Abstract
Precision medicine in cancer proposes that genomic characterization of tumors can inform personalized targeted therapies1,2,3,4,5. However, this proposition is complicated by spatial and temporal heterogeneity6,7,8,9,10,11,12,13,14. Here we study genomic and expression profiles across 127 multisector or longitudinal specimens from 52 individuals with glioblastoma (GBM). Using bulk and single-cell data, we find that samples from the same tumor mass share genomic and expression signatures, whereas geographically separated, multifocal tumors and/or long-term recurrent tumors are seeded from different clones. Chemical screening of patient-derived glioma cells (PDCs) shows that therapeutic response is associated with genetic similarity, and multifocal tumors that are enriched with PIK3CA mutations have a heterogeneous drug-response pattern. We show that targeting truncal events is more efficacious than targeting private events in reducing the tumor burden. In summary, this work demonstrates that evolutionary inference from integrated genomic analysis in multisector biopsies can inform targeted therapeutic interventions for patients with GBM.




Similar content being viewed by others
References
Hamburg, M.A. & Collins, F.S. The path to personalized medicine. N. Engl. J. Med. 363, 301–304 (2010).
Brennan, C.W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013).
Ceccarelli, M. et al. Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164, 550–563 (2016).
Cloughesy, T.F., Cavenee, W.K. & Mischel, P.S. Glioblastoma: from molecular pathology to targeted treatment. Annu. Rev. Pathol. 9, 1–25 (2014).
Frattini, V. et al. The integrated landscape of driver genomic alterations in glioblastoma. Nat. Genet. 45, 1141–1149 (2013).
Snuderl, M. et al. Mosaic amplification of multiple receptor tyrosine kinase genes in glioblastoma. Cancer Cell 20, 810–817 (2011).
Szerlip, N.J. et al. Intratumoral heterogeneity of receptor tyrosine kinases EGFR and PDGFRA amplification in glioblastoma defines subpopulations with distinct growth factor response. Proc. Natl. Acad. Sci. USA 109, 3041–3046 (2012).
Kim, H. et al. Whole-genome and multisector exome sequencing of primary and post-treatment glioblastoma reveals patterns of tumor evolution. Genome Res. 25, 316–327 (2015).
Sottoriva, A. et al. Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. Proc. Natl. Acad. Sci. USA 110, 4009–4014 (2013).
Patel, A.P. et al. Single-cell RNA–seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Kumar, A. et al. Deep sequencing of multiple regions of glial tumors reveals spatial heterogeneity for mutations in clinically relevant genes. Genome Biol. 15, 530 (2014).
Johnson, B.E. et al. Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343, 189–193 (2014).
Yates, L.R. et al. Subclonal diversification of primary breast cancer revealed by multiregion sequencing. Nat. Med. 21, 751–759 (2015).
Wang, J. et al. Clonal evolution of glioblastoma under therapy. Nat. Genet. 48, 768–776 (2016).
Ohka, F., Natsume, A. & Wakabayashi, T. Current trends in targeted therapies for glioblastoma multiforme. Neurol. Res. Int. 2012, 878425 (2012).
Kim, J. et al. Spatiotemporal evolution of the primary glioblastoma genome. Cancer Cell 28, 318–328 (2015).
Liu, Q. et al. Genetic, epigenetic, and molecular landscapes of multifocal and multicentric glioblastoma. Acta Neuropathol. 130, 587–597 (2015).
Singh, D. et al. Transforming fusions of FGFR and TACC genes in human glioblastoma. Science 337, 1231–1235 (2012).
Hartmann, C., Bartels, G., Gehlhaar, C., Holtkamp, N. & von Deimling, A. PIK3CA mutations in glioblastoma multiforme. Acta Neuropathol. 109, 639–642 (2005).
Sottoriva, A. et al. A Big Bang model of human colorectal tumor growth. Nat. Genet. 47, 209–216 (2015).
