Abstract
All haematopoietic cell lineages that circulate in the blood of adult mammals derive from multipotent haematopoietic stem cells (HSCs)1. By contrast, in the blood of mammalian embryos, lineage-restricted progenitors arise first, independently of HSCs, which only emerge later in gestation2,3. As best defined in the mouse, ‘primitive’ progenitors first appear in the yolk sac at 7.5 days post-coitum2,3. Subsequently, erythroid–myeloid progenitors that express fetal haemoglobin4, as well as fetal lymphoid progenitors5, develop in the yolk sac and the embryo proper, but these cells lack HSC potential. Ultimately, ‘definitive’ HSCs with long-term, multilineage potential and the ability to engraft irradiated adults emerge at 10.5 days post-coitum from arterial endothelium in the aorta-gonad-mesonephros and other haemogenic vasculature3. The molecular mechanisms of this reverse progression of haematopoietic ontogeny remain unexplained. We hypothesized that the definitive haematopoietic program might be actively repressed in early embryogenesis through epigenetic silencing6, and that alleviating this repression would elicit multipotency in otherwise lineage-restricted haematopoietic progenitors. Here we show that reduced expression of the Polycomb group protein EZH1 enhances multi-lymphoid output from human pluripotent stem cells. In addition, Ezh1 deficiency in mouse embryos results in precocious emergence of functional definitive HSCs in vivo. Thus, we identify EZH1 as a repressor of haematopoietic multipotency in the early mammalian embryo.
Similar content being viewed by others
Main
The differentiation of pluripotent stem cells to haematopoietic lineages generates robust erythroid–myeloid lineage-restricted progenitors but not HSCs. This pattern bears marked similarities to early haematopoietic ontogeny. We hypothesized that the same epigenetic factors actively repress multipotency in embryogenesis and differentiation from pluripotent stem cells. To identify these factors, we adopted a loss-of-function screen using lentivirally delivered short hairpin RNAs (shRNAs) that target 20 DNA- and histone-modifying factors (Extended Data Fig. 1a, Supplementary Table 1). Erythroid–myeloid progenitors differentiated from human pluripotent stem cells marked by CD34 and CD45 were expanded with five transcription factors (5F). They retained embryonic features, including lack of lymphoid potential7, and this enabled us to screen for reactivation of lymphoid potential as a measure of multipotency. 5F cells were transduced with individual shRNAs and screened for T cell potential on OP9-DL1 stromal cells (Fig. 1a). The knockdown of six factors independently enhanced CD4+CD8+ T cell potential from 5F cells (Fig. 1b, Extended Data Fig. 1b).
a, Scheme for human pluripotent stem-cell differentiation into haematopoietic progenitors. CD34+ cells were transduced with the five transcription factors (5F) HOXA9, ERG, RORA, SOX4 and MYB. 5F cells were then transduced with individual shRNAs (×4 each) that targeted each epigenetic modifier and seeded onto OP9-DL1 stromal cells to induce T cell differentiation. Dox, doxycycline; EB, embryoid body. b, Strictly standardized mean difference (SSMD) of CD4+CD8+ T cell frequencies across all four shRNAs targeting each epigenetic modifier in 5F cells using two iPS cell lines, CD45-iPS and MSC-iPS1, in two independent experiments. MT, methyltransferase. c, Prospective analysis of T and B cell frequencies from 5F cells plus shRNA targeting top candidates (n = 2 biological replicates). d, Flow analysis of CD4+CD8+ T cell development of 5F cells with shRNAs targeting luciferase (shLUC) or EZH1 (shEZH1) after 5 weeks of differentiation on OP9-DL1 stromal cells. e, Flow analysis of CD19+ B cell potential. f, Quantification of T cell potential of 5F plus shEZH1 cells compared to 5F plus shLUC cells pooled across two hairpins and five independent experiments (n = 10) using several iPS cell lines (CD34-iPS, CD45-iPS and MSC-iPS1). Individual values obtained for each hairpin are shown in the Source Data. ***P = 0.001 by unpaired two-tailed t-test. g, Quantification of colony-forming potential in three independent experiments. E, erythroid; GM, granulocyte, monocyte; M, monocyte; G, granulocyte; GEMM, granulocyte, erythroid, monocyte, megakaryocyte. h, i, Flow analysis of myeloid (CD11b+) (h) and erythroid (CD71+GLYA+) (i) potential. Experiments replicated at least twice. Data are mean ± s.e.m.
Prospective validation revealed that only EZH1 knockdown (using shEZH1) elicited robust T (16.3 ± 7.4%; mean ± s.e.m.) and B (22.5 ± 7.3%) cell potential (Fig. 1c–e), compared to shRNAs targeting a control luciferase gene (shLUC) (T cell 0.002 ± 0.002%; B cell 0.022 ± 0.006%) across multiple induced pluripotent stem (iPS) cell lines (Fig. 1f). EZH1-deficient cells retained erythroid–myeloid potential as shown by colony-forming assays (Fig. 1g) and flow cytometry (Fig. 1h, i). EZH1 knockdown also promoted lymphoid potential independently of the five transcription factors, as evidenced by robust T cell differentiation from naive CD34+ haemogenic endothelial cells (26.1 ± 16.5% shEZH1 versus 2.3 ± 0.4% shLUC) (Extended Data Fig. 1c). Further characterization was not possible owing to the limited proliferation of pluripotent stem and haemogenic endothelial cells. By contrast, 5F cells expanded exponentially (Extended Data Fig. 1d) and showed increased CD34+ progenitors after shEZH1 transduction (78.8 ± 14.2% versus 29.3 ± 10.0%) (Extended Data Fig. 1e). Taken together, these data show that EZH1 knockdown activates multipotency in lineage-restricted embryonic haematopoietic progenitors.
EZH1 is a component of the Polycomb repressive complex 2 (PRC2), which mediates epigenetic silencing of genes via methylation of lysine residue 27 of histone H38. To dissect the role of PRC2 in repressing haematopoietic multipotency, we assessed T cell differentiation upon depletion of each PRC2 subunit. In addition to EZH1, SUZ12 knockdown also enhanced T cell potential, albeit to a lesser extent. By contrast, knockdown of EED or EZH2 had no effect on T cell potential and dual EZH1 and EZH2 knockdown phenocopied that of EZH2 depletion (Fig. 2a, b). To determine whether the catalytic SET domain was required, we overexpressed full-length mouse Ezh1 or mutant Ezh1 lacking the SET domain (mEzh1ΔSET) (Fig. 2c). Overexpression of mouse Ezh1 completely abrogated T cell potential in shEZH1 cells, whereas the mutant mEzh1ΔSET did not (Fig. 2c, d, Extended Data Fig. 2d–g). Furthermore, overexpression of mouse Ezh2 failed to suppress T cell potential, despite the remarkable homology of the SET domains (Extended Data Fig. 2e, h, i). These data show that specific inhibition of EZH1, rather than antagonism of canonical PRC2, unlocks lymphoid potential and the catalytic SET domain is required for this function.
a, Representative flow plots of T cell potential of 5F cells with shRNAs targeting individual components of PRC2. b, Quantification of T cell potential of 5F cells plus shRNA targeting the indicated subunit in a, shown as using two hairpins across two independent experiments (n = 4). *P = 0.0457, **P = 0.0061 by unpaired two-tailed t-test. c, Representative flow analysis of T cell potential in 5F cells plus shEZH1, with co-expression of full-length mouse Ezh1 (mEzh1) or mutant mouse Ezh1 lacking the SET domain (mEzh1ΔSET, +ΔSET). d, Quantification of flow analysis in c (n = 3 biological replicates). *P = 0.0146, **P = 0.0011 by one-way ANOVA. All plots are gated on CD45+. Data are pooled across two independent experiments. Data are mean ± s.e.m. NS, not significant.
