Abstract
Infection with influenza virus induces antibodies to the viral surface glycoproteins hemagglutinin and neuraminidase, and these responses can be broadly protective. To assess the breadth and magnitude of antibody responses, we sequentially infected mice, guinea pigs and ferrets with divergent H1N1 or H3N2 subtypes of influenza virus. We measured antibody responses by ELISA of an extensive panel of recombinant glycoproteins representing the viral diversity in nature. Guinea pigs developed high titers of broadly cross-reactive antibodies; mice and ferrets exhibited narrower humoral responses. Then, we compared antibody responses after infection of humans with influenza virus H1N1 or H3N2 and found markedly broad responses and cogent evidence for 'original antigenic sin'. This work will inform the design of universal vaccines against influenza virus and can guide pandemic-preparedness efforts directed against emerging influenza viruses.
Similar content being viewed by others
Main
Infections with seasonal influenza virus cause substantial morbidity and mortality every year on a global scale1,2. In addition, influenza pandemics occur at irregular intervals and can claim millions of human lives3. The current vaccines against seasonal influenza virus are considered an efficacious countermeasure to prevent infection with this virus2. However, they usually induce strain-specific immune responses to the three to four strains included in the vaccine formulation. In contrast, infection with influenza virus can cause broader immune responses and longer lasting protection from re-infection by the same virus subtype4,5,6,7.
Protective humoral immune responses to influenza virus are usually associated with antibodies to its surface glycoproteins hemagglutinin (HA) and neuraminidase (NA). These proteins are readily accessible to antibodies on the virion itself or on infected cells, and antibodies that bind to them can often inhibit virus replication in vitro. The traditional correlate of protection provided by vaccines against seasonal influenza virus is based on antibodies that exhibit hemagglutination-inhibition (HI) activity. They block the interaction of the receptor-binding domain located on the HA head with its sialic-acid receptor8. Due to the high plasticity and ever-changing nature of the HA head domain, most antibodies that exhibit this function are relatively strain specific9,10. Antibodies to NA can block its enzymatic function (NA inhibition), and NA-inhibition-active antibodies interfere with viral release and possibly also block the efficient migration of the virus through mucosal fluids and contribute to protection from disease11,12. NA-reactive antibodies have been shown to be broadly reactive within a subtype but usually do not exhibit heterosubtypic activity13,14. A third species of antibodies that exerts in vitro neutralizing activity is HA-stalk-reactive antibodies. Due to the conserved nature of the HA stalk, these antibodies are often cross-reactive within and across HA subtypes. Most stalk-reactive antibodies, with rare exceptions, are restricted in binding to group 1 HAs (H1, H2, H5, H6, H8, H9, H11, H12, H13, H16, HA-like H17 and HA-like H18) or group 2 HAs (H3, H4, H7, H10, H14 and H15)15,16,17,18.
Notably, as a fourth antibody species, cross-reactive antibodies can also confer protection in vivo without showing neutralizing activity in vitro. Several mechanisms, including antibody-dependent cell-mediated cytotoxicity, antibody-dependent cellular phagocytosis and complement-dependent cytotoxicity, have been postulated to contribute to non-neutralizing cross-protection in vivo19,20,21,22,23. Antibody-dependent cell-mediated cytotoxicity has been shown to have a major role in the protective efficacy of HA-stalk-reactive antibodies as well24. These effector functions can be assessed through in vivo serum-transfer challenge experiments, for example, in the mouse model25,26.
Cross-reactive antibodies are potentially important for protection from infection with 'drifted' (seasonal) and 'shifted' (pandemic) influenza viruses, but their prevalence and functionality is not well understood. Their presence might offer some protection, including lowering morbidity and mortality, during pandemics. Better understanding of cross-reactive immunity in the human population is also important for the development of universal vaccine strategies against influenza virus that are designed to boost pre-existing antibodies to protective levels. Here we analyzed the titers and breadth of antibodies to the influenza virus surface glycoproteins HA and NA induced by infection in three animal models and in humans, as well as the prevalence of cross-reactive antibodies in the general human population. The resulting data sets represent the antibody cross-reactome against the influenza virus surface glycoproteins.
Results
Cross-reactive antibody profiles in animal models
To assess induction of cross-reactive antibodies, we sequentially infected mice, guinea pigs and ferrets with two divergent H1N1 or H3N2 strains of influenza virus (Supplementary Fig. 1). The viral strains were chosen with the intention of reflecting a consecutive exposure history consistent with strains that circulated in humans and because these strains replicate well in mice, guinea pigs and ferrets (Supplementary Fig. 2). Furthermore, the animal species were chosen because they are the most relevant and most widely used animal models for influenza-virus research. For H1N1, the pre-pandemic, 1999 seasonal strain A/New Caledonia/20/99 (NC99) was chosen as the primary infection, followed by infection with the antigenically distinct 2009 pandemic H1N1 (pH1N1) isolate A/Netherlands/602/09 (NL09), an isolate antigenically identical to the prototype pandemic H1N1 strain A/California/04/09 (Cal09). Primary infection of mice led to the induction of antibodies that targeted mainly the HA of the homologous strain but also bound to heterologous H1 HAs and other related group 1 HAs (Fig. 1a). Re-infection with the antigenically distinct NL09 H1N1 strain considerably boosted the broad antibody response to H1 and also increased reactivity to heterosubtypic group 1 HAs (Fig. 1b). Notably, reactivity was highest to the H1 FM47 HA (phylogenetically situated between the HAs of the two infecting strains) (Fig. 1b). In general, reactivity to group 2 HAs (H3 HAs) was low or absent (Fig. 1b).
ELISA of IgG antibodies to all HA subtypes (Anti-HA; key) in serum obtained from mice after a single infection with H1N1 (NC99) (1× H1N1) (a), after two consecutive infections with H1N1 (NC99 and NL09) (2× H1N1) (b), after a single infection with H3N2 (Phil82) (1× H3N2) (c) or after two consecutive infections with H3N2 (Phil82 and Vic11) (2× H3N2) (d), presented as reciprocal endpoint titers (geometric mean values; key), superimposed onto a phylogenetic tree based on differences in HA amino acids (scale bars, 4% difference); gray dashed lines divide the trees into group 1 influenza A virus HAs (top) and group 2 influenza A virus HAs (bottom). Left margin, results for the H1 stalk (top left) and H3 stalk (bottom left); bottom right, results for influenza B virus HA (Yam88), presented outside the trees because of the substantial difference between this and all influenza A virus HAs. Data are representative of two experiments with pooled samples (n = 10 per group) as technical duplicates.
For the H3N2 arm of the experiment, animals were infected with the 1982 isolate A/Philippines/2/82 (Phil82), followed by infection with a more recent 2011 isolate A/Victoria/361/11 (Vic11); these are two strains separated by 29 years of antigenic drift (Supplementary Fig. 1). Primary infection of mice with Phil82 induced an immune response focused on the Phil82 H3 HA, with moderate cross-reactivity to other H3 HAs and low reactivity to heterosubtypic group 2 HAs (Fig. 1c). Notably, we also detected low reactivity to several phylogenetically distant group 1 HAs, including H2 and H6 (Fig. 1c). Re-infection with the antigenically distinct Vic11 broadened the immune responses to heterologous H3 HAs and heterosubtypic group 2 HAs (Fig. 1d). The reactivity to specific group 1 HAs was also boosted (Fig. 1d). This cross-reactivity is noteworthy, since antibodies able to bind to the head domain of both H3 HAs and H2 HAs have previously been isolated from human donors27.
