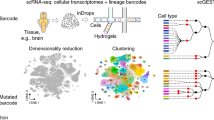
Abstract
Embryonic development is a crucial period in the life of a multicellular organism, during which limited sets of embryonic progenitors produce all cells in the adult body. Determining which fate these progenitors acquire in adult tissues requires the simultaneous measurement of clonal history and cell identity at single-cell resolution, which has been a major challenge. Clonal history has traditionally been investigated by microscopically tracking cells during development1,2, monitoring the heritable expression of genetically encoded fluorescent proteins3 and, more recently, using next-generation sequencing technologies that exploit somatic mutations4, microsatellite instability5, transposon tagging6, viral barcoding7, CRISPR–Cas9 genome editing8,9,10,11,12,13 and Cre–loxP recombination14. Single-cell transcriptomics15 provides a powerful platform for unbiased cell-type classification. Here we present ScarTrace, a single-cell sequencing strategy that enables the simultaneous quantification of clonal history and cell type for thousands of cells obtained from different organs of the adult zebrafish. Using ScarTrace, we show that a small set of multipotent embryonic progenitors generate all haematopoietic cells in the kidney marrow, and that many progenitors produce specific cell types in the eyes and brain. In addition, we study when embryonic progenitors commit to the left or right eye. ScarTrace reveals that epidermal and mesenchymal cells in the caudal fin arise from the same progenitors, and that osteoblast-restricted precursors can produce mesenchymal cells during regeneration. Furthermore, we identify resident immune cells in the fin with a distinct clonal origin from other blood cell types. We envision that similar approaches will have major applications in other experimental systems, in which the matching of embryonic clonal origin to adult cell type will ultimately allow reconstruction of how the adult body is built from a single cell.
Similar content being viewed by others
Main
The goal of our experiment is twofold: first, to link cells in the embryo to their corresponding clones in adult tissue (Fig. 1a); second, to quantify cell-type composition of these clones to determine the multipotency of embryonic progenitors. To reach the first goal, we need to uniquely label the cells in an embryo with permanent and heritable labels. For this, we use CRISPR–Cas9 technology, which induces a double-stranded break at the targeted genomic site that is repaired as insertions or deletions of different lengths at different positions (scars)16,17. To allow for multiple scarring in the same cell, we use a zebrafish line with eight in-tandem copies of a histone–green fluorescent protein (GFP) transgene17 (Methods). Scarring starts after injecting the yolk or cell of the zygote with Cas9 RNA or protein, and a single-guide RNA (sgRNA) that targets GFP (Fig. 1b).
a, Embryonic cells get permanent and unique labels that are transmitted to the clones in the adult. b, Zygote injection with Cas9 RNA or protein and sgRNA that targets GFP (GFP-sgRNA). H2A, histone 2A. c, Mean fraction of unscarred GFP as a function of time, computed over ten independently injected embryos (t > 6 h), and over three pools of ten embryos (t ≤ 6 h), in which GFP was PCR-amplified from gDNA. Error bars denote s.e.m. Unscarred GFP exponentially decreases at 0.064 ± 0.002 h−1 (0.294 ± 0.008 h−1) and is constant after 10 ± 1.0 h (3.1 ± 0.1 h) for RNA (protein) injections (solid lines, Supplementary Information section 1). d, ScarTrace protocol (Methods). IVT, in vitro transcription; RT, reverse transcription. cBC, cell-specific barcode for transcriptome; sBC, cell-specific barcode for scars.
We quantified the scarring rate by measuring the fraction of unscarred GFP in zebrafish embryos at different times after Cas9 delivery (Fig. 1c), which is five times faster in Cas9 protein than in RNA injections. Cas9 activity ceases at around 3 h for protein and at 10 h for RNA injections, when zebrafish embryos have about 1,000 and 8,000 cells, respectively2. We detect more than 1,000 distinct scars, the abundances and probabilities of which span several orders of magnitude13 (Supplementary Information sections 1 and 2).
To detect scars and transcriptome from single cells, we developed ScarTrace, which integrates a nested PCR step after transcriptome conversion to cDNA into the sorting and robot-assisted transcriptome sequencing (SORT-seq) protocol18 (Fig. 1d). Because the histone–GFP transgene is transcribed, scars can be detected from mRNA and genomic DNA (gDNA). Detection from gDNA is preferred because GFP expression might be tissue specific, vulnerable to silencing and scars might affect the half-life of the mRNA. We assessed the efficiency of scar detection from mRNA and gDNA by comparing scar patterns of single cells from the caudal fin obtained using ScarTrace with and without reverse transcription (Fig. 1d, step 1). We detected 3.3 ± 0.3 (mean ± s.e.m.) scars per clone on average, and approximately 25% of the cells remained unscarred and therefore do not contain clonal information (Extended Data Fig. 1a). Clone sizes from gDNA and gDNA–mRNA detection are very similar (Extended Data Fig. 1a–d), indicating that ScarTrace reliably detects scars from gDNA in single cells.
We next used ScarTrace to explore the clonal composition of haematopoietic cells isolated from the whole kidney marrow (WKM) of two protein-injected (P1 and P2) and two RNA-injected (R1 and R2) zebrafish. We found one and two major clones in P1 and P2, and eight and six in R1 and R2, respectively (Fig. 2a, b, Extended Data Fig. 2a, b, Extended Data Table 1). This is a direct result of the time window of Cas9 activity (Fig. 1c). The number of observed clones agrees with previous findings using GESTALT8, in which a similar Cas9-mediated approach is used to label embryonic clones in zebrafish, and with the number of clones (between 10.4 and 15.4) found for haematopoietic stem and progenitor cells at 10–14 hours post fertilization (hpf) using Zebrabow19.
a, b, Scar percentage per cell for fish P1 (a) and R1 (b) (for key, see Extended Data Fig. 2a). The bar above each panel indicates clones and corresponding P values. uGFP, unscarred GFP. c, d, Lineage trees for clones in P1 (c) and R1 (d). The root is an unscarred clone. Clones with corresponding cell fraction and scar copy number (Supplementary Information section 3) are at the tips. The statistical confidence of each branch is computed as its proportion among 10,000 bootstrapped tree replicates. Only clones with more than 2 cells are taken into account. e, t-distributed stochastic neighbour embedding (t-SNE) map of cells from fish R1 and R2 obtained with transcriptome data. Colours indicate the cell type (Extended Data Fig. 2). HSPCs, haematopoietic stem and progenitor cells. f, t-SNE map of cells in fish R1. Colours indicate the clone. g, Clonal cell fraction per cell type for fish R1.
