Abstract
Locomotion is a universal behaviour that provides animals with the ability to move between places. Classical experiments have used electrical microstimulation to identify brain regions that promote locomotion1,2,3,4,5, but the identity of neurons that act as key intermediaries between higher motor planning centres and executive circuits in the spinal cord has remained controversial6,7,8,9,10,11,12,13,14. Here we show that the mouse caudal brainstem encompasses functionally heterogeneous neuronal subpopulations that have differential effects on locomotion. These subpopulations are distinguishable by location, neurotransmitter identity and connectivity. Notably, glutamatergic neurons within the lateral paragigantocellular nucleus (LPGi), a small subregion in the caudal brainstem, are essential to support high-speed locomotion, and can positively tune locomotor speed through inputs from glutamatergic neurons of the upstream midbrain locomotor region. By contrast, glycinergic inhibitory neurons can induce different forms of behavioural arrest mapping onto distinct caudal brainstem regions. Anatomically, descending pathways of glutamatergic and glycinergic LPGi subpopulations communicate with distinct effector circuits in the spinal cord. Our results reveal that behaviourally opposing locomotor functions in the caudal brainstem were historically masked by the unexposed diversity of intermingled neuronal subpopulations. We demonstrate how specific brainstem neuron populations represent essential substrates to implement key parameters in the execution of motor programs.
Similar content being viewed by others
Main
To visualize neurons in the brainstem with direct access to circuits in the spinal cord, we retrogradely infected neurons with G-deleted rabies viruses expressing fluorescent proteins (rabies-FP)15 (Fig. 1a). Three-dimensional reconstructions in a brainstem model revealed bilaterally positioned neurons in the caudal medulla (Fig. 1b, c; Extended Data Fig. 1), residing in neighbouring subdomains16, in agreement with previous conventional tracing experiments in mice17. Regions include the LPGi, the alpha part of the gigantocellular nucleus (GiA), and the ventral part of the gigantocellular nucleus (GiV), a trio encompassing the previously characterized magnocellular nucleus (Mc)18, the more dorsally positioned gigantocellular nucleus (Gi), and the midline-resident raphe nucleus (Fig. 1b, c; Extended Data Fig. 1). We focused our analysis on the Mc and Gi regions given their possible roles in mediating midbrain locomotor region (MLR)-induced locomotion4.
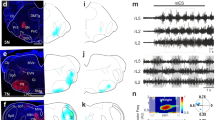
a, Retrograde labelling of spinally projecting neurons (PNs) in the brainstem medulla. b, c, Three-dimensional reconstruction of neurons projecting to the cervical spinal cord (bird’s eye and frontal view; c) and quantification (b) in raphe nucleus (blue), Gi (green) and Mc, subdivided into LPGi (magenta), GiA (cyan) and GiV (yellow). 7N, facial nucleus; 12N, hypoglossal nucleus; contra, contralateral; ipsi, ipsilateral; IO, inferior olive; pyr, pyramidal tract. 1 and 2 denote the rostral and caudal portion, respectively, of the 3D reconstruction shown in frontal view (right panels). d, e, Unconditional expression of ReaChR in the MLR or caudal brainstem neurons, and locomotor speed analysis in open field arena of stationary phase (before), upon triggering the laser (laser ON) and after laser offset (after), each during 1-s windows. Single-trial analysis for LPGi (d, top right; NS, not significant, one-way ANOVA/Bonferroni) and MLR (d, bottom right; *P < 0.05, ***P < 0.001, Friedman non-parametric test/Dunn) and speed analysis for LPGi, GiA, GiV, Gi and MLR (one circle per mouse; see Methods for laser intensity definitions; *P < 0.05, ***P < 0.001, one-way ANOVA/Bonferroni) (e). Data are mean ± s.e.m., for sample sizes see Supplementary Table 1.
To determine whether the stimulation of Mc subdomains or of Gi in mice can induce reliable full body locomotion, we injected adeno-associated viruses (AAVs) expressing the optogenetic activator ReaChR into a subset of mice (Fig. 1d). Unconditional light-induced activation of LPGi, GiA, GiV or Gi neurons failed to promote locomotion, but Gi stimulation induced ipsilateral head-turning (Fig. 1e; Extended Data Fig. 2a–d). MLR neuron stimulation produced reliable locomotion (Fig. 1d, e; Extended Data Fig. 2e). Together, these findings demonstrate that the unconditional Mc subdomain or Gi stimulation in mice fails to trigger full-body locomotion, as do medullary electrical microstimulation experiments6.
To determine whether locomotor properties may be masked by neuronal diversity, we used neurotransmitter identity to stratify neurons. We injected AAV-flex-fluorescent protein (FP) into vGlut2creGlyT2GFP mice (vGlut2 is also known as Slc17a6; GlyT2 is also known as Slc6a5) to reveal neuronal diversity and intraspinal injections of rabies-FP to mark spinal projection neurons (Fig. 2a). All subdomains showed intermingling of excitatory vGlut2-positive (vGlut2ON) and inhibitory GlyT2ON neurons containing spinal projection neurons (Fig. 2b), prompting us to assess separately the consequences of optogenetic activation of these diverse caudal medulla neurons.
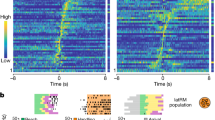
a, b, Neurotransmitter identity of spinal projection neurons in the LPGi, GiA and GiV (original magnification of images 20× objective). c–f, Locomotor speed and EMG analysis of mice expressing ReaChR in LPGi-vGlut2 neurons. c, Centre of body mass trajectories of single trials in open field arena during 1-s time windows: stationary phase (orange), laser application phase (cyan) and after laser offset (magenta). d, Speed versus time of single trials (grey lines) and the average (dotted black line) of one mouse. e, Triceps EMG analysis during laser-induced locomotion (right: latency analysis; mean and s.e.m. range are shown). f, Top, average speed analysis during and before laser application (*P < 0.05, **P < 0.01, ***P < 0.001, unpaired t-test). Middle, maximal speed analysis (*P < 0.05, Kruskal–Wallis/Dunn). Bottom, onset latency analysis (*P < 0.05, **P < 0.01, one-way ANOVA/Bonferroni). g, Average speed (left; **P < 0.01, ***P < 0.001, unpaired t-test) and latency to stop (right; *P < 0.05, unpaired t-test) after stimulation of LPGi-vGAT neurons during locomotion. h–j, Optogenetic stimulation of vGlut2 or inhibitory neurons in GiA (h), GiV (i) or Gi (j) as indicated (top left: single-trial locomotor trajectories for vGlut2; bar plots show average speed in open field 1 s before (orange dots) versus during (blue dots) laser application; **P < 0.01, ***P < 0.001, unpaired t-test). Data are mean ± s.e.m., for sample sizes see Supplementary Table 1.
We injected AAV-flex-ReaChR-YFP unilaterally, confined to specific Mc subdomains or Gi in vGlut2cre, vGATcre or GlyT2cre mice (Extended Data Figs 3, 4). Light pulses applied through optic fibres induced expression of Fos in ReaChRON neurons, providing evidence for neuronal activation (Extended Data Fig. 5a). Focusing first on excitatory neurons, we found that optogenetic stimulation of glutamatergic LPGi but not of GiA, GiV or Gi neurons caused initiation of forward-directed coordinated full-body locomotion in an open field arena (Fig. 2c, d, h–j; Extended Data Fig. 5; Gi-vGlut2 neuron stimulation induced ipsilateral head-turning; Supplementary Video 1). Notably, during induced locomotor episodes, mice were able to adjust trajectories and respect boundaries such as walls in the open field arena similar to natural locomotion (Extended Data Fig. 5i–k). Electromyographic (EMG) recordings in limb muscles of LPGi-vGlut2-ReaChR mice revealed short-latency onset signals (Fig. 2e; 20.1 ± 2.3 ms (mean ± s.e.m.); n = 3 mice), at latencies comparable to medullary electrical stimulation19,20, and EMGs as well as kinematics showed patterns similar to natural locomotion (Fig. 2e; Extended Data Fig. 5h, l, m). Moreover, increased laser intensities to LPGi-vGlut2-ReaChR neurons induced progressively higher speed locomotion initiated at shorter latencies (Fig. 2f; Extended Data Fig. 5d).