Koren, S. et al. PIK3CAH1047R induces multipotency and multi-lineage mammary tumours. Nature 525, 114–118 (2015).
Carlsson, G. Topology and data. Bull. Amer. Math. Soc. 46, 255–308 (2009).
Singh, G., Mémoli, F. & Carlsson, G.E. Topological methods for the analysis of high dimensional data sets and 3D object recognition. in SPBG (eds. Botsch, M. & Pajarola, R.) 91–100 (Citeseer, 2007).
Moiyadi, A., Syed, P. & Srivastava, S. Fluorescence-guided surgery of malignant gliomas based on 5-aminolevulinic acid: paradigm shifts but not a panacea. Nat. Rev. Cancer 14, 146 (2014).
Stummer, W. et al. Fluorescence-guided surgery with 5-aminolevulinic acid for resection of malignant glioma: a randomised controlled multicentre phase III trial. Lancet Oncol. 7, 392–401 (2006).
Ozawa, T. et al. Most human non-GCIMP glioblastoma subtypes evolve from a common proneural-like precursor glioma. Cancer Cell 26, 288–300 (2014).
Merzak, A., Koocheckpour, S. & Pilkington, G.J. CD44 mediates human glioma cell adhesion and invasion in vitro. Cancer Res. 54, 3988–3992 (1994).
Yoshida, T., Matsuda, Y., Naito, Z. & Ishiwata, T. CD44 in human glioma correlates with histopathological grade and cell migration. Pathol. Int. 62, 463–470 (2012).
Joo, K.M. et al. MET signaling regulates glioblastoma stem cells. Cancer Res. 72, 3828–3838 (2012).
Lemmon, M.A. & Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010).
Kinzler, K.W. & Vogelstein, B. Lessons from hereditary colorectal cancer. Cell 87, 159–170 (1996).
Verhaak, R.G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Lee, J. et al. Tumor stem cells derived from glioblastomas cultured in bFGF and EGF more closely mirror the phenotype and genotype of primary tumors than do serum-cultured cell lines. Cancer Cell 9, 391–403 (2006).
Clark, K. et al. The Cancer Imaging Archive (TCIA): maintaining and operating a public information repository. J. Digit. Imaging 26, 1045–1057 (2013).
Wangaryattawanich, P. et al. Multicenter imaging outcomes study of The Cancer Genome Atlas glioblastoma patient cohort: imaging predictors of overall and progression-free survival. Neuro-oncol. 17, 1525–1537 (2015).
Rios Velazquez, E. et al. Fully automatic GBM segmentation in the TCGA-GBM dataset: Prognosis and correlation with VASARI features. Sci. Rep. 5, 16822 (2015).
Giannopoulos, S. & Kyritsis, A.P. Diagnosis and management of multifocal gliomas. Oncology 79, 306–312 (2010).
Kyritsis, A.P., Levin, V.A., Yung, W.K. & Leeds, N.E. Imaging patterns of multifocal gliomas. Eur. J. Radiol. 16, 163–170 (1993).
Zinn, P.O. et al. Radiogenomic mapping of edema/cellular invasion MRI-phenotypes in glioblastoma multiforme. PLoS One 6, e25451 (2011).
Li, Y.M., Suki, D., Hess, K. & Sawaya, R. The influence of maximum safe resection of glioblastoma on survival in 1229 patients: can we do better than gross-total resection? J. Neurosurg. 124, 977–988 (2016).
Carter, S.L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Suzuki, H. et al. Mutational landscape and clonal architecture in grade II and III gliomas. Nat. Genet. 47, 458–468 (2015).
Li, B. & Dewey, C.N. RSEM: accurate transcript quantification from RNA–Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Iyer, M.K., Chinnaiyan, A.M. & Maher, C.A. ChimeraScan: a tool for identifying chimeric transcription in sequencing data. Bioinformatics 27, 2903–2904 (2011).