The average number of scars per clone equals 3.3 ± 0.3 for P1, 1.02 ± 0.01 for P2, 3.5 ± 0.3 for R1 and 3.0 ± 0.3 for R2, with a minimum of 1 scar and a maximum of 5 scars per clone, revealing that both Cas9 protein and RNA efficiently cause scarring. We determined the copy number for each scar in a clone by modelling the amplification and sequencing noise of ScarTrace as a branching process (Supplementary Information section 3). Typically, the resulting number of scars per clone is smaller than eight, as a consequence of two or more simultaneously Cas9-induced cuts in the same multi-copy tandem histone-GFP gene10. We computed the P value of a combination of scars to occur in a cell (Fig. 2a, b). Values obtained are commonly below 10−6, emphasizing that although identical scars might be independently introduced in different clones (for example, the yellow scar is present in one clone from fish R1 and four clones from fish R2), the chance of introducing the same combination of scars in independent clones is very small. Consistently, we do not find overlapping clones between different zebrafish. Using cell-to-cell variation in scar composition, we estimate a 90% scar detection efficiency (including unscarred GFP; Extended Data Fig. 1e, f). In addition, by assuming maximum parsimony for sequential scarring events, we build lineage trees for clones (Fig. 2c, d, Extended Data Fig. 2c, d, Supplementary Information section 4).
Using RaceID20 (Methods), we identify eight haematopoietic cell types in fish R1 and R2 (Fig. 2e). Gene expression profiles in the different cell types found for both fish are identical with the exception of erythrocytes, which show slight differences in the expression of characteristic markers (Extended Data Fig. 2e–h). After combining cell type and clonal information for single cells, we observe all clones in all cell types with similar proportions (Fig. 2f, g, Extended Data Fig. 2i, j), indicating that all clones contribute to the production of all blood cells. This is consistent with haematopoietic stem and progenitor cells specification (around 28 hpf), when scarring is already completed21.
Next, we used ScarTrace in the adult brain and eyes of two RNA-injected fish (R2 and R3), in which we identified different neuronal, glia and immune cells (Fig. 3a, Extended Data Fig. 3). To determine clonal enrichment or depletion in certain cell types quantitatively, we used Fisher’s exact test (Fig. 3b, Extended Data Fig. 4a). Here, several clones only generate neurons or retinal interneurons (Extended Data Fig. 4b, c). We observed that microglia share clones with the WKM, confirming that they originate from the WKM22.
a, t-SNE map of cells in fish R2. Colours indicate cell type (Extended Data Fig. 3). COPCs, committed oligodendrocyte progenitor cells; MFOLs, myelin-forming oligodendrocytes; OPCs, oligodendrocytes progenitors; RGC, retinal ganglion cell. b, Heat map of clonal cell fraction for cell types in fish R2 (COPC, OPC and MFOL clones merged as oligodendrocytes), and two-sided Fisher’s exact test for enriched and depleted clones per cell type with P < 0.05. The bars at the top depict organ, total number of cells, and P value for each clone. Bip., bipolar; RGC, retinal ganglion cell. c, Relative clone frequency in the left (L) and right (R) eye and midbrain for fish R2 and P1, and left and right eye and erythrocytes and immune cells for fish S1.
Upon the exclusion of WKM clones, we found that clones are not only cell-type specific, but also brain-region and eye specific (Extended Data Figs 4d–g, 5a–d). Although R2 and R3 left and right midbrains share a small fraction of clones, left and right eyes share none (Fig. 3c, Extended Data Fig. 4h). However, for fish P1, both midbrains share almost all clones whereas eyes share only one. To explore when this segregation is established, we injected one cell at the two-cell stage with Cas9–eScarlet fusion protein and sgRNA. We found Cas9–eScarlet protein present in only half of the embryo at dome stage (Extended Data Fig. 4i–l), approximately 3 h after Cas9 protein stops scarring. Therefore, scars only occur in one side of the embryo. However, ScarTrace on the left and right eyes of a 3-week-old-injected embryo (S1) reveals scars in both eyes. Upon removal of the clones found in immune cells and erythrocytes, the rest of the clones are specific to each eye (Fig. 3c). This indicates that both eyes get cellular contributions from both sides of the dome-stage embryo. To determine further when lateral commitment arises in eye progenitors, we built lineage trees for clones detected in the left and right eyes or midbrain for fish P1 and R2 (Extended Data Fig. 5e–h). In P1, no significant co-evolution is found among clones from the right (left) eye. By contrast, in R2 we observe a significant depletion of right eye clones evolving with left eye clones. This suggests that progenitors commit to the left or right eye shortly before the end of scarring with Cas9 protein. No significant co-evolution enhancement or depletion is found for clones detected in the left and right midbrain, indicating that cell mixing is important at 10 hpf. This is consistent with the processes of neurulation and neurogenesis23.
Next, we focused on zebrafish caudal fin ontogeny and regeneration. We performed ScarTrace on the primary, secondary and tertiary fins of fish R4, R5 and R6 (Fig. 4a). We identified four major cell types (osteoblasts, mesenchymal, epidermal and immune cells) and observed cell-type-restricted clones in all fish (Fig. 4b, c, Extended Data Figs 6, 7a–e). We found that mesenchymal and epidermal cells share clones, revealing a common developmental origin that is maintained during regeneration. Together with previous imaging-based studies24, this suggests that epidermal ancestors undergo epithelial-to-mesenchymal transition during gastrulation to generate mesenchymal cells in the caudal fin. Osteoblasts did not share clones with any other cell type in the primary fin and showed dorsal–ventral segregation, confirming their early lineage commitment during development13,25,26,27,28. We found lineage restriction of the different cell types as the main mechanism of fin regeneration, consistent with previous results25,28. However, after regeneration, we observed osteoblast-committed clones that generate a fraction (approximately 21% in R4, 44% in R6) of mesenchymal cells (Fig. 4d, Extended Data Fig. 7f). This suggests a certain degree of plasticity after injury, in which progenitors that produce osteoblasts during development can also give rise to mesenchymal cells during fin regeneration29.