To determine whether LPGi-vGlut2 neuron activation can lead to sustained locomotion and increase of natural locomotor speed, we used running wheels, on which continuous unobstructed locomotion can be easily observed (Extended Data Fig. 5n, o). We found that mice maintained running throughout the duration of laser application with characteristic oscillatory locomotor bouts (Extended Data Fig. 5o). Laser application also induced further acceleration when mice were already engaged in natural wheel locomotion (Extended Data Fig. 5p). Together, these findings provide evidence that optogenetic activation of LPGi-vGlut2 neurons is sufficient to trigger continuous locomotion.
By contrast, optogenetic activation of LPGi-vGAT or LPGi-GlyT2 neurons, but not of LPGi-Gad65 neurons, induced locomotor halt in mice engaged in open field exploration or wheel running, showing speed and latency changes that scaled with laser intensity (Fig. 2g; Extended Data Fig. 6; Supplementary Video 2). Despite behavioural arrest, mice kept body muscle tone (Supplementary Video 2). Optogenetic activation of GlyT2ON neurons in GiA, GiV or Gi also resulted in efficient locomotor arrest (Fig. 2h–j; Extended Data Fig. 6h). However, whereas GiA-GlyT2 neuron activation induced behavioural stalling similar to LPGi-GlyT2 stimulation, GiV-GlyT2 neuron stimulation provoked body collapse akin to behaviour observed during atonia, and Gi-vGAT neuron stimulation produced arrest associated with body collapse and spasms (Supplementary Video 3). These findings, together with published work21, suggest that neurons with locomotion-suppressing roles are more distributed and functionally diverse than locomotion-promoting neurons in the caudal brainstem, prompting us to focus on the more distinctive and locomotion-promoting LPGi-vGlut2 population for additional functional studies.
We next scored immediate early gene Fos expression as a proxy for monitoring neuronal activity in mice engaged in locomotion. We found that the percentage of NeuNON neurons expressing Fos protein was significantly increased in all Mc subregions and Gi after locomotion compared to mice kept in their home cages (Extended Data Fig. 7a–c). Because Fos immunoreactivity can correlate with neuronal firing22, we quantified Fos expression levels and found a Fos high-intensity population specifically within the ventral LPGi subdomain upon locomotion (Extended Data Fig. 7d, e), suggesting that these LPGi neurons might be the most recruited during locomotion.
To study the endogenous role of locomotion-promoting LPGi-vGlut2 neurons, we injected an AAV that conditionally expresses the human diphtheria toxin receptor (DTR)18 into the LPGi of vGlut2cre mice to ablate them (Fig. 3a, b; Extended Data Fig. 8). We coinjected AAV-flex-ReaChR in a subset of mice to assess the correct targeting for the LPGi locomotor region functionally, using optogenetic stimulation before neuronal ablation (Fig. 3a, c). Intraperitoneal injection of diphtheria toxin A (DTA) resulted in eradication of these responses 7 days thereafter (Fig. 3c). We confirmed efficient anatomical ablation of LPGi-vGlut2 neurons and injection confinement to the targeted area (Extended Data Fig. 8).
a–c, Conditional ablation of LPGi-vGlut2 neurons. a, Experimental strategy applied. b, Injection site analysis. Amb, nucleus ambiguus. c, Test of ablation by optogenetic stimulation. ***P < 0.001, paired t-test. d, Treadmill position for mouse at 60 cm s−1 speed before (pre-DTA) and after (post-DTA) LPGi-vGlut2 neuron ablation (dotted line at position 50 denotes the back-end treadmill position cut-off to determine time, during which mice do not keep up with speed). e–g, Analysis of the percentage of time mice cannot keep up with reinforced speed at 20, 40, 60 and 80 cm s−1 for all mice derived from the LPGi-vGlut2 experimental group (e), the LPGi-vGlut2 control group (f), and a GiA-vGlut2 experimental group (g). *P < 0.05, ***P < 0.001, two-way ANOVA/Sidak’s post-test (post-DTA versus pre-DTA mice for each group). Data are mean ± s.e.m., for sample sizes see Supplementary Table 1.
To assay locomotor performance at different speeds in LPGi-vGlut2-DTR mice, we monitored individual mice before and after DTA injection on a speed-reinforcement treadmill. Mice accomplishing this task proficiently run mostly towards the treadmill front end (Fig. 3d). They fall back towards the end of the belt when they cannot keep up with the imposed speed, but rapidly resume to maintain the reinforced speed (Fig. 3d). These observations prompted us to quantify the time mice spend in a restricted back-end zone as a proxy of running performance (Fig. 3d, e). A control group injected in LPGi but in which AAV-flex-DTR was omitted, showed increased time spent in the back with higher speeds (20–80 cm s−1) consistent with increased task difficulty, but we observed no statistical differences between pre- and post-DTA performance (Fig. 3f). By contrast, although LPGi-vGlut2-DTR mice performed indistinguishably from the control group before DTA injection, after neuronal ablation, there was a significant decay in locomotor performance at higher speeds (40–80 cm s−1), but no significant effects were seen at 20 cm s−1 (Fig. 3e). These were specific to LPGi-vGlut2 neurons, as mice with GiA-vGlut2 neuron ablation scored as controls (Fig. 3f, g; Extended Data Fig. 8).
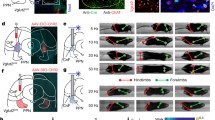
To determine how LPGi neuron subpopulations can elicit opposing locomotor effects, we next unravelled the broader circuitry into which these neurons are embedded. We observed axons of LPGi-vGlut2 and LPGi-GlyT2 neurons descending along the length of the spinal cord in predominantly ipsilateral trajectories (Fig. 4a–c; Extended Data Fig. 9a, b). Quantification of synaptic input to spinal grey matter circuits revealed an almost equal distribution to both spinal sides (Fig. 4b, c; Extended Data Fig. 9a, b). Both subpopulations targeted ventral spinal laminae in which rhythm- and pattern-generating interneurons reside23,24,25, but LPGi-vGlut2 synapses were concentrated mainly in the ventral central grey matter mostly avoiding motor neuron cell bodies, whereas LPGi-GlyT2 neurons contacted motor neurons directly (Fig. 4b–d; Extended Data Fig. 9a, b). LPGi injections into Gad65cre (also known as Gad2) mice failed to label spinally-projecting axons (data not shown), consistent with a lack of eliciting locomotor behaviour upon optogenetic activation, in agreement with recent work26.
a, Experimental strategy to map spinal output of LPGi neurons. INs, interneurons; MNs, motor neurons; NT, neurotransmitter. b, c, Mid-lumbar spinal cord sections depicting synaptic tag (SynTag) and ChAT labelling from LPGi-vGlut2 (b) or LPGi-GlyT2 (c) neurons (top), synaptic density and white matter axon tract position analysis (middle) and quantification (bottom). Percentages for ipsilateral spinal cords shown in pie charts are as follows: LPGi-vGlut2 (SynTag: 65.06 ± 0.40; white matter axons: 74.86 ± 10.75) and LPGi-GlyT2 (SynTag: 51.73 ± 2.32; white matter axons: 81.13 ± 8.68). d, Reconstruction (top) and quantification (bottom) of LPGi-vGlut2 and LPGi-GlyT2 synaptic input to motor neurons innervating hindlimb quadriceps (Q) or forelimb triceps (Tri) muscles. *P < 0.05, **P < 0.01, ***P < 0.001, unpaired t-test. e, Summary diagram displaying spinal cord output connectivity of LPGi-vGlut2 and LPGi-GlyT2 neurons. f–h, Mapping synaptic inputs from MLR-vGlut2 neurons to spinally projecting (SPN) Mc neurons. f, Experimental scheme. g, Reconstruction of synaptic input to LPGi neurons with neurotransmitter status. h, Left, synaptic input quantification. Right, positional map according to synaptic input density in the left panel, displayed by heat map (subdivision into dorsal and ventral LPGi is marked by line; x axis: distance from midline in mm, y axis: distance from ventral brainstem surface in mm). **P < 0.01, ***P < 0.001, one-way ANOVA/Bonferroni. i–k, LPGi-vGlut2 neuron ablation attenuates the effect of MLR locomotor signal. i, Experimental scheme. j, Speed evoked by MLR-vGlut2 neuron stimulation before and after LPGi-vGlut2 neuron ablation (group data, line denotes the mean, shaded area denotes the s.e.m.). k, Mean speed (left), maximal speed (middle) and reliability analysis (right) of locomotor response. **P < 0.01, ***P < 0.001, two-way ANOVA/Sidak. Data are mean ± s.e.m., for sample sizes see Supplementary Table 1.
