Abstract
When B cells encounter an antigen, they alter their physiological state and anatomical localization and initiate a differentiation process that ultimately produces antibody-secreting cells (ASCs). We have defined the transcriptomes of many mature B cell populations and stages of plasma cell differentiation in mice. We provide a molecular signature of ASCs that highlights the stark transcriptional divide between B cells and plasma cells and enables the demarcation of ASCs on the basis of location and maturity. Changes in gene expression correlated with cell-division history and the acquisition of permissive histone modifications, and they included many regulators that had not been previously implicated in B cell differentiation. These findings both highlight and expand the core program that guides B cell terminal differentiation and the production of antibodies.







Similar content being viewed by others
Accession codes
References
Nutt, S.L., Hodgkin, P.D., Tarlinton, D.M. & Corcoran, L.M. The generation of antibody-secreting plasma cells. Nat. Rev. Immunol. 15, 160–171 (2015).
Fujio, K., Okamura, T., Sumitomo, S. & Yamamoto, K. Regulatory cell subsets in the control of autoantibody production related to systemic autoimmunity. Ann. Rheum. Dis. 72 (suppl. 2) ii85–ii89 (2013).
Potter, M. Neoplastic development in plasma cells. Immunol. Rev. 194, 177–195 (2003).
Fairfax, K.A. et al. Different kinetics of blimp-1 induction in B cell subsets revealed by reporter gene. J. Immunol. 178, 4104–4111 (2007).
Radbruch, A. et al. Competence and competition: the challenge of becoming a long-lived plasma cell. Nat. Rev. Immunol. 6, 741–750 (2006).
Tarlinton, D. & Good-Jacobson, K. Diversity among memory B cells: origin, consequences, and utility. Science 341, 1205–1211 (2013).
Nutt, S.L., Taubenheim, N., Hasbold, J., Corcoran, L.M. & Hodgkin, P.D. The genetic network controlling plasma cell differentiation. Semin. Immunol. 23, 341–349 (2011).
Shapiro-Shelef, M. & Calame, K. Regulation of plasma-cell development. Nat. Rev. Immunol. 5, 230–242 (2005).
Shaffer, A.L. et al. Blimp-1 orchestrates plasma cell differentiation by extinguishing the mature B cell gene expression program. Immunity 17, 51–62 (2002).
Shaffer, A.L. et al. XBP1, downstream of Blimp-1, expands the secretory apparatus and other organelles, and increases protein synthesis in plasma cell differentiation. Immunity 21, 81–93 (2004).
Cocco, M. et al. In vitro generation of long-lived human plasma cells. J. Immunol. 189, 5773–5785 (2012).
Jourdan, M. et al. An in vitro model of differentiation of memory B cells into plasmablasts and plasma cells including detailed phenotypic and molecular characterization. Blood 114, 5173–5181 (2009).
Peperzak, V. et al. Mcl-1 is essential for the survival of plasma cells. Nat. Immunol. 14, 290–297 (2013).
Sciammas, R. et al. Graded expression of interferon regulatory factor-4 coordinates isotype switching with plasma cell differentiation. Immunity 25, 225–236 (2006).
Kallies, A. et al. Plasma cell ontogeny defined by quantitative changes in blimp-1 expression. J. Exp. Med. 200, 967–977 (2004).
Liao, Y., Smyth, G.K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
Liao, Y., Smyth, G.K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
O'Connor, B.P. et al. BCMA is essential for the survival of long-lived bone marrow plasma cells. J. Exp. Med. 199, 91–98 (2004).
Delogu, A. et al. Gene repression by Pax5 in B cells is essential for blood cell homeostasis and is reversed in plasma cells. Immunity 24, 269–281 (2006).
Wrammert, J., Kallberg, E., Agace, W.W. & Leanderson, T. Ly6C expression differentiates plasma cells from other B cell subsets in mice. Eur. J. Immunol. 32, 97–103 (2002).
Nakayama, T. et al. Cutting edge: profile of chemokine receptor expression on human plasma cells accounts for their efficient recruitment to target tissues. J. Immunol. 170, 1136–1140 (2003).
Hauser, A.E. et al. Chemotactic responsiveness toward ligands for CXCR3 and CXCR4 is regulated on plasma blasts during the time course of a memory immune response. J. Immunol. 169, 1277–1282 (2002).
Kunkel, E.J. et al. CCR10 expression is a common feature of circulating and mucosal epithelial tissue IgA Ab-secreting cells. J. Clin. Invest. 111, 1001–1010 (2003).
Cerutti, A., Cols, M. & Puga, I. Marginal zone B cells: virtues of innate-like antibody-producing lymphocytes. Nat. Rev. Immunol. 13, 118–132 (2013).
Jeong, H.D. & Teale, J.M. Contribution of the CD5+ B cell to D-proximal VH family expression early in ontogeny. J. Immunol. 145, 2725–2729 (1990).
