Abstract
Clostridium difficile disease has recently increased to become a dominant nosocomial pathogen in North America and Europe, although little is known about what has driven this emergence. Here we show that two epidemic ribotypes (RT027 and RT078) have acquired unique mechanisms to metabolize low concentrations of the disaccharide trehalose. RT027 strains contain a single point mutation in the trehalose repressor that increases the sensitivity of this ribotype to trehalose by more than 500-fold. Furthermore, dietary trehalose increases the virulence of a RT027 strain in a mouse model of infection. RT078 strains acquired a cluster of four genes involved in trehalose metabolism, including a PTS permease that is both necessary and sufficient for growth on low concentrations of trehalose. We propose that the implementation of trehalose as a food additive into the human diet, shortly before the emergence of these two epidemic lineages, helped select for their emergence and contributed to hypervirulence.
Similar content being viewed by others
Main
Whole-genome sequencing analysis of C. difficile ribotype 027 (RT027) strains demonstrated that two independent lineages emerged in North America from 2000 to 2003 (ref. 1). Comparison with historic, pre-epidemic, RT027 strains showed that both epidemic lineages acquired a mutation in the gyrA gene, leading to increased resistance to fluoroquinolone antibiotics. While the development of fluoroquinolone resistance has almost certainly played a role in the spread of RT027 strains, fluoroquinolone resistance has also been observed in non-epidemic C. difficile ribotypes and identified in strains dating back to the mid-1980s2,3. Thus, other factors probably contributed to the emergence of epidemic RT027 strains.
The prevalence of a second C. difficile ribotype, RT078, increased tenfold in hospitals and clinics from 1995 to 2007 and was associated with increased disease severity4. However, the mechanisms responsible for increased virulence remain unknown5,6,7,8. It is noteworthy that RT027 and RT078 lineages are phylogenetically distant from one another (Extended Data Fig. 1), indicating that the evolutionary changes leading to concurrent increases in epidemics and disease severity might have emerged by independent mechanisms9.
RT027 and RT078 strains grow on low trehalose
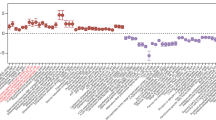
Ribotype 027 strains exhibit a competitive advantage over non-RT027 strains in vitro and in mouse models of C. difficile infection10. To investigate potential mechanisms for increased fitness, we examined carbon source utilization in an epidemic RT027 isolate (CD2015) using Biolog 96-well Phenotype MicroArray carbon source plates (see Methods and Extended Data Table 1). Out of several carbon sources identified that supported CD2015 growth, we found the disaccharide trehalose increased the growth yield of CD2015 by approximately fivefold compared with a non-RT027 strain. To examine the specificity of enhanced growth on trehalose across C. difficile lineages, 21 strains encompassing 9 ribotypes were grown on a defined minimal medium (DMM) supplemented with glucose or trehalose as the sole carbon source. All C. difficile strains grew robustly with 20 mM glucose; however, only epidemic RT027 (n = 8) and RT078 (n = 3) strains exhibited enhanced growth on an equivalent trehalose concentration (10 mM; Fig. 1). Increasing the trehalose concentration to 50 mM enabled growth in most ribotypes (Extended Data Fig. 2a).
Dashed grey line and band indicate mean growth and s.d. in DMM without a carbon source for all samples (n = 21). Solid lines are mean growth yield (absorbance at 600 nm, A600 nm) for groups: non-RT027/078 (n = 10), RT027 (n = 8), and RT078 (n = 3). All points represent biologically independent samples.
Molecular basis for RT027 growth on low trehalose
To identify the genetic basis for enhanced trehalose metabolism, we compared multiple C. difficile genomes. All C. difficile genomes encode a putative phosphotrehalase enzyme (TreA) preceded by a transcriptional repressor (TreR) (Fig. 2a). Phosphotrehalase enzymes metabolize trehalose-6-phosphate into glucose and glucose-6-phosphate. To test whether treA was essential for trehalose metabolism, we generated treA deletion mutants in the RT027 strain R20291 (R20291∆treA) and the RT012 strain CD630 (CD630∆treA) and grew them in DMM supplemented with 50 mM trehalose. The lack of treA prevented growth in both knockout strains, which could be complemented by plasmid expression of treA (Extended Data Fig. 2b). Thus, treA is required to metabolize trehalose.
a, Trehalose metabolism operon found in all C. difficile strains, consisting of a phosphotrehalase (treA) and its transcriptional regulator (treR). b, RT027 strains strongly induce treA at 50 μM trehalose and at a significantly higher level than non-RT027 strains (n = 4 biologically independent samples per trehalose concentration/strain). Bars are average fold increase, error bars are s.d.; P values derived from t-test (two-tailed) and Holm corrected for multiple comparisons. c, Structure of TreR monomer highlighting proximity of L172I mutation to trehalose-6-P binding pocket.