To understand the molecular changes upon EZH1 knockdown, we performed RNA sequencing (RNA-seq), assay for transposase-accessible chromatin using sequencing (ATAC–seq) and chromatin immunoprecipitation followed by sequencing (ChIP–seq). Upregulated genes after EZH1 knockdown were enriched for biological processes such as defence response (P = 6.8 × 10−9), immune response (P = 1.2 × 10−7) and T cell co-stimulation (P = 0.03) (Fig. 3a, b). Human haematopoieteic gene signatures9, such as of HSCs (stem), multi-lymphoid progenitors (MLP) (early lymphoid) and ProB, were highly enriched in shEZH1 cells, consistent with stem and lymphoid potential (Fig. 3c). We also performed RNA-seq and ATAC–seq on emergent haematopoietic stem and progenitor cells (HSPCs) at 10.5 days post-coitum10,11,12 from the yolk sac and aorta-gonad-mesonephros (AGM) of wild-type, Ezh1+/− and Ezh1−/− mouse embryos (Fig. 4a). Interestingly, in wild-type embryos, the expression of Ezh1 was lower in the AGM than in the yolk sac, whereas Ezh2 and Eed were higher in the AGM (Fig. 4b). Notably, Ezh1 deficiency in vivo also induced genes enriched for angiogenesis, haematopoietic/lymphoid development and immune system processes (Extended Data Fig. 3a–d).
a, Heat map of upregulated (104) and downregulated (49) genes (>2-fold; Benjamini–Hochberg corrected t-test, P < 0.1) from RNA-seq analysis of CD34+CD38− HSPCs 5F plus shEZH1 cells (n = 10 biological replicates) compared to 5F plus shLUC cells (n = 8 biological replicates). b, Gene Ontology (GO) analysis of biological processes associated with significantly upregulated genes in a, subdivided by GO hierarchical categories with P values labelled along the radius. c, Enrichment of human HSC and progenitor signatures by gene set enrichment analysis (GSEA) in 5F plus shEZH1 compared with 5F plus shLUC cells, overlaid on the map of human HSPC hierarchy. CMP, common myeloid progenitor; MEP, megakaryocyte–erythroid progenitor; GMP, granulocyte–monocyte progenitor; ETP, early thymic progenitor; NES, normalized enrichment score. d, Density map of upregulated and downregulated ATAC peaks by MAnorm29 in 5F plus shEZH1 compared to 5F plus shLUC cells (n = 2 biological replicates). e, GO terms of enriched biological processes of ATAC peaks in d by GREAT analysis30. f, Tracks of representative genes that acquire a significant ATAC peak upon EZH1 knockdown. g, ChIP–seq density map of EZH1 peaks within bivalent (B), repressed (R), active (A) or null (N) promoter groups (n = 2 biological replicates). K4, H3K4me4; K27, H3K27me3. h, Waterfall plot of CellNet31 predicted regulators of EZH1-bound bivalent gene networks. TF, transcription factor. i, Sitepro quantitative analysis32 of H3K27me3 levels at all upregulated genes around the transcription start site (TSS) upon EZH1 knockdown, relative to shLUC (n = 2 biological replicates). j, Left, Sankey diagram illustrating histone methylation changes of all bivalent genes in shLUC control cells and after EZH1 knockdown (n = 2 biological replicates). Right, genes that lose H3K27me3 (become activated) are specifically enriched in the HSC signature, whereas bivalent genes that are unchanged or inactivated are enriched in the ProB signature by Fisher’s exact test. k, ChIP–seq tracks of EZH1, H3K4me3 and H3K27me3 at representative HSC promoter regions in shLUC and shEZH1 cells. Experiments replicated at least twice.
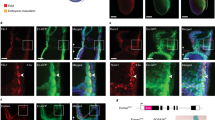
a, Representative images of E9.5 and E10.5 embryos (n > 50 embryos). b, Quantitative PCR (qPCR) of each PRC2 subunit in E10.5 wild-type yolk sac (YS) and AGM (n = 3 biological replicates). *P = 0.0439, ****P < 0.0001 by unpaired two-tailed t-test. Data are mean ± s.e.m. BM, bone marrow. c, ATAC density map of c-Kit+VE-cadherin+CD45+ HSPCs sorted from 30 pooled embryos of E10.5 wild-type (WT) and Ezh1+/− AGM. d, Significantly upregulated ATAC peaks were compared to HSPC, T and B cell networks and signatures of the human HSPC hierarchy45. *P < 0.05 by Fisher’s exact test. e, Left, engraftment of E10.5 AGM (3.5 ee) in sublethally irradiated adult NSG females. Donor chimaerism marked by CD45.2+ was measured in peripheral blood every 4 weeks up to 16 weeks post-transplantation. Each dot represents a single transplant recipient; lines denote mean values. Right, lineage distribution of engrafted mice showing T cell (T), B cell (B), and myeloid (M) contribution. f, Left, engraftment of E10.5 yolk sac (5 ee). Right, lineage distribution of engrafted mice. g, Left, engraftment of E9.5 PSP (10 ee). Right, lineage distribution of engrafted mice. h, Left, serial transplantation of whole bone marrow from primary recipients of E10.5 AGM cells in e. Secondary transplant (2°) was carried out after 24 weeks of primary transplant. Right, lineage distribution of engrafted mice (n ≥ 3 mice per group). *P < 0.05, ** P < 0.01, ***P < 0.0001 by unpaired two-tailed t-test. See Supplementary Information for exact P values per time point. Data are pooled across four (e–g) or three (h) independent experiments; experiment in c was performed once.
Regions of increased chromatin accessibility (1,610 ATAC peaks) in shEZH1 cells exhibited concomitantly increased gene expression upon EZH1 knockdown and were associated with T cell development and lymphocyte activation pathways, as well as HSC, HSC/MLP, B and T cell signatures (Fig. 3d, e, Extended Data Fig. 3e–g). EZH1 knockdown also increased accessibility to HSC/lymphoid transcription factors, such as HLF, FOXO1 and ARID5B13,14,15 (Fig. 3f). Downregulated peaks were enriched for alternative developmental processes and importantly, embryonic haematopoiesis (Fig. 3e, Extended Data Fig. 3e). In vivo, upregulated ATAC peaks in Ezh1-deficient AGM cells were enriched for immune response, T cell activation, lymphocyte differentiation pathways, as well as HSC and HSC/MLP signatures (Fig. 4a, c, d, Extended Data Fig. 3h, i); furthermore, Ezh1 deficiency increased accessibility to target genes of master haematopoietic transcription factors, including Runx1 (Extended Data Fig. 3k, l).