We next mapped synaptic input structures to spinally projecting excitatory and glycinergic LPGi neurons (Extended Data Fig. 9c). Both LPGi subpopulations receive synaptic inputs from many upstream regions, most of which were previously implicated in motor control including superior colliculus, hypothalamus, periaqueductal grey, deep cerebellar nuclei, red nucleus, zona incerta and motor cortex (data not shown). Because MLR-vGlut2 neurons were described as being sufficient to evoke locomotion27,28, we focused our attention on synaptic interactions between MLR and LPGi-vGlut2 neurons. Using retrograde transsynaptic technologies, we found that LPGi-vGlut2 neurons receive input from both pedunculopontine nucleus (PPN) and cuneiform (CnF) region neurons (Extended Data Fig. 9c–e), with more neurons located in the CnF region. Most LPGi-connected PPN neurons were non-cholinergic (Extended Data Fig. 9e). Overall, MLR input appeared biased to LPGi-vGlut2 neurons (Extended Data Fig. 9f), but as rabies-tracing experiments are not of sufficient resolution to determine synapse-level biases precisely, we quantified synaptic input density derived from the general population of MLR-vGlut2 neurons onto spinally projecting Mc neurons. We found preferential contacts to putatively glutamatergic Mc-GlyT2OFF neurons (Fig. 4f–h; Extended Data Fig. 10a–c). Notably, MLR inputs were highest to neurons residing within the ventral LPGi subdomain (Fig. 4h), consistent with our Fos expression intensity analysis.
These findings prompted us to assess whether MLR-vGlut2 neuron stimulation influences locomotor speed through LPGi-vGlut2 neurons. We found that optogenetic stimulation of MLR-vGlut2 axon terminals in the Mc induces reliable locomotor speed increase in stationary mice (Extended Data Fig. 10d, f–h), indicating that stimulation of MLR-vGlut2 neurons projecting to the Mc is sufficient to carry locomotor signals. Finally, we devised an experiment ablating LPGi-vGlut2 neurons together with optogenetic stimulation of MLR-vGlut2 neurons (Fig. 4i; Extended Data Fig. 10e, i, j). We found that MLR-vGlut2 neuron stimulation generated significantly reduced locomotor speed after LPGi-vGlut2 neuron ablation (Fig. 4j, k), demonstrating that locomotion-promoting MLR signals pass through LPGi-vGlut2 neurons.
Many brain regions encode locomotor parameters including speed29,30, yet the identification of neuronal populations linking higher brain centres to the execution of locomotor programs has been sparse. As hypothesized14, disentangling neuronal diversity in the caudal brainstem was essential for its successful functional characterization. We found that unilateral optogenetic stimulation of glutamatergic or glycinergic LPGi neurons alone was sufficient to elicit fully bilateralized behavioural responses of two functionally opposing motor output programs that should normally not be coactive. This suggests that LPGi neurons on either body side have potent access to regulatory circuits coordinately controlling the entire body. It is currently unclear precisely how identified LPGi populations interact with other LPGi neurons, the brainstem and/or broader circuits including the spinal cord, but we found them to exhibit differential input-output circuitries. Cross-repressive synaptic interactions between these pathways may explain the behavioural choice of a motor program and prevent the occurrence of unwanted conflicting programs, thereby possibly also explaining how unconditional optogenetic coactivation might lead to behavioural cancelation (see also Supplementary Discussion). Our study provides key insight into the identity of core neuronal circuits regulating important locomotor parameters at the intersection between higher brain centres and executive spinal circuits.
Methods
Mice
The following transgenic mouse lines were used and maintained on a mixed genetic background (129/C57BL6): vGlut2cre (RRID: IMSR_JAX:028863)31, GlyT2cre32, vGATcre (RRID: IMSR_JAX:028862)31, Gad65cre33, GlyT2GFP (RRID: IMSR_RBRC04708)34. Wild-type mice (C57BL6) were from Charles River. Mice of different litters but same genotype were used in individual experiments. No criteria were used to allocate mice to experimental groups, and mice had unique identifiers for blinding. For all behavioural experiments, we used heterozygous males aged 2–3 months backcrossed to C57BL6. For anatomical experiments, mice of both sexes were used. Mice used for behavioural experiments were housed in cages with horizontal running wheels. Housing, surgery procedures, behavioural experiments and euthanasia were performed in compliance with the Swiss Veterinary Law guidelines.
Virus production and injections
G-deleted, rabies-mCherry or rabies-eGFP (rabies-FP) coated with G protein or EnvA-coated variants were amplified and purified from local stocks following established protocols35,36,37. The following AAVs used in this study were previously described18,38,39,40,41 and are based on a backbone vector derived from Allen Brain (AAV-CAG-flex-tdTomato-WPRE-bGH): AAV-flex-SynGFP and AAV-flex-SynMyc (referred to as AAV-flex-SynTag), AAV-flex-TdTomato, AAV-flex-DTR, AAV-flex-H2B-GFP-T2A-TVA, AAV-flex-H2B-V5-2A-G-protein. AAV-ConFon-H2B-GFP-T2A-TVA was designed using a described strategy42, and AAV-flex-FLP-H2B-V5 was designed in analogy to above constructs with an FLP sequence successfully used before38. For non-conditional expression of tagged markers to assess injection specificity, we used AAV-H2B-10×Myc or AAV-TdTomato (AAV-marker). AAV-flex-ReaChR-YFP was resynthesized according to published sequences43 and inserted into the same vector backbone as the other AAVs used in this study, and for unconditional expression in wild-type mice coinjected with AAV-Cre. For infections targeting neuronal cell bodies but not axons, a serotype plasmid 2.9 was used as in previous studies18,38,39,40, whereas for targeting neurons retrogradely through axonal infection, a recently developed rAAV2-retro capsid plasmid44 was used for coating. AAVs in our study were of genomic titres greater than 1 × 1013, and production followed standard protocols.
To infect and visualize neurons in the brainstem with projections to the cervical or lumbar spinal cord segments harbouring the majority of neurons innervating limb muscles, we carried out unilateral injections of rabies-FP into spinal segments C4–C8 (cervical) or L1–L4 (lumbar). These injections were complemented with beads to confirm injection laterality and mice were euthanized 4 days after rabies injection. Analogous injections with AAVs confirmed laterality of infection with here-applied methodology (data not shown). Retrograde labelling of motor neurons was performed as previously described18,39. For targeted delivery of viruses to the brainstem by stereotaxic injections, we used high precision instruments (David Kopf) under isofluorane anaesthesia as previously described18. Coordinates to target studied brain regions used lambda as a reference point for anterior–posterior (AP), medio-lateral (ML) and dorso-ventral (DV), and were as follows (AP; ML; DV; in mm): LPGi (−2; ± .1; −5.4); GiV (−2.3; ± 0.5; −5); GiA (−1.6; ± 0.4; -5.2); Gi (2; ± 0.5 or ± 0.7; −4.7 or −4.8; variations in coordinates used to cover different subregions of Gi); MLR (lambda suture; ± 1.2; −2.9). For synaptic input mapping to spinally projecting vGlut2- or GlyT2-expressing brainstem neurons, we targeted these neurons bilaterally from the cervical spinal cord with retro-AAV-flex-FLP in vGlut2cre or GlyT2cre mice, respectively, combined with LPGi-targeted expression of AAV-ConFon-H2B-GFP-T2A-TVA and AAV-flex-H2B-V5-2A-G-protein (H2B targets expressed proteins to the nucleus), and followed by a secondary brainstem injection with EnvA-rabies-FP two weeks later. For loss-of-function experiments, AAV-flex-DTR was injected at least 10 days before pre-DTA behavioural time points were acquired, and DTA (Sigma) was injected intraperitoneally at 100 ng g−1 body weight to initiate neuronal ablation18. All brain injection sites were carefully analysed anatomically after termination of experiments using ChAT immunohistochemistry to visualize motor nuclei and the mouse brain atlas was used as reference for determining spatial injection specificity16. Only mice with confirmed anatomical precision to targeted regions were included in subsequent analysis. Specifically, we used only injections in which the co-injected beads were within the targeted region, and in which >80% of the infected neurons were confined to the area delineated by the atlas boundaries of the corresponding structure. For delineation of Mc subdomains, we followed the boundaries of the mouse brain atlas16, and division of LPGi into dLPGi and vLPGi for Fos expression and MLR synaptic input analysis followed the demarcation of LPGiE in this atlas as guideline. The MLR subregions included in Extended Data Fig. 9 were defined according to the mouse brain atlas16. For the area named pedunculo-tegmental nucleus in this atlas, we used the more common nomenclature PPN (pedunculo-pontine nucleus), and the joint region of precuneiform area and cuneiform nucleus were annotated as CnF region. For MLR injections, we used broad AAV injections according to a functional MLR definition used previously27, including sites with infected neurons residing in PPN and CnF (centre of injection sites are shown in Extended Data Fig. 10).
