Abstract
Viruses express several classes of non-coding RNAs1; the functions and mechanisms by which most of these act are unknown. Herpesvirus saimiri, a γ-herpesvirus that establishes latency in the T cells of New World primates and has the ability to cause aggressive leukaemias and lymphomas in non-natural hosts2, expresses seven small nuclear uracil-rich non-coding RNAs (called HSURs) in latently infected cells3,4,5. These HSURs associate with Sm proteins, and share biogenesis and structural features with cellular Sm-class small nuclear RNAs4,6. One of these HSURs (HSUR2) base-pairs with two host cellular microRNAs (miR-142-3p and miR-167) but does not affect their abundance or activity, which suggests that its interactions with them perform alternative functions. Here we show that HSUR2 also base-pairs with mRNAs in infected cells. We combined in vivo psoralen-mediated RNA–RNA crosslinking and high-throughput sequencing to identify the mRNAs targeted by HSUR2, which include mRNAs that encode retinoblastoma and factors involved in p53 signalling and apoptosis. We show that HSUR2 represses the expression of target mRNAs and that base-pairing between HSUR2 and miR-142-3p and miR-16 is essential for this repression, suggesting that HSUR2 recruits these two cellular microRNAs to its target mRNAs. Furthermore, we show that HSUR2 uses this mechanism to inhibit apoptosis. Our results uncover a role for this viral Sm-class RNA as a microRNA adaptor in the regulation of gene expression that follows precursor mRNA processing.
Similar content being viewed by others
Main
Since their discovery three decades ago8, the functions of the seven HSURs have remained elusive. HSUR1 and HSUR2 are conserved among all strains of Herpesvirus saimiri (HVS), and are the only HSURs that are also expressed by the related Herpesvirus ateles9. They resemble the 3′ untranslated regions of mRNAs, insofar as they contain adenylate-uridylate (AU)-rich elements (AREs)10,11,12 and binding sites for the host microRNAs (miRNAs) miR-27, miR-142-3p, and miR-167 (Extended Data Fig. 1a). The binding of HSUR1 to miR-27 results in the degradation of this miRNA and enhances the activated state of the infected T cell7,13. However, HSUR1 and HSUR2 do not affect the activity of ARE-binding proteins in virally transformed T cells10 or the abundance or activity of miR-142-3p and miR-167 (Extended Data Fig. 1b), which suggests that these HSURs do not function as inhibitors of these two miRNAs.
We hypothesized that HSUR2 regulates gene expression by base-pairing with target mRNAs, and by recruiting miR-142-3p, miR-16 and ARE-binding proteins to the mRNAs it targets. We first used in vivo psoralen-mediated crosslinking of RNA–RNA interactions14 to determine whether HSUR2 base-pairs with mRNAs in virally transformed marmoset T cells. Crosslinking with the psoralen derivative 4′-aminomethyltrioxalen (AMT), followed by polyadenylated (polyA+) RNA selection, showed that HSUR2 base-pairs with polyA+ RNA in vivo (Fig. 1a); we did not detect similar activity among cellular spliceosomal Sm-class RNAs. HSUR2 therefore probably targets mature mRNAs rather than precursor mRNAs (pre-mRNAs).
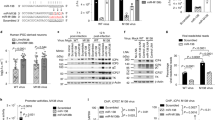
a, Northern blot analyses of polyA+ RNA from HVS-transformed marmoset T cells that were in vivo UV crosslinked in the absence (lanes 2 and 3) or presence (lanes 4 and 5) of psoralen (AMT). I, input (2%); S, supernatant (2%); P, pellet (100%). b–d, Northern blot analyses of several RNAs in cytoplasmic extracts from HVS-transformed marmoset T cells, left untreated (b, c) or incubated with puromycin or pactamycin (d). The extract in c was treated for 20 min on ice with 50 mM EDTA. I, input (20%). In each case, data shown are representative of three independent experiments.
miRNAs associate with actively translated mRNAs, which can be detected using sucrose-gradient co-sedimentation with polyribosomes15,16. Fractionation of cytoplasmic extracts from HVS-transformed marmoset T cells revealed that HSUR2 co-sediments with polyribosomes, mirroring the sedimentation of the HSUR2-binding miRNAs, miR-142-3p and miR-16. By contrast, spliceosomal U2 and U6 short nuclear RNAs (snRNAs) do not co-sediment with polyribosomes (Fig. 1b). The dissociation of polyribosomes into subunits using EDTA treatment shifted HSUR2, miRNAs, and β-actin mRNA to regions of the gradient indicative of lighter molecular weights; the presence of HSUR2 in the polyribosomal region of the gradient is therefore not a consequence of it associating with a high-molecular weight, EDTA-insensitive particle (Fig. 1c). To test whether HSUR2 associates with translationally active mRNAs, we treated cells with the drugs puromycin or pactamycin, which inhibit protein synthesis through different mechanisms. As expected, treatment with either of these drugs disassembled polyribosomes, increased levels of 80S ribosomes, and shifted miRNAs and mRNAs to a lighter-molecular weight region of the gradient (Fig. 1d). HSUR2 exhibited the same shift, indicating that it associates with translating mRNAs (Fig. 1d).
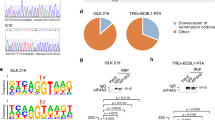
We developed a method to identify mRNAs directly bound by HSUR2, which we termed RNA–RNA interaction identification by crosslinking and capture (RICC). This combines in vivo psoralen-mediated crosslinking and specific HSUR2 capture, using biotinylated probe hybridization, with high-throughput sequencing (seq) to identify crosslinked RNAs (Fig. 2a and Extended Data Fig. 2a). By comparing pulldowns from RNA samples prepared from HVS-transformed marmoset T cells with those prepared from cells in which HSUR2 expression was knocked down before crosslinking by transient transfection with HSUR2 antisense oligonucleotide (ASO)7(Extended Data Fig. 2b), we were able to identify RNAs crosslinked to HSUR2. RICC–seq identified 74 transcripts that were significantly (P ≤ 0.05) enriched in HSUR2 pulldowns in all three replicates (Supplementary Table 1). Most transcripts targeted by HSUR2 correspond to mRNAs that code for cellular proteins, though a small subset of them encodes viral proteins (Fig. 2b). Ingenuity pathway analyses of HSUR2 targets, which include the Fas cell surface death receptor (FAS)17, the tumour suppressor retinoblastoma (RB1)18, and interferon-γ (IFN-γ)19, revealed enrichment of genes involved in cell cycle regulation, apoptosis, and immune response (Fig. 2c and Supplementary Table 2), all of which are relevant to HVS infection. These results suggest a possible role for HSUR2 in the regulation of these processes in virally transformed cells, and also provide confidence in RICC–seq as a method for detecting HSUR2 targets.
a, Diagram of the strategy used for purification of RNAs that base-pair with HSUR2 in vivo. b, Categories of transcripts identified by RICC–seq. c, Ingenuity pathway analysis showing selected pathways for HSUR2 targets. Dashed red line demarcates significance threshold (P < 0.05). d, HSUR2 target mRNA levels in HVS-transformed marmoset T cells (cj319-WT) after transfection with control or HSUR2 ASO (two-sided t-test). e, As in d, but in GSML cells sorted by fluorescence-activated cell sorting (FACS) for green fluorescent protein (GFP) after transient transfection with empty vector carrying GFP or transient transfection with a plasmid carrying GFP and HSUR2 driven by the HSUR2 promoter (HSUR2) or a plasmid carrying GFP and the HSUR2 promoter alone (ΔHSUR2, Extended Data Fig. 4b) (two-sided t-test). f, Western blot of HSUR2 targets in marmoset T cells transformed with wild-type HVS (cj137-WT) or with mutant HVS that lacks HSUR2 (cj137-ΔHSUR2). In each case, data shown are representative of three independent experiments. For d and e, dots represent mean values of independent experiments with error bars representing s.d. ****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; see Supplementary Table 7 for full description of statistics.