Cahoy, J.D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Bhasin, M. et al. Bioinformatic identification and characterization of human endothelial cell–restricted genes. BMC Genomics 11, 342 (2010).
Chtanova, T. et al. Identification of T cell–restricted genes, and signatures for different T cell responses, using a comprehensive collection of microarray datasets. J. Immunol. 175, 7837–7847 (2005).
Abbas, A.R. et al. Immune response in silico (IRIS): immune-specific genes identified from a compendium of microarray expression data. Genes Immun. 6, 319–331 (2005).
Huang, S. & Pang, L. Comparing statistical methods for quantifying drug sensitivity based on in vitro dose-response assays. Assay Drug Dev. Technol. 10, 88–96 (2012).
Zairis, S. et al. Moduli spaces of phylogenetic trees describing tumor evolutionary patterns. Lect. Notes Comput. Sci. 8609, 528–539 (2014).
Acknowledgements
This work was supported by a grant of the Korea Health Technology R&D project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (HI14C3418). R.R. acknowledges funding from the NIH (U54 CA193313, R01 CA185486, R01 CA179044). J.W. is supported by Precision Medicine Fellowship (UL1 TR000040). E.L. is supported by NIH (F99 CA212478) and Cancer Biology Training Program (T32 CA09503).
Author information
Authors and Affiliations
Contributions
J.-K.L., J.W., J.K.S., and E.L. are co-first authors. J.-K.L., J.W., J.K.S., and E.L. performed the majority of experiments and analyses. W.-Y.P., H.-O.L., K.-T.K., P.G.C., and P.v.N. performed experiments and analyses for the single-cell transcriptome. D.S.R., Z.L., A.B., A.C., Y.J.S., and S.S. conducted several experiments and analyses. D.-H.N., D.-S.K., H.J.S., C.-K.P., and J.-I.L. provided surgical specimens. S.W.J., S.W.C., and J.K. helped interpret clinical data. W.-Y.P., I.-H.L., Y. J.S., J.-M.O., and H.J.K. organized and processed specimens and genome data. J.-K.L., J.W., J.K.S., E.L., and P.G.C. wrote the manuscript with feedback from R.R., A.I., and D.-H.N. D.-H.N. and R.R. designed and supervised the entire project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–17, Supplementary Tables 3–6 and Supplementary Note (PDF 4975 kb)
Supplementary Table 1
Descriptive characteristics of multisector/longitudinal samples from the Samsung Medical Center (SMC) and TCGA cohorts. (XLSX 25 kb)
Supplementary Table 2
Somatic mutations and estimation of cancer cell frequency. (XLSX 4214 kb)
Supplementary Video 1
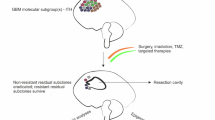
The truncal hypothesis. Illustrative Flash video for conceptual suggestion of the optimal therapeutic strategy for abolishing maximal tumor burden and preventing possible recurrence using multisector genomic information. (WMV 6697 kb)
Rights and permissions
About this article
Cite this article
Lee, JK., Wang, J., Sa, J. et al. Spatiotemporal genomic architecture informs precision oncology in glioblastoma. Nat Genet 49, 594–599 (2017). https://doi.org/10.1038/ng.3806
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3806
- Springer Nature America, Inc.
This article is cited by
-
Integrative multi-omics characterization reveals sex differences in glioblastoma
Biology of Sex Differences (2024)
-
Extracellular vesicles in glioblastoma: a challenge and an opportunity
npj Precision Oncology (2024)
-
High-throughput identification of repurposable neuroactive drugs with potent anti-glioblastoma activity
Nature Medicine (2024)
-
VC-resist glioblastoma cell state: vessel co-option as a key driver of chemoradiation resistance
Nature Communications (2024)
-
Spatially resolved transcriptomic profiles reveal unique defining molecular features of infiltrative 5ALA-metabolizing cells associated with glioblastoma recurrence
Genome Medicine (2023)