Abstract
The endoplasmic reticulum (ER) is a site of protein biogenesis in eukaryotic cells. Perturbing ER homeostasis activates stress programs collectively called the unfolded protein response (UPR). The UPR enhances production of ER-resident chaperones and enzymes to reduce the burden of misfolded proteins. On resolution of ER stress, ill-defined, selective autophagic programs remove excess ER components. Here we identify Sec62, a constituent of the translocon complex regulating protein import in the mammalian ER, as an ER-resident autophagy receptor. Sec62 intervenes during recovery from ER stress to selectively deliver ER components to the autolysosomal system for clearance in a series of events that we name recovER-phagy. Sec62 contains a conserved LC3-interacting region in the C-terminal cytosolic domain that is required for its function in recovER-phagy, but is dispensable for its function in the protein translocation machinery. Our results identify Sec62 as a critical molecular component in maintenance and recovery of ER homeostasis.








Similar content being viewed by others
Accession codes
Change history
18 November 2016
In the version of this Article originally published, the name of co-author Eduardo Cebollero Presmanes was coded wrongly resulting in it being incorrect when exported to citation databases. This has been corrected and the co-author's name now appears as 'Eduardo Cebollero' in the online versions of the Article.
References
Tannous, A., Pisoni, G. B., Hebert, D. N. & Molinari, M. N-linked sugar-regulated protein folding and quality control in the ER. Semin. Cell Dev. Biol. 41, 79–89 (2015).
Guerriero, C. J. & Brodsky, J. L. The delicate balance between secreted protein folding and endoplasmic reticulum-associated degradation in human physiology. Physiol. Rev. 92, 537–576 (2012).
Pisoni, G. B. & Molinari, M. Five questions (with their answers) on ER-associated degradation. Traffic 17, 341–350 (2016).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Ktistakis, N. T. & Tooze, S. A. Digesting the expanding mechanisms of autophagy. Trends Cell Biol. 26, 624–635 (2016).
Okamoto, K. Organellophagy: eliminating cellular building blocks via selective autophagy. J. Cell Biol. 205, 435–445 (2014).
Ogata, M. et al. Autophagy is activated for cell survival after endoplasmic reticulum stress. Mol. Cell Biol. 26, 9220–9231 (2006).
Jia, W., Pua, H. H., Li, Q. J. & He, Y. W. Autophagy regulates endoplasmic reticulum homeostasis and calcium mobilization in T lymphocytes. J. Immunol. 186, 1564–1574 (2011).
Pengo, N. et al. Plasma cells require autophagy for sustainable immunoglobulin production. Nat. Immunol. 14, 298–305 (2013).
Khaminets, A. et al. Regulation of endoplasmic reticulum turnover by selective autophagy. Nature 522, 354–358 (2015).
Mochida, K. et al. Receptor-mediated selective autophagy degrades the endoplasmic reticulum and the nucleus. Nature 522, 359–362 (2015).
Bernales, S., McDonald, K. L. & Walter, P. Autophagy counterbalances endoplasmic reticulum expansion during the unfolded protein response. PLoS Biol. 4, e423 (2006).
Schuck, S., Gallagher, C. M. & Walter, P. ER-phagy mediates selective degradation of endoplasmic reticulum independently of the core autophagy machinery. J. Cell Sci. 127, 4078–4088 (2014).
Bolender, R. P. & Weibel, E. R. A morphometric study of the removal of phenobarbital-induced membranes from hepatocytes after cessation of treatment. J. Cell Biol. 56, 746–761 (1973).
Feldman, D., Swarm, R. L. & Becker, J. Elimination of excess smooth endoplasmic reticulum after phenobarbital administration. J. Histochem. Cytochem. 28, 997–1006 (1980).
Conti, B. J., Devaraneni, P. K., Yang, Z. Y., David, L. L. & Skach, W. R. Cotranslational stabilization of Sec62/63 within the ER Sec61 translocon is controlled by distinct substrate-driven translocation events. Mol. Cell 58, 269–283 (2015).
Johnson, N. et al. The signal sequence influences post-translational er translocation at distinct stages. PLoS ONE 8, e75394 (2013).
Lang, S. et al. Different effects of Sec61 alpha, Sec62 and Sec63 depletion on transport of polypeptides into the endoplasmic reticulum of mammalian cells. J. Cell Sci. 125, 1958–1969 (2012).