We hypothesized that HSUR2-mediated recruitment of miR-142-3p and miR-16 would repress HSUR2 targets, primarily through mRNA destabilization20. Transient knockdown of HSUR2 in virally transformed marmoset T cells correlated with higher levels of HSUR2 target mRNAs (Fig. 2d). No changes were observed for the same target mRNAs when an HSUR2 ASO was transfected into marmoset T cells that had been transformed with a mutant version of HVS that lacks HSUR1 and HSUR27,21 (Extended Data Fig. 3), which suggests that HSUR2 is specifically involved in regulating these target mRNAs. Ectopic expression of HSUR2, together with other HSURs and HVS miRNAs, in GSML cells (a B cell line derived from a natural host of HVS, Saimiri sciureus) confirmed the repressive role of this viral snRNA on target mRNAs (Extended Data Fig. 4). Transient transfection of a vector expressing only the HSUR2 sequence, and not the stem-loop sequence encoding hvsa-miR-HSUR2 miRNAs (Extended Data Fig. 4b), showed that HSUR2 expression is necessary and sufficient to repress target mRNAs (Fig. 2e). Moreover, no changes were observed when cells were transfected with a vector carrying only HSUR2 transcriptional signals, suggesting that the downregulation of HSUR2 target mRNAs is not due to the titration of transcription factors. Similar results were obtained when HSUR2 was expressed in the human Burkitt’s lymphoma B cell line BJAB (Extended Data Fig. 5). Furthermore, when we analysed the levels of proteins encoded by mRNAs that are targeted by HSUR2 in marmoset T cells that were newly transformed with either wild-type HVS (cj137-WT cells) or a mutant HVS lacking only HSUR2 (cj137-ΔHSUR2 cells) (Extended Data Fig. 6), the absence of HSUR2 correlated with the upregulation of several of these proteins (Fig. 2f). These results indicate that RICC–seq successfully identified mRNAs repressed by HSUR2.
HSUR2 associates with argonaute (Ago) proteins7. Ago immunoprecipitation experiments (Fig. 3a) revealed that the transient knockdown of HSUR2 resulted in a lower association of HSUR2 target mRNAs with Ago proteins, supporting the hypothesis that interactions between HSUR2 and its target mRNAs result in the recruitment of Ago proteins specifically to these transcripts. Inhibiting miR-142-3p and miR-16 activity with locked nucleic acid (LNA) inhibitors did not affect the expression of HSUR2 (Extended Data Fig. 7a) or its ability to base-pair with mRNAs (Extended Data Fig. 7b), but did show that miR-142-3p activity is required for HSUR2-mediated repression of all tested target mRNAs; miR-16 activity is required for the repression of only a subset of HSUR2 targets (Fig. 3b). HSUR2 expression was not affected by mutations in its miRNA-binding sites (Extended Data Fig. 8a), though these mutations did affect its ability to destabilize target mRNAs (Fig. 3c and Extended Data Fig. 8b). For all tested target mRNAs, HSUR2-mediated repression required a direct interaction between miR-142-3p and HSUR2; cells transfected with a mutant version of HSUR2 that is unable to bind this miRNA (HSUR2Δ142-3p; Fig. 3c and Extended Data Fig. 8b) have levels of target mRNAs comparable to those of cells transfected with an empty vector. Similar experiments using an HSUR2 mutant that is unable to bind miR-16 (HSUR2Δ16) showed that a direct interaction between HSUR2 and miR-16 is required for HSUR2-mediated repression of some (for example, RB1), but not all, HSUR2 targets (Fig. 3b and Extended Data Fig. 8b). In the two ARE-like sequences, expression of a mutant version of HSUR2 with U → G substitutions (which are expected to interfere with ARE function11) had a weak, statistically non-significant effect on the levels of the RASA2 and DDX20 mRNAs targeted by HSUR2 (Extended Data Fig. 8b). This does not, however, rule out the possibility that the ARE sequences present in HSUR2 play an important role in regulating the stability and/or translation of some of the mRNAs it targets.
a, Enrichment of HSUR2 target mRNAs in Ago immunoprecipitates from extracts prepared from cj319-WT cells transfected with control or HSUR2 ASO (n = 5, two-sided t-test in all cases except for β-actin, which used Wilcoxon’s signed rank test, n = 4). b, qRT–PCR analyses of the HSUR2 targets neuroguidin (NGDN) and RB1 in GSML cells transiently co-transfected with empty vector carrying GFP or with a plasmid carrying GFP and HSUR2, in the presence of a control LNA inhibitor, or an LNA inhibitor with complementarity to miR-142-3p or miR-16. Sample size (n) is indicated below the chart in each case (two-sided t-test). c, As in b, after transient transfection with mutant versions of HSUR2. Partial sequences (and sequence mutations) of HSUR2 are shown (position indicated by number) with residues (bold) involved in base-pairing with miRNAs (pink) or in ARE-like sequences. Sample size (n) is indicated below the chart in each case (two-sided t-test). d, e, As in c, but transiently co-transfected with control miRNA or a mutant version of miR-142-3p (miR-142-3pmut; modified residues in blue) or miR-16 (miR-16mut; modified residues in blue) that has complementarity to HSUR2Δ142-3p or HSUR2Δ16, respectively (n = 3; two-sided t-test). For a–d, dots represent mean values of independent experiments with error bars representing s.d. ***P < 0.001; **P < 0.01; *P < 0.05; see Supplementary Table 7 for full description of statistics.
The ability of HSUR2Δ142-3p and HSUR2Δ16 to repress target mRNAs was restored by the co-transfection of mutant versions of miR-142-3p and miR-16, with seed regions modified to have complementarity to the corresponding mutant HSUR2 (miR-142-3pmut and miR-16mut; Fig. 3d, e and Extended Data Fig. 9). These results confirm that interactions between HSUR2 and these two cellular miRNAs are necessary for HSUR2-mediated mRNA repression. These results also suggest that HSUR2-mediated repression of target mRNAs is not a consequence of HSUR2 inhibiting miR-142-3p and miR-16 activities, favouring a model in which HSUR2 recruits these miRNAs to its target mRNAs.
HSUR2-mediated mRNA repression plays an important role in viral modulation of apoptosis. Staining cj137-WT and cj137-ΔHSUR2 cells with annexin V and propidium iodide showed that the absence of HSUR2 correlates with a higher percentage of cells undergoing apoptotic cell death (Fig. 4a). Marmoset T cells that lacked HSUR2 showed increased activity of the key effector caspase-3/7 compared with that observed in cells that expressed the viral snRNA, though both cell lines showed similar levels of activity of the initiator caspase-8 (Fig. 4b). In BJAB cells, transient expression of HSUR2 resulted in decreased caspase-3/7 activity on FAS-induced apoptosis (Fig. 4c) and on tumour necrosis factor alpha (TNFα)-induced apoptosis (Extended Data Fig. 10a), confirming a role for this viral snRNA in the regulation of apoptosis. To test whether HSUR2-mediated mRNA repression is necessary for the regulation of apoptosis, we expressed mutant versions of HSUR2 that were unable to bind miR-142-3p and/or miR-16 in BJAB cells, and measured caspase-3/7 activity on induction of apoptosis. Resistance to FAS-induced (Fig. 4c) and TNFα-induced (Extended Data Fig. 10a) apoptosis requires the interaction of HSUR2 with both miRNAs. Furthermore, co-transfection of miR-142-3pmut or miR-16mut restored the ability of HSUR2 mutants to confer resistance to apoptosis (Fig. 4d and Extended Data Fig. 10b). These results suggest that HSUR2 restricts apoptosis by recruiting host miR-142-3p and miR-16 to target mRNAs that encode key apoptotic factors, rather than by affecting the activity of the miRNAs themselves.