We hypothesized that these molecular changes upon EZH1 knockdown were mediated by bivalent, or poised, chromatin domains, often implicated in the control of developmentally regulated genes16. Consistent with previous reports, EZH1 was broadly associated with repressive (H3K27me3), bivalent (H3K27me3 and H3K4me3) and active (H3K4me3) histone methylation marks17,18 (Fig. 3gExtended Data Fig. 4a). Although active genes were associated with housekeeping functions (Extended Data Fig. 4b), EZH1-bound bivalent and repressed genes were enriched for developmental and morphogenic processes (Extended Data Fig. 4c, d). EZH1 knockdown increased the expression of bivalent genes, which were associated with HSC and early lymphoid lineages (Extended Data Fig. 4e, f). These genes included the targets of HSC transcription factors such as RUNX1T1 and SOX17, and NOTCH factors HES1, HEY1 and FOXC219 (Fig. 3h). EZH1 directly bound the promoters of HSC and ProB transcription factors including HLF, PRDM16, LMO2, ETS1, MEIS1, RUNX1 and HOX clusters (Extended Data Fig. 4e). We also observed a global reciprocal relationship between H3K27me3 and gene transcription (Fig. 3i, Extended Data Fig. 4g–k), with poised HSC genes exhibiting loss of H3K27me3 and increased expression upon EZH1 knockdown (Extended Data Fig. 4h, i). In total, 27 out of 29 of these activated HSC genes are direct targets of EZH1, including HOPX, HLF, MEIS1 and HES1 (P = 7.8 × 10−5; Fig. 3j, k).
EZH2 also bound activated HSC genes, consistent with its ability to target the same regions8 (Extended Data Fig. 4l); however, recent analysis of SET domain-swapping revealed context-specific sensitivity to an EZH2-specific inhibitor, further suggesting that although EZH1 and EZH2 can bind a common subset of HSC targets, these enzymes are likely to have distinct functions on chromatin20. Concordant with our observation that SUZ12 knockdown partially phenocopies EZH1 loss (Fig. 2a, b), we observed specific enrichment of EZH1 and SUZ12 at activated HSC and ProB genes, consistent with non-canonical targets of the EZH1–SUZ12 complex17 (Extended Data Fig. 4m–q). Similarly, upregulated ATAC peaks in Ezh1-deficient AGM were also enriched for SUZ12 binding, but not EZH2, indicating a conserved role for non-canonical PRC2 regulation in vivo (Extended Data Fig. 4r). These data suggest that in addition to the canonical function of EZH1–PRC2 in mediating H3K27me3 changes at poised HSC loci, EZH1 also regulates ProB genes through a complementary non-canonical EZH1–SUZ12 complex, highlighting an EZH1-specific function that is not phenocopied by EZH2.
The emergence of bona fide HSCs, defined by the capacity to repopulate irradiated adult recipients, marks the transition from embryonic to definitive haematopoiesis. We isolated AGM and yolk sac from embryonic day (E)10.5 wild-type, Ezh1+/− and Ezh1−/− embryos and transplanted adult non-obese diabetic (NOD)/severe combined immunodeficiency (SCID)/Il2rg−/− (NSG) recipients (Fig. 4a). We detected peripheral blood reconstitution from wild-type AGM in 3 out of 7 mice (11.9 ± 7.9%) at 4 weeks, but chimaerism decreased by 16 weeks (2 out of 7, 12.2 ± 8.1%); this corresponds to 1 repopulating unit in approximately 10.4 embryo equivalents (ee), consistent with HSCs being exceedingly rare at E10.510,21. By contrast, 5 out of 8 mice transplanted with Ezh1−/− AGM cells were engrafted at 4 weeks (39.2 ± 9.4%) and stabilized at 16 weeks (34.6 ± 14.6%). Notably, Ezh1+/− AGM transplant recipients had the highest initial chimaerism (41.2 ± 16.3%; 4 out of 5), which increased by 16 weeks (68.9 ± 17.8%), and was predominantly multilineage (3 out of 5) (Fig. 4e, Extended Data Fig. 5a, c). This corresponds to 1 repopulating unit in 3.6 Ezh1−/− and 2.2 Ezh1+/− ee, or an approximately fivefold increase in HSC frequency compared to wild type.
At E10.5, the yolk sac is thought to contain few, if any, HSCs21. We detected low-level engraftment of wild-type yolk sac cells in 5 out of 9 recipients at 4 weeks (3.4 ± 0.7%), and in 3 out of 9 mice at 16 weeks (4.3 ± 1.6%). Most Ezh1−/− (4.5 ± 0.9%, 6 out of 7 engrafted) and all of the Ezh1+/− yolk-sac-transplanted mice (5.4 ± 1.4%, 5 out of 5 engrafted) showed stable long-term engraftment at 16 weeks. The number of repopulating units calculated was similar to that of the AGM (about 1 in 12.3 ee wild-type mice; 1 in 2.6 ee Ezh1−/− mice, 1 in <2 ee Ezh1+/− mice). All engrafted mice were multilineage (Fig. 4f, Extended Data Fig. 5a, c). Importantly, up to 75% of peritoneal B cells in Ezh1+/− AGM-engrafted mice were of the adult-like B-2 phenotype, as opposed to the embryonic B-1 cells (Extended Data Fig. 6a). Moreover, up to 95% of donor-derived CD45.2+CD3+ T cells expressed adult-type TCRβ, as opposed to embryonic TCRγδ, in Ezh1−/− and Ezh1+/− AGM- and yolk-sac-engrafted mice (Extended Data Fig. 6b). These data provide compelling evidence that Ezh1 deficiency, and in particular haploinsufficiency, stimulates generation of definitive HSCs and adult-like lymphopoiesis.
The para-aortic splanchnopleura (PSP) at E9.5 lacks HSCs as determined by transplantation studies3. Transplantation of E9.5 wild-type PSP cells (Fig. 4a) failed to engraft adult recipients (0 out of 5)21,22; by contrast, we detected chimaerism in recipients of Ezh1−/− (3 out of 3, 1.6 ± 0.3%) and Ezh1+/− (4 out of 6 mice, 3.6 ± 1.3%) PSP at 4 weeks post-transplantation (Fig. 4g, Extended Data Fig. 5b). By 16 weeks, chimaerism increased in Ezh1−/− (3 out of 3, 9.4 ± 5.1%) and Ezh1+/− (5 out of 6, 13.1 ± 9.5%) recipients, and grafts were fully multilineage (Extended Data Fig. 5c). Thus, Ezh1 deficiency stimulates precocious generation of bona fide HSCs during embryogenesis.
To assess the self-renewal capacity of Ezh1-deficient HSCs, we performed secondary transplantation. No mice showed engraftment with E10.5 wild-type AGM (0 out of 4) or yolk sac (0 out of 7). By contrast, 4 out of 7 Ezh1−/− (4.4 ± 0.5%) and 9 out of 9 Ezh1+/− (57.8 ± 10.2%) AGM-derived secondary recipients were engrafted (Fig. 4h, Extended Data Fig. 5d). Of note, although no Ezh1−/− yolk sac recipients (0 out of 10) were engrafted, we observed secondary chimaerism from Ezh1+/− yolk sac cells (5 out of 7, 1.5 ± 0.3%), which increased by 16 weeks (6 out of 7, 5.1 ± 1.9%) (Extended Data Fig. 5d, e). All engrafted secondary recipients were multilineage, with no evidence of leukaemic transformation (Fig. 4h, Extended Data Fig. 5c, e). Taken together, these data indicate that genetic Ezh1 deficiency elicits precocious emergence of bona fide HSCs in vivo.