a, Primary (first week), secondary (fully regenerated after 3 weeks) and tertiary (fully regenerated after 6 weeks) fins are amputated. Tertiary fins are split dorsally and ventrally. b, t-SNE map of cells in fish R4, R5 and R6. Colours indicate cell type (Extended Data Fig. 6). c, Heat map of clonal cell fraction per cell type and fin in fish R4, and two-sided Fisher’s exact test for clones that are enriched and depleted with P < 0.05. The bars at the top show clones in the WKM, number of cells and P value per clone. d, t-SNE map of cells from fish R4, with cells from the primary (top) or regenerated (bottom) fin detected in osteoblast clones or remaining cell types. Percentages are mesenchymal cell fractions that share clones with osteoblasts. e, Magnified view of t-SNE map indicating immune cells with fin-specific clones, immune cells that share clones with the WKM and cells with no clone information (grey). Subpopulations of lymphoid cells and myeloid cells (numbered 1–4), and the percentage of RICs are indicated. f, Differential gene expression between the four subpopulations of myeloid cells (Supplementary Table 6). g, Cell type fraction for fin-specific clones detected in RICs in the primary fin for R4.
Finally, we investigated the clonal overlap of single cells from the WKM of fish R4, R5 and R6 with immune cells found in the fin. Clones detected in the WKM are enriched in the fin immune cells and depleted in the remaining cell types (Fig. 4c, e, Extended Data Fig. 7g). We found sub-populations of lymphoid and myeloid cells in all fins with different proportions of fin-specific clones, which we identify as resident immune cells (RICs). Differential gene expression analysis in myeloid cells revealed that subpopulation 4 expressed macrophage markers together with the epithelial marker epcam (Fig. 4f), which has been reported in resident macrophages in mice30. All RICs in the primary fin share clonality with epidermal and mesenchymal cells (Fig. 4g, Extended Data Fig. 7b, c). Therefore, our data indicate that RICs have a distinct origin from haematopoietic stem cells (Extended Data Fig. 7h), and arise either from epidermal and mesenchymal transdifferentiation, or from ectodermal ancestors similarly to mesenchymal cells.
We developed ScarTrace as a new method to quantify clonal origin and cell type simultaneously at single-cell resolution. This enabled us to investigate the embryonic origin of clones found in different organs of the adult zebrafish and their cell-type commitment during development and regeneration. CRISPR–Cas9 genome editing technology for lineage tracing purposes at the single cell level has recently also been used in zebrafish to investigate lineages and cell types in the vertebrate brain, and to unravel developmental lineages31,32. We anticipate many applications of ScarTrace in developmental and stem-cell biology, and similar approaches to study clonal selection in cancer models. Because ScarTrace provides a glimpse of the cellular past, it will be interesting to explore how this history is predictive of the current epigenetic and expression state.
Methods
Zebrafish Cas9 and sgRNA injections
Heterozygous zygotes of the transgenic zebrafish line Tg(h2afva:GFP)kca66:(h2afva:GFP)kca66 (ref. 17) were injected at the cell with 1 nl Cas9 protein (NEB; final concentration 1,590 ng μl−1) or at the yolk with 1 nl Cas9 RNA (300 ng μl−1) in combination with an sgRNA that targets GFP (25 ng μl−1, sequence: GGTGTTCTGCTGGTAGTGGT) (Fig. 1b). Cas9 RNA was in vitro transcribed from a linearized pCS2-nCas9n vector (Addgene plasmid 47929)16 using the mMESSAGE mMACHINE SP6 Transcription Kit (Thermo Scientific). The sgRNA was in vitro transcribed from a template using the MEGAscript. T7 Transcription Kit (Thermo Scientific). The sgRNA template was synthesized with T4 DNA polymerase (New England Biolabs) by partially annealing two single-stranded DNA oligonucleotides containing the T7 promotor and the GFP-binding sequence, and the tracrRNA sequence, respectively56. Male and female zebrafish were used, no randomization was done, no blinding was done and no animals were excluded from the analysis. No statistical methods were used to predetermine sample size. The age of the fish used in isolated organ spans 3–18 months. For sample sizes, see Extended Data Table 2. All animal experiments were performed in accordance with institutional and governmental regulations, and were approved by the Dier Experimenten Commissie of the Royal Netherlands Academy of Arts and Sciences and performed according to the guidelines.
Transgene copy number
To determine the number of integrations of the transgene, we performed whole-genome sequencing (NEBNext Ultra library preparation kit for Illumina (E7370S) and the NEB Multiplex Oligos for Illumina (E7500L)) on an homozygous Tg(h2afva:GFP)kca66:(h2afva:GFP)kca66 fish. Paired-end data were trimmed (TrimGalore-0.4.3) and mapped (bwa-0.7.10 mem) to the zebrafish reference genome (danRer10 from UCSC Genome Browser), and PCR and optical duplicates were removed (Picard-2.0.1) (Extended Data Fig. 8a, b). The copy number was extracted using FREEC-11.057 with default parameters. With a 1-kb window size, we find 19 copies of the transgene fragment, whereas with a 500-bp window size, we find 18 (Extended Data Fig. 8b). After correcting for reads due to endogenous copies, we estimate the number of copies of the transgene in a heterozygous fish to be 8 ± 1. This number agrees with single-cell data, because although we detected a maximum of 7 scars per clone (Extended Data Fig. 8c, d), we see that sometimes 6 of the scars in those clones represent approximately 12.5% of the scar content per cell, and one represents approximately 25% (Extended Data Fig. 8e). This again suggests that the number of integrations of the histone-GFP transgene is 1/0.125 = 8.