We next asked whether RT027 strains have altered regulation of the treA gene compared with other ribotypes. To test this hypothesis and determine the minimum level of trehalose required to activate treA expression, we grew CD2015 (RT027) and CD2048 (RT053) and exposed them to increasing amounts of trehalose. We found that the RT027 strain turned on treA expression at 50 μM trehalose, a concentration 500-fold lower than that required to turn on treA in RT053 (Fig. 2b). To confirm this phenotype, we took four RT027 strains and four non-RT027 strains and measured expression of treA in a single trehalose concentration. Again, RT027 strains exhibited significantly higher treA expression than all other ribotypes (P = 0.029; Extended Data Fig. 3). These results support the idea that RT027 strains are exquisitely sensitive to low concentrations of trehalose.
Sequence alignment of the trehalose operon across 1,010 sequenced C. difficile strains revealed a conserved single nucleotide polymorphism (SNP) within the treR gene of all RT027 strains (TreRRT027) (Extended Data Fig. 4a). The SNP encodes an L172I amino acid substitution near the predicted effector (trehalose-6-phosphate) binding pocket of TreR (Fig. 2c), a site that is highly conserved across multiple species (93.9% conservation; Extended Data Fig. 4b). This SNP is found not only in every RT027 strain sequenced so far, but also in a newly isolated fluoroquinolone sensitive ribotype (RT244) that has caused community-acquired epidemic outbreaks in Australia11,12 and other ribotypes very closely related to RT027, such as RT176 which has caused epidemic outbreaks in the Czech Republic and Poland13,14. Like RT027 strains, the RT244 strains DL3110 and DL3111 can grow on 10 mM trehalose (Extended Data Fig. 2c).
To determine the types of spontaneous mutation that lead to enhanced trehalose utilization, we cultivated several non-RT027/RT078 strains under low trehalose concentrations in minibioreactors10. After 3 days of continuous cultivation, 13 independent spontaneous mutants capable of growing on low concentrations (<10 mM) of trehalose were isolated. All 13 mutants contained either nonsense or missense mutations in the treR gene (Extended Data Table 2).
Effect of trehalose metabolism on disease severity
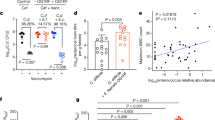
To test whether the ability of C. difficile RT027 strains to metabolize trehalose impacts disease severity, we performed two different experiments. In the first, humanized microbiota mice were challenged with 104 spores of either R20291 (RT027, n = 27) or R20291∆treA (n = 28). After infection, trehalose (5 mM) was provided ad libitum in the drinking water and disease progression monitored. The R20291∆treA mutant demonstrated a marked decrease in mortality (33.3% versus 78.6%) compared with R20291 (78% lower risk with R20291∆treA; hazard ratio 0.22; 95% confidence interval 0.09–0.59; P = 0.003, likelihood ratio test P = 0.002; Fig. 3a). In the second experiment, we infected two groups of humanized microbiota mice with RT027 strain R20291. One group received 5 mM trehalose in water as well as a daily gavage of 300 mM trehalose (n = 28) to mimic a dose expected in a meal for humans, whereas the control group (n = 27) received a water control. Trehalose addition was found to cause increased mortality compared with the RT027-infected mice without dietary trehalose (threefold increased risk with trehalose; hazard ratio, 3.20; 95% confidence interval 1.09–9.42; P = 0.035, likelihood ratio test P = 0.026; Fig. 3b). Combined, these results show that metabolism of dietary trehalose can contribute to disease severity of RT027 C. difficile strains.
a, Mice infected with R20291∆treA (n = 27 animals) have significantly attenuated risk of mortality compared with mice infected with R20291 (n = 28 animals) (78% lower risk with ∆treA mutant; hazard ratio 0.22; 95% confidence interval 0.09–0.59; P = 0.003). b, Mice infected with R20291 (RT027) have a significantly higher risk of mortality when trehalose is supplemented in the diet (n = 28 animals) than those with no trehalose supplementation (n = 27 animals) (threefold increased risk with trehalose; hazard ratio 3.20; 95% confidence interval 1.09–9.42; P = 0.035). Experiments were repeated twice. All statistical tests were two-sided.
To identify the cause of increased disease severity when trehalose is present, we challenged mice with either R20291 or R20291∆treA and provided 5 mM trehalose ad libitum in the drinking water. Forty-eight hours after challenge, C. difficile load and toxin levels were measured. Over two independent experiments, no significant difference in C. difficile numbers was observed; however, a significant increase in the relative levels of toxin B was detected (median 9.2 × 104, interquartile range (IQR) 5.1 × 104 to 1.0 × 105 versus median 4.1 × 104, IQR 2.3 × 104 to 4.6 × 104, P = 0.0268; Extended Data Fig. 5). This increased toxin production could contribute to increased disease severity.