Reactivity to NA after single infection with NC99 H1N1 in the mouse model was focused on the N1 subtype (Fig. 2a). Reactivity to N1 was boosted, but no broadening of the response was detected after the second infection (with NL09 H1N1) (Fig. 2b). The NA response after infections with H3N2 was as narrow as the one detected after infections with H1N1 (Fig. 2c,d).
ELISA of IgG antibodies to all NA subtypes (Anti-NA; key) in serum obtained from mice after a single infection with H1N1 (NC99) (a), after two consecutive infections with H1N1 (NC99 and NL09) (b), after a single infection with H3N2 (Phil82) (c) or after two consecutive infections with H3N2 (Phil82 and Vic11) (d), presented as reciprocal endpoint titers (geometric mean values; key), superimposed onto a phylogenetic tree based on differences in NA amino acids (scale bars, 8% difference); gray dashed lines divide the trees into group 1 influenza A virus NAs (top) and group 2 influenza A virus NAs (bottom). Bottom right, results for influenza B virus NA (Yam88) and the N10 bat isolate, presented outside the trees because of the substantial difference between this and all influenza A virus NA sequences. Data are representative of two experiments with pooled samples (n = 10 per group) as technical duplicates.
Next we compared the humoral cross-reactomes of mice, guinea pigs and ferrets (Supplementary Figs. 3 and 4). All three animal models are used for influenza-virus research, and intrinsic differences in each model's ability to induce cross-reactive antibodies might have important implications for the interpretation of pre-clinical data generated for candidate (universal) vaccines against influenza virus based on HA and NA. While heat-map–tree combinations are informative, they do not allow easy comparison of the cross-reactomes of different animal models. We thus plotted reactivity (presented as endpoint titers) against the percent difference in amino acids of the HAs compared with the sequence of the infection strain's HA. The resulting plots showed two-dimensional reactivity profiles that allowed visual comparison of the magnitude and breadth of responses in various animal models. While amino acid distances do not always correlate exactly with antigenic distances, we found it to be an adequate surrogate measure for the purposes of our data presentation (Fig. 3 and Supplementary Fig. 5).
ELISA of titers (geometric mean values; vertical axes) of antibodies to group 1 HA (key) in serum from mice (n = 10; pooled samples) (a), guinea pigs (n = 3; individual samples) (b) or ferrets (n = 2; individual samples) (c) after two consecutive infections with H1N1 (NC99 and NL09) (a–c) or of antibodies to group 2 HA (key) in serum from mice (n = 10; pooled samples) (d), guinea pigs (n = 2; individual samples) (e) or ferrets (n = 2; individual samples) (f) after two consecutive infections with H3N2 (Phil82 and Vic11) (d–f), plotted against the percent difference in amino acids (AA) versus the HA of the strain used for the second infection (phylogenetic distance; horizontal axes). Each symbol represents the titer of antibody to a single HA; results below a titer of 103 are plotted at 103. Solid black lines indicate a nonlinear fit (plateau followed by one-phase decay), to illustrate differences in the breadth of the antibody response; dashed lines indicate the titer (geometric mean value) of antibodies to all H1 HAs measured (dark blue) or all H3 HAs measured (dark red); dotted lines indicate the titer (geometric mean value) of antibodies to all group 1 non-H1 HAs (light blue) or all group 2 non-H3 HAs (light red). Data are representative of two or three experiments (technical duplicates (a,d) or biological replicates (b,c,e,f)).
For sequential infection with H1N1, our analysis showed that mice mounted high titers of antibodies to the homologous and related HAs from the same subtype but had much lower titers of antibodies to distantly related HAs (Fig. 3a). Guinea pigs exhibited a very broad plateau of cross-reactivity: titers of antibodies to the homologous H1 HA were as high as the titers of antibodies other distant group 1 HAs (Fig. 3b). Finally, ferrets induced lower titers and narrow responses (Fig. 3c). Data obtained by sequential infection with H3N2 largely echoed these findings: narrower trends were observed for the response to NA, with low cross-group reactivity of guinea pigs after infection with H3N2 (Fig. 2 and Supplementary Figs. 3 and 4).
Our data suggested that guinea pigs had the intrinsic ability to mount broader antibody responses to influenza virus HAs than did mice or ferrets. Mice showed an intermediate response breadth, and ferrets, the 'gold standard' animal model for influenza virology, show low titers and little cross-reactivity (Fig. 3 and Supplementary Fig. 5). The number of N-linked-glycosylation sites on the HAs used as substrate, which could influence cross-reactivity, did not seem to correlate with antibody titers (Supplementary Fig. 6).
Cross-reactive antibody profiles in infected humans
After mapping the antibody cross-reactome directed against the influenza virus surface glycoproteins that was induced by infection in animal models, we measured the human response to infection. We tested serum from human patients who were admitted to the hospital and were diagnosed as being infected with pandemic H1N1 or seasonal H3N2. The cohort included 11 males and 8 females, with five subjects below 18 years of age, six subjects between 18 and 59 years of age, and eight subjects above 59 years of age (Supplementary Table 1). Serum samples were obtained on the day of admission and 21 or 28 d after admission, and antibody induction was tested with a panel of recombinant HA and NA proteins. Notably, samples from patients infected with pandemic H1N1 induced a very broad response. The induction was greatest against group 1 HAs (ranging from tenfold to twentyfold), but the boost against group 2 HAs was almost as strong (ranging from sevenfold to ninefold) (Fig. 4a–c and Supplementary Figs. 7 and 8). Antibodies to influenza B virus HA were not boosted (Fig. 4a,b), which indicated that the measured increase in reactivity was specific to influenza A virus antigen.
(a) Individual antibody responses of human subjects (n = 9) after infection with H1N1, presented as geometric mean values relative to baseline (key), superimposed a phylogenetic tree of HA based on difference in amino acids (scale bar, 4% difference); gray dashed line divides the tree into group 1 influenza A virus HAs (top) and group 2 influenza A virus HAs (bottom). Left margin, results for the H1 stalk (top left) and H3 stalk (bottom left); bottom right, results for influenza B virus HA (Yam88), present outside the tree because of the substantial difference between this and all influenza A virus HAs. (b) Titers of antibodies to group 1 HAs (blue), group 2 HAs (red) or influenza B virus HA (green) in human subjects (n = 9) after infection with pH1N1, ranked from highest (left) to lowest (right); results are presented as geometric mean values relative to baseline. (c) Three dimensional surfaces of antibody titers before (gray) and after (blue (group 1 HAs) or red (group 2 HAs)) infection of human subjects (n = 9) with pH1N1, generated by plotting of the distances of HAs relative to each other (on the basis of differences in amino acids; x-axis and y-axis) against antibody titers (geometric mean values; z-axis). (d) Individual antibody responses of human subjects (n = 10) after infection with H3N2, presented as in a (scale bar, 4% difference). (e) Titers of antibodies to group 1 HAs, group 2 HAs or influenza B HA in human subjects (n = 10) after infection with H3N2, presented as in b. (f) Three dimensional surfaces of antibody titers before and after infection of human subjects (n = 10) with H3N2, presented as in c. Data are representative of nine (a–c) or ten (d–f) experiments (one donor in each as biological replicates) (error bars (b,e), 95% confidence intervals).
Samples from humans infected with H3N2 induced an antibody profile very different from that reported above. Reactivity to the homologous strain was induced (twentyfold), while induction to the next-closest H3 strain was significantly lower (eightfold) (Fig. 4d,e), which indicated that the infection induced mainly strain-specific antibodies. Induction was also observed against more distantly related group 2 HAs, while reactivity to group 1 HAs only increased marginally after infection with H3N2 (Fig. 4d–f). Again, antibodies to influenza virus B HA were not induced (Fig. 4d,e). When visualized as 'antigenic landscapes', these differences become very clear (Fig. 4c,f). In summary, these data suggested that in humans, infection with pandemic H1N1 virus induced a broader antibody response than did infection with a seasonal H3N2 virus.