ScarTrace protocol
Live single cells (based on DAPI exclusion and scatter properties) were sorted into 384-well plates (Sigma-Aldrich) containing 5 μl of mineral oil (Sigma-Aldrich), 50 nl of uniquely barcoded reverse transcription primers (Supplementary Table 1), dNTPs (Promega), Spike-in controls (Thermo Fisher) and RNase inhibitor (SUPERaseIn, Thermo Fisher). Plates were immediately spun down and stored at −80 °C. Cells were lysed at 65 °C for 5 min. Reverse transcription and second-strand synthesis mixes were dispensed into each well using the Nanodrop II and reactions were performed at 42 and 16 °C degrees, respectively (Fig. 1d, step 1). Genomic DNA was access by proteinase K treatment followed by a nested PCR strategy to amplify the scarred GFP region (Fig. 1d, step 2). In the second PCR, unique scar barcodes were introduced in each well (Supplementary Table 2). All cells were pooled and the aqueous phase was separated from the oil phase (Fig. 1d, step 3). The collected material was split for scar library and transcriptome library preparation (Fig. 1d, step 4). For transcriptome library preparation, the SORT-seq protocol18 was used (Fig. 1d, step 5a). For scar library preparation, a PCR introducing only Illumina TruSeq adapters was perfomed (Fig. 1d, step 5b). All libraries were sequenced paired-end at 75 bp read-length on the Illumina NextSeq platform. A detailed description of the protocol is available in Protocol Exchange33.
WKM isolation
The WKM was isolated as previously described34. A ventral midline incision was made to open the adult zebrafish body cavity. All internal organs were carefully removed to access the kidney. The WKM was collected in PBS supplemented with FCS. The tissue was aspirated through a 1 ml pipet tip several times to mechanically dissociate haematopoietic cells. After two consecutive filtering steps (using 70-μm and 40-μm cell strainers (VWR), cells were centrifuged and washed. The pellet of haematopoietic cells was resuspended in PBS and FCS supplemented with DAPI (Thermo Fisher) to assess cell viability
Brain parts and eye isolation
Brain and eyes were isolated from the zebrafish head and dissected in PBS. Optic nerves were removed. The forebrain (olfactory bulb and telencephalon) was isolated from the midbrain, followed by dissection of the hindbrain (rhombencephalon). The midbrain (mesencephalon) was dissected into left and right midbrain. The eyes lens was carefully removed. Brain parts and eyes were dissociated into single cells using a papain-based solution (Thermo Fisher, 88285) and washing solutions as previously described35. The washed cell pellet was resuspended in DMEM/F12 medium (Thermo Fisher, 11320033) and supplemented with DAPI (Thermo Fisher) to assess cell viability for FACS.
Fin amputation
Caudal fin amputations were performed as previously described36, after which fish were returned to 28 °C aquarium water. Once isolated, this tissue was immediately dissociated by moderately shaking at 30 °C for 1 h, with gentle trituration performed every 10 min with a p200 pipet, in a solution of 2 mg ml−1 collagenase A (Sigma-Aldrich) and 0.3 mg ml−1 protease (type XIV, Sigma-Aldrich) in Hanks solution. After 1 h, the solution was incubated for 5 min in 0.05% trypsin in PBS. The solution was strained using 70-μm and 40-μm cell strainers (Corning) and cells were washed in 2% FBS in Hanks solution. Before flow cytometry, cells were centrifuged and resuspended in PBS and FBS supplemented with DAPI (Thermo Fisher) to assess cell viability.
Transcriptome analysis
In transcriptome libraries, the first read contains cell barcode (Supplementary Table 1) and unique molecular identifier (UMI) information, and the second read contains biological information. Second reads with a valid cell barcode extracted from corresponding first reads are mapped using bwa mem-0.7.10 with default parameters to the reference zebrafish transcriptome (Danio rerio assembly Zv9, ensemble 74, extended with ERCC92). For each cell, the number of transcripts per gene was obtained as previously described37. We refer to transcripts as unique molecules based on UMI correction. We ran RaceID3 with different parameters for each organ under study (Supplementary Data 1) for cell filtering, normalization, gene filtering, cell clustering and differential gene expression analysis (in which P values are calculated using negative binomial distribution and corrected for multiple testing by the Benjamini–Hochberg method). The choice of filtering parameters was made to include the maximum number of cells in our analysis without losing cell type information. Supplementary Tables 3–6 provide results for the differentially expressed genes for each cell type compared with all other cells in the organ: WKM38,39,40,41 (90 dendritic cells, 76 eosinophils, 641 erytrhocytes, 516 haematopoietic stem and progenitor cells, 446 lymphocytes, 409 monocytes, 927 neutrophils and 76 thrombocytes), brain and eyes42,43,44,45,46,47,48,49,50,51 (250 bipolar and horizontal cells, 45 COPCs, 9 cones, 290 erythrocytes, 254 immune cells, 88 glia-like cells, 89 MFOLs, 66 microglia, 1,427 neurons, 10 OPCs, 31 RCL, 53 radial glia and 202 rods), caudal fin52,53,54 (144 epidermal cells, 2,834 fibroblasts, 1,784 immune cells, and 2,951 osteoblasts), and resident myeloid cell types in the fin (118 cells in subpopulation 1, 45 in subpopulation 2, 27 in subpopulation 3 and 133 in subpopulation 4).
Scar analysis
In scar libraries, the first read contains the cell barcode (Supplementary Table 2) and the forward primer used in the nested PCR and second read contains the sequence for the scar and the reversed primer. Scripts to extract scars and detect clones are provided as Supplementary Data 2, together with a reference manual (Supplementary Data 3). Bug fixes and updates of the scripts can be downloaded from https://github.com/anna-alemany/scScarTrace. Cells sharing an identical scar pattern are assumed to come from the same clone, independently of scar percentage. Cells with a detected scar pattern that can be assigned to another single clone by assuming that some scar was not sampled were pooled with that clone. Cells that according to their scar pattern can be ambiguously assigned to two or more other clones were removed from subsequent analysis. Clones with less than three cells were also removed.
Code availability
Transcriptome analysis was performed using RaceID3 available at https://github.com/dgrun/RaceID3_StemID2, with parameters summarized in Supplementary Data 1. Scripts for scar extraction and clone detection are provided in Supplementary Data 2, together with a reference manual (Supplementary Data 3). Bug fixes and updates of the scripts can be downloaded from https://github.com/anna-alemany/scScarTrace.
Data availability
The accession numbers for the RNA sequencing datasets reported in this paper have been deposited with the Gene Expression Omnibus (GEO) under accession GSE102990.