Molecular basis for RT078 growth on low trehalose
Unlike RT027, RT078 strains do not possess the TreR L172I substitution or other conserved SNPs in the treRA operon. To identify sequences of potential relevance to trehalose metabolism, we performed whole-genome comparisons. A four-gene insertion was found in all RT078 strains sequenced so far, annotated to encode a second copy of a phosphotrehalase (TreA2, sharing 55% amino acid identity with TreA), a potential trehalose specific PTS system IIBC component transporter (PtsT), a trehalase family protein that is a putative glycan debranching enzyme (TreX), and a second copy of a TreR repressor protein (TreR2, sharing 44% amino acid identity with TreR) (see Fig. 4a). Genomic comparison of publicly available C. difficile genomes revealed the four-gene insertion was present in RT078 and the closely related RT033, RT045, RT066 and RT126 ribotypes and absent from reference genomes of any other C. difficile lineage (Extended Data Fig. 6).
a, Structure of horizontally acquired trehalose metabolism module found in RT078 and closely related strains. b, Deletion of the trehalose transporter from a clinical RT078 strain (CD1015) ablates its ability to grow on 10 mM trehalose. Expression of ptsT from an inducible plasmid restores growth of CD1015∆ptsT on 10 mM trehalose (CD1015, n = 4; CD1015∆ptsT, n = 4; CD1015∆ptsT::ptsT, n = 5). c, Expressing ptsT from an inducible plasmid enables enhanced growth of CD630 (RT012) on 10 mM trehalose (CD630 n = 3; CD630::ptsT n = 3). d, The ptsT provides a competitive advantage in complex microbial communities. Dashed grey line (competitive index = 1) indicates equal fitness of the competing strains, points above this line represent out-competition by CD1015. All points (Fig. 4b–d) represent biologically independent samples, bars are mean, P values derived from t-test (two-tailed) and Holm corrected for multiple comparisons where appropriate.
To test whether the newly acquired transporter (ptsT) was responsible for enhanced trehalose metabolism, a ptsT deletion mutant was constructed in a RT078 (CD1015) strain. This strain was unable to grow on DMM supplemented with 10 mM trehalose (Fig. 4b), but retained the ability to grow in medium supplemented with 50 mM trehalose (Extended Data Fig. 2d). The growth defect in this deletion mutant (CD1015∆ptsT) was directly due to the lack of ptsT since expression of ptsT from an inducible promoter could complement growth on 10 mM trehalose (Fig. 4b).
We next tested whether ptsT was sufficient to confer enhanced trehalose utilization in a non-ribotype 078 strain, which fails to grow under low trehalose concentrations. To do this, ptsT was expressed from an inducible promoter in strain CD630 (RT012). Expression of ptsT was sufficient to allow growth of CD630 in DMM supplemented with 10 mM trehalose (Fig. 4c). Taken together, we conclude that ptsT is both necessary and sufficient to support growth on low concentrations of trehalose.
To test whether the expression of ptsT could confer a fitness advantage, CD1015 (RT078) was competed against its isogenic CD1015∆ptsT mutant in a human faecal minibioreactor model of C. difficile infection10. After clindamycin treatment of minibioreactor communities to enable infection, CD1015 and CD1015∆ptsT strains were added together to each reactor and levels monitored over time. Remarkably, the CD1015 strain was found to be significantly more efficient at competing in vivo in the presence of a complex microbiota than the CD1015∆ptsT mutant (mean competitive index of 246 on day 7). To ensure the CD1015∆ptsT loss was due to the absence of ptsT, CD1015 was competed against the CD1015∆ptsT mutant complemented with ptsT from an inducible vector. After 5 days of continuous competition, the wild-type RT078 had a mean competitive index of just 3.7 (Fig. 4d). Hence, ptsT provides a competitive fitness advantage to RT078 strains.
Trehalose is observed in the distal gut
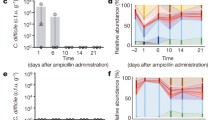
Despite the presence of a localized brush border trehalase enzyme in the small intestine, human studies suggest that high levels of trehalose consumption can result in significant amounts reaching the distal ileum and colon15,16,17. To demonstrate that a significant amount of dietary trehalose can survive transit through the small intestine, we gavaged mice with 100 μl (300 mM) trehalose (equivalent to the suggested concentration in ice cream) and measured trehalose levels in the caecum over time. Using clinical C. difficile strains as biosensors, we found the level of trehalose to be sufficient to activate treA gene expression in the RT027 strain CD2015 but not in RT053 strain CD2048 (Fig. 5a). To test whether we could detect a low dietary amount of trehalose, we gavaged antibiotic-treated mice with 100 μl (5 mM) trehalose and measured treA activation in these same strains. Again, the RT027 strain showed significant treA activation (Fig. 5b). Finally, to determine whether trehalose is bioavailable in humans at sufficient levels to be used by epidemic C. difficile isolates, we tested ileostomy effluent from three anonymous donors consuming their normal diets. In two of three samples, treA expression was strongly induced in the RT027 strain CD2015 but not in the RT053 strain CD2048 (Fig. 5c), supporting the notion that levels of trehalose found in food are sufficient to be used by epidemic C. difficile strains.
a, Twenty minutes after gavage, trehalose reaches high enough levels in the mouse caecum to turn on expression of treA in RT027 but not non-RT027 in both non-antibiotic- and antibiotic-treated mice (n = 3 animals per trehalose concentration/strain). b, Trehalose can be detected by RT027 but not non-RT027 in the caecum of antibiotic-treated mice gavaged with just 100 μl of 5 mM trehalose (n = 3 animals per group). c, RT027 strains can detect trehalose in two out of three human ileostomy fluid samples tested from patients eating a normal (no deliberate trehalose addition) diet. Points represent biologically independent replicates, bars are average fold increase, error bars are s.d.