Immunohistochemistry and microscopy
All mice used in this study were analysed by immunohistochemistry, including mice undergoing behavioural analysis to confirm injection site specificities. To prepare mice for immunohistochemistry, they were anaesthetized with a ketamine–xylazine solution and perfused transcardially with cold PBS, followed by a solution containing 4% paraformaldehyde (PFA) in PBS. Brains and spinal cords were isolated by dissection, post-fixed overnight in 4% PFA, and incubated in 30% sucrose (w/v) in PBS for cryopreservation for at least two days. All tissue was cut on a cryostat at 60–80-μm thickness and floating sections were collected in sequential order into individual wells (coronal for brain tissue and transverse for spinal cords). After 1-h incubation in blocking solution (1% BSA, 0.2% Triton X-100, PBS), primary antibodies were applied in blocking solution and incubated for 1–3 days at 4 °C. Fluorophore-coupled secondary antibodies (Jackson or Invitrogen) were applied to floating sections after extensive washing and incubated for 1 day at 4 °C. Sections were mounted after washing in anti-bleach preservative medium on slides in sequential order along the rostro-caudal axis. Primary antibodies used in this study were: chicken anti-GFP (Invitrogen), chicken anti-Myc (Invitrogen), goat anti-ChAT (Millipore), mouse anti-Myc (ATCC), mouse anti-NeuN (Millipore), mouse anti-V5 (Invitrogen), rabbit anti-Fos (Millipore), rabbit anti-RFP (Rockland). For low-resolution overview imaging, slides were scanned using an Axioscan light microscope (Zeiss, 5× objective). For higher resolution imaging, we used a FV1000 confocal microscope (Olympus) or a custom-made dual spinning disk microscope (Life Imaging Services GmbH).
Three-dimensional reconstructions of Mc and Gi
To assess the spatial distributions and quantitative contributions of spinally projecting neurons to the different subdivisions of the Mc, Gi and raphe nucleus, we acquired images of 80-μm coronal brainstem sections with the 20 × objective of a FV1000 confocal microscope using tiled mosaics (number of tiles varied according to the size of the medulla at different rostro-caudal levels) to cover the full medulla section (z-step = 4 μm). Images were stitched and the maximum intensity projection of each tile was used to produce a 3D model of the brain as previously described18. Cell body positions were assigned manually using IMARIS Spot function and following the Mc subdivision and Gi nomenclature of a widely used mouse brain atlas16.
Mapping synaptic input to LPGi subpopulations
Synaptic input mapping experiments to LPGi subpopulations were analysed 8 days after EnvA-rabies-FP injection. For identification of synaptic input structures to LPGi neurons, we used a common anatomy atlas16. To account for differences in spreading efficiency, we normalized the number of neurons located in the MLR to the one in the oral/caudal pontine reticular nucleus (PnO/C), for which the percentage of rabies-marked neurons of all analysed neurons was not significantly different between vGlut2cre and GlyT2cre mice. Moreover, only injection sites centred in LPGi were included in this analysis (n = 5 for each genotype).
Synaptic reconstructions of MLR input to spinally projecting Mc neurons
To quantify synaptic input of glutamatergic MLR neurons to spinally projecting Mc neurons, images were acquired ipsilateral to injection site using a custom-made dual spinning-disk microscope (60× objective; 0.2-μm steps) as previously described39. Complete cell bodies of spinally projecting rabies-FP neurons and proximal dendrites were reconstructed using Imaris with the manual surface module and glycinergic identity was assigned according to presence of GFP expression from the GlyT2GFP allele. Glutamatergic MLR synaptic appositions on the surface of spinally projecting Mc neurons were identified manually using Imaris spot detection function based on SynTag accumulation in contact with spinally projecting neurons (n = 3 mice). Images acquired with a 10× objective were used to identify the position of neurons scanned at high resolution.
Synaptic reconstructions of LPGi input to spinal motor neurons
Images were acquired ipsilateral to injection site using a custom-made dual spinning-disk microscope (60× objective; 0.2-μm steps). Density of synapses derived from LPGi-vGlut2 and LPGi-GlyT2 neurons in the spinal cord were quantified using Imaris spot detection function (n = 3 mice). Reconstruction of synaptic inputs to spinal motor neurons was performed as described39.
Treadmill and runway locomotion
We used a single lane treadmill with adjustable speed for our analysis (Panlab), set to zero slope and to shock delivery at the back-end at 0.6 mA. Before initiation of treadmill running sessions at different speed regimes, mice were handled on 3 days within 1 week for 10 min per day to acclimatize them to the room, treadmill and experimenter. On the first 2 days, mice were free to explore table and treadmill, and on the third day at the end of the session, mice were also put on the treadmill to run for 1 min at 20 cm s−1 speed to get used to the treadmill. During the subsequent sessions, mice were put on the treadmill for 1-min durations at each of the speed values of 20, 40, 60 and 80 cm s−1. Mice were allowed to recover from running by free exploration of the treadmill table for 1 min between different speed value sessions. Sessions were of shorter than 1-min duration in case mice reached a threshold criterion of >50 shock applications, or remained on the back-end grid of the treadmill for >5 s. Mice with poor running performance before DTA application, defined by values of >20% of testing time running within the 4.3 cm closest to the back-end grid location of the treadmill at 40 cm s−1, were not included in the analysis. Mice were tested three times at 1-week intervals before application of DTA (of which only the last two were included in the analysis), and 8, 15 and 22 days after DTA injection. During the testing sessions, the position of the mouse on the treadmill was tracked automatically, and the percentage of time (for each session) a mouse spent within the 4.3 cm closest to the back-end grid location of the treadmill was determined. For visualization purposes, the treadmill belt length was set to 450 units ( = 38 cm), and the cut-off for poor performance was at 50 units ( = 4.3 cm). For the analysis of mice for which sessions had to be terminated prematurely owing to poor performance, we added the remaining time of the trial as being within the defined back-end zone. The same treadmill was used for the analysis of Fos protein expression upon locomotion compared to home cage conditions. Mice were trained to run on the treadmill, and after several days of training as described above, they were put on the treadmill without application of shocks at speed values between 30 and 60 cm s−1 for 30 min. At the end of the treadmill session, mice were left undisturbed in their home cage, and perfused for analysis 90 min later. Kinematic recordings were carried out on a runway and monitored with a high-speed motion capture system (Vicon Motion Systems) as described40.
Running wheel experiments
Mice were left free to run on a standard horizontal running wheel, attached to a custom-made speedometer to analyse performance on the wheel. A similar wheel without speedometer is placed in home cages for at least 1 month before performance analysis in order for mice to voluntarily run smoothly on wheels. The output of the measuring device attached to the wheel is a continuous signal between 0.3 and 5 V with direct input to a Plexon system (Omniplex) for temporal synchronization with video acquisition and laser stimulation. In this assay, running wheel speed rather than actual animal speed is determined, also making it impossible to extract onset and offset latency of the animals according to real-time movement, as a result of the inertia of the wheel. Determination of running wheel speed values shown for experiments with 1-s laser application was restricted to a time window of 300 ms, to the phase just before laser onset (before) and the last 300 ms of laser application (ON). For experiments with 5-s laser application, we analysed the 300 ms before laser onset, individual seconds after laser onset during laser application over the entire time window, and the last 300 ms for the 1-s after laser offset. EMG recordings were carried out as previously described40, amplified (A-M Systems 1700, gain 100) and acquired with a Plexon system (Omniplex) at 1,000 Hz.
Open field assay
For optogenetic stimulation experiments, mice were analysed in a 35 × 35 cm open field arena and monitored with a top and side camera.