Accession codes
References
Sulston, J. E. & Horvitz, H. R. Post-embryonic cell lineages of the nematode, Caenorhabditis elegans. Dev. Biol. 56, 110–156 (1977)
Keller, P. J., Schmidt, A. D., Wittbrodt, J. & Stelzer, E. H. Reconstruction of zebrafish early embryonic development by scanned light sheet microscopy. Science 322, 1065–1069 (2008)
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007)
Behjati, S. et al. Genome sequencing of normal cells reveals developmental lineages and mutational processes. Nature 513, 422–425 (2014)
Reizel, Y. et al. Colon stem cell and crypt dynamics exposed by cell lineage reconstruction. PLoS Genet. 7, e1002192 (2011)
Sun, J. et al. Clonal dynamics of native haematopoiesis. Nature 514, 322–327 (2014)
Naik, S. H. et al. Diverse and heritable lineage imprinting of early haematopoietic progenitors. Nature 496, 229–232 (2013)
McKenna, A. et al. Whole-organism lineage tracing by combinatorial and cumulative genome editing. Science 353, aaf7907 (2016)
Guernet, A. et al. CRISPR-barcoding for intratumor genetic heterogeneity modeling and functional analysis of oncogenic driver mutations. Mol. Cell 63, 526–538 (2016)
Schmidt, S. T., Zimmerman, S. M., Wang, J., Kim, S. K. & Quake, S. R. Quantitative analysis of synthetic cell lineage tracing using nuclease barcoding. ACS Synth. Biol. 6, 936–942 (2017)
Kalhor, R., Mali, P. & Church, G. M. Rapidly evolving homing CRISPR barcodes. Nat. Methods 14, 195–200 (2017)
Frieda, K. L. et al. Synthetic recording and in situ readout of lineage information in single cells. Nature 541, 107–111 (2017)
Junker, J. P. et al. Massively parallel whole-organism lineage tracing using CRISPR/Cas9 induced genetic scars. Preprint at https://www.biorxiv.org/content/early/2016/06/01/056499 (2016)
Pei, W. et al. Polylox barcoding reveals haematopoietic stem cell fates realized in vivo. Nature 548, 456–460 (2017)
Tanay, A. & Regev, A. Scaling single-cell genomics from phenomenology to mechanism. Nature 541, 331–338 (2017)
Jao, L. E., Wente, S. R. & Chen, W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc. Natl Acad. Sci. USA 110, 13904–13909 (2013)
Pauls, S., Geldmacher-Voss, B. & Campos-Ortega, J. A. A zebrafish histone variant H2A.F/Z and a transgenic H2A.F/Z:GFP fusion protein for in vivo studies of embryonic development. Dev. Genes Evol. 211, 603–610 (2001)
Muraro, M. J. et al. A single-cell transcriptome atlas of the human pancreas. Cell Syst. 3, 385–394.e3 (2016)
Henninger, J. et al. Clonal fate mapping quantifies the number of haematopoietic stem cells that arise during development. Nat. Cell Biol. 19, 17–27 (2017)
Grün, D. et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 525, 251–255 (2015)
Jing, L. & Zon, L. I. Zebrafish as a model for normal and malignant hematopoiesis. Dis. Model. Mech. 4, 433–438 (2011).
Xu, J. et al. Temporal-spatial resolution fate mapping reveals distinct origins for embryonic and adult microglia in zebrafish. Dev. Cell 34, 632–641 (2015)
Schmidt, R., Strähle, U. & Scholpp, S. Neurogenesis in zebrafish — from embryo to adult. Neural Dev. 8, 3 (2013)
Lee, R. T., Knapik, E. W., Thiery, J. P. & Carney, T. J. An exclusively mesodermal origin of fin mesenchyme demonstrates that zebrafish trunk neural crest does not generate ectomesenchyme. Development 140, 2923–2932 (2013)
Tu, S. & Johnson, S. L. Fate restriction in the growing and regenerating zebrafish fin. Dev. Cell 20, 725–732 (2011)
Knopf, F. et al. Bone regenerates via dedifferentiation of osteoblasts in the zebrafish fin. Dev. Cell 20, 713–724 (2011)
Singh, S. P., Holdway, J. E. & Poss, K. D. Regeneration of amputated zebrafish fin rays from de novo osteoblasts. Dev. Cell 22, 879–886 (2012)
Tornini, V. A. et al. Live monitoring of blastemal cell contributions during appendage regeneration. Curr. Biol. 26, 2981–2991 (2016)
Tornini, V. A., Thompson, J. D., Allen, R. L. & Poss, K. D. Live fate-mapping of joint-associated fibroblasts visualizes expansion of cell contributions during zebrafish fin regeneration. Development 144, 2889–2895 (2017)
Gautier, E. L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–1128 (2012)
Raj, B. et al. Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nat. Biotechnol. https://doi.org/10.1038/nbt.4103 (2018).
Spanjaard, B. et al. Simultaneous lineage tracing and cell-type identification using CRISPR/Cas9-induced genetic scars. Nat. Biotechnol. https://doi.org/10.1038/nbt/4124 (2018)
Peterson-Maduro, J., Florescu, M., Baron, C. S., Alemany, A. & van Oudenaarden, A. Single-cell ScarTrace. Protoc. Exch. https://doi.org/10.1038/protex.2018.017 (2018)
Stachura, D. L. & Traver, D. Cellular dissection of zebrafish hematopoiesis. Methods Cell Biol. 101, 75–110 (2011)
Lopez-Ramirez, M. A., Calvo, C. F., Ristori, E., Thomas, J. L. & Nicoli, S. Isolation and culture of adult zebrafish brain-derived neurospheres. J. Vis. Exp. 108, 53617 (2016).