Discussion
Containing an α,α-1,1-glucoside bond between two α-glucose units, trehalose is a non-reducing and extremely stable sugar, resistant both to high temperatures and to acid hydrolysis. Although considered an ideal sugar for use in the food industry, the use of trehalose in the United States and Europe was limited before 2000 owing to the high cost of production (approximately US$700 per kilogram). The innovation of a novel enzymatic method for low-cost production from starch made it commercially viable as a food supplement (approximately US$3 per kilogram)18. Granted ‘generally recognized as safe’ status by the US Food and Drug Administration in 2000 and approved for use in food in Europe in 2001, reported expected usage ranges from concentrations of 2% to 11.25% for foods including pasta, ground beef, and ice cream. The widespread adoption and use of trehalose in the diet coincides with the emergence of both RT027 and RT078 outbreaks (Fig. 6).
Flags indicate reported outbreaks or first reports of RT027 (top) or RT078 (bottom) in PubMed. SM1 and SM2, outbreaks at Stoke Mandeville Hospital, Buckinghamshire, UK. CDI, C. difficile infection.
Several lines of evidence support the idea that dietary trehalose has participated in the spread of epidemic C. difficile ribotypes. First, the ability of RT027 and RT078 strains to metabolize trehalose was present before epidemic outbreaks. The earliest retrospectively recorded RT027 isolate was the non-epidemic strain CD196, isolated in 1985 in a Paris hospital19. Three years later in 1988, another non-epidemic strain RT027 (BI1) was isolated in Minneapolis, Minnesota. Both isolates, in addition to every RT027 strain sequenced so far, contain the L172I substitution in TreR. RT078 strains were also present in humans before 2001, but epidemic outbreaks were not reported until 2003 (ref. 4). Second, RT027 and RT078 lineages are phylogenetically distant clades of C. difficile, yet have convergently evolved distinct mechanisms to metabolize low levels of trehalose. Third, increased disease severity of a RT027 strain that can metabolize trehalose in our mouse model of C. difficile infection is consistent with increased virulence of RT027 and RT078 ribotypes observed in patients. Fourth, the ability to metabolize trehalose at lower concentrations confers a competitive growth advantage in the presence of a complex intestinal community. Finally, levels of trehalose in ileostomy fluid from patients eating a normal diet are sufficiently high to be detected by RT027 strains. On the basis of these observations, we propose that the widespread adoption and use of the disaccharide trehalose in the human diet has played a significant role in the emergence of these epidemic and hypervirulent strains20.
Methods
Bacterial strains and growth
A full list of strains can be found in Extended Data Table 3. Carbon source utilization of CD630 (RT012) and CD2015 (clinical RT027) was performed using Biolog Phenotypic Microarray plates. Growth studies were performed under anaerobic conditions (5% hydrogen, 90% nitrogen, 5% carbon dioxide). Strains were cultured overnight in BHI media (Difco) supplemented with 0.5% (w/v) yeast extract. Growth assays used a DMM as described previously21 supplemented with either trehalose or glucose as indicated. Anhydrous tetracycline was used at 500 ng ml−1 to induce expression of ptsT or treA from ectopic expression vectors.
Comparative genomics
To identify unique functional features in RT078 strains, we reviewed publicly available C. difficile genomes covering all phylogenetic lineages9 using a tool based on BLASTX comparisons of protein annotations22. The genomes included in the analysis were PCR RT012 (strain 630, lineage I), RT027 (R20291, lineage II), PCR RT017 (CF5, M68 lineage IV), and RT078 (QCD-23m63, CDM120 lineage V).
Genetic manipulation of C. difficile
Inactivation of treA in CD630 was accomplished by group-II intron-directed insertion as previously described23. Primers were designed to target intron to insert at base pair 177 of treA of CD630 (IBS1.2, EBS1, and EBS2; all primers are described in Extended Data Table 3). The resulting treA insertion–deletion mutant was verified by PCR using a primer pair (CR064–CR065) designed to flank the treA insertion site, resulting in a 350 bp product for the wild-type gene and a 2.4 kbp product for the gene knockout.
Clean deletions in R20291 and CD1015 were performed using a pyrE allelic exchange system as described previously24. This is the first case of the pyrE allelic exchange system being used in the RT078 lineage, which required generation of CD1015∆pyrE before further deletions. Complementation of treA and ptsT was performed using an anhydrous tetracycline-inducible system as described previously25. All plasmid conjugations into C. difficile strains were performed with Escherichia coli SD46. Cloning was accomplished with a combination of restriction digest and ligase cycling reactions as described previously26. Primers and detailed plasmid maps for construction of knockout strains are available at the links provided in Extended Data Table 3.