The HA of pandemic H1N1 differs substantially from the HA of seasonal H1N1 strains to which humans have previously been exposed, while seasonal H3N2 has been circulating in humans for several decades5,28. Our data therefore supported the hypothesis that secondary exposure to a highly divergent HA from the same HA group induced strong cross-reactive antibody responses. Notably, (homologous) responses to NA were stronger after infection with H3N2 than after infection with H1N1 (Supplementary Fig. 9).
Evidence for 'original antigenic sin' in humans
Assessing the immediate immune response to acute infection with influenza virus is very important. However, measuring the cross-reactive antibody titers in the general human population in the absence of any immunological perturbation provides better understanding of potential protection in the context of pandemic preparedness. In addition, the baseline titers of cross-reactive antibodies might substantially affect the efficacy of (universal) vaccines against influenza viruses29. Here we analyzed serum samples from 90 subjects from a sex-balanced US cohort.
To gain more insight into the changes of antibody titers over time due to exposure to influenza virus, we stratified the subjects into three age groups: 18–20 years of age ('young'), 33–44 years of age ('middle-aged'; born after the H2N2 subtype stopped circulating) and 49–64 years of age ('experienced'; potential prior exposure to the H2N2 subtype) (Fig. 5a). Subjects in the experienced age group were born when either a drifted version of the 1918 H1N1 virus or the H2N2 pandemic virus circulated. All subjects in this group should therefore have had prior exposure to H2N2 viruses. Subjects in the middle-aged group were born after H2N2 became extinct and was replaced by the group 2 H3N2 virus; therefore, this group had not been previously exposed to H2N2. The young cohort was used as a control group, with an exposure that was limited to recent group 1 and group 2 influenza A virus strains (Fig. 5a). Notably, the different exposure histories of the three age groups led to measurable differences in their cross-reactomes, including the breadth of the antibody response. The 'antigenic landscape' of the young group exhibited high titers of antibodies directed against recent H1 and H3, with low cross-reactivity to moderately distant group 1 HAs (H5) and group 2 HAs (H4 and H14) (Fig. 5b). The middle-aged group had higher reactivity to H1 and H3 HAs (Fig. 5c). Reactivity was particularly high to HAs that are similar to the H1 and H3 viruses that circulated during the childhood of these subjects and was lower to more recent H1 and H3 strains (Fig. 5c). This group also had considerable cross-reactivity to H2 and H5 (group 1) and H4 and H14 (group 2) (Fig. 5c). The experienced group showed medium reactivity to both H1 and H3 but had unexpectedly high reactivity to H2 HA, which is the subtype to which this group might have been first exposed to during childhood (Fig. 5d). This group also exhibited high reactivity to H5 and H18 (Fig. 5d), which suggested that exposure to H2 followed by H1 boosted broadly cross-reactive antibodies30 (endpoint titers, Supplementary Fig. 10). Titers of antibodies to NA were in general low and were confined mostly to the N1 and N2 subtype (Supplementary Fig. 10). An exception were the high titers of antibodies to N2 from the 1957 and 1968 pandemic viruses in the experienced group (Supplementary Fig. 10f). These data provided evidence that childhood exposure to influenza virus induced long-lasting immune responses in adults that we were able to measure by ELISA, which would support the hypothesis of 'original antigenic sin'.
(a) Allocation of subjects by age group (below) according to pre-exposure to the influenza viruses (top) that most likely circulated during the childhood of the subjects (middle bar, year). (b–d) Three-dimensional 'antigenic landscapes' of young subjects (n = 30) (b), middle-aged subjects (n = 30) (c) and experienced subjects (n = 30) (d), generated by plotting of titers of antibodies to HA (geometric mean values; z-axis) against the relative difference of all HAs to each other (based on differences in amino acid sequences; x-axis and y-axis). Data are representative of 30 experiments (one donor in each as biological replicates).
Functionality of human cross-reactive antibodies
Binding of antibodies to diverse HAs and NAs provides general information about the prevalence of cross-reactive antibodies in human serum. However, it is also important to study the biological activity of the antibody responses detected. Antibodies can protect via direct the inhibition or neutralization of virus and/or via effector functions mediated by the crystallizable fragment (Fc) of the antibody. Direct inhibition or neutralization can be assessed by micro-neutralization assays in vitro.
Here we used an assay setup that enhanced sensitivity by using multicycle viral growth in combination with purified immunoglobulin G (IgG) to minimize nonspecific inhibition. We investigated only HA-specific antibodies, which we achieved by using re-assortant viruses that express irrelevant NAs (Supplementary Fig. 10b,d,f). We created six re-assortant viruses on the backbone of influenza virus strain A/PR/8/34 for this purpose: H1N8 (H1 from NC99), H5N8 (H5 from VN04), H9N4 (H9 from gfHoKo99), H3N8 (H3 from Phil82), H4N8 (H4 from duckCzech56) and H7N8 (H7 from Shanghai13). These viruses were selected to determine if the antibody titers measured by ELISA translated into a functional assay. IgG antibodies from the 90 subjects described above were purified, reconstituted to the original concentration in serum and tested against all six viruses.
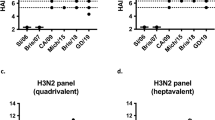
Neutralizing activity directed against H1N8 was high for all age groups, as expected; the frequency of samples that reached a titer of at least 1:40 was as follows: 100% for the young cohort, 93.3% for the middle-aged cohort and 80% for the experienced cohort (Fig. 6a,b). The neutralizing activity directed against H5N8 in samples from all cohorts was low, with only one subject in the middle-aged group reaching a titer of 1:40 (3.3%) (Fig. 6a,b). However, greater neutralizing activity directed against H9N4 was detected, with 23.3% of the young and middle-aged subjects and 33.3% of the experienced subjects developing titers of at least 1:40 (Fig. 6a,b). Similarly, neutralizing activity directed against H3N8 (group 2) was high, with 40% of the young subjects, 100% of the middle-aged subjects and 86.7% of the experienced subjects developing titers of 1:40 (Fig. 6c,d). Neutralizing activity directed against H4N8 (0%, 16.7% and 3.3% for young middle-aged and experienced subjects, respectively) and H7N8 (0%, 13.3% and 10% for young, middle-aged and experienced subjects, respectively) was considerably lower than that directed against H3N8 but was still detectable (Fig. 6c,d).
(a) Individual titers of neutralizing antibodies to group 1 influenza viruses (H1, H5 and H9; re-assortant viruses on the backbone of influenza virus strain A/PR/8/34) in serum from young subjects (blue circles; n = 30), middle-aged subjects (red squares; n = 30) and experienced subjects (teal triangles; n = 30) (age groups (as in Fig. 5a), horizontal axis). (b) Proportion of subjects in each age group as in a (horizontal axis; n = 30 per group) with a titer of 1:40 or higher for neutralization antibodies to group 1 viruses (H1, H5 and H9). (c) Individual titers of neutralizing antibodies to group 2 viruses (H3, H4 and H7; re-assortant viruses as in a) in serum from subjects as in a. (d) Proportion of subjects in each age group as in a (horizontal axis; n = 30 per group) with a titer of 1:40 or higher for neutralization antibodies to group 2 viruses (H3, H4 and H7). (e,f) Viral titers in lungs from mice (n = 5 per group per virus) given commercially available serum depleted of immunoglobulins (No Ig) or pooled serum from subjects of each age group as in a (horizontal axes; n = 30 donors per group), followed by challenge of the host mice with virus containing H3, H4 or H7 HAs and analysis of lungs by plaque assay on day 3 (e) and or day 6 (f) after challenge. PFU, plaque-forming units. Each symbol (a,c,e,f) represents an individual subject (a,c) or host mouse (e,f); bar tops indicate the geometric mean; gray dotted lines indicate the limit of detection.**P ≤ 0.01, ***P ≤ 0.001 and ****P ≤ 0.0001 (ordinary one-way analysis of variance followed by Tukey's multiple-comparisons test (a,c) or Dunnet's multiple-comparisons test (e,f)). Data are representative of 30 experiments per virus and age group (a–d; one donor in each, as biological replicates) or 5 experiments per virus and age group (e,f; five recipient mice per serum pool).
