Abstract
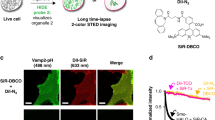
Imaging cellular structures and organelles in living cells by long time-lapse super-resolution microscopy is challenging, as it requires dense labeling, bright and highly photostable dyes, and non-toxic conditions. We introduce a set of high-density, environment-sensitive (HIDE) membrane probes, based on the membrane-permeable silicon-rhodamine dye HMSiR, that assemble in situ and enable long time-lapse, live-cell nanoscopy of discrete cellular structures and organelles with high spatiotemporal resolution. HIDE-enabled nanoscopy movies span tens of minutes, whereas movies obtained with labeled proteins span tens of seconds. Our data reveal 2D dynamics of the mitochondria, plasma membrane and filopodia, and the 2D and 3D dynamics of the endoplasmic reticulum, in living cells. HIDE probes also facilitate acquisition of live-cell, two-color, super-resolution images, expanding the utility of nanoscopy to visualize dynamic processes and structures in living cells.






Similar content being viewed by others
References
Hell, S.W. Far-field optical nanoscopy. Science 316, 1153–1158 (2007).
Huang, B., Babcock, H. & Zhuang, X. Breaking the diffraction barrier: super-resolution imaging of cells. Cell 143, 1047–1058 (2010).
Sahl, S.J. & Moerner, W.E. Super-resolution fluorescence imaging with single molecules. Curr. Opin. Struct. Biol. 23, 778–787 (2013).
Liu, Z., Lavis, L.D. & Betzig, E. Imaging live-cell dynamics and structure at the single-molecule level. Mol. Cell 58, 644–659 (2015).
Hess, S.T. et al. Dynamic clustered distribution of hemagglutinin resolved at 40 nm in living cell membranes discriminates between raft theories. Proc. Natl. Acad. Sci. USA 104, 17370–17375 (2007).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 5, 155–157 (2008).
Jones, S.A., Shim, S.H., He, J. & Zhuang, X. Fast, three-dimensional super-resolution imaging of live cells. Nat. Methods 8, 499–508 (2011).
Shim, S.H. et al. Super-resolution fluorescence imaging of organelles in live cells with photoswitchable membrane probes. Proc. Natl. Acad. Sci. USA 109, 13978–13983 (2012).
Huang, F. et al. Video-rate nanoscopy using sCMOS camera-specific single-molecule localization algorithms. Nat. Methods 10, 653–658 (2013).
Lukinavicˇius, G. et al. Fluorogenic probes for live-cell imaging of the cytoskeleton. Nat. Methods 11, 731–733 (2014).
Bottanelli, F. et al. Two-colour live-cell nanoscale imaging of intracellular targets. Nat. Commun. 7, 10778 (2016).
Lukinavicˇius, G. et al. A near-infrared fluorophore for live-cell super-resolution microscopy of cellular proteins. Nat. Chem. 5, 132–139 (2013).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Kiuchi, T., Higuchi, M., Takamura, A., Maruoka, M. & Watanabe, N. Multitarget super-resolution microscopy with high-density labeling by exchangeable probes. Nat. Methods 12, 743–746 (2015).
Fernández-Suárez, M. & Ting, A.Y. Fluorescent probes for super-resolution imaging in living cells. Nat. Rev. Mol. Cell Biol. 9, 929–943 (2008).
Toomre, D. & Bewersdorf, J. A new wave of cellular imaging. Annu. Rev. Cell Dev. Biol. 26, 285–314 (2010).
Rust, M.J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Heilemann, M. et al. Subdiffraction-resolution fluorescence imaging with conventional fluorescent probes. Angew. Chem. Int. Ed. Engl. 47, 6172–6176 (2008).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S.T., Girirajan, T.P.K. & Mason, M.D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Hell, S.W. & Kroug, M. Ground-state-depletion fluorescence microscopy—a concept for breaking the diffraction resolution limit. Appl. Phys. B 60, 495–497 (1995).