Lakkaraju, A. K. K. et al. Efficient secretion of small proteins in mammalian cells relies on Sec62-dependent posttranslational translocation. Mol. Biol. Cell 23, 2712–2722 (2012).
Meyer, H. A. et al. Mammalian Sec61 is associated with Sec62 and Sec63. J. Biol. Chem. 275, 14550–14557 (2000).
Jadhav, B. et al. Mammalian SRP receptor switches the Sec61 translocase from Sec62 to SRP-dependent translocation. Nat. Commun. 6, 10133 (2015).
Ampofo, E. et al. CK2 phosphorylation of human Sec63 regulates its interaction with Sec62. BBA-Gen. Subjects 1830, 2938–2945 (2013).
Birgisdottir, A. B., Lamark, T. & Johansen, T. The LIR motif—crucial for selective autophagy. J. Cell Sci. 126, 3237–3247 (2013).
Pirot, P. et al. Global profiling of genes modified by endoplasmic reticulum stress in pancreatic beta cells reveals the early degradation of insulin mRNAs. Diabetologia 50, 1006–1014 (2007).
Klionsky, D. J., Elazar, Z., Seglen, P. O. & Rubinsztein, D. C. Does bafilomycin A(1) block the fusion of autophagosomes with lysosomes? Autophagy 4, 849–850 (2008).
Yan, L. et al. Ube2g2-gp78-mediated HERP polyubiquitylation is involved in ER stress recovery. J. Cell Sci. 127, 1417–1427 (2014).
Rubinsztein, D. C. Cell biology receptors for selective recycling. Nature 522, 291–292 (2015).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222 (2016).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Wang, X. & Johnsson, N. Protein kinase CK2 phosphorylates Sec63p to stimulate the assembly of the endoplasmic reticulum protein translocation apparatus. J. Cell Sci. 118, 723–732 (2005).
Pisoni, G. B., Ruddock, L. W., Bulleid, N. & Molinari, M. Division of labor among oxidoreductases: TMX1 preferentially acts on transmembrane polypeptides. Mol. Biol. Cell 26, 3390–3400 (2015).
Bernasconi, R. & Molinari, M. ERAD and ERAD tuning: disposal of cargo and of ERAD regulators from the mammalian ER. Curr. Opin. Cell Biol. 23, 176–183 (2011).
Vittal, V., Stewart, M. D., Brzovic, P. S. & Klevit, R. E. Regulating the regulators: recent revelations in the control of E3 ubiquitin ligases. J. Biol. Chem. 290, 21244–21251 (2015).
Koenig, P. A. et al. The E2 ubiquitin-conjugating enzyme UBE2J1 is required for spermiogenesis in mice. J. Biol. Chem. 289, 34490–34502 (2014).
Linxweiler, M. et al. Sec62 bridges the gap from 3q amplification to molecular cell biology in non-small cell lung cancer. Am. J. Pathol. 180, 473–483 (2012).
Greiner, M. et al. Silencing of the SEC62 gene inhibits migratory and invasive potential of various tumor cells. Int. J. Cancer 128, 2284–2295 (2011).
Jung, V. et al. Genomic and expression analysis of the 3q25-q26 amplification unit reveals TLOC1/SEC62 as a probable target gene in prostate cancer. Mol. Cancer Res. 4, 169–176 (2006).
Weng, L. et al. Identification of cyclin B1 and Sec62 as biomarkers for recurrence in patients with HBV-related hepatocellular carcinoma after surgical resection. Mol. Cancer 11, 39 (2012).
Hagerstrand, D. et al. Systematic interrogation of 3q26 identifies TLOC1 and SKIL as cancer drivers. Cancer Discov. 3, 1044–1057 (2013).
Wemmert, S. et al. Initial evidence for Sec62 as a prognostic marker in advanced head and neck squamous cell carcinoma. Oncol. Lett. 11, 1661–1670 (2016).
Greiner, M. et al. Sec62 protein level is crucial for the ER stress tolerance of prostate cancer. Prostate 71, 1074–1083 (2011).
Zimmermann, R., Muller, L. & Wullich, B. Protein transport into the endoplasmic reticulum: mechanisms and pathologies. Trends Mol. Med. 12, 567–573 (2006).
