuropsychopharmacol 41, 3103–3113 (2016). https://doi.org/10.1038/npp.2016.144
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2016.144
- Springer Nature Switzerland AG
This article is cited by
-
tRNA epitranscriptomic alterations associated with opioid-induced reward-seeking and long-term opioid withdrawal in male mice
Neuropsychopharmacology (2024)
-
Histone Methyltransferase G9a Plays an Essential Role on Nicotine Preference in Zebrafish
Molecular Neurobiology (2024)
-
Structural variation of the coding and non-coding human pharmacogenome
npj Genomic Medicine (2023)
-
Aberrant Histone Modification of TNFAIP3, TLR4, TNIP2, miR-146a, and miR-155 in Major Depressive Disorder
Molecular Neurobiology (2023)
-
RNA methyltransferase NSun2 deficiency promotes neurodegeneration through epitranscriptomic regulation of tau phosphorylation
Acta Neuropathologica (2023)