a, Flow cytometry analysis of cj137-WT and cj137-ΔHSUR2 cells stained with annexin V-Alexa Fluor 647 (Annexin V A-647) and propidium iodide. Right, quantification of five independent experiments (viable and debris, unpaired two-sided t-test; necrotic and apoptotic, two-sided Mann–Whitney test). b, Caspase activity measured in cj137-WT and cj137-ΔHSUR2 cells with the Caspase-Glo assay system (n = 3; unpaired two-sided t-test). c, Caspase-3/7 activity measured as in b, in BJAB cells FACS-sorted after transient transfection with empty vector, or vectors expressing HSUR2 or HSUR2 mutants that cannot bind miR-142-3p and/or miR-16 (described in Fig. 3c) and incubated in the presence of soluble FAS ligand (n = 3; unpaired two-sided t-test). d, As in c, but transiently co-transfected with control miRNA, miR-142-3pmut, or miR-16mut (n = 3, unpaired two-sided t-test). e, Model of HSUR2 function. HSUR2 base-pairs with both the target mRNAs and the miRNAs (miR-142-3p and miR-16), tethering these miRNAs to target mRNAs. HSUR-2 mediated recruitment of miR-142-3p and miR-16 results in repression of target mRNA expression. For a–d, dots represent mean values of independent experiments with error bars representing s.d. ****P < 0.0001; *P < 0.05; see Supplementary Table 7 for full description of statistics.
We have demonstrated that HSUR2 snRNA interacts with specific mRNAs in virally transformed T cells (Fig. 1). Psoralen-dependent crosslinking indicates that these interactions occur through base-pairing (Fig. 1a). HSUR2 expression destabilizes target mRNAs (Fig. 2). Presumably, base-pairing with target mRNAs is required for HSUR2-mediated repression. Further investigation is required to elucidate the mechanism by which HSUR2 is recruited to its target mRNAs. HSUR2-mediated mRNA destabilization requires that HSUR2 binds to miR-142-3p and, in some cases, miR-16 (Fig. 3), suggesting that HSUR2 functions by recruiting these miRNAs to its target mRNAs (Fig. 4e). HSUR2-mediated tethering of miRNAs to target mRNAs leads to recruitment of Ago proteins (Fig. 3a), which promote translational repression and/or degradation, through de-adenylation of the target mRNAs22. As HSUR2 is not polyadenylated, its expression is unaffected by associations with Ago proteins7 (Extended Data Figs 7a, 8a). HSUR2 is more abundant than target mRNAs, but less abundant than miR-142-3p and miR-16 (Supplementary Table 3). This stoichiometry is probably important in ensuring effective miR-142-3p-dependent and miR-16-dependent HSUR2-mediated mRNA repression23. Disruption of HSUR2 ARE-like sequences did not have a statistically significant effect on target mRNA destabilization (Fig. 3c and Extended Data Fig. 8b). These sequences, as well as the recruited miRNAs, might modulate the translation of target mRNAs24.
Our results uncover a role for a viral Sm-class RNA in the regulation of gene expression after pre-mRNA processing, a role reminiscent of those performed by the bacterial Hfq-associated regulatory RNAs (structurally related to Sm-class RNAs) that regulate the stability and translation of target mRNAs25. Given that viruses often co-opt factors from their hosts, it is conceivable that similar non-coding RNAs exist in host cells, and act to regulate gene expression in a similar fashion26; further investigation is required to test these hypotheses.
Methods
Plasmids
For selection of transfected cells on the basis of expression of GFP by FACS, a fragment containing the human cytomegalovirus intermediate-early enhancer/promoter, the GFP coding region, and the polyadenylation signal of the simian virus 40 was generated by PCR using pEGFP–C2 (Clontech) as a template, and inserted between the KpnI and XhoI sites of pBluescript II SK+ (Stratagene) to generate the plasmid pBS-GFP (referred to as ‘empty vector’ in Figs 2e and 3, and Extended Data Figs 4d, 5, 8, and 9). For expression of HSURs and HVS miRNAs, a genomic fragment including positions +1,444 and +3,504 of the genome of HVS A11 (GenBank: X64346.1) was inserted between the SalI and BamHI sites of pBS-GFP to generate the plasmid pBS-GFP-H1-5 for expression of HSUR1, HSUR2, and HSUR5, together with hvsa-miR-HSUR2 and hvsa-miR-HSUR5 miRNAs (HSURs1–5, Extended Data Fig. 4a). A genomic fragment including positions +4,613 and +6,945 of the genome of HVS A11 was inserted between the SalI and BamHI sites of pBS-GFP to generate the plasmid pBS-GFP-H3-7 for expression of HSUR3, HSUR4, HSUR6, and HSUR7, together with hvsa-miR-HSUR4 miRNAs (HSURs3–7, Extended Data Fig. 4a). For expression of HSUR2 only, and not hvsa-miR-HSUR2 miRNAs, a fragment of HVS A11 genome including positions +2,344 to +2,726 was inserted between the EcoRI and BamHI sites of pBS-GFP to generate the plasmid pBS-GFP-HSUR2 (Extended Data Fig. 4b). The plasmid pBS-GFP-ΔHSUR2 (provided by S. Camolotto) containing only the HSUR2 promoter and enhancer sequences, and not the HSUR2 sequence (Extended Data Fig. 4b), was generated by inserting an HVS A11 genomic fragment including positions +2,486 to +2,726 between the EcoRI and BamHI sites of pBS-GFP. Mutant versions of HSUR2 (plasmids pBS-HSUR2Δ142-3p, pBS-HSUR2Δ16, pBS-HSUR2ΔARE, and pBS-HSUR2Δ142-3p/16) were obtained by site-directed mutagenesis using a QuickChange site-directed mutagenesis kit (Stratagene), and confirmed by sequencing.
Recombinant HSV
The bacmid constructs HVS-BAC-GFP-WT and HVS-BAC-GFP-ΔHSUR2 were generated by en passant mutagenesis27,28. We generated a plasmid that contained the first ~ 7 kb of the HVS A11 genome by inserting an HVS A11 genomic fragment generated by overlapping PCR with two fragments (one fragment including positions +22 to +2,354 of the HVS A11 genome, and a second fragment including positions +2,547 to +7,249). The generated fragment, which lacked both HSUR2 and the HSUR2 promoter sequence required for expression of hvsa-miR-HSUR2 miRNAs29, was inserted between the SalI and BamHI sites of pBluescript II SK+ to generate the plasmid pBS-7.4ΔHSUR2. A unique NcoI site was generated at a position corresponding to +3,664 of the HVS A11 genome, by performing site-directed mutagenesis on both pBS-7.4ΔHSUR2 and pBS-7.429 to generate the plasmids pBS-7.4ΔHSUR2-Nco and pBS-7.4-Nco. A PCR amplicon containing the kanamycin resistance gene, the I-SceI restriction site and a 50-bp duplication of the HVS A11 genomic sequence was generated using the pEPkanS2 vector27 (provided by K. Tischer), and inserted into the NcoI site of pBS-7.4ΔHSUR2-Nco and pBS-7.4-Nco to generate the plasmids pBS-7.4ΔHSUR2-Nco-Kan and pBS-7.4-Nco-Kan, respectively. PCR-amplified fragments encompassing positions +65 to +4,637 of the HVS A11 genome were generated using pBS-7.4ΔHSUR2-Nco-Kan and pBS-7.4-Nco-Kan as templates (Extended Data Fig. 6a), and recombined into the bacmid HVS-BAC-GFP30 (provided by A. Whitehouse) in GS1783 cells (a gift from G. Smith) as previously described28. The bacmid HVS-BAC-GFP carries the SacI deletion31 that removes part of the saimiri transforming protein-A coding region, together with the genes encoding HSUR1, HSUR2, and HSUR5, hvsa-miR-HSUR2 and hvsa-miR-HSUR5 miRNAs, and part of viral dihydrofolate reductase (vDHFR) (Extended Data Fig. 6a). Viruses generated with HVS-BAC-GFP are therefore unable to transform cells30,31. Recombinants were selected on chloramphenicol-kanamycin plates. The scarless removal of the kanamycin cassette was achieved by induction of I-SceI expression during the second Red recombination step, through arabinose induction; this generated the bacmids HVS-BAC-GFP-WT and HVS-BAC-GFP-ΔHSUR2. Positive clones were screened by PCR, and analysed by restriction endonuclease (Extended Data Fig. 6b) and sequencing.