Shcherbakova, D.M., Sengupta, P., Lippincott-Schwartz, J. & Verkhusha, V.V. Photocontrollable fluorescent proteins for superresolution imaging. Annu. Rev. Biophys. 43, 303–329 (2014).
Dempsey, G.T., Vaughan, J.C., Chen, K.H., Bates, M. & Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 8, 1027–1036 (2011).
Zhang, M. et al. Rational design of true monomeric and bright photoactivatable fluorescent proteins. Nat. Methods 9, 727–729 (2012).
Wäldchen, S., Lehmann, J., Klein, T., van de Linde, S. & Sauer, M. Light-induced cell damage in live-cell super-resolution microscopy. Sci. Rep. 5, 15348 (2015).
Uno, S.N. et al. A spontaneously blinking fluorophore based on intramolecular spirocyclization for live-cell super-resolution imaging. Nat. Chem. 6, 681–689 (2014).
Devaraj, N.K., Hilderbrand, S., Upadhyay, R., Mazitschek, R. & Weissleder, R. Bioorthogonal turn-on probes for imaging small molecules inside living cells. Angew. Chem. Int. Ed. Engl. 49, 2869–2872 (2010).
Erdmann, R.S. et al. Super-resolution imaging of the Golgi in live cells with a bioorthogonal ceramide probe. Angew. Chem. Int. Ed. Engl. 53, 10242–10246 (2014).
Wu, M.M. et al. Organelle pH studies using targeted avidin and fluorescein-biotin. Chem. Biol. 7, 197–209 (2000).
Munro, S. & Pelham, H.R.B. A C-terminal signal prevents secretion of luminal ER proteins. Cell 48, 899–907 (1987).
Storrie, B. et al. Recycling of Golgi-resident glycosyltransferases through the ER reveals a novel pathway and provides an explanation for nocodazole-induced Golgi scattering. J. Cell Biol. 143, 1505–1521 (1998).
Puhka, M., Joensuu, M., Vihinen, H., Belevich, I. & Jokitalo, E. Progressive sheet-to-tubule transformation is a general mechanism for endoplasmic reticulum partitioning in dividing mammalian cells. Mol. Biol. Cell 23, 2424–2432 (2012).
Greenfield, J.J.A. & High, S. The Sec61 complex is located in both the ER and the ER-Golgi intermediate compartment. J. Cell Sci. 112, 1477–1486 (1999).
van Meer, G., Voelker, D.R. & Feigenson, G.W. Membrane lipids: where they are and how they behave. Nat. Rev. Mol. Cell Biol. 9, 112–124 (2008).
Ishigaki, M. et al. STED super-resolution imaging of mitochondria labeled with TMRM in living cells. Mitochondrion 28, 79–87 (2016).
Yano, M. et al. Visualization of mitochondrial protein import in cultured mammalian cells with green fluorescent protein and effects of overexpression of the human import receptor Tom20. J. Biol. Chem. 272, 8459–8465 (1997).
Nemoto, Y. & De Camilli, P. Recruitment of an alternatively spliced form of synaptojanin 2 to mitochondria by the interaction with the PDZ domain of a mitochondrial outer membrane protein. EMBO J. 18, 2991–3006 (1999).
Michelle, T.Z. & Spence, I.D.J. The Molecular Probes Handbook: A Guide to Fluorescent Probes and Labeling Technologies (Life Technologies Corporation, 2010).
Leung, W.Y.H., Haugland, R.P. & Mao, F. Lipophilic cyanine dyes with enhanced aqueous solubility. US Patent 6004536A (1999).
Stone, D.M. et al. The tumour-suppressor gene patched encodes a candidate receptor for Sonic hedgehog. Nature 384, 129–134 (1996).
Kukic, I., Rivera-Molina, F. & Toomre, D. The IN/OUT assay: a new tool to study ciliogenesis. Cilia 5, 23 (2016).