Neutralization titers in general correlated with ELISA titers and pre-exposure history of the specific age groups (Figs. 5 and 6). All subjects were probably exposed to the NC99 H1N1 strain, but titers inversely correlated with age: titers of antibodies directed against H1N8 were slightly higher in the young subjects and were lowest in the experienced subjects. The young subjects lacked a strong neutralizing response to H3N8, which expresses an HA (Phil82) to which they have not been previously exposed. Titers of antibodies directed against H3 were highest in the middle-aged subjects, who were probably exposed to a strain similar to Phil82 H3N2 early in life and were slightly lower in the experienced subjects, who were likely to have been first exposed to H1N1 and/or H2N2 viruses.
In vitro neutralization assays do not capture mechanisms of protection mediated by Fc and its receptor FcR, which might enhance the potency of antibodies in vivo24,26. To explore the in vivo potency of serum from the three cohorts described above, we performed serum-transfer-challenge experiments in the mouse model (Supplementary. Fig. 11). Upon serum transfer, mice were challenged with the H3N8, H4N8 or H7N8 viruses used in the neutralization assay above. Lungs were harvested on day 3 and day 6 after infection, and viral titers in the lungs were assessed by plaque assay. For H3N8, only slightly lower viral titers were seen on day 3 after infection in mice passively immunized with serum relative to the viral titers in control mice that received serum depleted of immunoglobulins (Fig. 6e). However, significantly lower viral titers were observed for all three groups on day 6 relative to the viral titers in the control mice, with the lowest viral titers in mice that received serum from the middle-aged cohort (Fig. 6f). In this case, the reduction in viral titers in the lungs inversely correlated with the measured neutralization titers (Fig. 6c,f). For H4N8, serum transfer reduced viral titers slightly but significantly on day 3 (Fig. 6f). On day 6 after infection, the greatest effect on viral titers was seen for serum from the middle-aged cohort (Fig. 6e,f). This reduction correlated with reactivity to H4 measured by ELISA but not with the findings of the neutralization assay (Figs. 5 and 6). This indicated that these antibodies measured by ELISA mediated their effects via FcR mechanisms. Serum transfer also had a slight effect on the replication of H7N8 both on day 3 and day 6 after infection, but the reduction in lung viral titers never reached significance (Fig. 6e,f). In summary, we demonstrated that cross-reactive antibodies in the general human population were functional both through direct neutralizing activity and through effector function.
Discussion
It has been observed that infection of humans with influenza virus induces immune responses of greater quality, quantity and longevity than does vaccination against influenza virus6,7. Cross-reactive and cross-protective antibodies have become the focus of influenza-virus research, since these antibodies can guide efforts to design broadly protective vaccines and therapeutics directed against influenza virus31. Here we used ELISAs to measure the antibody cross-reactome directed against an extensive panel of influenza virus HAs and NAs (including all know subtypes) induced by infection in animal models and in humans. ELISAs are a very sensitive and useful tool with which to study antibody binding and cross-reactivity. However, they provide only limited insight into the biological activity of the measured responses. Using this technique, we found significant differences in both the breadth and the magnitude of the antibody response in models commonly employed for influenza-virus research.
For all animal models, the immune response to HAs was focused mainly on either group 1 (after infection with H1N1) or group 2 (after infection with H3N2). Antibodies to NA were mostly subtype specific. Mice and guinea pigs induced high titers of antibody, while the responses measured for ferrets were lower. Guinea pigs exhibited an exceptionally broad response relative to that of the other animal models. Ferrets had a low and relatively narrow immune response when assessed by ELISA, even though they are capable of inducing high HI responses to homologous viruses. It has been noted that this HI response is usually focused on specific epitopes and is not reflective of the broad HI response of adult humans32. The extent of viral replication, which differs depending on viral strain and animal model, did not seem to be a major influence on the titers of cross-reactive antibodies. The differences observed should be considered in choosing animal models for research on (universal) vaccines against influenza virus, since the model chosen might strongly influence the outcome of pre-clinical studies. Notably, none of these models accurately reflects the immune response in humans with pre-existing immunity and complicated histories of exposure to influenza virus. This highlights the need for human clinical trials for broadly protective and/or universal vaccines against influenza virus.
In the human population, infection with influenza virus induced substantial antibody cross-reactivity in terms of both magnitude and breadth. The reactivity was narrow after infection with H3N2 but was unexpectedly broad after infection with pandemic H1N1 virus. Also unexpectedly, the induction to group 2 HAs was almost as strong after infection with the pandemic H1N1 virus as after infection with H3N2 (with the exception of the matched H3 HA). This phenomenon might be explained by the greater phylogenetic and antigenic distance between the pandemic H1N1 strain and the pre-pandemic seasonal H1N1 strains (especially the variable head domain of the HAs) to which humans have pre-existing immunity33. It has been noted before that infection with and vaccination against the pandemic H1N1 virus induce antibodies directed against the stalk in humans because they present a novel head domain to the immune system, which then refocuses the antibody response toward the more conserved stalk domain to which memory exists4,5,34. However, the extent of this response has remained unknown so far. Also, an alternative hypothesis for this finding could be the small number of N-linked glycans on the HA of 2009 pandemic H1N1 viruses, in contrast to the large number of N-linked glycans on the HA of recent (e.g., Vic11) H3N2 viruses. While we cannot test this hypothesis in humans, we did not find any clear evidence of an influence of the number of putative N-linked glycosylation sites of the subtrate HAs on cross-reactivity. Only small amounts of serum were available from these infected subjects, which prevented our studying the functionality of these antibodies measured by ELISA. It is therefore not known whether the cross–group-reactive antibodies elicited by infection are also functional in vivo. Another limitation of our study is that the 'pre-infection' serum samples were obtained after the onset of symptoms. Therefore, the true pre-infection titers might be even lower and the antibody induction might be even higher than reported here.
While high titers of cross-reactive antibodies were detected in humans after natural infection, we wanted to further explore the breadth of the cross-reactome in the human population. This information is important, since high baseline reactivity to a specific subtype might ameliorate disease and limit viral spread during a future pandemic with this subtype. We therefore studied the cross-reactome of the general human population by selecting 90 subjects that we grouped by three different age ranges (18–20, 33–44 and 49–64 years of age) and based our analysis on their putative history of exposure to influenza virus.
We found that the cross-reactivity profiles depended on the pre-exposure history and were influenced by the viral strains first encountered during childhood. The young cohort, of subjects who grew up when H1N1 and H3N2 viruses circulated together, showed high reactivity to very recent isolates of these two subtypes, with limited cross-reactivity. The middle-aged cohort, of subjects who were first exposed to H3N2, had high titers of antibodies to H3 and other group 2 HAs but low titers of antibodies to H1 (which they encountered only later in life). Finally, the subjects in the experienced cohort were first exposed to H1N1 or H2N2 and exhibited very high titers of antibodies to H2, H1 and related subtypes such as H5. These findings provide evidence for the phenomenon in which the first HA subtype to which a person is exposed leaves an immunological imprint that will substantially affect the antibody cross-reactome that this person develops ('original antigenic sin'). The high titers of cross-reactive antibodies to group 1 HA in the experienced population might contribute to protection against new pandemic viruses that express group 1 HAs (such as H5, H6 or others). In general, we found that baseline titers of antibodies to H11, H12, H13 and others (group 1) and H7, H10 and H15 (group 2) were low. Of note, H7 influenza viruses35 and, to some extent, also H10 influenza viruses36, have infected humans in Asia with a high case-fatality rate.