Suzuki, H. et al. Structural basis of the autophagy-related LC3/Atg13 LIR complex: recognition and interaction mechanism. Structure 22, 47–58 (2014).
Molinari, M. et al. Contrasting functions of calreticulin and calnexin in glycoprotein folding and ER quality control. Mol. Cell 13, 125–135 (2004).
Soldà, T., Garbi, N., Hammerling, G. J. & Molinari, M. Consequences of ERp57 deletion on oxidative folding of obligate and facultative clients of the calnexin cycle. J. Biol. Chem. 281, 6219–6226 (2006).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Mizushima, N., Ohsumi, Y. & Yoshimori, T. Autophagosome formation in mammalian cells. Cell Struct. Funct. 27, 421–429 (2002).
Soding, J. Protein homology detection by HMM-HMM comparison. Bioinformatics 21, 951–960 (2005).
Raveh, B., London, N. & Schueler-Furman, O. Sub-angstrom modeling of complexes between flexible peptides and globular proteins. Proteins 78, 2029–2040 (2010).
London, N., Raveh, B., Cohen, E., Fathi, G. & Schueler-Furman, O. Rosetta flexpepdock web server-high resolution modeling of peptide-protein interactions. Nucleic Acids Res. 39, 249–253 (2011).
Hilpert, K., Winkler, D. F. H. & Hancock, R. E. W. Peptide arrays on cellulose support: SPOT synthesis, a time and cost efficient method for synthesis of large numbers of peptides in a parallel and addressable fashion. Nat. Protoc. 2, 1333–1349 (2007).
Wilm, M. et al. Femtomole sequencing of proteins from polyacrylamide gels by nano-electrospray mass spectrometry. Nature 379, 466–469 (1996).
Cox, J. & Mann, M. 1D and 2D annotation enrichment: a statistical method integrating quantitative proteomics with complementary high-throughput data. BMC Bioinformatics 13, S12 (2012).
Vizcaino, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Acknowledgements
We are indebted to D. Ron for having suggested the use of CPA. We thank A. Helenius (Institute of Biochemistry, ETH Zurich, Zurich, Switzerland), A. Smith, M. Komatsu (Department of Cardiovascular Medicine, Tohoku University Graduate School of Medicine, Sendai, Japan), T. Johansen (Department of Molecular Cell Biology, Institute for Cancer Research, Oslo University Hospital, Montebello, Oslo, Norway), N. Mizushima (Department of Bioregulation and Metabolism, Tokyo Metropolitan Institute of Medical Science, Tokyo, Japan), P. Paganetti, F. Reggiori, C. Tacchetti and C. Guerra for gifts of reagents, discussions and critical reading of the manuscript. We are grateful to the ALEMBIC facility at San Raffaele Scientific Institute, Milan, Italy for help with electron microscopy analysis and to E. Neuhaus and T. Möhlmann at Technical University, Kaiserslautern, Germany for giving us access to their Biacore system. M.M. is supported by Signora Alessandra, the Foundation for Research on Neurodegenerative Diseases, the Swiss National Science Foundation (SNF) and the Comel and Gelu Foundations. M.M. and M.P. are recipients of a Sinergia grant of the SNF, and M.P. is supported by the ETH and an ERC advanced fellowship. L.V. is supported by SNF and Oncosuisse. R.Z. and K.H. are supported by the German Research Foundation (DFG).