Virus cultures, marmoset T cell immortalization and cell lines
Recombinant viruses were generated by transfection of bacmids HVS-BAC-GFP-WT and HVS-BAC-GFP-ΔHSUR2 into owl monkey (Aotus trivirgatus) kidney cells (ATCC, OMK-637-69) using Lipofectamine 2000 (ThermoFisher Scientific), according to the manufacturer’s instructions. Complete cytopathic effects were observed around 14 days after transfection. High-titre virus stocks were generated by infecting 60–80% confluent owl monkey kidney cells in T-75 flasks with low multiplicity of infection (MOI of 0.1). Virus titres were estimated by the limited dilution protocol as previously described32. In vitro immortalization of marmoset T cells without interleukin-2 was performed according to established protocols33. Preservative-free heparinized marmoset blood samples from three animals (cj137, cj290 and cj323) were obtained from PrimateBiologicals (http://primatebiologicals.com/). Although all samples were successfully immortalized, only cell lines derived from animal cj137 (cj137-WT and cj137-ΔHSUR2; Figs 2f, 4a, b and Extended Data Fig. 6) continued to grow after six months of cultivation. Cell line cj137-WT, cell line cj137-ΔHSUR2, and marmoset T cell lines derived from a single marmoset that were transformed in parallel with wild-type HVS A11, or HVS A11 Δ2a21 deleted for the HSUR1 and HSUR2 genes10 (cj319-WT and cj319-Δ2a; Figs 1, 2d, and Extended Data Figs 1, 3), were grown in RPMI-1640 medium (ThermoFisher Scientific) supplemented with 20% fetal bovine serum, 100 units per ml penicillin, 100 μg ml−1 streptomycin, 2 mM glutamax (ThermoFisher Scientific), 1 mM sodium pyruvate (ThermoFisher Scientific) and 0.001% β-mercaptoethanol (Sigma Aldrich). Epstein-Barr virus (EBV)-transformed B-lymphoblast GSML cells34 (ATCC-CRL-2699) and BJAB cells (a gift from J. Steitz) were not authenticated and were grown in RPMI-1640 medium supplemented with 10% fetal bovine serum, penicillin, streptomycin, sodium pyruvate and glutamax. BJAB cells were used for apoptosis assays (Fig. 4 and Extended Data Fig. 10) because they express miR-142-3p and are not transformed by Epstein-Barr virus. All cell lines were routinely tested for the presence of mycoplasma.
Antibodies
Polyclonal antibodies against STAG1 (catalogue number: LS-C161537, lot number: 49409) and NGDN (catalogue number: 16524-1-AP) were purchased from Life Span Biosciences and Proteintech Group, respectively. Antibodies against DDX5 (catalogue number: 4387S), Rock2 (catalogue number: 8236S), Rac1/2/3 (catalogue number: 2465S), N-Wasp (catalogue number: 4848S), Fas (catalogue number: 8023S) and retinoblastoma (catalogue number: 9309S) were obtained from Cell Signaling Technology. Polyclonal antibodies against GRLF1 (catalogue number: A301-736A-M) were obtained from Bethyl Laboratories, and mouse monoclonal antibody against α-tubulin (catalogue number: SC-23948, lot number: K0812) was obtained from Santa Cruz Biotechnology. Monoclonal antibodies against Ago proteins (PAN-Ago, clone 2A8) were a gift from Z. Mourelatos.
Polysome gradients
Soluble extracts from cj319-WT cells left untreated or treated for 16–18 h with 200 μg ml−1 puromycin (Invivogen) or 2.5 μg ml−1 pactamycin (Sigma Aldrich) were analysed on 12-ml linear 15–50% sucrose gradients prepared in polysome buffer (20 mM Tris-HCl, pH 7.5, 150 mM KCl, 10 mM MgCl2 and 100 μg ml−1 cycloheximide (Sigma Aldrich)), and equilibrated at 4 °C for 3 h. In brief, cells were collected, centrifuged, and suspended in 20 ml complete RPMI-1640 medium with 100 μg ml−1 cycloheximide. Cells were incubated at 37 °C and 5% CO2 for 25 min and centrifuged. Cells were suspended in four volumes of hypotonic lysis buffer (5 mM Tris-HCl, pH 7.5, 1.5 mM KCl, 10 mM MgCl2, 5 mM DTT, 0.5% Triton X-100, 0.5% sodium deoxycholate, 10 μg ml−1 digitonin, 10 mM ribonucleoside vanadyl complex (New England Biolabs) preheated at 65 °C and 100 μg ml−1 cycloheximide). Cells were vortexed and passed three times through a 25-gauge needle and then three times through a 27.5-gauge needle. The extracts were clarified at 17,000g for 10 min at 4 °C and heparin (ThermoFisher Scientific) was added to a final concentration of 1 mg ml−1. Where indicated, EDTA was added to the clarified extract to a final concentration of 50 mM and incubated on ice for 20 min. Extract samples of 300 μl were layered on top of duplicate sucrose gradients and centrifuged at 150,000g on a Beckman SW41 rotor for 2 h at 4 °C. The sedimentation profile of one of the gradients was determined using a 65% sucrose pump at a constant flow rate of 1.5 ml min−1 linked to an ISCO UA-6 254-nm ultraviolet detector equipped with a chart recorder set at 60 cm h−1 and 1 absorbance unit full scale. The second gradient was fractionated (0.5 ml per fraction) and the RNA was purified by acid phenol-chloroform extraction and 3 M sodium acetate-ethanol precipitation, and then analysed by northern blot.
Northern blotting
RNA from each fraction of the sucrose gradients was suspended in 20 μl 1× DNase reaction mixture: 2 μl 10× DNase reaction buffer, 0.4 units RNase-free DNase I (New England Biolabs) and 17.8 μL H2O, and incubated at 37 °C for 30 min. The volume of each sample was taken to 0.5 ml with H2O, and the RNA was re-purified by acid phenol-chloroform extraction and sodium acetate-ethanol precipitation. RNA samples were then suspended in 20 μl 2 mM Tris-HCl, pH 7.5, 8 M urea and 20 mM EDTA and heated at 65 °C before loading onto a 14% denaturing polyacrylamide gel. Gels were run at 10 W in 1× TBE and transferred onto a Zeta-Probe nylon membrane (Bio-Rad) for 30 min at 1 A using a semi-dry blotting unit (Fisher Biotech). The membranes were pre-hybridized at 42 °C in ExpressHyb hybridization solution (Clontech) for at least 1 h. Hybridization of 32P-labelled probes to HSUR2, U2 and U6 was performed in ExpressHyb solution overnight at 42 °C. Hybridization of 32P-labelled probes to miR-142-3p, miR-16 and miR-21 was performed overnight at 30 °C. Blots were washed once with 2× saline sodium citrate (SSC) buffer, 0.1% sodium dodecyl sulfate (SDS) for 15 min at room temperature, followed by either a wash with 1× SSC, 0.1% SDS for 15 min at room temperature (miR-142-3p, miR-16, and miR-21 probes) or a wash with 0.5× SSC, 0.1% SDS for 15 min at room temperature (HSUR2, U2, and U6 probes). Membranes were wrapped in Saran Wrap and exposed to a phosphorImager screen (GE Healthcare). To detect β-actin mRNA in the polysome gradient fractions, RNA was suspended in 25 μl of 20 mM 3-(N-morpholino)propanesulfonic acid (MOPS), pH 7, 15% formaldehyde, 50% deionized formamide and 20 mM EDTA, heated at 95 °C for 15 min, placed on ice and loaded onto 1.2% agarose-formaldehyde gels. The gels were run at 100 V in 20 mM MOPS, pH 7, 2.6% formaldehyde and transferred overnight onto nylon membranes through capillarity. The membranes were stained with 0.02% methylene blue in 0.4 M sodium acetate and 0.4 M acetic acid, and destained in H2O. Blots were prehybridized in ExpressHyb solution at 42 °C for 4 h and hybridized with 32P-labelled probe to β-actin mRNA overnight at 80 °C. The membranes were washed with 2× SSC, 0.1% SDS for 15 min at room temperature followed by a wash with 0.5× SSC, 0.1% SDS for 15 min at room temperature. Membranes were wrapped in Saran Wrap and exposed to a phosphorImager screen.