Axelrod, D. Carbocyanine dye orientation in red cell membrane studied by microscopic fluorescence polarization. Biophys. J. 26, 557–573 (1979).
Fricker, M., Hollinshead, M., White, N. & Vaux, D. Interphase nuclei of many mammalian cell types contain deep, dynamic, tubular membrane-bound invaginations of the nuclear envelope. J. Cell Biol. 136, 531–544 (1997).
Schermelleh, L. et al. Subdiffraction multicolor imaging of the nuclear periphery with 3D structured illumination microscopy. Science 320, 1332–1336 (2008).
Nixon-Abell, J. et al. Increased spatiotemporal resolution reveals highly dynamic dense tubular matrices in the peripheral ER. Science 354, aaf3928 (2016).
Bai, X. et al. Novel septin 9 repeat motifs altered in neuralgic amyotrophy bind and bundle microtubules. J. Cell Biol. 203, 895–905 (2013).
Lin, Y. et al. Quantifying and optimizing single-molecule switching nanoscopy at high speeds. PLoS One 10, e0128135 (2015).
MathWorks. Global image threshold using Otsu's method – MATLAB graythresh http://www.mathworks.com/help/images/ref/graythresh.html (2016).
Carlson, J.C.T., Meimetis, L.G., Hilderbrand, S.A. & Weissleder, R. BODIPY-tetrazine derivatives as superbright bioorthogonal turn-on probes. Angew. Chem. Int. Ed. Engl. 52, 6917–6920 (2013).
Rivera-Molina, F. & Toomre, D. Live-cell imaging of exocyst links its spatiotemporal dynamics to various stages of vesicle fusion. J. Cell Biol. 201, 673–680 (2013).
Juette, M.F. . et al. Three-dimensional sub-100 nm resolution fluorescence microscopy of thick samples. Nat. Methods 5, 527–529 (2008).
Huang, B., Wang, W., Bates, M. & Zhuang, X. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319, 810–813 (2008).
Acknowledgements
We thank the following people for plasmids and reagents: F. Bottanelli, L. Schroeder, S. Baguley, Z. Xi and I. Kukic (Yale University School of Medicine), E.T. Spiliotis (Drexel University), M. Davidson (Florida State University), and C.G. Giraudo (Children's Hospital of Philadelphia). We thank J. Coleman and S.C. Alexander for advice and help with liposomes preparations. This work was supported by a Wellcome Trust Foundation grant (095927/A/11/Z, D.T., J.B., J.E.R.), the NIH (R01GM83257 to A.S., RO1GM118486 to D.T. and J.B., and P30 DK45735) and partly by grants AMED Brain/MINDS and JSPS Core-to-Core Program, A. Advanced Research Networks to Y.U., and by a MEXT KAKENHI grant number JP15H05951 to M.K. H.T. was supported by a JSPS postdoctoral fellowship for research abroad. R.S.E. was supported by an Advanced Postdoc Mobility fellowship from the Swiss National Science Foundation. A.D.T. was supported by the National Institutes of Health Ruth L. Kirschstein NRSA (F31GM119259). S.U. was supported by a JSPS postdoctoral fellowship.
Author information
Authors and Affiliations
Contributions
H.T., Y.Z., R.S.E., A.D.T., J.B., A.S., J.E.R. and D.T. designed experiments. H.T., Y.Z., R.S.E., A.D.T., F.R.-M. and Y.L. performed the imaging experiments. A.D.T. designed and synthesized RhoB-TCO, DiI-TCO and DiI-(C6-)TCO. R.S.E. synthesized Cer-TCO. B.M. performed computer simulations. F.R.-M. generated the COPβ1-mEos3.2 stable HeLa cell line. S.U., M.K. and Y.U. developed and synthesized HMSiR-Tz. H.T., R.S.E., A.D.T., J.B., A.S. and D.T. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.B. discloses financial interest in Bruker Corp. and Hamamatsu Photonics. R.S.E., A.D.T., D.T. and A.S. disclose a pending patent for lipid probes.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–20, Supplementary Table 1 and Supplementary Data. (PDF 14429 kb)
Monte Carlo simulations support the hypothesis that combining a high density labeling with an ultra-low ON fraction improves apparent photostability.