Herzenberg, L.A., Baumgarth, N. & Wilshire, J.A. B-1 cell origins and VH repertoire determination. Curr. Top. Microbiol. Immunol. 252, 3–13 (2000).
Metcalf, D. et al. Gigantism in mice lacking suppressor of cytokine signalling-2. Nature 405, 1069–1073 (2000).
Hasbold, J., Corcoran, L.M., Tarlinton, D.M., Tangye, S.G. & Hodgkin, P.D. Evidence from the generation of immunoglobulin G–secreting cells that stochastic mechanisms regulate lymphocyte differentiation. Nat. Immunol. 5, 55–63 (2004).
Hodgkin, P.D., Lee, J.H. & Lyons, A.B. B cell differentiation and isotype switching is related to division cycle number. J. Exp. Med. 184, 277–281 (1996).
Rush, J.S., Liu, M., Odegard, V.H., Unniraman, S. & Schatz, D.G. Expression of activation-induced cytidine deaminase is regulated by cell division, providing a mechanistic basis for division-linked class switch recombination. Proc. Natl. Acad. Sci. USA 102, 13242–13247 (2005).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Whyte, W.A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Chu, V.T. & Berek, C. The establishment of the plasma cell survival niche in the bone marrow. Immunol. Rev. 251, 177–188 (2013).
Kurtz, J.E. & Dufour, P. Adecatumumab: an anti-EpCAM monoclonal antibody, from the bench to the bedside. Expert Opin. Biol. Ther. 10, 951–958 (2010).
Lu, R. & Wang, G.G. Tudor: a versatile family of histone methylation 'readers'. Trends Biochem. Sci. 38, 546–555 (2013).
Xiao, W. et al. U19/Eaf2 knockout causes lung adenocarcinoma, B-cell lymphoma, hepatocellular carcinoma and prostatic intraepithelial neoplasia. Oncogene 27, 1536–1544 (2008).
Korthuis, P.M. et al. CITED2-mediated human hematopoietic stem cell maintenance is critical for acute myeloid leukemia. Leukemia 29, 625–635 (2015).
Knosp, C.A. et al. SOCS2 regulates T helper type 2 differentiation and the generation of type 2 allergic responses. J. Exp. Med. 208, 1523–1531 (2011).
Zak, D.E., Tam, V.C. & Aderem, A. Systems-level analysis of innate immunity. Annu. Rev. Immunol. 32, 547–577 (2014).
Basso, K. et al. Reverse engineering of regulatory networks in human B cells. Nat. Genet. 37, 382–390 (2005).
Yosef, N. et al. Dynamic regulatory network controlling TH17 cell differentiation. Nature 496, 461–468 (2013).
Perfetti, V. et al. Membrane CD22 defines circulating myeloma-related cells as mature or later B cells. Lab. Invest. 77, 333–344 (1997).
Zhang, J.A., Mortazavi, A., Williams, B.A., Wold, B.J. & Rothenberg, E.V. Dynamic transformations of genome-wide epigenetic marking and transcriptional control establish T cell identity. Cell 149, 467–482 (2012).
Ritchie, M.E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, doi:10.1093/nar/gkv007 (2015).
Law, C.W., Chen, Y., Shi, W. & Smyth, G.K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Smyth, G.K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
McCarthy, D.J. & Smyth, G.K. Testing significance relative to a fold-change threshold is a TREAT. Bioinformatics 25, 765–771 (2009).
Wu, D. et al. ROAST: rotation gene set tests for complex microarray experiments. Bioinformatics 26, 2176–2182 (2010).
Acknowledgements
We thank D. Emslie, T. Kratina, M. Everest and S. Heinzel for technical assistance and J. Vasiliadis, J. Leahy and T. Camilleri for animal care. Socs2−/− mice were a gift from W. Alexander (The Walter and Eliza Hall Institute, Parkville, Victoria, Australia). The institutional flow cytometry facility provided essential services. Supported by the National Health and Medical Research Council (NHMRC) of Australia (NHMRC IRIISS grant 361646; Program Grants 575500 and 1054925 to D.M.T., P.D.H., S.L.N. and L.M.C.; Program Grant 1054618 to G.K.S.; Project Grant 1023454 to G.K.S. and W.S.; and fellowships to G.K.S., D.M.T., P.D.H. and L.M.C.), the Multiple Myeloma Research Foundation (Senior Award to S.L.N.), the Australian Research Council (ARC Future Fellowship to S.L.N.) and the Victorian State Government through an Operational Infrastructure Support grant.
Author information
Authors and Affiliations
Contributions
W.S., Y.L., G.K.S. and M.I. did the bioinformatic and statistical analyses; S.N.W. and N.T. did experiments relating to in vitro differentiation and cell-division studies; S.L.N., D.M.T., P.D.H. and L.M.C. contributed to the experimental design and analysis, and provided mouse models; L.M.C. carried out many of the experiments; W.S., S.L.N. and L.M.C. wrote the manuscript; all authors had editorial input.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Fraction of absolutely expressed genes in each cell population.