Probe radiolabelling
Radiolabelled probes were prepared in 20-μl reactions, containing 10 pmoles ASO to HSUR2, U2, U6, miR-142-3p, miR-16 or miR-21, 10 units of T4 polynucleotide kinase (New England Biolabs) and 151.5 μCi of [γ-32P]ATP (6,000 Ci mmol−1, PerkinElmer) and incubated at 37 °C for 1 h. Unincorporated isotope was removed by centrifugation using Mini Quick G-25 gel filtration columns, according to the manufacturer’s protocol (Roche). Radiolabelled probes were eluted in a volume of 50 μl H2O, and 10 μl of eluted radiolabelled probe was used in each hybridization experiment. A β-actin probe was labelled using 50 μCi [α-32P]dCTP and Klenow DNA polymerase for 15 min, using the DecaPrime II kit according to the manufacturer’s instructions (Ambion). A 10-μl aliquot of the radiolabelled probe was taken to a volume of 100 μl with 10 mM EDTA, pH 8 and incubated at 95 °C for 10 min. The labelled probe was then added directly without further purification to the prehybridized membranes.
Immunoblotting
Marmoset cj137-WT and cj137-ΔHSUR2 cells were collected and stained with 100 ng ml−1 2-(4-amidinophenyl)-1H-indole-6-carboxamidine (DAPI) and FACS-sorted in a BD FACSAria-II cell sorter (BD Biosciences) equipped with a UV laser, at the University of Utah Flow Cytometry Core. Live DAPI-negative cells were collected and total cell lysates were prepared in Glo Lysis buffer (Promega) containing complete protease inhibitors (Roche), 10 mM N-ethylmaleimide (Sigma Aldrich) and 1 mM phenylmethylsulfonyl fluoride (Fluka BioChemika). Cj319-WT and cj319-Δ2a cells were collected, washed with phosphate-buffered saline (PBS) and directly lysed for analysis as described above. The protein concentration of the lysates was determined with the Coomassie Plus Protein Assay reagent (ThermoFisher Scientific). Lysates were boiled with 1× SDS–PAGE sample buffer, and 15 or 20 μg total protein was separated on 10% high Tris polyacrylamide gels (0.75 M Tris base) using high Tris running buffer (50 mM Tris base, 193 mM gycine, 0.1% SDS). Proteins were transferred onto nitrocellulose membranes, probed with the indicated antibodies and visualized by enhanced chemiluminescence, according to standard protocols.
Transfections
Ten million cj319-WT or cj319-Δ2a cells were nucleofected with control (complementary to GFP coding sequence) or HSUR2 chimaeric ASO (see RICC–seq described below) (Integrated DNA Technologies) as described above, using Amaxa’s human T-cell kit (Lonza) and program X-001 in a nucleofector 2b device. Twenty-one hours after transfection the cells were collected, washed with PBS and stored in TRIzol (ThermoFisher Scientific) at −80 °C until used. GSML and BJAB cells were nucleofected using 5 × 106 and 3 × 106 cells, respectively, per nucleofection, with Amaxa’s kit V (Lonza) and program T-020, combined with 3.4 μg pBS-GFP, 1 μg vector that encoded pBS-GFP-HSUR2 (and mutant variants of HSUR2), or 1 μg pBS-GFPΔHSUR2. In experiments in which GSML cells or BJAB cells were co-transfected with miR-142-3pmut or miR-16mut (Integrated DNA Technologies), 50 pmoles mutant miRNA or 50 pmoles control siRNA-A (Santa Cruz Biotechnology) were added to the nucleofection reaction. To test the effect of LNA miRNA inhibitors on HSUR2-mediated repression of HSUR2 targets, pBS-GFP or pBS-GFP-HSUR2 plasmid was co-transfected with: 100 pmoles control LNA (miRCURY LNATM microRNA inhibitor negative control A); 50 pmoles miRCURY LNA miRNA inhibitor against hsa-miR-142-3p (Exiqon), with 50 pmoles control LNA; 50 pmoles miRCURY LNA miRNA inhibitor against hsa-miR-16 (Exiqon), with 50 pmoles control LNA; or 50 pmoles each of both miR-142-3p and miR-16 LNA inhibitors. Transfections of GSML cells with pBS-GFP-H1-5 and pBS-GFP-H3-7 were performed using 5 μg of plasmid, with 3.4 μg pBS-GFP or 5 μg each of both pBS-GFP-H1-5 and pBS-GFP-H3-7. In all the experiments described above, transfected GSML cells or BJAB cells were grown for 18–21 h after nucleofection and FACS-sorted on a BD FACSAria-II cell sorter equipped with a 488-nm laser (525/50 filter set) for GFP-positive cells. Cells that were collected expressed a mid-range level of GFP (arbitrary fluorescence of 103–105 within the GFP window; DAPI staining in initial experiments showed that cells expressing GFP beyond 105 were necrotic, and these were therefore discarded and excluded from further analysis). Immediately after sorting, cells were centrifuged, suspended in 100 μl of culture medium and stored in TRIzol at −80 °C until used.
Quantitative RT–PCR (qRT–PCR)
Total RNA was isolated from cj319-WT, cj319-Δ2a, GSML, or BJAB cells with TRIzol according to the manufacturer’s instructions. The TRIzol-purified total RNA was suspended in 85 μl of water and 10 μl of 10× DNase I buffer, treated with 10 units RNase-free DNase I (New England Biolabs) at 37 °C for 30 min and re-purified with acid phenol-chloroform extraction, followed by sodium acetate-ethanol precipitation. cDNA was synthesized in 20-μl reactions using the High Capacity cDNA Reverse Transcription Kit with MultiScribe Reverse Transcriptase (Applied Biosystems) using 600–900 ng total RNA treated with DNase I, and random primers. Subsequently, quantitative PCR (qPCR) was performed in 8-μl reactions using primers (Supplementary Table 4) at 0.5 μM, cDNA diluted at ratios 1:3–1:7.5 (depending upon the input of total RNA used in the cDNA synthesis reaction as described above) and KAPA SYBR green (KAPA Biosystems) in a Roche 480 LightCycler (95 °C for 5 min, 1 cycle; 95 °C for 10 s, 60 °C for 10 s and 72 °C for 10 s, 55 cycles followed by a melting curve of the reaction product from 65 °C to 97 °C with a ramp rate of 0.11 °C s−1). qPCR primers were designed using Primer3Plus and tested using cDNA prepared from 900 ng DNase-treated total RNA and diluted at ratios 1:5–1:5,000. All reactions were performed at least four times in each independent experiment. Ct values were determined using the ‘Abs quant/2nd derivative max’ function of the LightCycler. Relative mRNA levels of HSUR2 targets were calculated using the  method35, dividing the
method35, dividing the  of the HSUR2 target in the treated sample (combined with HSUR2 and so on) by the
of the HSUR2 target in the treated sample (combined with HSUR2 and so on) by the  of the HSUR2 target in control sample (combined with empty vector or ΔHSUR2). For normalization, the Ct values of 18S ribosomal RNA were subtracted from the Ct values of HSUR2 targets to calculate ΔCt.
of the HSUR2 target in control sample (combined with empty vector or ΔHSUR2). For normalization, the Ct values of 18S ribosomal RNA were subtracted from the Ct values of HSUR2 targets to calculate ΔCt.