Quantitative PCR with reverse transcription
Strains were grown overnight and subcultured 1:50 into DMM supplemented with 20 mM succinate. Upon reaching an A600 nm of 0.2–0.3, indicated concentrations of trehalose were added to the culture. After incubation for 30 min, C. difficile cells were collected by centrifugation, resuspended in RNALater solution (Invitrogen), and stored at −80 °C. Cells were resuspended in 1 ml RLT buffer (Qiagen RNeasy Kit) and lysed by bead beating (2 × 1 min) at 4 °C followed by RNA extraction according to the manufacturer’s instructions. cDNA was synthesized using Invitrogen Superscript III reverse transcriptase following the recommended protocol. Quantitative PCR reactions were performed in triplicate using Power SYBR Green PCR Master Mix (ABI) with either C. difficile 16 s (JP048–JP049)- or treA (CR045–CR046)-specific primers. Standard curves of cDNA were run to determine primer efficiencies and calculated as in ref. 27. Expression of treA was determined using an average of triplicate CT values from each biological sample.
Mouse model of C. difficile infection
Humanized microbiota mice were derived from an initial population of germ-free C57BL/6 mice stably colonized with human gut microbiota and validated for use as a model of C. difficile infection28. Humanized microbiota mice aged 6–8 weeks of both sexes were treated with a five-antibiotic cocktail consisting of kanamycin (0.4 mg ml−1), gentamicin (0.035 mg ml−1), colistin (850 U ml−1), metronidazole (0.215 mg ml−1), and vancomycin (0.045 mg ml−1) administered ad libitum in drinking water for 4 days. Water was switched to antibiotic-free sterile water and 24 h later mice were administered an intraperitoneal injection of clindamycin (10 mg per kg (body weight)). After a further 24 h, mice were challenged with 104 C. difficile spores by oral gavage. Sterile drinking water containing 5 mM trehalose was provided ad libitum (for the with or without trehalose study, mice were administered an additional 100 μl oral gavage of 300 mM trehalose daily) and mice were monitored for signs of disease.
In a separate experiment to determine C. difficile colonization load and toxin production, mice were euthanized 48 h after challenge with either R20291 or R20291∆treA. C. difficile levels in caecal contents were determined by qPCR of toxin genes10. Relative toxin levels were assessed using a Vero Cell rounding assay10. Sample sizes for all experiments were determined using power analysis based upon previous experimental data. No randomization of animals was performed; however, all groups were checked to ensure no significant difference in the age, weight, or sex of mice between groups before starting experiments. All animal use was approved by the Animal Ethics Committee of Baylor College of Medicine (protocol number AN-6675). The investigators were not blinded to allocation during experiments and outcome assessment.
Detection of trehalose in caecal contents and human ileostomy fluid
Antibiotic-treated groups were pre-treated with the five-antibiotic cocktail for 3 days. Mice were gavaged with 100 μl of 5 mM trehalose, 300 mM trehalose, or water. Twenty minutes after gavage, mice were euthanized, and caecal contents harvested and vigorously mixed with two volumes/weight ice cold DMM (no carbohydrate). Supernatant was separated by centrifugation, filter sterilized, and reduced in an anaerobic chamber overnight before use. Ileostomy effluent from three anonymous donors was self-collected into sterile containers and stored at −20 °C until thawed, filter sterilized, and used for assay.
Strains were grown overnight and subcultured 1:50 into DMM supplemented with 20 mM succinate. Upon reaching an A600 nm of 0.2–0.3, cells were collected by centrifugation and resuspended in approximately 300 μl caecal or ileostomy fluid and incubated anaerobically for 30 min. Cells were then centrifuged and resuspended in RNALater (Invitrogen) before qRT–PCR analysis.
Bioreactor model for RT078 ΔptsT competition
Faecal communities were established in continuous-flow minibioreactor arrays as previously described10 using bioreactor defined medium29 without starch (BDM4). Communities were disrupted by addition of clindamycin (250 μg ml−1) continuously supplied in the medium for 4 days. After clindamycin treatment, communities were supplied BDM4 without clindamycin supplemented with trehalose (5 mM final concentration, BDM4tre). After 1 day of growth in BDM4tre, to allow washout of clindamycin, communities were challenged with a mixture of exponentially growing CD1015 strains (RT078 wild type and ΔptsT). The competitive index was determined by dividing the proportion of wild-type cells at the end of the competition by the proportion at the start. The competitive index of wild type:ΔptsT strains was determined by qPCR. The competitive index of wild type versus CD1015∆ptsT::ahTCptsT was calculated by selective plating.