A detection P value cutoff of ≤0.05 was applied for calling absolutely expressed genes. 26k NBCI RefSeq genes were included in the analysis. See the Online Methods for a description of the algorithm used for calling absolutely expressed genes.
Supplementary Figure 2 Expression profiles of selected genes encoding chemokines, cytokines and their receptors in ex vivo B cell and ASC populations.
Three groups of genes are highlighted here: (a) Sphingosine-1-phosphate receptor, (b) chemokines and chemokine receptors and (c) cytokine receptors. Normalized FPKM values of genes are shown.
Supplementary Figure 3 Patterns of gene expression change during the cell-division process.
This figure is the same as Fig. 6f in the main text, except that gene expression values are shown instead of expression fold changes. The k-means clustering algorithm was used to classify the genes, which had ≥2.5-fold changes at the division stage (2,095 genes), into nine clusters based on stepwise expression changes from Div 0 to Div 7. These nine clusters were then further grouped into six distinct patterns. Numbers shown in upper corners of the figures indicate the number of genes included in each cluster.
Supplementary Figure 4 Chromatin changes during B cell differentiation.
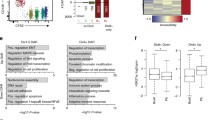
(a) Correlation of gene expression changes and the prevalence of H3K4me3 and H3K4me1 in the A20 (GC-like B lymphoma) and MPC-11 (plasmacytoma) cell lines. The top 500 genes most differentially expressed between A20 and MPC11 were included in the analysis. Values of Pearson correlation coefficients calculated with these genes are shown at the top left corner of each graph. The red diagonal lines indicate the slope of linear regression of histone mark changes on gene expression changes. (b) Correlation of the H3K4me3 and H3K4me1 data from the A20 and MPC-11 cell lines with gene expression changes between their primary cell counterparts, GC B cells and BMPCs, respectively. Here the top 500 most differentially expressed genes between GC B cells and BMPCs were included in the graphs. (c) Barcode plot showing that super-enhancer–associated genes (756 genes) tend to be upregulated in FoBs relative to BMPCs. The horizontal axis shows TREAT t-statistics for all genes comparing FoBs to BMPCs, and the vertical bars show the ranks of the super-enhancer genes. The worm shows enrichment of super-enhancer genes relative to random ordering. Roast P values are shown here and for d. (d) Barcode plot showing that super-enhancer–associated genes tend to be upregulated in GC B cells relative to BMPCs.
Supplementary Figure 5 Expression profiles of genes encoding transcription factors and chromatin-associated proteins that are specifically expressed in B cells.
(a–d) Normalized FPKM values of genes in ex vivo B cell and ASC populations: (a) B cells including FoBs, MZBs, B1 cells and GC B cells; (b) GC B cells; (c) MZBs; and (d) B1 cells.
Supplementary Figure 6 Expression profiles of genes encoding transcription factors and chromatin-associated proteins that are specifically expressed in ASCs.
Shown are normalized FPKM values of genes in ex vivo B cell and ASC populations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 794 kb)
Supplementary Table 1
The list of ASC signature genes discovered in the study (XLSX 83 kb)
Supplementary Table 2
Lists of genes that are shared between or unique to different ASC populations on the basis of comparison with FoBs. These gene lists correspond to different sets of genes included in Fig. 4e,f. (XLSX 488 kb)
Supplementary Table 3
Lists of genes that are shared between or unique to TI and TD plasmablasts on the basis of comparison with B cell blasts. These gene lists correspond to different sets of genes included in Fig. 5a,b. (XLSX 327 kb)
Supplementary Table 4
Summary of library sizes, read mapping percentages and read quantification results for RNA-seq data. Column "% mapped" gives the percentage of mapped reads. Column "% assigned" gives the percentage of fragments assigned to NCBI RefSeq genes, out of all the fragments included in each library. In each paired-end library, a fragment represents a pair of reads. In each single-end library, a fragment represents a read. (XLSX 51 kb)
Rights and permissions
About this article
Cite this article
Shi, W., Liao, Y., Willis, S. et al. Transcriptional profiling of mouse B cell terminal differentiation defines a signature for antibody-secreting plasma cells. Nat Immunol 16, 663–673 (2015). https://doi.org/10.1038/ni.3154
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3154
- Springer Nature America, Inc.
This article is cited by
-
Gene expression profiling in NOD mice reveals that B cells are highly educated by the pancreatic environment during autoimmune diabetes
Diabetologia (2023)
-
Dysregulated B cell differentiation towards antibody-secreting cells in neuromyelitis optica spectrum disorder
Journal of Neuroinflammation (2022)
-
Heterogeneous plasma cells and long-lived subsets in response to immunization, autoantigen and microbiota
Nature Immunology (2022)
-
The secret to longevity, plasma cell style
Nature Immunology (2022)
-
Distinct B cell subsets in Peyer’s patches convey probiotic effects by Limosilactobacillus reuteri
Microbiome (2021)