It has long been a curiosity that haematopoietic ontogeny progresses in reverse order, with haematopoietic progenitors appearing first in embryonic development independently of HSCs2,3. We propose that EZH1 represses definitive loci in primitive blood progenitors differentiated from human pluripotent stem cells and in mouse embryos, which precludes precocious HSC emergence during gestation. EZH1 deficiency promotes multipotency in lineage-restricted blood progenitors and enables precocious emergence of HSCs. Although PRC2 is a well-characterized HSC regulator, our data contribute compelling evidence for the distinct molecular functions of EZH1 and EZH2, and suggest a putative role for non-canonical PRC2, involving EZH1 and SUZ12. Homozygous loss of Suz12 in mice impairs HSC function and lymphopoiesis, but heterozygosity for Suz12 or Eed enhances HSC self-renewal23,24. Consistent with this, our data reinforce the concept that HSCs are exquisitely sensitive to PRC2 dosage, with partial reduction or increase affecting function23,24,25,26. Interestingly, Runx1 haploinsufficiency also promotes premature HSC generation27. Our data unify these observations; EZH1 marks many transcription factor-binding sites, whereas Ezh1 deficiency enhances accessibility to targets of key HSC transcription factors, including Runx1, to promote HSC emergence (Extended Data Fig. 3k, l). We identify Ezh1 as a molecular regulator of lineage-restricted potential of the first blood progenitors in the mammalian embryo, which accounts in part for why early embryonic progenitors lack multipotency. Beyond developmental implications, our findings suggest that resolution of EZH1-marked domains may be essential for physiological specification of HSCs from pluripotent stem cells, as a complementary approach to the synthetic reactivation of stem-cell programs by HSC transcription factors7,28.
Methods
A step-by step protocol can be found at the Protocol Exchange33.
Human iPS cell culture
All experiments were performed using MSC-iPS134, CD34-iPS and CD45-iPS cells, obtained from the Boston Children’s Hospital Human Embryonic Stem Cell Core (hESC) and verified by immunohistochemistry for pluripotency markers, teratoma formation and karyotyping. All cells were routinely tested for mycoplasma contamination. Human iPS cells were maintained on mouse embryonic fibroblast (GlobalStem) feeders in DMEM/F12 plus 20% KnockOut-Serum Replacement (Invitrogen), 1 mM l-glutamine, 1 mM non-essential amino acids (NEAA), 0.1 mM β-mercaptoethanol and 10 ng ml−1 bFGF. Medium was changed daily, and cells were passaged 1:4 onto fresh feeders every 7 days using standard clump passaging with collagenase IV.
Embryoid body differentiation
Differentiation of embryoid bodies was performed as previously described35. In brief, human pluripotent stem cell colonies were scraped into non-adherent rotating 10 cm plates at the ratio of 2:1. Embryoid body medium was KO-DMEM plus 20% FBS (Stem Cell Technologies), 1 mM l-glutamine, 1 mM NEAA, 1% penicillin–streptomycin, 0.1 mM β-mercaptoethanol, 200 μg ml−1 human transferrin and 50 μg ml−1 ascorbic acid. After 24 h, medium was changed by allowing embryoid bodies to settle by gravity, and replaced with embryoid body medium supplemented with growth factors: 50 ng ml−1 BMP4 (R&D Systems), 200 ng ml−1 SCF, 200 ng ml−1 FLT3, 50 ng ml−1 G-CSF, 20 ng ml−1 IL-6 and 10 ng ml−1 IL-3 (all Peprotech). Medium was changed on days 5 and 10. Embryoid bodies were dissociated on day 14 by digesting with collagenase B (Roche) for 2 h, followed by treatment with enzyme-free dissociation buffer (Gibco), and filtered through an 80-μm filter. Dissociated embryoid bodies were frozen in 10% DMSO, 40% FBS freezing solution.
Progenitor sorting
Dissociated embryoid body cells were thawed following the Lonza Poietics protocol and resuspended at 1 × 106 per 100 μl staining buffer (PBS plus 2% FBS). CD34+ cells were sorted from bulk embryoid body culture using human CD34 microbeads (Miltenyi Biotec) and run through a magnetic column separator (MACS) as per the manufacturer’s instructions.
Lentiviral and shRNA library plasmids
The 5F lentiviral plasmids HOXA9, ERG, RORA, SOX4 and MYB were cloned into pInducer-21 doxycycline-inducible lentiviral vector. The shRNA library targeting 20 epigenetic modifiers36 was obtained from the Broad Institute RNAi Consortium in pLKO.1 or pLKO.5 lentiviral vectors. Lentiviral particles were produced by transfecting 293T-17 cells (ATCC) with the lentiviral plasmids and third-generation packaging plasmids. Viruses were harvested 24 h after transfection and concentrated by ultracentrifugation at 64,965g for 3 h using the Beckman Coulter SW 32 Ti rotor. All viruses were titred by serial dilution on 293T cells.
5F gene transfer and 5F culture
MACS-separated CD34+ embryoid body progenitors were seeded on retronectin-coated (10 μg cm−2) 96-well plates at a density of 2 × 104–5 × 104 cells per well. The infection medium was SFEM (StemCell Technologies) with 50 ng ml−1 SCF, 50 ng ml−1 FLT3, 50 ng ml−1 TPO (all R&D Systems), 50 ng ml−1 IL-6 and 10 ng ml−1 IL-3 (both from Peprotech). Lentiviral infections were carried out in a total volume of 150 μl. The multiplicity of infection (MOI) for each factor was as follows: ERG MOI = 5, HOXA9 MOI = 5, RORA MOI = 3, SOX4 MOI = 3, MYB MOI = 3, and MOI = 2 for shRNA. Virus was concentrated onto cells by centrifuging the plate at 924g for 30 min at room temperature. Infections were carried out for 24 h. After gene transfer, 5F cells were cultured in SFEM with 50 ng ml−1 SCF, 50 ng ml−1 FLT3, 50 ng ml−1 TPO, 50 ng ml−1 (all R&D Systems) IL-6, and 10 ng ml−1 IL-3 (Peprotech). Doxycycline (Dox) was added at 2 μg ml−1 (Sigma). Puromycin was added at 0.3 μg ml−1 (ThermoFisher Scientific). Cultures were maintained at a density of <1 × 106 cells ml−1, and the medium was changed every 3–4 days.
T cell differentiation
After 14 days of respecification, 1 × 105 5F cells were plated in OP9-DL1 stromal co-culture37. Cells were cultured in α-MEM (Gibco), 1% penicillin–streptomycin, 20% FBS (Gemini), and 1 mM l-glutamine with 30 ng ml−1 SCF, 5 ng ml−1 FLT3, 5 ng ml−1 IL-7 (all R&D Systems) for 20 days with 2 μg ml−1 Dox followed by Dox removal. Cells were collected by mechanical dissociation and filtered through a 40-μm filter and passaged onto fresh stroma every 5–7 days. T cell development was assessed after 35 days using CD45, CD7, CD3, CD4 and CD8.
B cell differentiation
After 14 days of respecification, 5 × 104 5F cells were plated into a single well of MS-5 stroma in a 6-well NUNC plate. Cells were cultured in Myelocult H5100 (Stem Cell Technologies) supplemented with 50 ng ml−1 SCF, 10 ng ml−1 FLT3, 25 ng ml−1 IL7, 25 ng ml−1 TPO (all R&D Systems) and 1% penicillin–streptomycin for 10 days with 2 μg ml−1 Dox followed by Dox removal.