Isolation of spontaneous treR mutants
C. difficile strains were inoculated into continuous-flow minibioreactor arrays as previously described10 using bioreactor defined medium29 without starch (BDM4) supplemented with 5 mM trehalose (BDM4tre). Every 24 h after the start of the experiment, 200 μl PBS containing 100 mM trehalose was spiked into each minibioreactor. The reactors were sampled daily, serially diluted, and plated to DMM agar supplemented with 10 mM trehalose. Resulting colonies were streak purified, and the ability to grow on low trehalose (10 mM) verified on plates and in broth culture. The treR gene was sequenced and compared with the isogenic parent strain.
Statistics
Statistical analyses were performed using R (version 3.3.2). A Student’s two-sample t-test (two-tailed) was used for comparisons of continuous variables between groups with similar variances; Welch’s two-sample t-test (two-tailed) was used for comparisons of continuous variables between groups with dissimilar variances. P values from multiple comparisons were corrected using the Holm method30. A Wilcoxon rank-sum test with continuity correction was used for the toxin assay where data were non-normal. Fold-change data from treA gene expression experiments were log-normalized before statistical analysis. Data were visualized using individual data points and group means. Cox proportional hazards models and likelihood ratio tests were used to test significant differences in survival distributions among C. difficile-challenged groups of animals.
Collection of human bio-specimens
For faecal samples, live participants who were self-described as healthy and had not consumed antibiotics within the previous 2 months were recruited to provide faecal samples for human faecal bioreactor experiments. Informed consent was obtained before collection of samples and no identifying information was obtained along with the sample. Faecal samples were collected in sterile containers, transported to the laboratory on ice in the presence of anaerobic gas packs (BD Biosciences) within 16 h of collection, manually homogenized in an anaerobic environment, aliquoted into anaerobic tubes, and sealed and stored at −80 °C until use. Participants who had ileostomies placed owing to previous, undisclosed illnesses were recruited to provide ileostomy effluents. Informed consent was obtained before collection of samples and no identifying information was obtained with the sample. After transfer from the ostomy bag to a sterile collection container, ileostomy samples were transported to the laboratory on ice within 12 h of collection. Upon receipt, samples were stored at −20 °C. Ileostomy donors were recruited through the Ostomy Association of Greater Lansing, and were most probably residents of Lansing, Michigan, USA, and its surrounding counties. Samples were stored at −80 °C or −20 °C for 3–4 years before use. Samples were randomly selected for testing from a bank of available samples. Samples were collected according to a protocol approved by the Institutional Review Board of Michigan State University (protocol number 10-736SM).
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. Source data for Figs 1, 2, 3, 4, 5 and Extended Data Figs 2, 3 and 5 are provided with the paper.
References
He, M. et al. Emergence and global spread of epidemic healthcare-associated Clostridium difficile. Nat. Genet. 45, 109–113 (2013)
Spigaglia, P. et al. Fluoroquinolone resistance in Clostridium difficile isolates from a prospective study of C. difficile infections in Europe. J. Med. Microbiol. 57, 784–789 (2008)
Spigaglia, P., Barbanti, F., Dionisi, A. M. & Mastrantonio, P. Clostridium difficile isolates resistant to fluoroquinolones in Italy: emergence of PCR ribotype 018. J. Clin. Microbiol. 48, 2892–2896 (2010)
Jhung, M. A. et al. Toxinotype V Clostridium difficile in humans and food animals. Emerg. Infect. Dis. 14, 1039–1045 (2008)
Goorhuis, A. et al. Emergence of Clostridium difficile infection due to a new hypervirulent strain, polymerase chain reaction ribotype 078. Clin. Infect. Dis. 47, 1162–1170 (2008)
Gupta, A. & Khanna, S. Community-acquired Clostridium difficile infection: an increasing public health threat. Infect. Drug Resist. 7, 63–72 (2014)
Limbago, B. M. et al. Clostridium difficile strains from community-associated infections. J. Clin. Microbiol. 47, 3004–3007 (2009)
Walker, A. S. et al. Relationship between bacterial strain type, host biomarkers, and mortality in Clostridium difficile infection. Clin. Infect. Dis. 56, 1589–1600 (2013)
He, M. et al. Evolutionary dynamics of Clostridium difficile over short and long time scales. Proc. Natl Acad. Sci. USA 107, 7527–7532 (2010)
Robinson, C. D., Auchtung, J. M., Collins, J. & Britton, R. A. Epidemic Clostridium difficile strains demonstrate increased competitive fitness compared to nonepidemic isolates. Infect. Immun. 82, 2815–2825 (2014)
Lim, S. K. et al. Emergence of a ribotype 244 strain of Clostridium difficile associated with severe disease and related to the epidemic ribotype 027 strain. Clin. Infect. Dis. 58, 1723–1730 (2014)