Notably, our findings also translated into in vitro and in vivo functional assays. In a virus-neutralization assay, we found that the group of middle-aged subjects with the highest titers of antibodies to H3 Phil82, as measured by ELISA, also most effectively neutralized an H3N8 virus based on this HA. Similarly, the group of young subjects had high titers of neutralizing antibodies to an H1N8 virus based on NC99, which is an HA from a strain that circulated during their childhood. Furthermore, these antibodies also conferred protection in an in vivo serum-transfer model.
In conclusion, we have created antigenic landscapes that describe the antibody cross-reactome directed against the glycoproteins of influenza virus in animal models and humans recently infected with influenza viruses. We found that the prevalence and breadth of the antibody cross-reactome of the general human population varied largely and depended on the individual history of exposure to influenza viruses. These data provide information for pandemic preparedness and the choice of animal models for the development of broadly protective vaccines against influenza virus. Finally, the wide prevalence of cross-reactive antibodies in humans would suggest that future strategies of universal vaccines that target the HA head, stalk or NA might be successful in boosting these antibody levels to protective titers.
Methods
Cells, viruses and proteins.
Madin-Darby canine kidney (MDCK) cells and 293T human embryo kidney cells were maintained in complete Dulbecco's modified Eagle's medium (cDMEM, Gibco) containing 10% FBS (FBS, Gibco) and pen-strep mix (100 units/ml of penicillin and 100 μg/ml of streptomycin, Gibco). Influenza viruses were grown in 8- to 10-day-old embryonated chicken eggs (Charles River Laboratories). Re-assortants were rescued using a previously described protocol37. A full list of viruses and abbreviations is in Supplementary Table 2. Sf9 cells were maintained in TNM-FH medium (Gemini Bio-Products) in the presence of 10% FBS and pen-strep mix. BTI-TN5B1-4 cells38 were maintained in SFX medium (HyClone) containing Pen-strep mix. Recombinant proteins were expressed as described in detail before13,39,40. All HAs were expressed as ectodomains fused to a C-terminal fold on trimerization domain and a hexahistidine tag for purification. All NAs were expressed as ectodomains with an N-terminal hexahistidine tag and vasodilator-stimulated phosphoprotein tetramerization domain. The design of the constructs were identical to designs reported to produce high-quality recombinant HAs and NAs for crystallographic studies41,42. All proteins were purified using standard operation procedures that have been established in the laboratory to guarantee consistent quality of the recombinant proteins40. A list of recombinant HA and NA proteins used in this study is in Supplementary Table 3.
Animal infection.
All of the animal experiments described here were conducted with protocols approved by the Icahn School of Medicine at Mount Sinai Institutional Animal Care and Use Committee. Infections and blood draws were performed on animals anesthetized with ketamine-xylazine. Ferrets received 0.45 ml of ketamine-xylazine mixture intramuscularly; guinea pigs received 200 l of 30 mg of ketamine per kg body weight (mg/kg) and 5 mg/kg of xylazine intramuscularly; and mice received 0.1 ml of 0.15 mg/kg of ketamine and 0.03 mg/kg of xylazine intraperitoneally. Animals were sequentially infected with sublethal doses of two divergent viruses of the same subtype 6 weeks apart. For H1N1, the first infection was performed with influenza virus strain A/New Caledonia/20/99 (1 × 105 plaque-forming units (PFU) per mouse in 50 μl, n = 10; 1 × 106 PFU per guinea pig in 300 μl, n = 3; 1 × 106 PFU per ferret in 1000 μl, n = 2) followed by an infection with influenza virus strain A/California/04/09 (1 × 104 PFU per mouse in 50 μl; 1 × 106 PFU per guinea pig in 300 μl; 1 × 106 PFU per ferret in 1000 μl). For H3N2, the first infection was performed with influenza virus strain A/Philippines/2/82 (1 × 105 PFU per mouse in 50 μl, n = 10; 1 × 106 PFU per guinea pig in 300 μl, n = 3; 1 × 106 PFU per ferret in 1000 μl, n = 2) followed by influenza virus strain A/Victoria/361/11 (5 × 105 PFU per mouse in 50 μl; 1 × 106 PFU per guinea pig in 300 μl; 1 × 106 PFU per ferret in 1000 μl). Serum was collected on day 42 (after the first infection) and day 84 (after the second infection). Serum pools of naive animals were used to establish background reactivity in ELISA. Samples for ferrets and guinea pigs were analyzed individually, and geometric mean values for titers are reported. Serum from one guinea pig given repeated infection with H3N2 could not be used, and geometric mean value of the titers of the remaining two animals are reported. Mouse samples for each time point and group were pooled and analysis was performed in technical duplicates.
Serum samples from subjects infected with influenza A virus.
Clinical–epidemiological data, along with a blood samples, were collected after informed written consent was obtained under protocol 11-116, reviewed and approved by the Scientific Ethics Committee of the School of Medicine at Pontificia Universidad Católica de Chile before the start of sample collection. Matched human serum samples, from day 1 and days 21 or 28 after infection, were obtained and archived from 19 hospitalized patients infected with human influenza A virus between years 2011 and 2013 in Santiago, Chile (Supplementary Table 1). All personal information about samples was removed of any before blinded analysis (IRB exempt, HS#: 1500125). Infection with influenza A virus was confirmed by qRT-PCR of viral RNA extracted from nasopharyngeal samples by the Clinical Virology Laboratory at Pontificia Universidad Católica de Chile (PUC). Positive samples were subtyped as influenza A strain H1N1pdm09 or H3N2 by qRT-PCR analysis and/or confirmed by hemagglutinin-inhibition (HI) assays. Serum volumes for two patients were not sufficient to complete all testing. For patient p3/2011 (infected with pH1N1), reactivity to N8 and H3 HK68 could not be tested. For patient p36/2012, reactivity to H2 Jap57, H3 seMass11, H10, H11, H12, H13, H16, H18, cH5/3, N1 Texa91, N2 Vic11, N2 Sing57, N3, N4, N5, N6, N7, N9, N10 and B NA could not be tested.
Human cohort serum.
Serum samples from 90 humans were ordered as research reagents from Innovative Research. The subjects were chosen to be in three separate age groups (18–20, 33–44 and 49–64 years of age) and to obtain similar male/female ratios for each group (Supplementary Table 4). All samples were collected between August 2014 and November 2014 and were received without personal information about the donors, except for age, race and sex.
Enzyme linked immunosorbent assay (ELISA).
ELISAs were performed as previously described25. In short, 96-well high-binding, flat-bottomed plates were coated with 50 μl/well of recombinant protein at a concentration of 2 μg/ml in coating buffer (50 mM sodium carbonate and 50 mM sodium hydrogen carbonate, pH 9.4) and were incubated overnight at 4 °C. The coating buffer was removed and plates were incubated for 1 h at 25 °C with 220 μl of blocking solution (mouse: PBS with 0.5% Tween-20 and 3% milk powder; human, ferret and guinea pig: PBS with 0.5% Tween-20, 3% goat serum and 0.5% milk powder). Serum was serially diluted threefold in blocking solution and plates were incubated for two hours at 25 °C. Plates were washed three times with PBS-T (PBS with 0.5% Tween-20) and 50 μl of IgG-specific secondary antibody diluted in blocking solution added to each well (mouse: 1:3,000, Sigma #A9044; ferret: 1:5,000, Alpha Diagnostic International #70530; guinea pig: 1:3,000, Millipore #AP108P; human: 1:3,000, Sigma #A0293). After 1 h of incubation at 25 °C, plates were washed three times with PBS-T. Plates were developed with 100 μl of SigmaFast OPD (Sigma) and this was stopped after 10 min by the addition of 3 M hydrochloric acid. Plates were then read at a wavelength of 490 nm. For mice, guinea pigs and ferrets, the cut-off for endpoint titers was calculated as the mean and three standard deviations of all blank wells. For human samples, a standard cut-off of 0.07 was used. Plates were discarded if the background and three standard deviations exceeded the standard cut-off.