Author information
Authors and Affiliations
Contributions
F.F. and J.N. performed biochemical and transcriptional analyses, IF, image quantifications and MS analysis; T.J.B. analysed protein associations in living cells, performed qPCRs and analysed MS data; G.B.P. and E.C. prepared DNA constructs and performed initial biochemical analyses; E.F. performed IF and analysis of MS and biochemical data; I.F. performed biochemical analysis, toxicity assays and analysis of MS data; C.G. prepared plasmids for Sec62 and Sec62LIRm expression, and generated and characterized inducible cell lines; C.G. and M.L. performed and analysed isopycnic density gradients, collected and analysed MS data, contributed to the generation of the FlipIn T-rex CRISPR62 and CRISPR63 cell lines; T.S. generated and characterized the FlipIn T-rex CRISPR62 and CRISPR63 cell lines; R.D’A. developed CellProfiler pipelines; L.S. and L.V. designed and performed computational simulations and structural analysis; A.R. performed CLEM and electron microscopy on murine cell lines; M.J. and A.M. synthesized peptides and used them in LC3 interaction analysis; S.S. and F.F. carried out protein translocation assay; A.S. purified human LC3B; E.C. and C.W.-Z. performed yeast experiments that influenced the design of this study; O.Z., L.V. and L.S. performed NMR experiments and analysed data; M.P. supervised purification of LC3B and was involved in writing of the manuscript; K.H. performed the bioinformatics analyses to identify the Sec62 LIR and predicted the autophagy role of Sec62; M.Q. carried out mass spectrometry measurements, protein identification and quantification on autolysosome fractions; R.Z. supervised protein transport and peptide binding experiments and was involved in writing of the manuscript; M.M. designed the study, analysed data and wrote the manuscript. All the authors discussed the results and the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 ER stress induction with CPA and DTT.
(a) Amplification with appropriate primers of unspliced and spliced XBP1 transcripts in mock-treated cells and in cells exposed for 12 h to 10 μM CPA-induced ER stress. (b) Incorporation of 35S-radiolabeled amino acids (2 h pulse with 35S-methionine and-cysteine) in total MEF proteome after 8 h of Mock- or 10 μM CPA-treatment. TCE, total cell extract. (c) Same as (b) for the incorporation of 35S-radiolabeled amino acids in the ER marker proteins Cnx, Grp94 and BiP as assessed by autoradiography of the respective Immunoprecipitates. (d) Quantification of (b) and (c), data from a single experiment. (e) Dithiothreitol (DTT) induces ER stress by modifying redox homeostasis. At 1 mM, DTT caused a milder stress in MEF compared to non-toxic concentrations of CPA as determined by lower induction of BiP transcription, data from a single experiment (f) Like CPA, the wash out of DTT at the end of the stress phase was followed by rapid return of BiP transcripts at or below their pre-stress levels as determined by qPCR. For CPA 10 μM, mean of two independent experiments. For others data from a single experiment. (g) Like CPA, on DTT wash out (but not in untreated cells (Mock) or during the stress (ER stress)), excess ER chaperones did accumulate in Lamp1-positive autolysosomes in cells exposed to the lysosomal inhibitor BafA1. Studies were continued with CPA, which proved less toxic to cells. Scale bars, 10 μm. Images from a single experiment. Details of statistics in the Statistics and Reproducibility section. Statistics source data in Supplementary Table 4.
Supplementary Figure 2 Reversible block of ER marker proteins in autolysosomes obtained by cell exposure to the lysosomotropic drug BafA1.
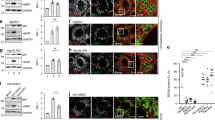
(a) After the CPA-induced stress, MEF were grown in the presence of BafA1 for 12 h and analyzed in IF (upper panels) or analyzed in IF 4 h after BafA1 wash out (lower panels). Scale bars, 10 μm. (b) Quantification of the Cnx-containing puncta in the upper and lower panels. n = 206 and 112 single cells for +BafA1 and BafA1 wash out, respectively. Mean ± SEM shown. ∗∗∗∗P < 0.0001, unpaired two-tailed t-test. Scale bars, 10 μm. (c) On inhibition of recovery from ER stress with BafA1 (+BafA1), Cnx accumulates in autolysosomes displaying GFP-Rab7 and Lamp1 in the limiting membrane. Scale bars, 10 μm. (d) Same as (c) for Sec62 (also refer to Fig. 2). Scale bars, 10 μm. Details of statistics in the Statistics and Reproducibility section. Statistics source data in Supplementary Table 4.
Supplementary Figure 3 Generation of CRISPR62 and CRISPR63 cell lines and characterisation of ER stress recovery.