Colony assays
After 14 days of respecification, 5 × 104 cells were plated into 3 ml of complete methylcellulose H3434 (StemCell Technologies) supplemented with 10 ng ml−1 IL-6 (Peprotech), 10 ng ml−1 FLT3 (R&D) and 50 ng ml−1 TPO (R&D) without 2 μg ml−1 Dox. The mixture was distributed into two 60-mm dishes and maintained in a humidified chamber for 14 days.
Mouse transplantation
NOD/SCID/Il2rg−/− (NSG) (Jackson Laboratory) mice were bred and housed at the Boston Children’s Hospital animal care facility. Animal experiments were performed in accordance with institutional guidelines approved by Boston Children’s Hospital Animal Care Committee. At least three animals were used per cohort, based on previous transplantation studies. Mice were assigned randomly to groups and blinding was not used. In brief, 8–12-week-old mice were irradiated (2.75 Gy) 24 h before transplant. To ensure consistency between experiments, only female mice were used. Sublethally irradiated adult NSG females were transplanted intravenously with 3.5 ee of whole E10.5 AGM, 5 ee of whole E10.5 yolk sac or 10 ee of whole E9.5 PSP. Mice were bled retroorbitally every 4 weeks to monitor donor chimaerism up to 16 weeks post-transplantation. Twenty-four weeks after primary transplantation, primary recipients from each group were euthanized and 4 × 106 whole bone marrow cells were transplanted into 1–3 secondary recipients. Cells were transplanted in a 200 μl volume using a 28.5-gauge insulin needle. Sulfatrim was administered in drinking water to prevent infections after irradiation. Data points were combined from all independent experiments and outliers were not excluded.
Flow cytometry
The following antibodies were used for human cells: CD45 allophycocyanin (APC)-conjugated Cy7 (557833, BD Biosciences), CD4 phycoerythrin (PE)-conjugated Cy5 (IM2636U, Beckman Coulter Immunotech), CD8–BV421 (RPA-T8, BD Horizon), CD5–BV510 (UCHT2, BD Biosciences), TCRγδ–APC (555718, BD Biosciences), TCRαβ–BV510 (T10B9.1A-31, BD Biosciences), CD3–PE–Cy7 (UCHT1, BD Pharmigen), CD7–PE (555361, BD Pharmigen), CD1a–APC (559775, BD Pharmigen) for T cell staining. For B cell staining: CD45–PE–Cy5 (IM2652U, Beckman Coulter Immunotech), CD19–PE (4G7, BD Biosciences), CD56–V450 (B159, BD Biosciences), CD11b–APC–Cy7 (557754, BD Biosciences), For HSC/progenitor sorting: CD34–PE–Cy7 (8G12, BD Biosciences), CD45–APC–Cy7 (557833, BD Biosciences), CD38–PE–Cy5 (IM2651U, Beckman Coulter) and DAPI. For myeloid and erythroid staining: CD11b–APC–Cy7 (557754, BD Biosciences), GLYA–PE–Cy7 (A71564, Beckman Coulter), CD71–PE (555537, BD Biosciences), CD45–PE–Cy5 (IM2652U, Beckman Coulter Immunotech). All staining was performed with <1 × 106 cells per 100 μl staining buffer (PBS plus 2% FBS), with a 1:100 dilution of each antibody, for 30 min at room temperature in the dark. Compensation was performed by automated compensation with anti-mouse Igκ and negative beads (BD Biosciences). All acquisitions were performed on a BD Fortessa or BD Aria cytometer.
The following antibodies were used for mouse cells: CD45.2–PE–Cy7 (104, eBioscience), CD45.1–FITC (A20, eBioscience), B220–PB (RA3-6B2, BD Biosciences), Ter119–PE–Cy5 (Ter 119, eBioscience), GR1 (RB6-8C5, BD Bioscience), CD3–APC (145-2C11, eBioscience), CD19–APC–Cy7 (1D3, BD Bioscience), MAC1–AF700 (M1/70, BD Bioscience) for engraftment analyses. For B cell staining: CD45.2–APC–Cy7 (104, BioLegend), CD23–PE–Cy7 (B3B4, eBioscience), Ter119–PE–Cy5 (Ter 119, eBioscience), MAC1–A700 (M1/70, BD Bioscience), CD5–BV510 (53-7.3, BD Biosciences), IgM–eFluor660 (II/41, eBioscience). For T cell staining: CD45.2–PE–Cy7 (104, eBiosciences), TCRβ–PE–Cy5 (H57-597, BD Biosciences), CD8–APC–EF780 (53-6.7, eBioscience), CD4–APC (GK1.5, eBioscience), CD3–AF700 (17A2, BioLegend), TCRγδ–FITC (GL3, BD Biosciences). For HSPC sorting: CD16/32 (93, Biolegend), Ter119–biotin (Ter119, eBioscience), Gr-1–biotin (RB6-8C5, eBioscience), CD3–biotin (17A2, eBioscience), CD5–biotin (53-7.3, eBioscience), CD8–biotin (53-6.7, eBioscience), CD19–biotin (eBio1D3, eBioscience), streptavidin–eFluor450 (eBioscience), CD45–PerCP-Cy5.5 (30-F11, eBioscience), CD144–eFluor660 (eBioBV13, eBioscience), CD117–APC–eFluor 780 (2B8, eBioscience), CD41–PE–Cy7 (eBioMWReg30, eBioscience). All staining was performed with <1 × 106 cells per 100 μl staining buffer (PBS plus 2% FBS), with a 1:100 dilution of each antibody, for 30 min on ice in the dark. Compensation was performed by automated compensation with anti-rat and anti-hamster Igκ and negative beads (BD Biosciences). All acquisitions were performed on a BD Fortessa or BD Aria cytometer.
RNA-seq
Human cells were stained and sorted using CD34–PE–Cy7 (8G12, BD Biosciences), CD38–PE–Cy5 (IM2651U, Beckman Coulter) and DAPI (Beckman Coulter). RNA-seq libraries were prepared using the NEB Ultra (PolyA) kit as per the manufacturer’s protocol with 50 ng input RNA. Mouse cells were stained and sorted using the ‘HSPC stain’ (see ‘Flow cytometry’). RNA-seq libraries were prepared using the Clontech SMARTer Universal Low Input kit as per the manufacturer’s protocol with 10 ng input RNA. Libraries were sequenced using the 200 cycle paired-end kit on the Illumina HiSeq2500 system. RNA-seq reads were analysed with Tuxedo Tools following a standard protocol38. Reads were mapped with TopHat version 2.1.0 and Bowtie2 version 2.2.4 with default parameters against build hg19 of the human genome, and build hg19 of the RefSeq human genome annotation. Samples were quantified with the Cufflinks package version 2.2.1. Differential expression was performed using Cuffdiff with default parameters.
ATAC–seq
ATAC–seq was performed as previously described39. 5 × 103–50 × 103 cells were used for each tagmentation using Tn5 transposases. The resulting DNA was isolated, quantified and sequenced on an Illumina NextSeq500 system. The raw reads were aligned to the human genome assembly hg19 using Bowtie40 with the default parameters, and only tags that uniquely mapped to the genome were used for further analysis. ATAC peaks were identified using MACS41.