Microneutralization assay.
For increased sensitivity, without nonspecific inhibition of viruses by other serum proteins, IgG from human cohort serum was purified with protein G columns and was reconstituted to the original serum volume in PBS. MDCK cells were seeded in 96-well tissue culture plates at a density of 1.5 × 104 to 1.8 × 104 cells/well in 100 μl of cDMEM and were incubated at 37 °C overnight. IgG was serially diluted twofold in infection medium (minimal essential medium containing trypsin (treated with tosyl phenylalanyl chloromethyl ketone) at a concentration of 1 μg/ml), starting with a dilution of 1:2. 50 μl of serially diluted IgG was incubated for one hour at 25 °C with 50 μl of virus diluted to 200 PFU per 50 μl. Cells were washed once with PBS, then 100 μl of virus–antibody mixture was transferred to each well and plates were incubated at 37 °C for 1 h. Cells were washed once with PBS, then 50 μl of serially diluted antibody at the original concentration, as well as 50 μl of infection medium, were added to each well. Plates were incubated for 48–72 h and were read by hemagglutination assay. Wells containing virus only without antibodies served as a positive control.
Serum-transfer experiments in mice.
Serum samples from humans of each age group were pooled separately and filtrated in sterile way. Commercially available immunoglobulin-depleted human serum (BBI Solutions, #SF5050-2) was used as negative control. For each virus tested, 10 mice per serum group were given intraperitoneal injection of 250 μl of serum. 2 h later, mice anesthetized with ketamine-xylazine were infected intranasally with 1 × 105 PFU of virus in 50 μl (diluted in PBS). Five mice each per serum group were euthanized on day 3and day 6 after infection. Lungs were extracted and homogenized in 600 μl of PBS, and the cell debris removed by centrifugation. Aliquots were stored at −80 °C until the viruses were titered by plaque assay as previously described25.
Graph presentation.
All data graphs, except for the three-dimensional antibody landscapes, were generated in GraphPad Prism 7. To visualize antibody titers of human serum samples against different HA subtypes, three-dimensional 'antibody landscapes'43 were constructed. In these 'landscapes', the distance between points in the horizontal plane (x-y coordinates) represents the amino-acid-sequence distance among strains, and the height (z coordinate) represents the titers of antibodies (geometric mean values) to corresponding strains on the horizontal plane. The horizontal plane was constructed by multi-dimensional scaling of the amino-acid-sequence distance44,45. The sequence distances among strains were defined as the total number of amino acids that were different between corresponding HAs in the multiple sequence alignment of all the HAs used in this study. The sum of squared errors between the Euclidean distance in the two-dimensional plane and the HA sequence distance was minimized by the SMACOF algorithm45. The HA sequences were divided into two HA groups: one included H1, H2, H5, H6, H8, H9, H11, H12, H13, H16, H17 and H18, and the other included H3, H4, H7, H10, H14 and H15. For each HA group, the surface of the 'antibody landscape' was approximated from antibody titers (geometric mean values) using multilevel B-splines. We used the mba.surf function implemented in the Multilevel B-spline Approximation package in R version 3.2.5.
Statistical analysis.
Statistical analysis was performed with GraphPad Prism 7. Microneutralization titers were compared by ordinary one-way ANOVAs followed by Tukey's multiple comparisons tests. The viral lung titers were compared by ordinary one-way ANOVAs followed by Dunnett's multiple comparison tests with the no-Ig group serving as the control group. The nonlinear regression function 'Plateau followed by one phase decay' was used to create nonlinear fit curves for the ELISA data.
Data availability.
The data that support the findings of this study are available from the corresponding author upon request.
References
Jayasundara, K., Soobiah, C., Thommes, E., Tricco, A.C. & Chit, A. Natural attack rate of influenza in unvaccinated children and adults: a meta-regression analysis. BMC Infect. Dis. 14, 670 (2014).
Krammer, F. & Palese, P. Advances in the development of influenza virus vaccines. Nat. Rev. Drug Discov. 14, 167–182 (2015).
Horimoto, T. & Kawaoka, Y. Influenza: lessons from past pandemics, warnings from current incidents. Nat. Rev. Microbiol. 3, 591–600 (2005).
Wrammert, J. et al. Broadly cross-reactive antibodies dominate the human B cell response against 2009 pandemic H1N1 influenza virus infection. J. Exp. Med. 208, 181–193 (2011).
Pica, N. et al. Hemagglutinin stalk antibodies elicited by the 2009 pandemic influenza virus as a mechanism for the extinction of seasonal H1N1 viruses. Proc. Natl. Acad. Sci. USA 109, 2573–2578 (2012).
Margine, I. et al. H3N2 influenza virus infection induces broadly reactive hemagglutinin stalk antibodies in humans and mice. J. Virol. 87, 4728–4737 (2013).
Yu, X. et al. Neutralizing antibodies derived from the B cells of 1918 influenza pandemic survivors. Nature 455, 532–536 (2008).
Ohmit, S.E., Petrie, J.G., Cross, R.T., Johnson, E. & Monto, A.S. Influenza hemagglutination-inhibition antibody titer as a correlate of vaccine-induced protection. J. Infect. Dis. 204, 1879–1885 (2011).
Heaton, N.S., Sachs, D., Chen, C.J., Hai, R. & Palese, P. Genome-wide mutagenesis of influenza virus reveals unique plasticity of the hemagglutinin and NS1 proteins. Proc. Natl. Acad. Sci. USA 110, 20248–20253 (2013).
Yewdell, J.W., Webster, R.G. & Gerhard, W.U. Antigenic variation in three distinct determinants of an influenza type A haemagglutinin molecule. Nature 279, 246–248 (1979).
Wohlbold, T.J. & Krammer, F. In the shadow of hemagglutinin: a growing interest in influenza viral neuraminidase and its role as a vaccine antigen. Viruses 6, 2465–2494 (2014).
Yang, X. et al. A beneficiary role for neuraminidase in influenza virus penetration through the respiratory mucus. PLoS One 9, e110026 (2014).
Wohlbold, T.J. et al. Vaccination with adjuvanted recombinant neuraminidase induces broad heterologous, but not heterosubtypic, cross-protection against influenza virus infection in mice. MBio 6, e02556 (2015).
Easterbrook, J.D. et al. Protection against a lethal H5N1 influenza challenge by intranasal immunization with virus-like particles containing 2009 pandemic H1N1 neuraminidase in mice. Virology 432, 39–44 (2012).
Ekiert, D.C. et al. Antibody recognition of a highly conserved influenza virus epitope. Science 324, 246–251 (2009).
Ekiert, D.C. et al. A highly conserved neutralizing epitope on group 2 influenza A viruses. Science 333, 843–850 (2011).
Sui, J. et al. Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses. Nat. Struct. Mol. Biol. 16, 265–273 (2009).
Dreyfus, C. et al. Highly conserved protective epitopes on influenza B viruses. Science 337, 1343–1348 (2012).