Monte Carlo simulations were performed as described in Methods (section 2.3). The movies shown illustrate the time-lapse image expected using various labeling densities and ON fractions. (i) Images expected when labeling density = 10 (low) and ON fraction = 1% (low); (ii) Images expected when labeling density = 500 (high) and ON fraction = 1% (low); (iii) Images expected when labeling density is 10 (low) and ON fraction is 0.01% (ultra-low); (iv) Images expected when labeling density is 500 (high) and ON fraction is 0.01% (ultra-low). Images (i) and (iv) represent the parameters expected for HMSiR-conjugated proteins and lipids, respectively. Selected frames from these movies are shown in Supplementary Fig. 6. (MOV 4482 kb)
Time-lapse movie of the ER in a live HeLa cell labeled with Cer-HMSiR.
Live HeLa cells were labeled with Cer-TCO followed by HMSiR-Tz and imaged live on a custom 2D wide-field SMS microscope (see Fig. 2 c-f). The movie was recorded at 400 frames per second, reconstructed with a 2 s temporal resolution (800 frames per movie frame) and Kalman-filtered using default parameters. Laser intensity: 9.9 kW/cm2. Scale bar: 1 μm. (MOV 24600 kb)
Zoom-in view of 2D ER dynamics in a live HeLa cell labeled with Cer-HMSiR.
The movie illustrates the area of interest labeled d in Fig. 2c; individual frames are shown as Fig. 2d. The yellow arrow identifies a loop growing out of a single ER tubule. The blue and pink arrows identify the fission and fusion of two separate ER tubules. The green arrow identifies the elongation of an ER tubule and fusion with a neighboring tubule. Scale bar: 500 nm. (MOV 16909 kb)
Comparison of movie quality at different temporal resolutions.
Five movies generated from the same raw data but with different temporal resolution. The number above each panel represents the temporal resolution (time per frame) of that movie. The movie with a 2.0 s temporal resolution corresponds to the single image shown as Fig. 2c; stills are shown in Supplementary Fig. 9. Scale bar: 1 μm. (MOV 46692 kb)
Direct comparison between long time-lapse SMS and diffraction-limited images of the ER in a live HeLa cell labeled with Cer-HMSiR.
Side-by-side comparison of ER dynamics observed under SMS and diffraction limited conditions. Scale bar: 1 μm. (MOV 10738 kb)
Direct comparison between SMS images obtained using HMSiR conjugated to a lipid (Cer-HMSiR) or a protein (Sec61β-HMSiR) within the ER in live HeLa cells.
Live HeLa cells were labeled with Cer-TCO and HMSiR-Tz or transfected Halo-Sec61β and labeled with HMSiR-CA, and subsequently imaged live on a custom 2D wide-field SMS microscope. The data from this movie was used to prepare the plot shown as Fig. 2g. Scale bar: 1 μm. (MOV 1689 kb)
Time-lapse movie of mitochondria in a live HeLa cell labeled with RhoB-HMSiR.
Live HeLa cells were labeled with RhoB-TCO followed by HMSiR-Tz and imaged live on a custom 2D wide-field SMS microscope. The movie was recorded at 400 frames per second, reconstructed with a 2 s temporal resolution (800 frames per movie frame) and Kalman-filtered with default parameters. This movie corresponds to the time-colored image shown in Fig. 3c. Laser intensity: 4.4 kW/cm2. Scale bar: 1 μm. (MOV 12047 kb)
Time-lapse movie of the plasma in a live HeLa cell labeled with DiI-HMSiR.