ChIP–seq
ChIP experiments were performed as previously described42 using the antibodies for H3K4me3 (04-745, Millipore) and H3K27me3 (07-449, Millipore) in 5F cells. For bioChIP analysis of EZH1 or EZH2 occupancy, Flag–biotin-tagged EZH1 or EZH2 was stably expressed in 5F cells. The chromatin was isolated and immunoprecipiated with streptavidin Dynabeads (Life Technologies) as previously described43. ChIP–seq libraries were generated using NEBNext ChIP–seq Library Prep Master Mix following the manufacturer’s protocol (New England Biolabs), and sequenced on an Illumina NextSeq500 system. ChIP–seq raw reads were aligned to the human genome assembly hg19 using Bowtie40 with the default parameters; only tags that uniquely mapped to the genome were used for further analysis. ChIP–seq peaks were identified using MACS41.
Bioinformatics and statistical analysis
All statistical calculations were performed using GraphPad Prism. Tests between two groups used a two-tailed unpaired Student’s t-test. Data are presented as mean ± s.e.m. Where indicated, ANOVA was used, with P < 0.05 considered significant. GSEA and GO were run according to default parameters in their native implementations. Statistical enrichment of gene lists was performed using Fisher’s exact test. No statistical methods were used to predetermine sample size.
Data availability
All RNA-seq, ATAC–seq and ChIP–seq data have been deposited to the Gene Expression Omnibus (GEO) database under the accession number GSE89418.
Accession codes
References
Orkin, S. H. & Zon, L. I. Hematopoiesis: an evolving paradigm for stem cell biology. Cell 132, 631–644 (2008)
Medvinsky, A., Rybtsov, S. & Taoudi, S. Embryonic origin of the adult hematopoietic system: advances and questions. Development 138, 1017–1031 (2011)
Dzierzak, E. & Speck, N. A. Of lineage and legacy: the development of mammalian hematopoietic stem cells. Nat. Immunol. 9, 129–136 (2008)
McGrath, K. E. et al. Distinct sources of hematopoietic progenitors emerge before HSCs and provide functional blood cells in the mammalian embryo. Cell Rep. 11, 1892–1904 (2015)
Böiers, C. et al. Lymphomyeloid contribution of an immune-restricted progenitor emerging prior to definitive hematopoietic stem cells. Cell Stem Cell 13, 535–548 (2013)
Cedar, H. & Bergman, Y. Epigenetics of haematopoietic cell development. Nat. Rev. Immunol. 11, 478–488 (2011)
Doulatov, S. et al. Induction of multipotential hematopoietic progenitors from human pluripotent stem cells via respecification of lineage-restricted precursors. Cell Stem Cell 13, 459–470 (2013)
Shen, X. et al. EZH1 mediates methylation on histone H3 lysine 27 and complements EZH2 in maintaining stem cell identity and executing pluripotency. Mol. Cell 32, 491–502 (2008)
Laurenti, E. et al. The transcriptional architecture of early human hematopoiesis identifies multilevel control of lymphoid commitment. Nat. Immunol. 14, 756–763 (2013)
North, T. E. et al. Runx1 expression marks long-term repopulating hematopoietic stem cells in the midgestation mouse embryo. Immunity 16, 661–672 (2002)
Rybtsov, S. et al. Hierarchical organization and early hematopoietic specification of the developing HSC lineage in the AGM region. J. Exp. Med. 208, 1305–1315 (2011)
Boisset, J. C. et al. In vivo imaging of haematopoietic cells emerging from the mouse aortic endothelium. Nature 464, 116–120 (2010)
Riddell, J. et al. Reprogramming committed murine blood cells to induced hematopoietic stem cells with defined factors. Cell 157, 549–564 (2014)
Papaemmanuil, E. et al. Loci on 7p12.2, 10q21.2 and 14q11.2 are associated with risk of childhood acute lymphoblastic leukemia. Nat. Genet. 41, 1006–1010 (2009)
Tothova, Z. et al. FoxOs are critical mediators of hematopoietic stem cell resistance to physiologic oxidative stress. Cell 128, 325–339 (2007)
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006)
Xu, J. et al. Developmental control of Polycomb subunit composition by GATA factors mediates a switch to non-canonical functions. Mol. Cell 57, 304–316 (2015)
Margueron, R. et al. Ezh1 and Ezh2 maintain repressive chromatin through different mechanisms. Mol. Cell 32, 503–518 (2008)
Jang, I. H. et al. Notch1 acts via Foxc2 to promote definitive hematopoiesis via effects on hemogenic endothelium. Blood 125, 1418–1426 (2015)
Hohmann, A. F. et al. Sensitivity and engineered resistance of myeloid leukemia cells to BRD9 inhibition. Nat. Chem. Biol. 12, 672–679 (2016)
Müller, A. M., Medvinsky, A., Strouboulis, J., Grosveld, F. & Dzierzak, E. Development of hematopoietic stem cell activity in the mouse embryo. Immunity 1, 291–301 (1994)
Yoder, M. C. et al. Characterization of definitive lymphohematopoietic stem cells in the day 9 murine yolk sac. Immunity 7, 335–344 (1997)
Lee, S. C. et al. Polycomb repressive complex 2 component Suz12 is required for hematopoietic stem cell function and lymphopoiesis. Blood 126, 167–175 (2015)
Xie, H. et al. Polycomb repressive complex 2 regulates normal hematopoietic stem cell function in a developmental-stage-specific manner. Cell Stem Cell 14, 68–80 (2014)
Majewski, I. J. et al. Polycomb repressive complex 2 (PRC2) restricts hematopoietic stem cell activity. PLoS Biol. 6, e93 (2008)
Kamminga, L. M. et al. The Polycomb group gene Ezh2 prevents hematopoietic stem cell exhaustion. Blood 107, 2170–2179 (2006)
Cai, Z. et al. Haploinsufficiency of AML1 affects the temporal and spatial generation of hematopoietic stem cells in the mouse embryo. Immunity 13, 423–431 (2000)
Sugimura, R. et al. Haematopoietic stem and progenitor cells from human pluripotent stem cells. Nature 545, 432–438 (2017)
Shao, Z., Zhang, Y., Yuan, G. C., Orkin, S. H. & Waxman, D. J. MAnorm: a robust model for quantitative comparison of ChIP–seq data sets. Genome Biol. 13, R16 (2012)
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010)
Cahan, P. et al. CellNet: network biology applied to stem cell engineering. Cell 158, 903–915 (2014)
Shin, H., Liu, T., Manrai, A. K. & Liu, X. S. CEAS: cis-regulatory element annotation system. Bioinformatics 25, 2605–2606 (2009)
Vo, L. T., Doulatov, S. & Daley, G. Q. Haematopoietic progenitor and lymphoid differentiation from human pluripotent stem cells. Protoc. Exch. http://doi.org/10.1038/protex.2017.160 (2018)
Park, I. H. et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature 451, 141–146 (2008)
Chadwick, K. et al. Cytokines and BMP-4 promote hematopoietic differentiation of human embryonic stem cells. Blood 102, 906–915 (2003)
Onder, T. T. et al. Chromatin-modifying enzymes as modulators of reprogramming. Nature 483, 598–602 (2012)
Holmes, R. & Zúñiga-Pflücker, J. C. The OP9-DL1 system: generation of T-lymphocytes from embryonic or hematopoietic stem cells in vitro. Cold Spring Harb. Protoc. 2009, http://doi.org/10.1101/pdb.prot5156 (2009)
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46–53 (2013)
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008)
Huang, J. et al. Dynamic control of enhancer repertoires drives lineage and stage-specific transcription during hematopoiesis. Dev. Cell 36, 9–23 (2016)
Kim, J., Cantor, A. B., Orkin, S. H. & Wang, J. Use of in vivo biotinylation to study protein–protein and protein–DNA interactions in mouse embryonic stem cells. Nat. Protoc. 4, 506–517 (2009)
Ditadi, A. et al. Human definitive haemogenic endothelium and arterial vascular endothelium represent distinct lineages. Nat. Cell Biol. 17, 580–591 (2015)
Doulatov, S. et al. Revised map of the human progenitor hierarchy shows the origin of macrophages and dendritic cells in early lymphoid development. Nat. Immunol. 11, 585–593 (2010)
Acknowledgements
We thank T. Jenuwein for sharing the Ezh1 mutant mice, which were generated at the Research Institute of Molecular Pathology (IMP, Vienna) in 2000 by D. O’Carroll (laboratory of T. Jenuwein) with the help of M. Sibilia (laboratory of E. Wagner). We also thank T. Schlaeger and the hESC Core Facility at Boston Children’s Hospital for providing pluripotent stem-cell lines, R. Mathieu from BCH Flow Cytometry Core, and M. J. Chen for technical advice. This work was supported by grants from the NIH NIDDK (R24-DK092760, R24-DK49216) and NHLBI Progenitor Cell Biology Consortium (U01-HL100001); NHLBI R01HL04880 and NIH R24OD017870-01. L.T.V. is supported by the NSF Graduate Research Fellowship. M.A.K. is supported by T32 NIH Training Grant from BWH Hematology. M.C. is supported by a fellowship from the Leukemia and Lymphoma Society. S.D. is supported by K99 NIH NHLBI award (1K99HL123484). S.H.O. is an Investigator of the Howard Hughes Medical Institute. J.X. is supported by NIH grants (K01DK093543 and R01DK111430) and a Cancer Prevention and Research Institute of Texas (CPRIT) New Investigator award (RR140025). G.Q.D. was supported by the Howard Hughes Medical Institute, and is an associate member of the Broad Institute and an investigator of the Manton Center for Orphan Disease Research.