Optogenetic activation experiments
Mice in which we performed optogenetic activation of brainstem neurons received an implantation of optic fibres (diameter: 200 μm: MFC_200/230-0.48_Xmm_ZF1.25_FLT Mono Fibreoptic Cannula; X refers to fibre length according to stereotaxic coordinates; Doric lenses) placed 200–300 μm above the stereotaxic coordinates used for AAV injection, except for axon terminal stimulation of glutamatergic MLR neurons in the Mc, in which fibres were placed above Mc at the following coordinates: −1.8 mm, 1.5 mm, 5.2 mm with a lateral angle of 10°. Optic fibre implantations were carried out at least 1 week before the first stimulation experiments. To trigger optogenetic activation of brainstem neurons in the open field arena or on the running wheel, we used a PlexBright Optogenetic Stimulation System (Plexon), and monitored behaviour of mice simultaneously with two Pike cameras (top-down: TAMRON 8 mm, 018203; side-view: running wheel: model RICOH FL-HC6Z0810-VG; open field arena: model RICOH FL-HC0612A-VG) at 100 frames per second. The laser we used was a Cobolt 06-MLD; 473 nm; 100 mW. As control experiments for optogenetic activation, we also injected mice with AAV-flex-H2B-GFP-T2A-TVA (n = 3), with identical fibre implantation to the optogenetic activation group (Extended Data Fig. 5f). Laser onset was trigger manually at different laser intensities and frequencies, but when frequency is not otherwise specified, we used 100 Hz (5 ms-ON; 5 ms-OFF). The laser intensity was measured at the beginning of every testing day at the tip of an optic fibre of the same length as the one implanted to calibrate the system. As a control experiment with a few experimental mice, before termination of experiments and to assess whether fibre placement was accurate and suitable for stimulation of AAV-flex-ReaChR infected neurons, we stimulated neurons with 20 Hz (10 ms-ON; 40 ms-OFF), 15 mW pulsed blue light for 10 min, left the mice undisturbed in their home cages for 90 min, before perfusion and analysis of Fos expression as well as assessment of accuracy of injection and fibre placement. For MLR optogenetic stimulation experiments, we used 10 Hz square-pulses for 1 s and we assessed behaviour at progressively increasing laser intensities (0.1, 0.2, 0.5, 0.75, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 12.5, 15, 17.5 and 20 mW), each of them repeated for 10–20 trials until an intensity was reached at which mice run fast and reliably but not uncontrollably, ending the laser intensity ramp session.
Analysis of open field data and optogenetic stimulation experiments
Mice in the open field arena were tracked using the recordings of a top-down camera with Plexon and saved in avi format. Mice on the treadmill were tracked using the recordings of a side-view camera (Basler Camera; 200 Hz). All videos were subsequently cropped in size to regions of interest, divided into multiple shorter files using a MATLAB script, and processed using the machine learning software Ilastik (version 1.1.5). Specifically, for every acquisition day, a computational training session with refinement via machine learning was first used to instruct the software in order to detect the mouse from the background. The features used for this purpose were colour/intensity (Gaussian smoothing), edge (Gaussian gradient magnitude, difference of Gaussians) and texture (structure tensor eigenvalues, hessian of Gaussian eigenvalues). For each of the features, the probability was calculated using a sigma of 0.3, 1, 3.5 and 10 pixels. This training was used to create a probability map with the positional information of the mouse for each video controlling Ilastik in headless mode from the MATLAB environment.
Analysis of open field data and optogenetic stimulation experiments
We calculated the centre of body mass (COBM) of a mouse using the probability output file from Ilastik software with a custom-made MATLAB script. The COBM is the centroid of the filled area of the animal encoded in the probability file. The MLR analysis used the object contour tracking function from Cineplex Studio (Plexon; MLR analysis). Next, instantaneous animal speed in the open field was calculated from the extracted x–y position in the open field without smoothening and values were averaged for the analysed bins. For speed-versus-time plots, data were smoothened by averaging a moving window of 100 ms. To determine whether optogenetic stimulation or ablation of brainstem neurons affects locomotor parameters, COBM values were the input for our analysis. For analysis of optogenetic stimulation experiments, laser onset time stamps were extracted from plx files recorded by the Plexon system and imported into MATLAB for temporal alignment with the positional tracking information stored as COBM. We then analysed the instantaneous speed of a mouse based on its COBM position one second before, one second during and one second after laser stimulation. We quantified either the average speed or the maximum speed during the 1-s time window as specified in the corresponding figure legends. For every mouse assayed, at least 10 trials per laser intensity and frequency were analysed. To compare laser intensity effects across mice (Fig. 1e) or pre- versus post-DTA conditions (Fig. 4k), we determined the lowest laser intensity eliciting locomotion upon MLR neuron stimulation with 100% reliability as the maximum laser intensity used for experimental analysis. The minimum laser intensity included in our analysis was defined as the lowest laser intensity at which locomotion-promoting effects were detected for MLR neuron stimulation. Between these boundary values, we included either one (Fig. 1e) or two (Fig. 4k) additional in-between laser intensities in our analysis. For Mc subdomain and Gi stimulation (Fig. 1e), we used 5, 10 and 20 mW for these three categories. The same laser intensities were used for stimulation before and after ablation of LPGi-vGlut2 neurons described in Fig. 4i–k. To control for potential variability across days, mice were tested three times before DTA administration and three times after at 7, 10 and 14 days after DTA injection (Fig. 4i–k). To visualize the effect of laser stimulation of brainstem neurons on locomotion, we aligned the position of analysed trials of an experiment to the same point in space, allowing us to display the 2D trajectories of mice travelled before, during and after the 1-s laser window graphically. To determine the latency of movement onset, the duration of induced locomotion and the latency to movement termination from laser offset in optogenetic activation experiments of glutamatergic neurons, we manually analysed side-view videos recorded in the open field arena of laser-induced locomotion for at least 10 individual trials in several animals. Latency to start was defined as the duration between laser onset and the first detectable movement of the paw; latency to stop was defined as the time between laser offset and when the four paws are back on the ground; duration of induced locomotion was calculated as the time between the first movement of the paw and the time when the four paws are back on the ground for each individual trial. For experiments with optogenetic activation of inhibitory neurons, latency was defined as the duration between laser onset and the termination of the last step reaching behavioural arrest with all four limbs on ground.
Locomotion directional analysis in the open field
Our analysis was restricted to locomotor bouts defined as events in which animals maintain a speed above 5 cm s−1 for more than 200 ms, for laser-induced trials or natural locomotor bouts. To quantify the directionality of laser-induced locomotion, we compared the trajectory angles with the ones of freely moving animals in the open field (Extended Data Fig. 5i–k). We used the COBM to determine the frame-by-frame position of a mouse and calculated the trajectory angle at each individual frame with respect to the previous one (Extended Data Fig. 5i, α2−α1). The frequency of angles is given by the difference between consecutive angles. Normalization was applied to compare this distribution among locomotor bouts of different length. We also calculated the ratio between the length of each locomotor trajectory and the shortest distance between the starting and end position of a locomotor bout (Extended Data Fig. 5i). We calculated this ratio for both natural and laser-induced locomotion and find comparable values (Extended Data Fig. 5k).
Treadmill analysis
Mice on the treadmill were tracked using a side-view camera (Basler Camera; 200 Hz) and subsequently analysed as described above for the open field data. The Ilastik probability file was used to extract the COBM of a mouse for each video frame using a custom-made MATLAB script, defining the boundaries of the treadmill belt (front and back). This information was then used to calculate the time a mouse spent below the defined threshold for every trial (see above).
Quantification of Fos data
Images were acquired using an Olympus FluoView FV1000 confocal microscope with a 20× objective and z-steps of 1 μm. Tiled z-stack mosaics encompassing the medulla were used to automatically detect NeuNON and FosON cells using a custom-built workflow in KNIME45. In brief, z-stack pictures were merged in a maximal intensity projection to define Mc subdivisions and Gi manually according to common nomenclature16. The coordinates of these files with marked regions of interest were then used to crop the original z-stack into defined subregions. The number and positions of NeuNON and FosON cells were subsequently extracted upon manual confirmation of accurate detection. We performed intensity measurements for Fos by determining average pixel intensity within a defined nucleus. We used a threshold of 2,000 as gate for high-intensity Fos neurons in our analysis, selected because 5% of Fos and NeuN neurons in cage control mice were at or above this value. Automatic spot detection performance for Fos was validated manually on every section. Our analysis included sections at rostro-caudal levels depicted in the mouse brain atlas16 on figures 79–90.
Statistics and data availability
Significance levels indicated are as follows: *P < 0.05, **P < 0.01, ***P < 0.001. All data are presented as mean ± s.e.m. For each dataset, normal distribution was checked using the D’Agostino–Pearson, the Shapiro–Wilk or the Kolmogorov–Smirnov normality tests, or normality was tested using quantile plots to choose the appropriate statistical test. In case the requirements for none of these tests were fulfilled, we used a non-parametric Kruskal–Wallis test for non-matched data or a Friedman test for repeated measures. All statistical tests used were two-tailed. The experiments were not randomized and no statistical methods were used to predetermine sample size. All further statistical tests used in this study are spelled out in the corresponding figure legends, and details on all statistical tests can also be found in Supplementary Table 1. We used GraphPad PRISM version 6.04, R 3.3.2 or MATLAB 2015b for generation of graphs and statistics. Custom-made scripts described in this manuscript are available upon request. All other data are available from the corresponding author upon reasonable request.