Live HeLa cells were labeled with DiI-TCO followed by HMSiR-Tz and imaged live on a custom 2D wide-field SMS microscope. A still image of an early movie frame is shown in Fig. 4b. Time-stamped snapshots from the movie are shown in Fig. 4d. The movie was recorded at 400 frames per second, reconstructed with a 2 s temporal resolution (800 frames per movie frame) and Kalman-filtered with default parameters. Snapshots from this movie are shown in Supplementary Fig. 16. Laser intensity: 9.9 kW/cm2. Scale bar: 1 μm. (MOV 8730 kb)
Extra-long time-lapse movie of the plasma membrane in a live HeLa cell labeled with DiI-C6-HMSiR.
Live HeLa cells were labeled with DiI-C6-TCO followed by HMSiR-Tz and imaged live on a custom 2D wide-field SMS microscope. The movie was recorded at 400 frames per second, reconstructed with a 2 s temporal resolution (800 frames per movie frame) and Kalman-filtered with default parameters. Data from this movie was used to generate Fig. 4e. Snapshots from this movie are shown in Supplementary Fig. 16. Laser intensity: 9.9 kW/cm2. Scale bar: 1 μm. (MOV 23786 kb)
Multi-perspective 3D zoom-in view of ER tubule network in a live HeLa cell.
Live HeLa cells were labeled with Cer-TCO followed by HMSiR-Tz and imaged live on a custom 3D wide-field SMS microscope. Movie shows multiple 3D perspectives of the area of interest shown in the pink box in Fig. 6a and b; stills from this movie are shown as Fig. 6c and d. A cartoon schematic of this region is shown as Fig. 6e. Movie is depth-colored. Laser intensity 9.7 kW/cm2. Scale bar: 1 μm. (MOV 1830 kb)
Long time-lapse, depth-colored 3D movie of the ER in a live HeLa cell labeled with Cer-HMSiR.
Live HeLa cells were labeled with Cer-TCO followed by HMSiR-Tz and imaged live on a custom 3D wide-field SMS microscope. A depth-colored image of an early frame of this movie is shown as Fig. 6a. Laser intensity 9.7 kW/cm2. Scale bar: 1 μm. (MOV 20423 kb)
Long time-lapse, 3D movie of the ER in a live HeLa cell labeled with Cer-HMSiR (for 3D glasses).
Supplementary Movie 11 rendered for observation with 3D glasses. (MOV 20504 kb)
2D ER dynamics in a live HeLa cell labeled with Cer-HMSiR.
Live HeLa cells were labeled with Cer-TCO followed by HMSiR-Tz and imaged live on a custom 3D wide-field SMS microscope. Pink arrow shows tubule elongation spanning different z-layers and fusion with a neighboring tubule. Images from this movie are shown as Fig. 6f. Movie is depth-colored. Laser intensity 9.7 kW/cm2. Scale bar: 4 μm. (MOV 8345 kb)
Rights and permissions
About this article
Cite this article
Takakura, H., Zhang, Y., Erdmann, R. et al. Long time-lapse nanoscopy with spontaneously blinking membrane probes. Nat Biotechnol 35, 773–780 (2017). https://doi.org/10.1038/nbt.3876
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3876
- Springer Nature America, Inc.
This article is cited by
-
The endoplasmic reticulum connects to the nucleus by constricted junctions that mature after mitosis
EMBO Reports (2024)
-
Long-term super-resolution inner mitochondrial membrane imaging with a lipid probe
Nature Chemical Biology (2024)
-
Membrane topography and the overestimation of protein clustering in single molecule localisation microscopy – identification and correction
Communications Biology (2024)
-
MAO–SiR is a tool for visualizing the inner workings of mitochondria
Nature Chemical Biology (2024)
-
Bioorthogonal Chemistry in Cellular Organelles
Topics in Current Chemistry (2024)