Jegaskanda, S., Weinfurter, J.T., Friedrich, T.C. & Kent, S.J. Antibody-dependent cellular cytotoxicity is associated with control of pandemic H1N1 influenza virus infection of macaques. J. Virol. 87, 5512–5522 (2013).
Jegaskanda, S. et al. Cross-reactive influenza-specific antibody-dependent cellular cytotoxicity antibodies in the absence of neutralizing antibodies. J. Immunol. 190, 1837–1848 (2013).
Laidlaw, B.J. et al. Cooperativity between CD8+ T cells, non-neutralizing antibodies, and alveolar macrophages is important for heterosubtypic influenza virus immunity. PLoS Pathog. 9, e1003207 (2013).
DiLillo, D.J., Palese, P., Wilson, P.C. & Ravetch, J.V. Broadly neutralizing anti-influenza antibodies require Fc receptor engagement for in vivo protection. J. Clin. Invest. 126, 605–610 (2016).
Terajima, M. et al. Complement-dependent lysis of influenza a virus-infected cells by broadly cross-reactive human monoclonal antibodies. J. Virol. 85, 13463–13467 (2011).
DiLillo, D.J., Tan, G.S., Palese, P. & Ravetch, J.V. Broadly neutralizing hemagglutinin stalk-specific antibodies require FcγR interactions for protection against influenza virus in vivo. Nat. Med. 20, 143–151 (2014).
Nachbagauer, R. et al. Induction of broadly reactive anti-hemagglutinin stalk antibodies by an H5N1 vaccine in humans. J. Virol. 88, 13260–13268 (2014).
Henry Dunand, C.J. et al. Both neutralizing and non-neutralizing human H7N9 influenza vaccine-induced monoclonal antibodies confer protection. Cell Host Microbe 19, 800–813 (2016).
Krause, J.C. et al. Human monoclonal antibodies to pandemic 1957 H2N2 and pandemic 1968 H3N2 influenza viruses. J. Virol. 86, 6334–6340 (2012).
Smith, G.J. et al. Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459, 1122–1125 (2009).
Krammer, F. & Palese, P. Universal influenza virus vaccines: need for clinical trials. Nat. Immunol. 15, 3–5 (2014).
Miller, M.S. et al. Neutralizing antibodies against previously encountered influenza virus strains increase over time: a longitudinal analysis. Sci. Transl. Med. 5, 198ra107 (2013).
Krammer, F. & Palese, P. Influenza virus hemagglutinin stalk-based antibodies and vaccines. Curr. Opin. Virol. 3, 521–530 (2013).
Li, Y. et al. Immune history shapes specificity of pandemic H1N1 influenza antibody responses. J. Exp. Med. 210, 1493–1500 (2013).
Garten, R.J. et al. Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science 325, 197–201 (2009).
Li, G.M. et al. Pandemic H1N1 influenza vaccine induces a recall response in humans that favors broadly cross-reactive memory B cells. Proc. Natl. Acad. Sci. USA 109, 9047–9052 (2012).
Gao, R. et al. Human infection with a novel avian-origin influenza A (H7N9) virus. N. Engl. J. Med. 368, 1888–1897 (2013).
Chen, H. et al. Clinical and epidemiological characteristics of a fatal case of avian influenza A H10N8 virus infection: a descriptive study. Lancet 383, 714–721 (2014).
Martínez-Sobrido, L. & García-Sastre, A. Generation of recombinant influenza virus from plasmid DNA. J. Vis. Exp. 42, 2057 (2010).
Krammer, F. et al. Trichoplusia ni cells (High Five) are highly efficient for the production of influenza A virus-like particles: a comparison of two insect cell lines as production platforms for influenza vaccines. Mol. Biotechnol. 45, 226–234 (2010).
Krammer, F. et al. A carboxy-terminal trimerization domain stabilizes conformational epitopes on the stalk domain of soluble recombinant hemagglutinin substrates. PLoS One 7, e43603 (2012).
Margine, I., Palese, P. & Krammer, F. Expression of functional recombinant hemagglutinin and neuraminidase proteins from the novel H7N9 influenza virus using the baculovirus expression system. J. Vis. Exp. 81, e51112 (2013).
Xu, X., Zhu, X., Dwek, R.A., Stevens, J. & Wilson, I.A. Structural characterization of the 1918 influenza virus H1N1 neuraminidase. J. Virol. 82, 10493–10501 (2008).
Stevens, J. et al. Structure of the uncleaved human H1 hemagglutinin from the extinct 1918 influenza virus. Science 303, 1866–1870 (2004).
Fonville, J.M. et al. Antibody landscapes after influenza virus infection or vaccination. Science 346, 996–1000. (2014).
Ito, K. et al. Gnarled-trunk evolutionary model of influenza A virus hemagglutinin. PLoS One 6, e25953 (2011).
Borg, I. & Groenen, P. Modern Multidimensional Scaling: Theory and Applications (Springer, 2005).
Acknowledgements
We thank L. Aguado for cloning of several of the HA expression vectors; J. Runstadler (Massachusetts Institute of Technology) for making several avian influenza viruses available to us; F. Amanat for IgG purification; P. E. Leon (Icahn School of Medicine at Mount Sinai) for the H3N8 and H9N4 viruses; C. Marizzi for reviewing the manuscript; BEI Resources for sequences of influenza virus HA and NA; and R. Fouchier (Erasmus MC) for depositing plasmids encoding HA and NA at BEI Resources. Supported by the US National Institutes of Health (U19 AI109946-01 to P.P. and F.K.; and R01 AI117287-01A1 to F.K. and N.M.B.), Centers of Excellence in Influenza Virus Research and Surveillance of the US National Institutes of Health (HHSN272201400008C to A.G.S., P.P., F.K., R.A.M. and N.M.B.), Comisión Nacional de Investigación Científica y Tecnológica (FONDECYT 1121172 and 1161791 to R.A.M.; and PIA ACT 1408 to R.A.M.), and the Chilean Ministry of Economy, Development and Tourism (P09/016-F to R.A.M.).
Author information
Authors and Affiliations
Contributions
R.N., A.C., A.H., I.M. and F.K. performed serological experiments. R.N., A.C., I.M., N.M.B., R.A.A. and F.K. performed animal experiments. A.B., M.F. and R.A.M. acquired clinical samples and performed all necessary experiments to characterize the clinical samples. R.N., S.I., A.G.S., K.I., R.A.M. and F.K. performed data analysis. R.N., R.A.A., P.P., A.G.S. and F.K. contributed to the overall concept, experimental design and hypothesis and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Infection strategy.
To test the antibody responses against influenza viruses, animals were sequentially infected with two divergent strains of the same subtype. For H1N1 infections, animals were primed with a human seasonal strain from 1999 (NC99, A/New Caledonia/20/1999), followed by the 2009 human pandemic H1N1 strain (NL09, A/Netherlands/602/2009) six weeks later. These viruses express divergent HA head domains, but similar HA stalk domains. For H3N2 infection animals were primed with a human seasonal strain from 1982 (Phil82, A/Philippines/2/1982), followed by a 2011 human seasonal strain (Vic11, A/Victoria/361/2011). These viruses are separated by almost 30 years of antigenic drift.
Supplementary Figure 2 Viral titers on day 2 after infection in animal models.
Mice (A, n=5/virus), Guinea pigs (B, n=2/virus) and ferrets (C, n=2/virus) were infected with NC99, NL09, Phil82 and Cal09 and viral titers were measured on day 2 post infection from nasal wash (guinea pigs and ferrets) or nasal turbinates (mice).
Supplementary Figure 3 Heat-map presentation of guinea pig antibody titers after infection.