Eyre, D. W. et al. Emergence and spread of predominantly community- onset Clostridium difficile PCR ribotype 244 infection in Australia, 2010 to 2012. Euro Surveill. 20, 21059 (2015)
Polivkova, S., Krutova, M., Petrlova, K., Benes, J. & Nyc, O. Clostridium difficile ribotype 176 – a predictor for high mortality and risk of nosocomial spread? Anaerobe 40, 35–40 (2016)
Rupnik, M. et al. Distribution of Clostridium difficile PCR ribotypes and high proportion of 027 and 176 in some hospitals in four South Eastern European countries. Anaerobe 42, 142–144 (2016)
Bergoz, R. Trehalose malabsorption causing intolerance to mushrooms. Report of a probable case. Gastroenterology 60, 909–912 (1971)
Bergoz, R., Bolte, J. P. & Meyer zum Bueschenfelde, K.-H. Trehalose tolerance test. Its value as a test for malabsorption. Scand. J. Gastroenterol. 8, 657–663 (1973)
Oku, T. & Nakamura, S. Estimation of intestinal trehalase activity from a laxative threshold of trehalose and lactulose on healthy female subjects. Eur. J. Clin. Nutr. 54, 783–788 (2000)
Higashiyama, T. Novel functions and applications of trehalose. Pure Appl. Chem. 74, 1263–1269 (2002)
Stabler, R. A. et al. Comparative genome and phenotypic analysis of Clostridium difficile 027 strains provides insight into the evolution of a hypervirulent bacterium. Genome Biol. 10, R102 (2009)
Leffler, D. A. & Lamont, J. T. Clostridium difficile infection. N. Engl. J. Med. 372, 1539–1548 (2015)
Theriot, C. M. et al. Antibiotic-induced shifts in the mouse gut microbiome and metabolome increase susceptibility to Clostridium difficile infection. Nat. Commun. 5, 3114 (2014)
Knetsch, C. W. et al. Genetic markers for Clostridium difficile lineages linked to hypervirulence. Microbiology 157, 3113–3123 (2011)
Bouillaut, L., Self, W. T. & Sonenshein, A. L. Proline-dependent regulation of Clostridium difficile Stickland metabolism. J. Bacteriol. 195, 844–854 (2013)
Ng, Y. K. et al. Expanding the repertoire of gene tools for precise manipulation of the Clostridium difficile genome: allelic exchange using pyrE alleles. PLoS ONE 8, e56051 (2013)
Fagan, R. P. & Fairweather, N. F. Clostridium difficile has two parallel and essential Sec secretion systems. J. Biol. Chem. 286, 27483–27493 (2011)
de Kok, S. De et al. Rapid and reliable DNA assembly via ligase cycling reaction. ACS Synth. Biol. 3, 97–106 (2014)
Pfaffl, M. W. in Real-time PCR (ed. Dorak, T. ) 63–82 (Taylor & Francis, 2006)
Collins, J., Auchtung, J. M., Schaefer, L., Eaton, K. A. & Britton, R. A. Humanized microbiota mice as a model of recurrent Clostridium difficile disease. Microbiome 3, 35 (2015)
Auchtung, J. M., Robinson, C. D., Farrell, K. & Britton, R. A. in Clostridium difficile: Methods and Protocols (eds Roberts, A. P. & Mullany, P. ) 235–258 (Springer, 2016)
Holm, S. A simple sequentially rejective multiple test procedure. Scand. J. Stat. 6, 65–70 (1979)
Griffiths, D. et al. Multilocus sequence typing of Clostridium difficile. J. Clin. Microbiol. 48, 770–778 (2010)
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011)
Roca, A. I., Abajian, A. C. & Vigerust, D. J. ProfileGrids solve the large alignment visualization problem: influenza hemagglutinin example. F1000 Res. 2, 2 (2013)
Dingle, T. C., Mulvey, G. L. & Armstrong, G. D. Mutagenic analysis of the Clostridium difficile flagellar proteins, FliC and FliD, and their contribution to virulence in hamsters. Infect. Immun. 79, 4061–4067 (2011)
Popoff, M. R., Rubin, E. J., Gill, D. M. & Boquet, P. Actin-specific ADP-ribosyltransferase produced by a Clostridium difficile strain. Infect. Immun. 56, 2299–2306 (1988)
Smith, C. J., Markowitz, S. M. & Macrina, F. L. Transferable tetracycline resistance in Clostridium difficile. Antimicrob. Agents Chemother. 19, 997–1003 (1981)
Acknowledgements
This work was supported by the National Institutes of Health (U01AI124290-01 and 5U19AI09087202). We thank the members of the Ostomy Association of Greater Lansing for anonymously donating samples, and D. Lyras for the RT244 strains. We thank V. Young, D. Mills, J. Walter and M. Costa-Mattioli for comments on the manuscript.
Author information
Authors and Affiliations
Contributions
Concept and design of study: R.A.B., J.M.A., J.C. and C.R. Experiments: C. difficile growth, J.C. and C.R.; identification of L172I SNP and comparative analysis, C.R. and J.C.; treA RT–qPCR, H.D.; mouse infection model, J.C.; genetic manipulation of C. difficile strains, J.C. and C.R.; identification of RT078 trehalose insertion, C.W.K., H.C.L. and T.D.L.; faecal minibioreactor competitions, J.M.A.; spontaneous C. difficile mutant identification, H.D.; analysis, J.C., C.R., H.D., J.M.A. and R.B. The manuscript was drafted by J.C., J.M.A., and R.A.B., and revised by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks J. Ballard, E. Pamer and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Phylogenetic organization of C. difficile MLST profiles.