Poss, K. D. et al. Roles for Fgf signaling during zebrafish fin regeneration. Dev. Biol. 222, 347–358 (2000)
Grün, D., Kester, L. & van Oudenaarden, A. Validation of noise models for single-cell transcriptomics. Nat. Methods 11, 637–640 (2014)
Kobayashi, I. et al. Comparative gene expression analysis of zebrafish and mammals identifies common regulators in hematopoietic stem cells. Blood 115, e1–e9 (2010)
Moore, F. E. et al. Single-cell transcriptional analysis of normal, aberrant, and malignant hematopoiesis in zebrafish. J. Exp. Med. 213, 979–992 (2016)
Macaulay, I. C. et al. Single-cell RNA-sequencing reveals a continuous spectrum of differentiation in hematopoietic cells. Cell Reports 14, 966–977 (2016)
Carmona, S. J. et al. Single-cell transcriptome analysis of fish immune cells provides insight into the evolution of vertebrate immune cell types. Genome Res. 27, 451–461 (2017)
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008)
Nelson, S. M., Frey, R. A., Wardwell, S. L. & Stenkamp, D. L. The developmental sequence of gene expression within the rod photoreceptor lineage in embryonic zebrafish. Dev. Dyn. 237, 2903–2917 (2008)
Zhang, H., Copara, M. & Ekstrom, A. D. Differential recruitment of brain networks following route and cartographic map learning of spatial environments. PLoS ONE 7, e44886 (2012)
Hickman, S. E. et al. The microglial sensome revealed by direct RNA sequencing. Nat. Neurosci. 16, 1896–1905 (2013)
Di Donato, V., Auer, T. O., Duroure, K. & Del Bene, F. Characterization of the calcium binding protein family in zebrafish. PLoS ONE 8, e53299 (2013)
Sunkin, S. M. et al. Allen Brain Atlas: an integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res. 41, D996–D1008 (2013)
La Manno, G . et al. Molecular diversity of midbrain development in mouse, human, and stem cells. Cell 167, 566–580 (2016)
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016)
Chen, R., Wu, X., Jiang, L. & Zhang, Y. Single-cell RNA-seq reveals hypothalamic cell diversity. Cell Reports 18, 3227–3241 (2017)
Oosterhof, N. et al. Identification of a conserved and acute neurodegeneration-specific microglial transcriptome in the zebrafish. Glia 65, 138–149 (2017).
Marie, P. J. Transcription factors controlling osteoblastogenesis. Arch. Biochem. Biophys. 473, 98–105 (2008)
Akerberg, A. A., Stewart, S. & Stankunas, K. Spatial and temporal control of transgene expression in zebrafish. PLoS ONE 9, e92217 (2014)
Smyth, I. et al. The extracellular matrix gene Frem1 is essential for the normal adhesion of the embryonic epidermis. Proc. Natl Acad. Sci. USA 101, 13560–13565 (2004)
Scott, D. On optimal and data-based histograms. Biometrika 66, 605–610 (1979)
Woo, K. & Fraser, S. E. Order and coherence in the fate map of the zebrafish nervous system. Development 121, 2595–2609 (1995)
Boeva, V. et al. Control-free calling of copy number alterations in deep-sequencing data using GC-content normalization. Bioinformatics 27, 268–269 (2011)
Acknowledgements
This work was supported by a European Research Council Advanced grant (ERC-AdG 742225-IntScOmics), Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO) TOP award (NWO-CW 714.016.001), and the Foundation for Fundamental Research on Matter, financially supported by NWO (FOM-14NOISE01). This work is part of the Oncode Institute which is partly financed by the Dutch Cancer Society. We thank M. Sen for help with sequencing, R. der Linden for cell sorting, and B. de Barbanson for help with programming, and all the other members of the A.v.O. laboratory for discussions and input. In addition, we thank B. Artegiani and J. Bakkers for discussions, P. Shang and N. Geijsen for sharing the Cas9–eScarlet fusion protein, the Hubrecht Sorting Facility, and the Utrecht Sequencing Facility, subsidized by the University Medical Center Utrecht, Hubrecht Institute and Utrecht University.
Author information
Authors and Affiliations
Contributions
A.v.O. conceived and designed the project. J.P.-M. developed the experimental protocol, with support from A.A., M.F. and C.S.B. C.S.B. performed WKM-related experiments; M.F. performed brain- and eye-related experiments; and C.S.B. and J.P.-M. performed fin-related experiments. A.A. developed the computational methods and modelling. A.A., C.S.B. and A.v.O. analysed WKM-related data; A.A. and M.F. analysed brain- and eye-related data; A.A., C.S.B. and J.P.-M. analysed fin-related data. All authors discussed and interpreted results, and wrote the manuscript. A.A., M.F., C.S.B. and J.P.-M. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks L. Zon and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 gDNA versus gDNA–mRNA detection of scars.
a, Scar percentage per cell (top bar indicates clones), and pie chart of fraction of cells per clone (colours matching histograms’ top bar) detected via gDNA (ScarTrace without step 1) and gDNA–mRNA detection (full protocol). b, Number of detected cells per clone in gDNA versus gDNA–mRNA detection and Pearson’s correlation coefficient computed using the 20 different clones identified. Dot sizes are proportional to the total number of cells found taking together the two detection strategies. c, Fraction of cells detected per clone in gDNA (green) and gDNA–mRNA (purple) detection, in which clone ‘0’ represents unscarred cells. d, Top, one-sided Fisher’s exact test on a contingency table made of the number of cells detected for the given scar clone x for each detection strategy (a and b, respectively), and the number of cells taking together all other clones (c and d, respectively) found in gDNA detection (n = 147 cells in total) and gDNA–mRNA detection (n = 128 cells in total). Bottom, heat map shows the one-sided P value of each scar clone to be enriched in the gDNA or mRNA/gDNA detection protocol. No enrichment is found with P <0.05, therefore results found for the two detection protocols are compatible. e, Normalized histograms and corresponding fit noise model function (grey line; Supplementary Information section 4) for the scar percentage detected for 1 clone found in fish P1. Scar detection efficiency is defined as the area above 5% scar content (vertical red line). Efficiency of detection of unscarred molecules is assumed to be the same as for scarred molecules. f, Normalized histogram of the scar detection efficiencies found after pooling all clones from all organs for all fish (in total, n = 371 detected clones; Extended Data Table 1). The vertical black line and the grey area indicate the mean scar detection efficiencies and s.e.m., respectively.