A) Anti-HA titers of guinea pigs (n=3) after one H1N1 (NC99) infection. B) Anti-HA titers of guinea pigs (n=3) after sequential infection with two divergent H1N1 (NC99, NL09) strains. C) Anti-HA titers of guinea pigs (n=3) after one H3N2 (Phil82) infection. D) Anti-HA titers of guinea pigs (n=2) after sequential infection with two divergent H3N2 (Phil82, Vic11) strains. E-H) Anti-NA titers of the corresponding sera to panels A-D.
Supplementary Figure 4 Heat-map presentation of ferret antibody titers after infection.
A) Anti-HA titers of ferrets (n=2) after one H1N1 (NC99) infection. B) Anti-HA titers of ferrets (n=2) after sequential infection with two divergent H1N1 (NC99, NL09) strains. C) Anti-HA titers of ferrets (n=2) after one H3N2 (Phil82) infection. D) Anti-HA titers of ferrets (n=2) after sequential infection with two divergent H3N2 (Phil82, Vic11) strains. E-H) Anti-NA titers of the corresponding sera to panels A-D.
Supplementary Figure 5 Profiles of the titers of cross-reactive antibodies after a single infection in animal models.
Antibody titers measured by ELISA after single influenza virus infection were plotted on the y-axis and the percent amino acid difference to the HA of the strain used for the (later) second infection was plotted on the x-axis. Each point represents the geometric mean titer measured against a single HA (dark blue dot for H1 HAs, light blue dot for other group 1 HAs, dark red triangle for H3 HAs, light red triangle for other group 2 HAs). A non-linear fit (plateau followed by one phase decay) was performed to illustrate the differences in the breadth of the antibody response in all animals. Points for HAs with titers lower than 103 are plotted at 103. A) Mouse group 1 HA titers after H1N1 (NC99) infection. ELISAs were performed on pooled sera of 10 mice and geometric mean titers of technical duplicates are shown. B) Guinea pig group 1 HA titers after H1N1 (NC99) infection. ELISAs were performed on individual sera of 3 guinea pigs and geometric mean titers are shown. C) Ferret group 1 HA titers after H1N1 (NC99) infection. ELISAs were performed on individual sera of 2 ferrets and geometric mean titers are shown. D) Mouse group 2 HA titers after H3N2 (Phil82) infection. ELISAs were performed on pooled sera of 10 mice and geometric mean titers of technical duplicates are shown. E) Guinea pig group 2 HA titers after H3N2 (Phil82) infection. ELISAs were performed on individual sera of 3 guinea pigs and geometric mean titers are shown. F Ferret group 2 HA titers after H3N2 (Phil82) infection. ELISAs were performed on individual sera of 2 ferrets and geometric mean titers are shown.
Supplementary Figure 6 Glycosylation versus antibody titers after infection.
A-C) Antibody titers post 2x infection with either H1N1 (circles) or H3N2 (triangles) against H1 (blue) or H3 (red) HAs were plotted against the number of corresponding glycosylation sites of each HA for all animal models. Mouse samples show the geometric mean of technical duplicates of pooled sera from 10 mice per virus. Guinea pigs show the geometric mean titers of individual animals (H1N1: n=3, H3N2: n=2). For ferrets, the geometric mean titers of individual animals (n=2/virus) is shown. D) Human antibody titers (geometric mean) post pH1N1 (Cal09, n=9) or H3N2 (Vic11, n=10) infection were plotted against the number of glycosylation sites of the corresponding HAs. Circles indicate titers for individuals infected with pH1N1 and triangles indicate titers for individuals infected with H3N2. Symbols for titers measured against H1 HAs are colored red and symbols for titers measured against H3 HAs are colored blue.
Supplementary Figure 7 Profiles of the titers of cross-reactive antibodies after infection of humans with influenza virus.
A) Geometric mean antibody titers of human individuals post pH1N1 infection (n=9) were plotted on the y-axis and the percent amino acid difference to the HA of the infection strain was plotted on the x-axis. Each point represents the geometric mean titer of all individuals measured against a single HA. Dots illustrate group 1 HAs, triangles group 2 HAs and squares influenza B HA. The post-infection titers are shown in color (blue for group 1, red for group 2 and green for influenza B) and the corresponding pre-infection titers are plotted in gray. A non-linear fit (plateau followed by one phase decay) was performed to illustrate the breadth of the antibody response. B) Geometric mean antibody titers of human individuals post H3N2 infection (n=10) were plotted in the same manner as panel A.
Supplementary Figure 8 ELISA of the titers of antibody to HA in humans before and after infection.
A) Heat map representation of pre-pH1N1 infection ELISA antibody titers against HA (geometric mean of 9 individuals). B) Heat map representation of post-pH1N1 infection ELISA antibody titers against HA (geometric mean of 9 individuals). C) Heat map representation of pre-H3N2 infection ELISA antibody titers against HA (geometric mean of 10 individuals). B) Heat map representation of post-H3N2 infection ELISA antibody titers against HA (geometric mean of 10 individuals).
Supplementary Figure 9 Human reactivity to NA before and after infection.
Heat map representations of human geometric mean ELISA antibody titers against NA pre- and post-pH1N1 (A, B, n=9) or H3N2 (C, D, n=10) infections. E-F) Heat map representations of geometric mean fold-induction of antibody titers against NA post H1N1 (E, n=9) or H3N2 (F, n=10) infection in humans. G-H) Geometric mean fold-induction of antibody titers against NA post-pH1N1 (n=9) or post-H3N2 (n=10) infection in humans were sorted by highest induction in a bar graph. Blue bars represent group 1 NAs, red bars group 2 NAs, green influenza B NA and grey the N10 bat isolate. Error bars indicate the 95% confidence intervals.
Supplementary Figure 10 Heat-map presentation of ELISA of titers of human antibodies to HA and NA (geometric mean values), separated by age group.
A) Geometric mean anti-HA titers of 18-20 year olds (n=30). B) Geometric mean anti-NA titers of 18-20 year olds (n=30). C) Geometric mean anti-HA titers of 33-44 year olds (n=30). D) Geometric mean anti-NA titers of 33-44 year olds (n=30). E) Geometric mean anti-HA titers of 49-64 year olds (n=30). F) Geometric mean anti-NA titers of 49-64 year olds (n=30).
Supplementary Figure 11 Serum-transfer experiment.
For each virus challenge, serum samples of 30 individuals per age group were pooled, sterile filtrated and 250 μl of serum intraperitoneally injected into 10 mice for each group. An equal number of mice received immunoglobulin depleted serum as a negative control group. Two hours after the serum transfer, mice were anesthetized and infected with 1 × 105 PFU of virus diluted in PBS (50 μl total volume) intranasally. Five mice each were euthanized on days 3 and 6, their lungs extracted and viral titers measured by plaque assay.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Tables 1–4 (PDF 2148 kb)
Rights and permissions
About this article
Cite this article
Nachbagauer, R., Choi, A., Hirsh, A. et al. Defining the antibody cross-reactome directed against the influenza virus surface glycoproteins. Nat Immunol 18, 464–473 (2017). https://doi.org/10.1038/ni.3684
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3684
- Springer Nature America, Inc.
This article is cited by
-
Bat-borne H9N2 influenza virus evades MxA restriction and exhibits efficient replication and transmission in ferrets
Nature Communications (2024)
-
Immunization with matrix-, nucleoprotein and neuraminidase protects against H3N2 influenza challenge in pH1N1 pre-exposed pigs
npj Vaccines (2023)
-
Protein engineering strategies for rational immunogen design
npj Vaccines (2021)
-
AS03-adjuvanted H7N9 inactivated split virion vaccines induce cross-reactive and protective responses in ferrets
npj Vaccines (2021)
-
Influenza virus infection expands the breadth of antibody responses through IL-4 signalling in B cells
Nature Communications (2021)