Estimation of the number of molecules of HSUR2 and HSUR2 target mRNAs in wild-type and ΔHSUR2 marmoset T cells
Sequences of select HSUR2 targets were amplified from marmoset T-cell cDNA by PCR using Taq polymerase and pairs of primers (listed in Supplementary Table 4), and introduced into the pCR2.1-TOPO vector by TA-cloning (Invitrogen). Plasmids encoding sequences of HSUR2 targets were purified, quantified and used to prepare 1:10 serial dilutions in H2O from 100 pg μl−1 to 100 ag μl−1. Two-microlitre samples of each dilution were then used in qPCR reactions, using KAPA SYBR green as described above to generate standard curves (log (number of molecules) versus Ct) for each target. A standard curve for HSUR2 was generated using 1:10 serial dilutions of pBS-HSUR2ΔmiR in H2O as template. Total RNA from 4 × 106 wild-type or 3.05 × 106 ΔHSUR2 marmoset T cells was isolated with TRIzol, and 900 ng were used to synthesize cDNA as described above. The resulting cDNAs were diluted 1:5 for analysis of HSUR2 targets, and diluted 1:1,000 for analysis of HSUR2 in H2O. Subsequently, qPCR was performed as described above. The resulting Ct values were used to estimate the number of molecules for HSUR2 and for HSUR2 targets, using the previously generated standard curves.
Analysis of mRNAs co-immunoprecipited with Ago proteins
Thirty to forty million cj319-WT cells were transfected with control or HSUR2 ASO as described above and used to prepare whole-cell extracts in 0.35–0.4 ml NET-2 buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 0.05% Nonidet-40), containing complete protease inhibitors, 1 mM PMSF and 5 μl of RNaseOUT (ThermoFisher Scientific). Extracts were sonicated on ice three times for 10 s each. Lysates were cleared by centrifugation at 16,000g for 10 min at 4 °C. Aliquots equal to 10% of each lysate were saved and immediately stored in 1 ml TRIzol. Rabbit anti-mouse IgG (ThermoFisher Scientific) was immobilized on Protein A Sepharose 4 Fast Flow beads (GE Healthcare) in bulk overnight at 4 °C and aliquoted into corresponding tubes before the last wash to guarantee that all samples were incubated with the same amount of antibody. The remaining 90% fraction of each lysate was incubated with 5 μl anti-PAN-Ago antibodies, or normal mouse serum (Sigma Aldrich), and 30 μl rabbit anti-mouse IgG, with continuous rotation for 4 h at 4 °C. Samples were washed four times with NET-2 buffer containing protease inhibitors, and the RNA immunoprecipitation (RIP) fractions were collected by centrifugation and stored immediately in TRIzol at −80 °C.
For RNA purification and cDNA synthesis, the total RNA in TRIzol from input and RIP fractions was isolated according to the manufacturer’s instructions using 20 μg glycogen during the precipitation step. Total RNA was resuspended in 15 μl water and used for cDNA synthesis as described above, except that 1.5 μl of each input fraction (1% of the total) and 14.2 μl of each RIP fraction was used per 20-μl reaction.
For immunoprecipitated mRNAs, qPCR of HSUR2 targets was performed as described above using cDNAs from input and RIP fractions, diluted 1:5 in H2O. The Ct values of HSUR2 targets were normalized to the input RNA fraction Ct values according to the following equation: ΔCt [normalized RIP] = (Ct [RIP] − (Ct [input] − log2(input dilution factor))). As 1% of each input fraction was used to synthesize cDNA, log2100 = 6.644. The normalized RIP-fraction Ct value was then adjusted for the normalized background (normal mouse serum reaction (NS)) fraction Ct value as follows: ΔCt [RIP/NS] = ΔΔCt [normalized RIP] − ΔCt [normalized NS]. The fold enrichment, above the sample specific background, of each HSUR2 target in the RIP fraction was then calculated as  . The fold enrichment of the control ASO relative to HSUR ASO was calculated as
. The fold enrichment of the control ASO relative to HSUR ASO was calculated as  (Control ASO) divided by
(Control ASO) divided by  (HSUR2 ASO).
(HSUR2 ASO).
Apoptosis assays
Growing cultures of cj137-WT and cj137-ΔHSUR2 cells were collected and layered on top of 10-ml cushions of Histopaque-1077 (Sigma Aldrich) to remove cell debris. The cells were centrifuged at 400g for 30 min at room temperature, and cells at the interface were collected, washed, suspended in complete medium, and counted. Cells were seeded at a density of 500,000 cells per ml and grown for two weeks. Cells were then collected, washed in ice-cold PBS, centrifuged and suspended at a density of 106 cells per ml in 100 μl annexin-binding buffer (10 mM HEPES, pH 7.4, 140 mM NaCl and 2.5 mM CaCl2). Cells were stained for 15 min at room temperature with Alexa Fluor 647 annexin V conjugate (1:20 dilution, Life Technologies) and 1 μg ml−1 propidium iodide (Life Technologies). Samples were diluted with 400 μl annexin-binding buffer and placed on ice. Data were acquired on a BD FACSCanto (BD Biosciences) flow cytometer using the 488-nm (685/40 filter set) and 640-nm (670/30 filter set) lasers. Alternatively, cj137-WT and cj137-ΔHSUR2 cells were stained with 100 ng ml−1 DAPI, and live DAPI-negative cells were FACS-sorted directly into a 96-well plate at a density of 20,000 cells per well. Caspase activity was then measured with the Caspase-Glo 3/7 and Caspase-Glo 8 reagents, according to the manufacturer’s instructions (Promega). Caspase 3/7 activity was also measured for BJAB cells transiently expressing HSUR2 or HSUR2 variants (Fig. 4 and Extended Data Fig. 10).These BJAB cells were transiently transfected with pBS-GFP-HSUR2, pBS-GFP-ΔHSUR2, pBS-GFP-HSUR2Δ142-3p, pBS-GFP-HSUR2Δ16, or pBS-GFP-HSUR2Δ142-3p/16, and GFP-positive cells were FACS-sorted and suspended in culture medium to a density of 400,000 cells per ml; 100 μl of this suspension was seeded per well onto a 96-well plate. The cells were then incubated with 100 ng ml−1 recombinant soluble Fas ligand (PeproTech) or 100 ng ml−1 TNFα (Sigma Aldrich) for 19 h at 37 °C. Caspase 3/7 activity was then assayed as described above.
