Abstract
A computational challenge raised by chromosome conformation capture (3C) experiments is to reconstruct spatial distances and three-dimensional genome structures from observed contacts between genomic loci. We propose a two-step algorithm, ShRec3D, and assess its accuracy using both in silico data and human genome-wide 3C (Hi-C) data. This algorithm avoids convergence issues, accommodates sparse and noisy contact maps, and is orders of magnitude faster than existing methods.



Similar content being viewed by others
References
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Science 295, 1306–1311 (2002).
Lieberman-Aiden, E. et al. Science 326, 289–293 (2009).
Marti-Renom, M.A. & Mirny, L.A. PLOS Comput. Biol. 7, e1002125 (2011).
Baù, D. & Marti-Renom, M.A. Methods 58, 300–306 (2012).
Duan, Z. et al. Nature 465, 363–367 (2010).
Hu, M. et al. PLOS Comput. Biol. 9, e1002893 (2013).
Nagano, T. et al. Nature 502, 59–64 (2013).
Rousseau, M. et al. BMC Bioinformatics 12, 414 (2011).
Trieu, T. & Cheng, J. Nucleic Acids Res. 42, e52 (2014).
Varoquaux, N., Ay, F., Noble, W.S. & Vert, J.P. Bioinformatics 30, i26–i33 (2014).
Zhang, Z. et al. J. Comput. Biol. 20, 831–846 (2013).
Sippl, M.J. & Scheraga, H.A. Proc. Natl. Acad. Sci. USA 82, 2197–2201 (1985).
Torgerson, W.S. Psychometrika 17, 401–419 (1952).
Havel, T.F., Kuntz, I. & Crippen, G.M. Bull. Math. Biol. 45, 665–720 (1983).
Fraser, J. et al. Genome Biol. 10, R37 (2009).
Hajjoul, H. et al. Genome Res. 23, 1829–1838 (2013).
Dixon, J.R. et al. Nature 485, 376–380 (2012).
Kalhor, R. et al. Nat. Biotechnol. 30, 90–98 (2012).
Burton, J.N. et al. Nat. Biotechnol. 31, 1119–1125 (2013).
Kaplan, N. & Dekker, J. Nat. Biotechnol. 31, 1143–1147 (2013).
Cournac, A. et al. BMC Genomics 13, 436 (2012).
Schoenberg, I.J. Ann. Math. 36, 724–732 (1935).
Young, G. & Householder, A.S. Psychometrika 3, 19–22 (1938).
Kruskal, J.B. & Wish, M. Sage University papers series on quantitative applications in the social sciences (no. 07-011) (SAGE Publications, Newbury Park, 1978).
Burton, J.N. et al. G3, 4, 1339–1346 (2014).
Franzke, B. & Kosko, B. Phys. Rev. E 84, 041112 (2011).
Acknowledgements
The authors thank D. Arendt for the online-available implementation of the Floyd-Warshall algorithm. They acknowledge funding from UPMC (Université Pierre et Marie Curie, Sorbonne-Universités), grant CONVERGENCE2011, project CVG1110 (J.M.), from the French National Cancer Institute, grant INCa_5960 (A.L.), from the French National Research Agency (ANR), grant ANR-13-BSV5-0010-03 (A.L.) and from the French National Research Agency (ANR), grant ANR-09-PIRI-0024 (A.C.).
Author information
Authors and Affiliations
Contributions
J.M., A.L. and J.R. designed the algorithm. J.M. implemented it. J.R., P.R., A.C. and J.M. tested its validity. A.C. analyzed experimental data sets. J.M. and A.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Comparison of ShRec3D with the simple MDS-inverse-frequency approach.
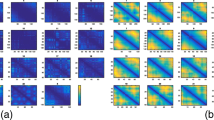
The top panels display the spectrum (eigenvalue histogram) of the metric matrix, Eq. 3, derived from our simulated contact map (with here N = 2,600) using (a) our shortest-path distance and (b) the simple distance equal to the inverse contact frequency (both distances are dimensionless: the units on the abscissa axis depend on the chosen normalization for the contact frequencies). The three rightmost black arrows underline the first three eigenvalues and the leftmost red arrow underlines the fourth one, demonstrating the presence of a significant spectral gap in a, and the absence of spectral gap in b. The bottom panels presents a scatter plot of the original distances in our simulated benchmark (horizontal axis, simulation unit equal to 10 nm, N = 2,600) and the distances reconstructed from the corresponding contact map using as a preliminary step either (a) our shortest-path distance or (b) distances obtained as the inverse contact frequencies, followed in both cases by the MDS procedure described in Figure 1a, step 5 followed by step 4 (vertical axis, dimensionless distances). Spearman rank correlation coefficient R is indicated in inset.
Supplementary Figure 2 Polymer connectivity in ShRec3D reconstruction.
Normalized histogram of the reconstructed distances Di, i+1 between neighbors along the genome (light blue peaked curves), for (a) our simulated benchmark (here N = 2,600 points) and (b) genome-wide real Hi-C data18, compared to the normalized histogram of all distances taken as a reference (dark blue broad curves).
Supplementary Figure 3 Visualization of human autosomal chromosomes using ShRec3D.
Color labeling of the different chromosomes: 1: blue, 2: red, 3: grey, 4: orange, 5: yellow, 6: gold, 7: silver, 8: green, 9: pink, 10: cyan, 11: purple, 12: lime, 13: mauve, 14: ochre, 15: ice blue, 16: black, 17: light green,18: light cyan, 19: violet,20: magenta, 21: dark red, 22: light orange. (Hi-C data in lymphoblastoid cells)18.
Supplementary Figure 4 Annotation of human chromosome 1 structure.
(a) Overlay of human chromatin partition in two compartments2 (highlighted by the color code on the structure and boxes; yellow indicates gene rich, GC rich regions, on the left; red indicates gene poor, AT rich regions, on the right). (b) Overlay of two histone H3 modifications on lysine 9. In cyan regions harboring a high level of acetylation, in pink regions harboring a high level of tri-methylation and in purple region harboring both modifications.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 (PDF 890 kb)
Rights and permissions
About this article
Cite this article
Lesne, A., Riposo, J., Roger, P. et al. 3D genome reconstruction from chromosomal contacts. Nat Methods 11, 1141–1143 (2014). https://doi.org/10.1038/nmeth.3104
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3104
- Springer Nature America, Inc.
This article is cited by
-
3D Genome Reconstruction from Partially Phased Hi-C Data
Bulletin of Mathematical Biology (2024)
-
Three-dimensional genome landscape comprehensively reveals patterns of spatial gene regulation in papillary and anaplastic thyroid cancers: a study using representative cell lines for each cancer type
Cellular & Molecular Biology Letters (2023)
-
ParticleChromo3D: a Particle Swarm Optimization algorithm for chromosome 3D structure prediction from Hi-C data
BioData Mining (2022)
-
Imputation-free reconstructions of three-dimensional chromosome architectures in human diploid single-cells using allele-specified contacts
Scientific Reports (2022)
-
Reconstruct high-resolution 3D genome structures for diverse cell-types using FLAMINGO
Nature Communications (2022)