Maximum likelihood tree based upon concatenated multi locus sequence typing genes of the 399 current profiles available at https://pubmlst.org/cdifficile/31. Stars indicate position of strains used in this study, with red stars indicating sequence types possessing either the TreR L172I amino acid substitution (ST1, ST41) or four-gene insertion (ST11). Tree constructed using MEGA7 (ref. 32).
Extended Data Figure 2 Growth of C. difficle strains.
a, The majority of strains can grow on 50 mM trehalose. Dashed grey line and band indicate mean growth and s.d. in DMM without a carbon source. Solid lines indicate mean growth yield (A600 nm) for groups: non-RT027/078 (n = 10), RT027 (n = 8), and RT078 (n = 3). b, Deletion of treA ablates the ability of both CD630 (RT12) and R20291 (RT027) to grow on trehalose. This phenotype can be restored by supplying treA on an inducible plasmid (n = 3 for each strain/group). c, RT244 strains (DL3110 and DL3111) possessing the TreR L172I mutation are capable of growth on 10 mM trehalose (n = 3 for each strain/group). d, CD1015∆ptsT can metabolize 50 mM trehalose (n = 4 for each strain/group). For a–d, points represent biologically independent samples, solid bars are means.
Extended Data Figure 3 RT027 strains express treA at a significantly higher level than non-RT027 strains in the presence of 25 mM trehalose.
Each data point (n = 4 ribotypes per group) represents gene expression from a different, biologically independent, strain and is an average from two to five independent experiments. P = 0.029, Mann–Whitney–Wilcoxon test (two-sided). Bar indicates mean expression.
Extended Data Figure 4 RT027 strains have an L172I mutation at a highly conserved site.
a, The treR genes from available C. difficile whole-genome sequencing files on the NCBI database (accessed 11 May 2017) were identified by tblastn and translated to protein sequences. Sequence fragments shorter than 240 amino acids were discarded and the remaining 1,010 sequences aligned with Clustal Omega33. All 191 sequences containing the L172I SNP also contained the thyA gene, a marker for the RT027 lineage; thyA was not found in any other genomes. Numbers indicate the number of sequences with a corresponding amino acid in that position. Multiple sequence alignment visualization generated with ProfileGrid34. b, The TreR protein sequence from RT027 strain R20291 was blasted against non-C. difficile sequences in the NCBI database and the top 99 matches (along with R20291treR) aligned with Clustal Omega. The leucine at position 172 was found to be conserved in 93 of 99 non-C. difficile sequences. To confirm the importance of this residue, TreR was blasted against all non-clostridial sequences in the NCBI database and the top 500 hits saved. After removal of duplicate species, 191 sequences were aligned with Clustal Omega. The leucine at residue 172 was conserved in 83% of sequences (data not shown).
Extended Data Figure 5 A treA knockout strain decreased toxin production 48 h after infection.
Mice were gavaged with 104 spores of either R20291 or R20291∆treA and provided with 5 mM trehalose in drinking water. Points represent toxin levels from individual mice (R20291 n = 10, R20291∆treA n = 11) euthanized 48 h after infection. Bars are means. Mice gavaged with R20291∆treA had significantly lower toxin levels (P = 0.0268; Wilcoxon–Mann–Whitney test (two-sided), median 40,960, IQR 23,040–46,080 versus 92,160, IQR 51,200–102,400).
Extended Data Figure 6 The four-gene trehalose insertion is only present in the RT078 lineage.
Artemis comparison tool displaying pairwise comparisons between C. difficile RT078 genome (M120) sequence and genome sequences from other C. difficile ribotypes (ribotypes indicated on the left). Numbers between grey bars indicate the genomic region where the trehalose four-gene insert is located (3231169–3237057). Regions of sequence homology are displayed in red. The trehalose four-gene insert of RT078 (indicated by the arrow on the top) was observed in RT078, but was absent in other ribotypes.
Supplementary information
Rights and permissions
About this article
Cite this article
Collins, J., Robinson, C., Danhof, H. et al. Dietary trehalose enhances virulence of epidemic Clostridium difficile. Nature 553, 291–294 (2018). https://doi.org/10.1038/nature25178
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25178
- Springer Nature Limited
This article is cited by
-
Non-canonical start codons confer context-dependent advantages in carbohydrate utilization for commensal E. coli in the murine gut
Nature Microbiology (2024)
-
Quantifying the adaptive landscape of commensal gut bacteria using high-resolution lineage tracking
Nature Communications (2024)
-
Elucidating human gut microbiota interactions that robustly inhibit diverse Clostridioides difficile strains across different nutrient landscapes
Nature Communications (2024)
-
Chronic Consumption of Trehalose Reduces Lifespan in Drosophila Model
Plant Foods for Human Nutrition (2023)
-
Particular genomic and virulence traits associated with preterm infant-derived toxigenic Clostridium perfringens strains
Nature Microbiology (2023)