Author information
Authors and Affiliations
Contributions
L.T.V., S.D. and G.Q.D. conceived the project. L.T.V. designed all experiments, performed all pluripotent stem-cell and mouse transplantation studies and interpreted data. M.A.K. analysed RNA-seq, ChIP–seq and ATAC–seq data, performed all network analyses and interpreted data. X.L. performed ChIP–seq and ATAC–seq experiments. Y.Z. and Z.S. analysed ChIP–seq and ATAC–seq data. J.B. performed and analysed qPCR and western blot validations, assisted with tissue culture, animal dissections and mouse transplantation studies. P.M.S. assisted with timed matings, animal dissections and mouse transplantation studies. D.K.J. performed western blot validations, cloned the Ezh2-mCherry overexpression construct, assisted with ChIP–seq optimization and interpreted data. M.C. assisted with ChIP–seq optimization. A.H. assisted with RNA-seq analysis. T.E.N., S.H.O., S.D., J.X. and G.Q.D. supervised research, interpreted data and participated in project planning. L.T.V., T.E.N., S.D. and G.Q.D. wrote the manuscript with input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks B. Gottgens, H. Mikkola and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 EZH1 knockdown activates lymphoid potential from pluripotent stem cells.
a, List of all candidate epigenetic modifiers in loss-of-function shRNA screen. b, Representative flow plots of CD4+CD8+ T cell potential across top six candidates from four independent hairpins in two independent experiments (n = 8). See Fig. 1. c, CD34+ cells were isolated after 9 days of embryoid body (EB) differentiation (top left), transduced with shLUC or shEZH1 and cultured under conditions that promote endothelial-to-haematopoietic transition44. After 6 days, rounded haematopoietic cells (top right) were collected and co-cultured on OP9-DL1 stroma. Bottom, flow cytometric analysis of T cell potential in shLUC and shEZH1 cells without 5F is shown for two independent iPS lines (34-iPS and MSC-iPS1) in one experiment (n = 2 biological replicates). PSC-HE, pluripotent stem-cell-derived haemogenic endothelium. d, Expansion and differentiation potential of 5F plus shEZH1 cells after long-term in vitro culture. 5F plus shEZH1 cells were maintained in cultures containing doxycycline for 14 days respecification (approximately 100-fold expansion), plus an additional 6 weeks (approximately 1,000-fold expansion) and then plated into OP9-DL1 stromal cells for T cell differentiation. Representative flow cytometric analyses of T cell potential of 5F plus shLUC and 5F plus shEZH1 cells after 13 weeks of expansion and differentiation (n = 2 biological replicates). e, Flow cytometric analysis (left) and quantification (right) of the proportion of CD34+ and CD34− haematopoietic progenitors in doxycycline-containing suspension culture at day 25 (n = 2 biological replicates).
Extended Data Figure 2 Ezh1, but not Ezh2, suppresses T cell potential and requires its catalytic domain.
a, qPCR of PRC2 expression (human genes EZH1, EED, EZH2 and SUZ12), during the course of differentiation from human pluripotent stem cell-derived CD34+ cells, respecification (5F), expansion, OP9-DL1 co-culture and CD4+CD8+ T cells (n = 2 biological replicates in one experiment). b, qPCR of mRNA knockdown efficiency of individual shRNAs for PRC2 genes (n = 2 biological replicates). See also Fig. 2a, b. c, Western blot for EZH1 and GAPDH protein levels. d, Scheme for rescue experiments. GFP+ 5F cells were transduced with shRNAs and selected with puromycin. 5F plus shRNA cells were then transduced with full-length mouse Ezh1 open-reading frame (mEzh1) or mutant mouse Ezh1 with the catalytic SET domain deleted (mEzh1ΔSET), marked by mCherry fluorescence. Triple-transduced (GFP+, puromycin-resistant, mCherry+) cells were sorted and seeded onto OP9-DL1. T cells were analysed by flow cytometry after 5 weeks of differentiation. See also Fig. 2c. e, Expression of full-length mouse Ezh1, catalytic-deleted mEzh1ΔSET, or full-length mouse Ezh2 in shLUC and shEZH1 cells by qPCR. f, Western blot validation of expression of mouse Ezh1 or mutant mEzh1ΔSET in shLUC and shEZH1 cells. g, Top, representative flow cytometry plots of T cell potential for 5F plus shLUC cells for rescue experiments in d (n = 3 biological replicates). Bottom, CD4+CD8+ T cells were verified for mCherry fluorescence. See also Fig. 2c. All plots are gated on CD45+. h, 5F plus shRNA cells were transduced with full-length mouse Ezh2 open-reading frame (mEzh2) marked by mCherry fluorescence. Triple-transduced (GFP+, puromycin-resistant, mCherry+) cells were sorted and seeded onto OP9-DL1 stromal cells. T cells were analysed by flow cytometry after 5 weeks of differentiation. Representative flow plots for two biological replicates in one experiment. i, Quantification of data in h. Data are mean ± s.e.m.