References
Shik, M. L. & Orlovsky, G. N. Neurophysiology of locomotor automatism. Physiol. Rev. 56, 465–501 (1976)
Skinner, R. D. & Garcia-Rill, E. The mesencephalic locomotor region (MLR) in the rat. Brain Res. 323, 385–389 (1984)
Mori, S., Matsuyama, K., Mori, F. & Nakajima, K. Supraspinal sites that induce locomotion in the vertebrate central nervous system. Adv. Neurol. 87, 25–40 (2001)
Noga, B. R., Kriellaars, D. J., Brownstone, R. M. & Jordan, L. M. Mechanism for activation of locomotor centers in the spinal cord by stimulation of the mesencephalic locomotor region. J. Neurophysiol. 90, 1464–1478 (2003)
Jordan, L. M., Liu, J., Hedlund, P. B., Akay, T. & Pearson, K. G. Descending command systems for the initiation of locomotion in mammals. Brain Res. Rev. 57, 183–191 (2008)
Drew, T. & Rossignol, S. Functional organization within the medullary reticular formation of intact unanesthetized cat. I. Movements evoked by microstimulation. J. Neurophysiol. 64, 767–781 (1990)
Kinjo, N. et al. Medioventral medulla-induced locomotion. Brain Res. Bull. 24, 509–516 (1990)
Noga, B. R., Kettler, J. & Jordan, L. M. Locomotion produced in mesencephalic cats by injections of putative transmitter substances and antagonists into the medial reticular formation and the pontomedullary locomotor strip. J. Neurosci. 8, 2074–2086 (1988)
Garcia-Rill, E. & Skinner, R. D. The mesencephalic locomotor region. I. Activation of a medullary projection site. Brain Res. 411, 1–12 (1987)
Ross, G. S. & Sinnamon, H. M. Forelimb and hindlimb stepping by the anesthetized rat elicited by electrical stimulation of the pons and medulla. Physiol. Behav. 33, 201–208 (1984)
Hajnik, T., Lai, Y. Y. & Siegel, J. M. Atonia-related regions in the rodent pons and medulla. J. Neurophysiol. 84, 1942–1948 (2000)
Mori, S. Integration of posture and locomotion in acute decerebrate cats and in awake, freely moving cats. Prog. Neurobiol. 28, 161–195 (1987)
Takakusaki, K., Chiba, R., Nozu, T. & Okumura, T. Brainstem control of locomotion and muscle tone with special reference to the role of the mesopontine tegmentum and medullary reticulospinal systems. J. Neural Transm. (Vienna) 123, 695–729 (2016)
Orlovsky, G. N ., Deliagina, T. G. & Grillner, S. Neuronal Control of Locomotion: from Mollusc to Man Ch. 12, 205–214 (Oxford Univ. Press, 1999)
Wickersham, I. R., Finke, S., Conzelmann, K. K. & Callaway, E. M. Retrograde neuronal tracing with a deletion-mutant rabies virus. Nat. Methods 4, 47–49 (2007)
Franklin, K. B. & Paxinos, G. The Mouse Brain in Stereotaxic Coordinates 3rd edn (Elsevier, 2007)
Liang, H., Paxinos, G. & Watson, C. Projections from the brain to the spinal cord in the mouse. Brain Struct. Funct. 215, 159–186 (2011)
Esposito, M. S., Capelli, P. & Arber, S. Brainstem nucleus MdV mediates skilled forelimb motor tasks. Nature 508, 351–356 (2014)
Drew, T. Functional organization within the medullary reticular formation of the intact unanesthetized cat. III. Microstimulation during locomotion. J. Neurophysiol. 66, 919–938 (1991)
Drew, T. & Rossignol, S. Functional organization within the medullary reticular formation of intact unanesthetized cat. II. Electromyographic activity evoked by microstimulation. J. Neurophysiol. 64, 782–795 (1990)
Bouvier, J. et al. Descending command neurons in the brainstem that halt locomotion. Cell 163, 1191–1203 (2015)
Schoenenberger, P., Gerosa, D. & Oertner, T. G. Temporal control of immediate early gene induction by light. PLoS One 4, e8185 (2009)
Grillner, S. Biological pattern generation: the cellular and computational logic of networks in motion. Neuron 52, 751–766 (2006)
Kiehn, O. Decoding the organization of spinal circuits that control locomotion. Nat. Rev. Neurosci. 17, 224–238 (2016)
Goulding, M. Circuits controlling vertebrate locomotion: moving in a new direction. Nat. Rev. Neurosci. 10, 507–518 (2009)
Weber, F. et al. Control of REM sleep by ventral medulla GABAergic neurons. Nature 526, 435–438 (2015)
Lee, A. M. et al. Identification of a brainstem circuit regulating visual cortical state in parallel with locomotion. Neuron 83, 455–466 (2014)
Roseberry, T. K. et al. Cell-type-specific control of brainstem locomotor circuits by basal ganglia. Cell 164, 526–537 (2016)
Kropff, E., Carmichael, J. E., Moser, M. B. & Moser, E. I. Speed cells in the medial entorhinal cortex. Nature 523, 419–424 (2015)
Fuhrmann, F. et al. Locomotion, theta oscillations, and the speed-correlated firing of hippocampal neurons are controlled by a medial septal glutamatergic circuit. Neuron 86, 1253–1264 (2015)
Vong, L. et al. Leptin action on GABAergic neurons prevents obesity and reduces inhibitory tone to POMC neurons. Neuron 71, 142–154 (2011)
Foster, E. et al. Targeted ablation, silencing, and activation establish glycinergic dorsal horn neurons as key components of a spinal gate for pain and itch. Neuron 85, 1289–1304 (2015)
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuron 71, 995–1013 (2011)
Zeilhofer, H. U. et al. Glycinergic neurons expressing enhanced green fluorescent protein in bacterial artificial chromosome transgenic mice. J. Comp. Neurol. 482, 123–141 (2005)
Osakada, F. & Callaway, E. M. Design and generation of recombinant rabies virus vectors. Nat. Protocols 8, 1583–1601 (2013)
Stepien, A. E., Tripodi, M. & Arber, S. Monosynaptic rabies virus reveals premotor network organization and synaptic specificity of cholinergic partition cells. Neuron 68, 456–472 (2010)
Wickersham, I. R. et al. Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53, 639–647 (2007)
Pivetta, C., Esposito, M. S., Sigrist, M. & Arber, S. Motor-circuit communication matrix from spinal cord to brainstem neurons revealed by developmental origin. Cell 156, 537–548 (2014)
Basaldella, E., Takeoka, A., Sigrist, M. & Arber, S. Multisensory signaling shapes vestibulo-motor circuit specificity. Cell 163, 301–312 (2015)
Takeoka, A., Vollenweider, I., Courtine, G. & Arber, S. Muscle spindle feedback directs locomotor recovery and circuit reorganization after spinal cord injury. Cell 159, 1626–1639 (2014)
Satoh, D., Pudenz, C. & Arber, S. Context-dependent gait choice elicited by EphA4 mutation in Lbx1 spinal interneurons. Neuron 89, 1046–1058 (2016)
Fenno, L. E. et al. Targeting cells with single vectors using multiple-feature Boolean logic. Nat. Methods 11, 763–772 (2014)
Lin, J. Y., Knutsen, P. M., Muller, A., Kleinfeld, D. & Tsien, R. Y. ReaChR: a red-shifted variant of channelrhodopsin enables deep transcranial optogenetic excitation. Nat. Neurosci. 16, 1499–1508 (2013)
Tervo, D. G. et al. A designer AAV variant permits efficient retrograde access to projection neurons. Neuron 92, 372–382 (2016)
Dietz, C. & Berthold, M. R. KNIME for Open-Source Bioimage Analysis: A Tutorial. Adv. Anat. Embryol. Cell Biol. 219, 179–197 (2016)
Acknowledgements
We are grateful to M. Sigrist, M. Mielich, M. Pinto, J. Alonso, C. Hartmann and M. Theodore for experimental help, P. Tovote for advice with optogenetic technologies, S. Valencia Garcia and E. Arteaga Bracho for advice on EMG recordings, E. Jankowska for advice on cat brainstem nomenclature, A. Karpova for sharing the rAAV2-retro capsid plasmid before publication, R. Thierry and J. Eglinger from the FMI imaging facility and N. Ehrenfeuchter from the Biozentrum imaging facility for help and advice with image acquisition and analysis, P. Argast and P. Buchmann from the FMI mechanical workshop for building devices for behavioural experiments, M. Stadler and D. Gaidatzis for help with statistical analysis, and P. Caroni for discussions and comments on the manuscript. All authors were supported by an ERC Advanced Grant, the Swiss National Science Foundation, Kanton Basel-Stadt and Novartis Research Foundation, and the Louis Jeantet Prize for Medicine. M.S.E. was also supported by an HFSP long-term postdoctoral fellowship, Synapsis Foundation Grant and NARSAD Young Investigator Grant by the Brain and Behavior Foundation.