Extended Data Figure 2 Transcriptome analysis of the zebrafish WKM.
a, b, Scar percentage per cell for fish P2 (a) and R2 (b). The bar above each panel indicates clones and corresponding P values. c, d, Lineage trees for clones detected in P2 (c) and R2 (d) obtained as described in Fig. 2. e, t-SNE map of cells from fish R1 and R2. Colours and numbers indicate RaceID clusters. f, Heat map of the Pearson correlation between cells sorted according to RaceID clusters. Cluster numbers are indicated on the x and y axes. g, t-SNE map for the WKM of R1 and R2 coloured according to fish of origin (R1 in pink and R2 in green). All cell types intermingle well, except erythrocytes. Even though erythrocytes appear separately on the t-SNE space, they belong to the same RaceID cluster. h, t-SNE maps for R1 and R2, coloured according to the number of unique transcripts per single cell for each marker38,39,40,41. A full list of marker genes for each cell type is available in Supplementary Table 3. i, t-SNE map for fish R2 with cells coloured according to clone. j, Clonal cell fraction per cell type for fish R2.
Extended Data Figure 3 Cell types and batch effects in the brain and eyes for fish R2 and R3.
a, t-SNE map obtained with RaceID of pooled cells with a minimum of 100 total transcripts from fish R2 and R3 (isolated from the right and left eyes, right and left midbrain, forebrain and hindbrain). Different symbols indicate cells with different minimal total transcript counts. Cells are coloured according to the assigned cell type using the lowest cut off (that is, taking into account cells with at least 100 transcripts). We do not lose any cell type cluster when applying higher transcript cut offs, nor do we generate new clusters of low transcript cells when applying lower cut offs. The fraction of cells that would be termed a different cell type with a higher cut off is very low (<1%). Low transcript cells cluster mainly around the clusters formed by high transcript cells. b, c, t-SNE maps as in a, in which cells are coloured according to organ (b) and fish (c) of origin. d, t-SNE map as in a, but showing only cells from fish R3, with corresponding cell types indicated. e, t-SNE maps for fish R2 and R3 coloured according to the number of unique transcripts per single cell42,43,44,45,46,47,48,49,50,51. A full list of marker genes for each cell type is available in Supplementary Table 4.
Extended Data Figure 4 ScarTrace in the zebrafish brain and eyes.
a, Heat map of the fraction of cells per clones for cell types in fish R3 (COP, OPC and MFOL clones merged as oligodendrocytes), and two-sided Fisher’s exact test for enriched (magenta upwards triangle) and depleted (blue downwards triangle) clones per cell type with P < 0.05. The bars at the top depict organ and corresponding total number of cells. All clones have P values < 10−5. b, c, t-SNE map of fish R2 and R3 cells showing different colours for enriched clones detected in glia cells, neurons or retinal interneurons. Other cells are shown in grey. d, e, Scar percentage per cell for clones found in the WKM, forebrain, hindbrain, left and right eyes, and left and right midbrain for R2 and R3. In all panels, each colour represents the same scar (for example, the yellow scar is the same for R2 and R3), and unscarred GFP is shown in green. f, g, Heat maps of the fraction of cells per clones for each organ for R2 (f) and R3 (g). Enriched (magenta upwards triangles) and depleted (blue downwards triangles) scar clones per organ are determined by a two-sided Fisher’s exact test with P <0.05. The bar above each panel depicts the number of cells and P value for each clone. h, Histograms of the relative clone frequency in the left (blue) and right (red) midbrain and eye for R3. i, Image of dome-stage embryo injected with Cas9–eScarlet in one cell at the two-cell stage (n > 10 embryos showed similar patterns). BF, bright-field. j, Scarring efficiency shown as the percentage of unscarred GFP in S1, P1 and P2 for the left and right eyes, and the WKM. k, t-SNE map of cells isolated from the left and right eyes of S1, in which cells are coloured according to their cell type. l, Heat map of the fraction of cells per clones for each cell type in S1. Enriched (magenta upwards triangle) and depleted (blue downwards triangle) scar clones per cell type are determined from a two-sided Fisher’s exact test with P < 0.05. The bars at the top depict the total number of cells and the fraction of cells found in the right eye for each clone. All P values are below 10−5.
Extended Data Figure 5 Clones for fish P1 and P2 and lineage tree of the eyes and midbrain.
a, b, Scar percentage per cell for clones found in the WKM, forebrain, hindbrain, left and right eyes, and left and right midbrain for P1 (a) and P2 (b). Each colour represents a different scar, in which unscarred GFP is always shown in green. Colour legend per scars is different between panels (yellow scar in a is not yellow scar in b). c, d, Heat maps of the fraction of cells per clones for each organ for P1 (c) and P2 (d). Enriched (magenta upwards triangles) and depleted (blue downwards triangles) scar clones per organ are determined from a two-sided Fisher’s exact test with P <0.05. The bar above each panel depicts the number of cells and P value for each clone. e–h, Lineage trees obtained assuming the principle of maximum parsimony as described in Supplementary Information section 5 for clones detected in the right and the left eyes (e, g) and right and left midbrain (f, h) of fish P1 (e, f) and R2 (g, h). The root of the trees is set as an unscarred clone, with eight copies of the GFP transgene. In the tips there are the detected clones. The statistical confidence of each branch is computed as the proportion of each branch among 10,000 tree replicates constructed by bootstrapping scars present in all clones. To assess statistically whether clones from the left or the right side co-evolve together, we randomized the clones at the tips of the tree and checked how many times, randomly, clones from the right or the left were found to be sisters with other clones from the right or the left. This allowed us to build a distribution of co-evolution (histograms in each tree) of clones for the null hypothesis and check whether the number of times we saw clones from one side together was statistically significant or not. The vertical dashed line in each histogram indicates the number of times we see clones from one side together as sisters in the reference tree. When such line is found at the right-hand (left-hand) side of the maximum, we assume that the coevolution of the clones is enhanced (depleted). In the heat maps, we indicate the degree of co-evolution of clones in the right or the left eye or midbrain, computed as the fraction of the area of the histogram at the right- or the left-hand side (that is, enhanced or depleted co-evolution, respectively) of the vertical line divided by the corresponding area of the histogram at the right- or left-hand side of maximum of the distribution.
Extended Data Figure 6 Transcriptome analysis of the zebrafish caudal fin.
a, t-SNE maps obtained by pooling together cells from primary and regenerated fins from fish R4, R5 and R6. In each panel, single cells are coloured according to the number of unique transcripts observed for a given gene. The corresponding cell type is indicated in parenthesis52,53,54. A complete list of marker genes used for each cell type is available in Supplementary Table 5. b, t-SNE maps for the caudal fin of R4, R5 and R6 coloured based on fish (left) and fin version (right) of origin. All cells are present in all fins and fin version. No batch effects are observed.