Extended Data Figure 3 Ezh1 regulates haematopoietic and lymphoid programs in vitro and in vivo.
a, Representative images of E10.5 embryo (top), yolk sac (middle) and AGM (bottom) from n > 30 embryos. Lin−c-Kit+VE-cadherin+CD45+CD41+ cells from E10.5 yolk sac and AGM were FACS-sorted followed by RNA-seq analysis. See also Fig. 4a, c, d. b, Genes upregulated and downregulated by more than twofold in Ezh1+/− or Ezh1−/− yolk sac and AGM compared to those from wild-type mice. c, d, GO term annotations of upregulated genes in Ezh1+/− and Ezh1−/− yolk sac and AGM compared to those from wild-type mice. e, GO analysis of enriched pathways of the 1,033 nearest neighbour genes associated with upregulated ATAC peaks (top) and the nearest 1,012 neighbour genes associated with downregulated ATAC peaks (bottom). See also Fig. 3d, e. f, Comparison of upregulated ATAC peaks in 5F plus shEZH1 cells with HSPC hierarchy signatures45 (top) and HSPC B and T cell networks (bottom). See also Fig. 3d, e. g, Box plot of expression of genes associated with upregulated and downregulated ATAC peaks. *P < 0.05 by one-way ANOVA. h, ATAC density map of c-Kit+VE-cadherin+CD45+ HSPCs sorted from approximately 30 embryos of E10.5 wild-type and Ezh1−/− AGM (top) from one experiment. Significantly upregulated ATAC peaks were compared to HSPC, T, B cell networks and signatures of the human HSPC hierarchy (bottom). See also Fig. 4c, d. i, GO terms of enriched pathways of regions associated with significantly upregulated ATAC peaks annotated by GREAT analysis in Ezh1+/− AGM (top) and Ezh1−/− AGM (bottom) compared to wild type. See also Fig. 4c, d. j, GO terms of enriched pathways of regions associated with significantly downregulated ATAC peaks annotated by GREAT analysis30 in Ezh1+/− AGM (top) and Ezh1−/− AGM (bottom) compared to wild type. See also Fig. 4c. k, Transcription factor binding to genes with upregulated ATAC peaks in Ezh1+/− (left) and Ezh1−/− (right) AGM from i compared to wild-type AGM.
Extended Data Figure 4 Genome-wide chromatin occupancy reveals EZH1 enrichment at bivalent HSC genes and non-canonical active lymphoid genes.
a, Breakdown of EZH1 binding at promoter regions and associated histone marks. b–d, GO term analysis of EZH1-bound active (b), bivalent (c) and repressed (d) genes. e, Distribution of EZH1-bound genes across the haematopoietic hierarchy (left) and their associated histone marks (right). A, active (H3K4me3-marked); B, bivalent (H3K4me3 and H3K27me3-marked); R, repressed (H3K27me3-marked). f, GSEA analysis of EZH1-bound genes correlated with RNA-seq data upon EZH1 knockdown. g, Sankey diagram showing genome-wide changes in histone methylation status upon EZH1 knockdown. h, Upregulated genes exhibit reciprocal decreases in H3K27me3 levels, as quantified by EpiChIP software. K4, H3K4me3; K27, H3K27me3. See also Fig. 3i. i, Activated (formerly bivalent) HSC genes exhibit increased gene expression upon EZH1 knockdown and loss of H3K27me3. See also Fig. 3j. j, Correlations between changes in H3K27me3 and gene expression levels upon EZH1 knockdown, subdivided by subgroups corresponding to methylation changes. N, null. k, Breakdown of bivalent–bivalent (left), bivalent–repressed (centre) and bivalent–null (right) genes upon EZH1 knockdown across the haematopoietic hierarchy. l, Overlap of EZH1- and EZH2-enriched peaks and the distribution of all EZH1-enriched, EZH2-enriched or common genes across the hierarchy (left), or specifically bivalent genes that become activated upon EZH1 knockdown (middle) and active genes, marked by H3K4me3 in shLUC (right). m, SUZ12 binding (from the ChEA database) across the haematopoietic hierarchy. n, Canonical and non-canonical previously identified targets17 across the haematopoietic hierarchy. o, p, Breakdown of histone marks on non-canonical ProB genes (o) and the genome-wide distribution from CEAS analysis32 (p). q, Changes in actively marked, non-canonical ProB genes (green bar in o), upon EZH1 knockdown. r, SUZ12 and EZH2 binding (ChEA database) at ATAC peaks in Ezh1+/− and Ezh1−/− AGM. *P < 0.05 by one-way ANOVA.
Extended Data Figure 5 Ezh1 deficiency enhances embryonic HSPC engraftment.
a, Whole E10.5 AGM and yolk sac were transplanted intravenously into sublethally irradiated NSG adult females. Chimaerism was monitored by retroorbital bleeding every 4 weeks. Representative flow plots are shown for analysis after 4 weeks in n ≥ 3 mice. See also Fig. 4e, f. b, Whole E9.5 PSP was transplanted intravenously into sublethally irradiated NSG adult females. Chimaerism was monitored via retroorbital bleeding every 4 weeks. Representative flow plots are shown for analysis after 8 weeks in n ≥ 3 mice. See also Fig. 4g. c, Representative flow plots of lineage analysis in E10.5 AGM Ezh1+/− and Ezh1−/− primary transplant recipients after 24 weeks, and in E9.5 PSP Ezh1+/− primary transplant recipient after 16 weeks (n ≥ 3 mice per group). See also Fig. 4e, g. d, Primary recipients in a were euthanized 24 weeks post-transplantation and 4 × 106 whole bone marrow was transplanted into sublethally irradiated adult NSG females intravenously. Chimaerism was monitored via retroorbital bleeding. Representative flow plots of E10.5 AGM and yolk sac secondary transplants after 4 weeks in n ≥ 3 mice. See also Fig. 4h. e, Left, secondary transplantation of E10.5 yolk sac primary recipients (Fig. 4f). Right, lineage distribution of E10.5 secondary recipients. Data are pooled across three independent experiments. *P < 0.05, ** P < 0.01 by unpaired two-sided t-test; see Supplementary Information for exact P values per time point.
Extended Data Figure 6 Ezh1-deficient embryonic HSPCs contribute to adult-type lymphopoiesis in vivo.
a, Flow analysis of B1 and B2 progenitors in the peritoneal cavity of engrafted primary recipients (n = 1 mouse per group). b, Flow analysis of TCRβ and TCRγδ frequencies of donor-derived peripheral CD3+ T cells from engrafted primary recipients (n = 1 mouse per group). See also Fig. 4e, f.
Supplementary information
Supplementary Information
This file contains the uncropped gels and a list of sequences used in the manuscript. (PDF 1176 kb)
Rights and permissions
About this article
Cite this article
Vo, L., Kinney, M., Liu, X. et al. Regulation of embryonic haematopoietic multipotency by EZH1. Nature 553, 506–510 (2018). https://doi.org/10.1038/nature25435
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25435
- Springer Nature Limited
This article is cited by
-
Gain and loss of function variants in EZH1 disrupt neurogenesis and cause dominant and recessive neurodevelopmental disorders
Nature Communications (2023)
-
Generation and clinical potential of functional T lymphocytes from gene-edited pluripotent stem cells
Experimental Hematology & Oncology (2022)
-
Evolutionary adaptation of the Polycomb repressive complex 2
Epigenetics & Chromatin (2022)
-
Integrative epigenomic and transcriptomic analysis reveals the requirement of JUNB for hematopoietic fate induction
Nature Communications (2022)
-
Continuous human iPSC-macrophage mass production by suspension culture in stirred tank bioreactors
Nature Protocols (2022)