Author information
Authors and Affiliations
Contributions
All authors were involved in the design of experiments. P.C. together with C.P. carried out most experiments, acquired and analysed data. M.S.E. carried out experiments related to MLR and MLR–Mc interactions, was involved in EMG and kinematic experiments, acquired and analysed data. S.A. initiated the project, designed experiments, analysed data and wrote the manuscript. All authors discussed the experiments and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Subdivision of lumbar projection neurons in caudal medulla.
Views of three-dimensional reconstruction of rabies-FP-marked cell body positions of neurons projecting to the lumbar spinal cord, depicted with a colour code for different regions as described in Fig. 1b, c.
Extended Data Figure 2 Injection site location for unconditional optogenetic stimulation of brainstem neurons.
a–e, Analysis of injection and corresponding optic fibre positions targeting LPGi (a), GiA (b), GiV (c), Gi (d) and MLR (e) in wild-type mice is shown on brain atlas sections aligning with sites identified in corresponding experiments. Crosses depict centre of injections and rectangles show optic fibre tip positions, with each colour representing a different mouse included in the analysis shown in Fig. 1e. Bottom row shows representative pictures of YFP fluorescence after injections of AAV-flex-ReaChR-YFP. The mouse brain atlas images in this figure have been reproduced with permission from Elsevier.
Extended Data Figure 3 Injection site location for optogenetic stimulation of glutamatergic Mc and Gi neurons.
a–d, Analysis of injection and corresponding optic fibre positions targeting LPGi (a), GiA (b), GiV (c) and Gi (d) in vGlut2cre mice is shown on brain atlas sections aligning with sites identified in corresponding experiments. Crosses depict centre of injections and rectangles show optic fibre tip positions, with each colour representing a different mouse included in the analysis shown in Fig. 2 and Extended Data Fig. 5. Top row shows representative pictures of YFP fluorescence after injections of AAV-flex-ReaChR-YFP, and neighbouring motor nuclei (red). The mouse brain atlas images in this figure have been reproduced with permission from Elsevier.
Extended Data Figure 4 Injection site location for optogenetic stimulation of inhibitory Mc and Gi neurons.
a–d, Analysis of injection and corresponding optic fibre positions targeting LPGi (a), GiA (b), GiV (c) and Gi (d) in mice expressing Cre recombinase from loci of inhibitory neurotransmitters is shown on brain atlas sections aligning with sites identified in corresponding experiments. Crosses, circles and stars depict centre of injections in different genotypes and rectangles show optic fibre tip positions (legend bottom left), with each colour representing a different mouse included in the analysis shown in Fig. 2 and Extended Data Fig. 6. The mouse brain atlas images in this figure have been reproduced with permission from Elsevier.
Extended Data Figure 5 Stimulation of glutamatergic LPGi neurons elicits locomotion.
a, Top, stereotaxic injection of AAV-flex-ReaChR into LPGi, GiA, GiV or Gi of vGlut2cre mice. Bottom, blue laser stimulation of infected neurons induces Fos expression as proxy for neuronal activation in LPGi or GiA neurons. b, Centre of body mass trajectories of single trials in open field arena during two 1-s time windows: stationary phase (before, orange) and laser application phase (laser ON, cyan) centred to the starting position (top; 0.1 and 2 mW laser power). c, d, Average speed of single trials with 2 mW laser intensity (c), maximal speed during laser ON (d; left) and latency to initiate locomotion after laser onset (d; right) at different laser intensities for one representative example mouse. *P < 0.05, ***P < 0.001, one-way ANOVA/Bonferroni. e, Example of instantaneous speed of single mice (grey lines) and average of 4 mice (magenta) during 1-s optogenetic stimulation of LPGi-vGlut2 neurons (blue box), as well as 1 s before and after laser stimulation. f, A group of mice was injected with AAV-flex-GFP in LPGi as a control experiment (analysis as shown in Fig. 2h–j). g, Histogram of duration (left) and latency to stop (right; 0 s denotes laser offset) of individual locomotor bouts pooled from 3 mice induced by optogenetic stimulation of LPGi-vGlut2 neurons. h, Triceps EMG analysis during natural running wheel locomotion of LPGi-vGlut2 mice (compare to Fig. 2e). i–k, Direction of natural (NL) or laser-induced locomotion analysis in the open field arena (see Methods for analysis details). i, Experimental scheme to describe applied analysis (i, left) and frequency plot of angles (shown angle range covers 99% of all analysed trials) for a representative animal (i, middle) and over 3 animals (i, right). j, Percentage of angles in the range between −6 and +6°. k, Ratio of shortest to real distance travelled calculated as indicated in i. l, m, Kinematic analysis of natural and laser-induced locomotion of vGlut2cre mouse injected with AAV-flex-ReaChR-YFP in LPGi at a 40–50 cm s−1 speed, including principle component analysis (l; grey denotes swing phase; black denotes stance phase; n.s., not significant, non parametric t-test) and hindlimb (HL) and forelimb (FL) oscillations (m; grey box denotes swing phase of left (L) hindlimb). n–p, Analysis of running wheel (RW) locomotion in vGlut2cre mice with ReaChR expression targeted to LPGi neurons. n, o, In stationary mice, the application of laser light triggers locomotion during 1-s (n) or 5-s (o) laser stimulation (single trials are in grey, the average of analysed trials is in magenta). AU, arbitrary units. Plots at the bottom show the increasing running wheel speed for higher laser intensities of a single animal (left; **P < 0.01, ***P < 0.001, paired t-test) or group data (right; ***P < 0.001, two-way ANOVA/Bonferroni) upon 1-s stimulation (n), and maintained locomotion throughout 5 s of stimulation (o). p, For a LPGi-vGlut2-ReaChR mouse already running on the wheel, the application of a 5-mW laser can further increase speed. ***P < 0.001, paired t-test. q, Single trial average speed profiles to determine the effects of laser stimulation of vGlut2ON GiA (top), GiV (middle) or Gi (bottom) neurons on locomotor speed (1 s before, during and after laser). *P < 0.05, one-way ANOVA/Bonferroni). Data are mean ± s.e.m., for sample sizes see Supplementary Table 1.
Extended Data Figure 6 Stimulation of inhibitory Mc and Gi neurons.
a, b, Locomotor speed analysis of mice expressing ReaChR in LPGi-vGAT (a) or LPGi-GlyT2 (b) neurons in the open field arena. Left, trajectories of centre of body mass tracking for individual trials at 5 mW laser intensity 1 s before (orange), 1 s during (cyan), and 1 s after (magenta) laser application are shown aligned to a central point. Right, representative single trial speed analysis 1 s before and during laser application. ***P < 0.001, paired t-test. c, Top, example of single trials (grey lines) and average (dotted black line) of one mouse. Bottom, group data depicting averages of single mice (grey lines) and of 3 mice (magenta) during 1-s optogenetic stimulation of LPGi-GlyT2 neurons (blue box), as well as 1 s before and after laser stimulation. d, Average speed before (orange) and during (cyan) 1-s laser application at different laser intensities upon stimulation of LPGi-GlyT2 neurons for different mice analysed. *P < 0.05, **P < 0.01, unpaired t-test comparing data before and during laser application at each intensity. e, Locomotor speed analysis of mice expressing ReaChR in LPGi-Gad65 neurons in the open field arena. Left, trajectories of centre of body mass tracking for individual trials at 20 mW laser intensity 1 s before (orange) and 1 s during (cyan) laser application are shown aligned to a central point. Right, speed analysis 1 s before and during laser application. Not significant, one-way ANOVA/Bonferroni. f, g, Analysis of running wheel locomotion experiment in GlyT2cre (f) or Gad65cre (g) mice with ReaChR expression targeted to LPGi neurons. *P < 0.05, **P < 0.01, two-way ANOVA/Bonferroni. The application of laser light to GlyT2cre but not to Gad65cre mice running on the wheel triggers speed decrease, and higher laser intensities have a stronger impact. h, Representative single trial and group data speed analysis in the open field arena for optogenetic activation of GiA-GlyT2 (left), GiV-GlyT2 (middle) and Gi-vGAT (right) neurons 1 s before and during laser application. **P < 0.01, ***P < 0.001, unpaired t-test. Data are mean ± s.e.m., for sample sizes, see Supplementary Table 1.