RICC–seq: in vivo RNA–RNA crosslinking
Two sets of three independent samples of 6 × 107 cj319 cells per sample were treated with chimaeric ASOs (the oligonucleotides are 20-nt long, contain backbone phosphorothioate linkages to increase stability, and have five nucleotides on each end substituted with 2′-O-methoxyethyl ribonucleotides7,36) directed to GFP (5′- UCACCTTCACCCTCTCCACU -3′) (control HSUR2-positive sample), or with an ASO directed to HSUR2 (5′- AAGCGATACCTCGTGUGUGA -3′), which specifically targets HSUR2 for degradation (HSUR2-negative sample)7 (Extended Data Fig. 2). Cells were transfected (107 cells per nucleofection) with 1 nmol ASO by applying three 1,700-V pulses for 10 ms each (Neon System, ThemoFisher Scientific). Cells were then plated at a density of 1 × 106 cells per ml, grown at 37 °C and 5% CO2 for 24 h, and collected. Cells from each sample were pooled, washed with PBS and suspended in 1 ml PBS containing 200 μg ml−1 4′-aminomethyl-4,5′,8-trimethylpsoralen (Cayman Chemical), crosslinked as described7, collected, and fractionated in 100-μl samples. One hundred microlitres of 6 M guanidinium hydrochloride was added to each sample followed by 20 μl of a 20 mg ml−1 solution of RNase-free proteinase K (Ambion) and 5 μl of 20% SDS. Samples were incubated at 65 °C for 1 h. One millilitre of TRIzol was added to each sample and then stored at −80 °C.
RICC–seq: polyA+ RNA purification
Total RNA was purified from control HSUR2-positive and HSUR2-negative samples in TRIzol, according to the manufacturer’s protocol. PolyA+ RNA was isolated using oligo d(T)25 magnetic beads (New England Biolabs). In brief, purified total RNA was suspended in 600 μl H2O and heated to 65 °C for 5 min. One volume of 2× binding buffer (40 mM Tris-HCl, pH 7.5, 1 M NaCl, 1% SDS, 2 mM EDTA, 10 mM DTT) was added to each sample and mixed with 250 μl of oligo d(T)25 magnetic beads. Samples were incubated with continuous rotation for 30 min at room temperature. The beads were washed twice for 1 min with 1 ml of wash buffer 1 (20 mM Tris-HCl, pH 7.5, 500 mM NaCl, 0.1% SDS, 1 mM EDTA, 5 mM DTT) and then once with 1 ml of wash buffer 2 (20 mM Tris-HCl, pH 7.5, 500 mM NaCl, 1 mM EDTA). The beads were then incubated with 1 ml of low salt buffer (20 mM Tris-HCl, pH 7.5, 200 mM NaCl, 1 mM EDTA) for 1 min at room temperature. PolyA+ RNA was eluted from the beads with 500 μl of elution buffer (20 mM Tris-HCl, pH 7.5, 1 mM EDTA) at 50 °C for 5 min with continuous agitation. PolyA+ RNA was precipitated by adding 50 μl of 3 M sodium acetate, pH 5.2, 5 μl of a 10 mg ml−1 solution of glycogen and 1 ml of 100% ethanol, and incubated overnight at −20 °C.
RICC–seq: capture of HSUR2 with biotinylated ASO
PolyA+ from HSUR2-positive or HSUR2-negative samples was suspended in 500 μl H2O, heated to 95 °C for 3 min and placed on ice. Five hundred pmoles of a biotinylated HSUR2 ASO (5′-GGTTTTTAAATATGTACACCC-3′-Bio) was added to each sample. The samples were incubated at 65 °C for 5 min and 500 μl of 2× binding buffer (20 mM Tris-HCl, pH 7.5, 1 M NaCl, 2 mM EDTA, 0.1% SDS) heated at 65 °C was added to each sample. One hundred microlitres from each sample were saved as ‘PolyA+ input’ samples and stored in 1 ml TRIzol at −80 °C until used. The remaining 900 μl from each sample was incubated for 4 h at room temperature with continuous rotation. After annealing was complete, 125 μl of MPG streptavidin magnetic beads (PureBiotech) was added to each sample and incubated overnight at 4 °C. The beads were washed twice with 1× binding buffer (10 mM Tris-HCl, pH 7.5, 500 mM NaCl, 1 mM EDTA, 0.05% SDS) and twice with 1× wash buffer (10 mM Tris-HCl, pH 7.5, 200 mM NaCl, 1 mM EDTA, 0.05% SDS). Bound RNA was recovered by adding 1 ml TRIzol to the beads. RNA was purified from both the input and HSUR2 ASO-treated samples, according to the manufacturer’s protocol.
RICC–seq: library preparation and high-throughput sequencing
Purified RNA samples were suspended in 25 μL H2O and placed on a glass plate covered with parafilm on ice. The AMT crosslinks were reversed by directly irradiating the samples at 254 nm for 10 min. The samples were collected and taken to a volume of 300 μl with H2O. RNA was precipitated by adding 40 μg glycogen, 30 μl of 3 M sodium acetate, pH 5.2 and 900 μl of 100% ethanol. Samples were incubated overnight at −20 °C. The RNA was recovered by centrifugation, washed and suspended directly in 19.5 μl ‘Fragment, Prime, Finish’ (FPF) mix from the TruSeq Stranded mRNA LT kit, to prepare libraries for deep sequencing according to the manufacturer’s protocol (Illumina). HiSeq 50 Cycle single-read sequencing was performed by the High Throughput Genomics Core Facility at the Huntsman Cancer Institute (University of Utah) on an Illumina HiSeq 2500 instrument.
RICC–seq: bioinformatic analyses
A reference sequence was created using a marmoset genomic sequence (calJac3 assembly), Saimiriine herpesvirus 2 (GenBank: X64346.1) and splice junction sequences. The splice junction sequences were generated using USeq (v8.8.8) MakeTranscriptome using default settings. CalJac3 Ensembl build 80 and X64346.1 annotations were used in MakeTranscriptome. Reads were aligned to the reference sequence using Novoalign (2.08.03) allowing up to 50 alignments and removing adaptor sequences. The resulting alignments were then run through USeq SamTranscriptomeParser using default settings to convert splice alignments back to genomic space and to remove low quality reads. Reads aligning to multiple locations in the genome were removed. USeq DefinedRegionDifferentialSeq was used to count the number of aligned reads to each gene. The only reads that were counted were those aligning to the opposite strand of the annotation. Genes with fewer than ten reads across all replicates were thrown out. The resulting read counts were then run in DESeq (v1.4.5) to determine which genes were differentially expressed in each condition, using default settings. Genes that were significantly enriched (Benjamini–Hochberg adjusted P value < 0.05, log2 fold change > 0) in the HSUR2-positive pulldown versus HSUR2-negative pulldown, and in the HSUR2-positive pulldown versus HSUR2-positive input, were used in further analyses (Supplementary Table 1). Significantly enriched genes were analysed by Ingenuity Pathway Analysis version 24718999 (IPA, Qiagen) using common name, and compared against the ingenuity knowledge base. All pathways that showed Fisher’s Exact Test P values < 0.05 (not adjusted for multiple hypothesis testing) are shown in Supplementary Table 2, whereas significant pathways that are relevant for HVS infection are shown in Fig. 2c.
Statistical analyses
No statistical methods were used to predetermine sample size, nor were the experiments randomized or the investigators blinded to sample allocation during experiments and evaluation of experimental results. ‘Biological replicates’ (n), indicated in figures and figure legends, refers to the number of independent experiments performed. The number of independent experiments was chosen to allow for statistical significance. Statistical analysis was performed using Graphpad Prism 7. Data were analysed for the presence of outliers using the ROUT method (Q = 10%). Statistical analysis of biological replicates shown in Fig. 2d and Extended Data Fig. 3 was performed with Student’s t-tests corrected for multiple comparisons with the Holm–Sidak method (alpha = 0.05), after using Bartlett’s test (https://home.ubalt.edu/ntsbarsh/business-stat/otherapplets/BartletTest.htm) to determine whether the variance of the different groups was significantly different. Two-sided P values of biological replicates shown in Figs 2e, 3, and Extended Data Figs 4d, 5, 8, and 9 were obtained with Student’s t-tests, compared to the control samples set at 1.0. When outliers were present, the samples were compared with Wilcoxon’s signed rank test. The pairwise comparisons made between experimental groups and controls shown in Fig. 4 and Extended Data Fig. 10 were performed using unpaired Student’s t-test, with or without Welch’s correction for unequal variances, after using Bartlett’s test to determine whether the variance of the different groups was significantly different. When outliers were present (ROUT method, Q = 10%), the paired groups were compared using the non-parametric Mann–Whitney test. For all data presented, see Supplementary Table 7 for the specific test used, the exact number of replicates and exact P values.