Extended Data Figure 7 Clonal analysis in the caudal fin.
a, Scar percentage per cell in clones detected in fish R4 (left), R5 (middle) and R6 (right). The corresponding organ is indicated above each barplot (WKM or fin version). Spatial information (dorsal or ventral) is indicated when available. The bars at the top indicate clones. Each colour represent a scar, the same colour scheme is used for all panels. b, c, Heat maps of the fraction of cells per clones for each cell type and fin in fish R5 (b) and R6 (c). Enriched (magenta upwards triangle) and depleted (blue downwards triangle) scar clones per cell type per primary, secondary and tertiary fin obtained from two-sided Fisher’s exact test with P < 0.05. Top bars depict clones found in the WKM of the same fish, the corresponding number of cells, and the P value for each clone. d, t-SNE map of R6, in which cells with clone 24 (as a representative example of clones shared between mesenchymal and epidermal cells) are highlighted in red. e, t-SNE map of all cells detected in the caudal fin, in which cells from clone 19 (as a representative example of clones shared between mesenchymal and epidermal cells) in R4 are highlighted in red. f, t-SNE map of R6 primary (left) and regenerated (right) caudal fin cells (grey circles), in which cells from osteoblast clones are highlighted in red. Dashed lines represent mesenchymal cells (Fig. 4b, Extended Data Fig. 6). The percentages indicate the fraction of mesenchymal cells that share clones with osteoblasts. g, Magnified view of the t-SNE maps of R4, R5 and R6 for immune cells (dashed line on Fig. 4b). Cells are coloured based on fish (left) and fin version (right) of origin. Subpopulations of lymphoid (dashed circles) and myeloid (solid circles) are found in all fish and fin versions. h, Scar percentage for cells detected in the WKM (top) and RICs (bottom) for R4 (left), R5 (middle) and R6 (right) in the primary fins reveals the absence of common scars between the two. The bar above each panel indicates the different clones.
Extended Data Figure 8 The histone-GFP transgene and scar characterization.
a, Scheme of one copy of the h2afva:GFP transgene as previously described18. b, Copy number of the transgene. Top, average number of reads in bin sizes of 1 kb and sliding window of 200 bp obtained in whole-genome sequencing data. Bottom, copy number extracted using FREEC-11.0 with default parameters (Methods). c, Number of clones detected with a given number of scars, obtained by pooling all data from all fish used in this study. d, Scar percentage per cell in a clone in which seven different scars are detected. e, Probability density function (normalized histogram) of the fraction of scars detected in the clone depicted in d (colour code as in d). f, Average fraction of a scar per embryo computed over ten independently injected embryos (for times larger than 6 h) and over three pools of ten embryos (for times lower or equal to 6 h) detected from gDNA as a function of time for Cas9 RNA injections, for the six most observed scars (described with CIGAR strings). Error bars denote s.e.m. Solid green lines are the fit to equation (5) in Supplementary Information section 2. g, Top 100 observed scars, sorted according to their probability, and corresponding position of deletions (red) and insertions (blue) along the GFP coordinate. h, Scar probabilities as a function of sorted scars. Error bars denote s.e.m. from the fit (in f). i, Probabilities of measuring the percentages xi for three scars with copy numbers 2 (top), 1 (middle) and 1 (bottom), in which the expected percentages are fi = 50%, 25% and 25%, respectively, present in the same cell with four surviving integrations of GFP. The probability has been obtained by independently simulating 1,000 times the ScarTrace protocol for ε = 0.50 (left) and ε = 0.85 (right). Solid green lines are the fit to equation (7) in Supplementary Information section 3. j, Scar percentage for cells from the same clone made of three scars, in which each scar is represented with a different colour. k, Corresponding probability density functions (normalized histogram) for the fraction of each scar per cell (colour code as in j). Black lines denote the best fit for each scar to equation (7) (see Supplementary Information). l, Violin plot showing the distribution of measured P values obtained using a Gaussian kernel density with bandwidth determined using the Scott method55, for all clones with a given estimated NGFP. Labels indicate the number of clones observed for NGFP.
Supplementary information
Supplementary Data 1
This file summarizes the RaceID parameters used for transcriptome analysis. (PDF 37 kb)
Supplementary Data 2
This file contains a zipped file with scripts to extract extract scars and detect cones in single cells. (ZIP 74 kb)
Supplementary Data 3
The reference manual for the scripts provided in Supplementary Data 2 to extract clones. (PDF 1978 kb)
Supplementary Information
This file contains Supplementary Sections 1-4. (PDF 392 kb)
Supplementary Table 1
Cell specific barcodes used in transcriptome library preparation. (XLSX 25 kb)
Supplementary Table 2
Cell specific barcodes used in scar library preparation. (XLSX 18 kb)
Supplementary Table 3
Differently expressed genes detected in the different hematopoietic cell types. (XLSX 149 kb)
Supplementary Table 4
Differently expressed genes detected in the cell types detected in brain and eyes. (XLSX 15794 kb)
Supplementary Table 5
Differently expressed genes detected in the cell types detected in the caudal fin. (XLSX 527 kb)
Supplementary Table 6
Differently expressed genes detected in subgroups of myeloid cells. (XLSX 191 kb)
Rights and permissions
About this article
Cite this article
Alemany, A., Florescu, M., Baron, C. et al. Whole-organism clone tracing using single-cell sequencing. Nature 556, 108–112 (2018). https://doi.org/10.1038/nature25969
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25969
- Springer Nature Limited
This article is cited by
-
The implications of single-cell RNA-seq analysis in prostate cancer: unraveling tumor heterogeneity, therapeutic implications and pathways towards personalized therapy
Military Medical Research (2024)
-
Gene-expression memory-based prediction of cell lineages from scRNA-seq datasets
Nature Communications (2024)
-
Single-cell lineage capture across genomic modalities with CellTag-multi reveals fate-specific gene regulatory changes
Nature Biotechnology (2024)
-
Dissecting the spatiotemporal diversity of adult neural stem cells
Molecular Systems Biology (2024)
-
PhyloVelo enhances transcriptomic velocity field mapping using monotonically expressed genes
Nature Biotechnology (2024)