Extended Data Figure 7 Fos expression levels in Mc subdomains and Gi after treadmill locomotion.
a, Schematic depiction of experimental strategy. Cage control mice are compared to mice running on a treadmill for 30 min, after which they are left undisturbed in the home cage before termination of the experiment and analysis of Fos expression. b, Representative example for Fos expression in cage control (left) compared to run (right) mice shown at similar rostro-caudal brainstem level at which optogenetic stimulation of LPGi-vGlut2 neurons elicits locomotion. c, Quantification of Fos-expressing neurons normalized to NeuN-expressing neurons on the same sections demonstrates a higher percentage of neurons in Mc subregions and Gi upon running. **P < 0.01, ***P < 0.001, unpaired t-test. d, e, Quantification of Fos intensity of analysed neurons, showing example neurons with corresponding pixel average values (d; left), and the percentage of Fos neurons with intensity values greater than 2,000, normalized to NeuNON neuronal number (d, right; n as in c; ***P < 0.001, two-way ANOVA/Bonferroni), and frequency distribution of Fos expression for different caudal brainstem regions analysed (e). dLPGi, dorsal LPGi; vLPGi, ventral LPGi. The cut-off used for the bar plot in d is shown in magenta (see Methods). Data are mean ± s.e.m., for sample sizes see Supplementary Table 1.
Extended Data Figure 8 Injection site location for loss-of-function experiments.
a–c, Analysis of injection positions targeting LPGi (a, b) or GiA (c) for loss-of-function experiments. b, For quantification of ablation efficiency, we used the injection of an AAV-flex marker (known to target most vGlut2-expressing neurons at the injection site) in mice without (left) compared to with (middle) coinjection of AAV-flex-DTR two weeks after DTA application. Note that 96.9% of targeted neurons are ablated with this strategy compared to corresponding reference control injections (right). Crosses in panels a and c depict centre of injections on brain atlas sections aligning with sites identified in corresponding experiments, with each colour representing a different mouse included in the analysis shown in Fig. 3. Data are mean ± s.e.m., for sample sizes see Supplementary Table 1. The mouse brain atlas images in this figure (a, c) have been reproduced with permission from Elsevier.
Extended Data Figure 9 Input and output circuitry of LPGi subpopulations.
a, b, Representative spinal cord sections depicting SynTag and ChAT labelling from LPGi-vGlut2 (a) or LPGi-GlyT2 (b) neurons for motor neurons (top), synaptic density analysis and white matter axon tract position (middle), and quantification of ipsilateral and contralateral synapse and axon fractions (bottom) at caudal cervical spinal cord levels. Percentages for ipsilateral spinal cords shown in the pie charts are as follows: LPGi-vGlut2 (SynTag: 59.97 ± 0.64; white matter axons: 74.80 ± 0.96), and LPGi-GlyT2 (SynTag: 54.15 ± 1.55; white matter axons: 79.21 ± 1.69). c, Schematic diagram of experimental strategy to map synaptic input to spinally projecting LPGi-vGlut2 and LPGi-GlyT2 neurons in vGlut2cre and GlyT2cre mice. NT, neurotransmitter. In a first injection, LPGi neurons are infected retrogradely from the spinal cord by a retro-AAV-flex-FLP, and locally by coinjection of AAV-ConFon-TVA and AAV-flex-G. In a second injection, LPGi is injected locally with EnvA-rabies-FP. d, e, Distribution of neurons within the MLR connected to LPGi-vGlut2 neurons. d, Images show example pictures of CnF neurons (not co-expressing ChAT) and PPN neurons, a minority of which is cholinergic (pie chart in e; percentage: 21.8 ± 7.2). e, Representative distribution of visualized neurons in CnF and PPN neurons overlaid on atlas section. f, Quantification demonstrating connectivity bias of MLR neurons to LPGi-vGlut2 compared with LPGi-GlyT2 neurons. *P < 0.05, unpaired t-test. Data are mean ± s.e.m., for sample sizes see Supplementary Table 1. The mouse brain atlas image in this figure has been reproduced with permission from Elsevier.
Extended Data Figure 10 Glutamatergic LPGi neurons mediate MLR motor commands.
a, b, Representative injection site (a) and projection pattern (b) of MLR-vGlut2 neurons labelled with SynTag. c, Single plane image of a spinally projecting vLPGi-vGlut2 dendrite with opposing MLR-vGlut2 synapses. d, e, Analysis of injection and corresponding optic fibre positions performed to optogenetically stimulate MLR-vGlut2 neurons (d, left) or their synaptic terminals in the Mc (d, right), and to determine the effect of LPGi-vGlut2 neuron ablation in response to MLR-vGlut2 light-induced locomotion (e), shown on brain atlas sections aligning with sites identified in corresponding experiments. Crosses depict centre of injections and rectangles show optic fibre tip positions, with each colour representing a different mouse included in the analysis shown in Fig. 4 and in g–j. f, Experimental scheme to stimulate MLR-vGlut2 neurons or their synaptic terminals optogenetically in the Mc using two optic fibres. g, Single trial speed analysis of stationary mouse for 1 s before laser onset (orange) compared to 1 s during laser application (cyan) for stimulations in the MLR (left) and terminals over the Mc (right). ***P < 0.001, paired t-test. h, Correlation between laser power and evoked speed upon stimulation of MLR-vGlut2 cell bodies (magenta) or axon terminals (cyan) over the Mc (different symbols indicate different mice). Note that cell body stimulation elicits stronger locomotor responses, but in both cases, higher laser intensities elicit higher speed locomotion (least square linear regression through origin followed by extra sum-of-squares F-test comparison; MLR slope: 42.41 ± 9.60, Mc slope: 3.70 ± 0.66, P < 0.0001). i, Speed versus time traces for single trials (grey lines) and average (dotted black line) of one representative mouse. j, Histogram of latency to start (left), duration of individual locomotor bouts (middle), and latency to stop (right) induced by MLR-vGlut2 neuron stimulation at the minimum laser intensity needed to evoke maximum reliability. Data are mean ± s.e.m., for sample sizes see Supplementary Table 1. The mouse brain atlas images in this figure have been reproduced with permission from Elsevier.
Supplementary information
Supplementary Information
This file contains additional discussion to the work presented in this study. It also contains Supplementary Table 1 which shows detailed statistical tests and sample sizes used in this study described in the accompanying main and extended data figures. (PDF 683 kb)
Stimulation of excitatory neurons in the caudal brainstem
Representative examples of video sequences showing the behavioral effects of mice in the open field arena upon laser application to LPGi-vGlut2, GiA-vGlut2, GiV-vGlut2, and Gi-vGlut2 neurons. (MP4 23824 kb)
Stimulation of inhibitory LPGi neuron subpopulations
Representative examples of video sequences showing the behavioral effects of mice in the open field arena upon laser application to LPGi-vGAT, LPGi-GlyT2 and LPGi-Gad65 neurons. (MP4 6267 kb)
Stimulation of glycinergic GiA, GiV and Gi neuron subpopulations
Representative examples of video sequences showing the behavioral effects of mice in the open field arena upon laser application to GiA-GlyT2, GiV-GlyT2 and Gi-vGAT neurons. (MP4 8811 kb)
Rights and permissions
About this article
Cite this article
Capelli, P., Pivetta, C., Soledad Esposito, M. et al. Locomotor speed control circuits in the caudal brainstem. Nature 551, 373–377 (2017). https://doi.org/10.1038/nature24064
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24064
- Springer Nature Limited
This article is cited by
-
Functional plasticity of glutamatergic neurons of medullary reticular nuclei after spinal cord injury in mice
Nature Communications (2024)
-
An output-null signature of inertial load in motor cortex
Nature Communications (2024)
-
Basal ganglia–spinal cord pathway that commands locomotor gait asymmetries in mice
Nature Neuroscience (2024)
-
Multimodal cortical neuronal cell type classification
Pflügers Archiv - European Journal of Physiology (2024)
-
A Graded Speed Control Method for Cyborg Rats Based on Electrical Stimulation of the Cuneiform Nucleus
Journal of Bionic Engineering (2024)