(a) Control of Sec62 (upper panel) and Sec63 (lower panel) KO in HEK293 cells inducible for Sec62 expression. *, non-specific cross-reacting bands. (b) Induction of Sec62-HA or Sec62LIRm-HA in CRISPR62 cells exposed to increasing (0–150 ng ml−1) Tet concentrations. WB representative of 2 independent experiments. (c) CRISPR62 HEK293 cells as well as the same cells on induction of Sec62 (upper panel) or Sec62LIRm expression (lower panel) for 17 and 41 h do not activate the UPR. As positive control, cell exposure to CPA (30 μM) or tunicamycin (5 μg ml−1) induces an UPR. Data from a single experiment. (d) Reversibility of the transcriptional UPR induced in CRISPR cells on cell exposure to 30 μM CPA in the absence or presence of 100 nM BafA1. Note that the presence of BafA1 does not affect the rate of BiP and sXbp1 transcripts reduction on CPA wash out. Data from a single experiment. (e) Deletion of Sec62 (CRISPR62 HEK293 cells) prevents delivery of ER marker proteins to Lamp1-positive autolysosomes during recovery from ER stress. Scale bars, 10 μm. (f) Deletion of Sec63 (CRISPR63) leads to delivery of ER protein markers to Lamp1-positive autolysosomes even at steady state. Scale bars, 10 μm. (g) Deletion of Sec63 does not interfere with delivery of ER protein markers to Lamp1-positive autolysosomes during recovery from ER stress. Scale bars, 10 μm. Statistics source data in Supplementary Table 4.
Supplementary Figure 4 ERj3 depends on Sec62:Sec63 for post-translational translocation in the mammalian ER.
In a complementation or rescue assay, the Sec62 and Sec62LIRm equally support ERj3 translocation into the ER of Sec62 silenced HeLa cells (please also refer to Fig. 6f for KO cells generated with the CRISPR-Cas9 gene editing technology). Expression (Sec62) and transport (ERj3) efficiencies were calculated as described in the Methods section. n = 4 independent experiments, mean ± SEM. We note that SEC62 plasmids (lanes 2–4 and 6–8) lack the untranslated region (UTR) of endogenous mRNA and, therefore, are resistant to UTR-targeting siRNA.
Supplementary Figure 5 The cytosolic LIR motif of Sec62 is required and sufficient for delivery of ER portions to autolysosomes.
(a) At steady state, Crt is delivered to Lamp1-positive autolysosomes in MEF expressing ER membrane-associated GFP displaying the 145 C-terminal residues of the cytosolic tail of Sec62 (GFPTM62) containing the LIR. Scale bars, 10 μm. (b) Crt is not delivered to autolysosomes in cells expressing GFPTM62LIRm. Scale bars, 10 μm. Please also refer to Fig. 7d, e for Cnx.
Supplementary Figure 6 Separation of AV and ER in isopycnic density gradients.
(a) Schematic representation of the experiments. At the end of the ultracentrifugation, AV float at the top of the density gradient, the ER has higher density (Fig. 8a). (b) Extracts of WT and Atg7 KO MEF recovering from ER stress were prepared and loaded at the bottom of an isopycnic density gradient (as in Fig. 8a and Methods). Fractions floating at the top of the gradient (AV) and fractions containing the ER were separated in SDS-PAGE and probed for the presence of Cnx, Sec62 and LC3 by WB. WB representative of 2 independent experiments.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1371 kb)
Supplementary Table 1
Supplementary Information (XLS 30 kb)
Supplementary Table 2
Supplementary Information (XLS 30 kb)
Supplementary Table 3
Supplementary Information (XLS 33 kb)
Supplementary Table 4
Supplementary Information (XLSX 193 kb)
Rights and permissions
About this article
Cite this article
Fumagalli, F., Noack, J., Bergmann, T. et al. Translocon component Sec62 acts in endoplasmic reticulum turnover during stress recovery. Nat Cell Biol 18, 1173–1184 (2016). https://doi.org/10.1038/ncb3423
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3423
- Springer Nature Limited
This article is cited by
-
Selective autophagy in cancer: mechanisms, therapeutic implications, and future perspectives
Molecular Cancer (2024)
-
ER-to-lysosome-associated degradation acts as failsafe mechanism upon ERAD dysfunction
EMBO Reports (2024)
-
TMX4-driven LINC complex disassembly and asymmetric autophagy of the nuclear envelope upon acute ER stress
Nature Communications (2023)
-
The function of ER-phagy receptors is regulated through phosphorylation-dependent ubiquitination pathways
Nature Communications (2023)
-
Decreased syntaxin17 expression contributes to the pathogenesis of acute pancreatitis in murine models by impairing autophagic degradation
Acta Pharmacologica Sinica (2023)