Data availability
Source data for all figures in this article are included in its Supplementary Information. The described RNA-seq data have been deposited in the Gene Expression Omnibus under accession number GSE79082. Further data that support the findings of this study are available from the corresponding author upon reasonable request.
Accession codes
References
Tycowski, K. T. et al. Viral noncoding RNAs: more surprises. Genes Dev. 29, 567–584 (2015)
Ensser, A. & Fleckenstein, B. T-cell transformation and oncogenesis by gamma2-herpesviruses. Adv. Cancer Res. 93, 91–128 (2005)
Biesinger, B., Trimble, J. J., Desrosiers, R. C. & Fleckenstein, B. The divergence between two oncogenic Herpesvirus saimiri strains in a genomic region related to the transforming phenotype. Virology 176, 505–514 (1990)
Lee, S. I., Murthy, S. C., Trimble, J. J., Desrosiers, R. C. & Steitz, J. A. Four novel U RNAs are encoded by a herpesvirus. Cell 54, 599–607 (1988)
Wassarman, D. A., Lee, S. I. & Steitz, J. A. Nucleotide sequence of HSUR 5 RNA from herpesvirus saimiri. Nucleic Acids Res. 17, 1258 (1989)
Golembe, T. J. et al. Lymphotropic Herpesvirus saimiri uses the SMN complex to assemble Sm cores on its small RNAs. Mol. Cell. Biol. 25, 602–611 (2005)
Cazalla, D., Yario, T. & Steitz, J. A. Down-regulation of a host microRNA by a Herpesvirus saimiri noncoding RNA. Science 328, 1563–1566 (2010)
Murthy, S., Kamine, J. & Desrosiers, R. C. Viral-encoded small RNAs in herpes virus saimiri induced tumors. EMBO J. 5, 1625–1632 (1986)
Albrecht, J. C. Primary structure of the Herpesvirus Ateles genome. J. Virol. 74, 1033–1037 (2000)
Cook, H. L., Mischo, H. E. & Steitz, J. A. The Herpesvirus saimiri small nuclear RNAs recruit AU-rich element-binding proteins but do not alter host AU-rich element-containing mRNA levels in virally transformed T cells. Mol. Cell. Biol. 24, 4522–4533 (2004)
Fan, X. C., Myer, V. E. & Steitz, J. A. AU-rich elements target small nuclear RNAs as well as mRNAs for rapid degradation. Genes Dev. 11, 2557–2568 (1997)
Myer, V. E., Lee, S. I. & Steitz, J. A. Viral small nuclear ribonucleoproteins bind a protein implicated in messenger RNA destabilization. Proc. Natl Acad. Sci. USA 89, 1296–1300 (1992)
Guo, Y. E., Riley, K. J., Iwasaki, A. & Steitz, J. A. Alternative capture of noncoding RNAs or protein-coding genes by herpesviruses to alter host T cell function. Mol. Cell 54, 67–79 (2014)
Calvet, J. P. & Pederson, T. Base-pairing interactions between small nuclear RNAs and nuclear RNA precursors as revealed by psoralen cross-linking in vivo. Cell 26, 363–370 (1981)
Maroney, P. A., Yu, Y., Fisher, J. & Nilsen, T. W. Evidence that microRNAs are associated with translating messenger RNAs in human cells. Nat. Struct. Mol. Biol. 13, 1102–1107 (2006)
Nottrott, S., Simard, M. J. & Richter, J. D. Human let-7a miRNA blocks protein production on actively translating polyribosomes. Nat. Struct. Mol. Biol. 13, 1108–1114 (2006)
Fouqué, A., Debure, L. & Legembre, P. The CD95/CD95L signaling pathway: a role in carcinogenesis. Biochim. Biophys. Acta 1846, 130–141 (2014)
Di Fiore, R., D’Anneo, A., Tesoriere, G. & Vento, R. RB1 in cancer: different mechanisms of RB1 inactivation and alterations of pRb pathway in tumorigenesis. J. Cell. Physiol. 228, 1676–1687 (2013)
Platanias, L. C. Mechanisms of type-I- and type-II-interferon-mediated signalling. Nat. Rev. Immunol. 5, 375–386 (2005)
Eichhorn, S. W. et al. mRNA destabilization is the dominant effect of mammalian microRNAs by the time substantial repression ensues. Mol. Cell 56, 104–115 (2014)
Murthy, S. C., Trimble, J. J. & Desrosiers, R. C. Deletion mutants of herpesvirus saimiri define an open reading frame necessary for transformation. J. Virol. 63, 3307–3314 (1989)
Fabian, M. R., Sonenberg, N. & Filipowicz, W. Regulation of mRNA translation and stability by microRNAs. Annu. Rev. Biochem. 79, 351–379 (2010)
Bosson, A. D., Zamudio, J. R. & Sharp, P. A. Endogenous miRNA and target concentrations determine susceptibility to potential ceRNA competition. Mol. Cell 56, 347–359 (2014)
Kim, H. H. et al. HuR recruits let-7/RISC to repress c-Myc expression. Genes Dev. 23, 1743–1748 (2009)
Storz, G., Vogel, J. & Wassarman, K. M. Regulation by small RNAs in bacteria: expanding frontiers. Mol. Cell 43, 880–891 (2011)
Lu, Z., Guan, X., Schmidt, C. A. & Matera, A. G. RIP-seq analysis of eukaryotic Sm proteins identifies three major categories of Sm-containing ribonucleoproteins. Genome Biol. 15, R7 (2014)
Tischer, B. K., von Einem, J., Kaufer, B. & Osterrieder, N. Two-step red-mediated recombination for versatile high-efficiency markerless DNA manipulation in Escherichia coli. Biotechniques 40, 191–197 (2006)
Tischer, B. K., Smith, G. A. & Osterrieder, N. En passant mutagenesis: a two step markerless red recombination system. Methods Mol. Biol. 634, 421–430 (2010)
Cazalla, D., Xie, M. & Steitz, J. A. A primate herpesvirus uses the integrator complex to generate viral microRNAs. Mol. Cell 43, 982–992 (2011)
White, R. E., Calderwood, M. A. & Whitehouse, A. Generation and precise modification of a herpesvirus saimiri bacterial artificial chromosome demonstrates that the terminal repeats are required for both virus production and episomal persistence. J. Gen. Virol. 84, 3393–3403 (2003)
Desrosiers, R. C., Burghoff, R. L., Bakker, A. & Kamine, J. Construction of replication-competent Herpesvirus saimiri deletion mutants. J. Virol. 49, 343–348 (1984)
Fickenscher, H. & Fleckenstein, B. in Methods in Microbiology Vol. 32 (eds Kaufmann, S. & Kabelitz, D. ) 824 (Elsevier, 2002)
Desrosiers, R. C., Silva, D. P., Waldron, L. M. & Letvin, N. L. Nononcogenic deletion mutants of Herpesvirus Saimiri are defective for in vitro immortalization. J. Virol. 57, 701–705 (1986)
Scammell, J. G., Wright, J. L. & Tuck-Muller, C. M. The origin of four squirrel monkey cell lines established by karyotype analysis. Cytogenet. Cell Genet. 93, 263–264 (2001)
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protocols 3, 1101–1108 (2008)
Ideue, T., Hino, K., Kitao, S., Yokoi, T. & Hirose, T. Efficient oligonucleotide-mediated degradation of nuclear noncoding RNAs in mammalian cultured cells. RNA 15, 1578–1587 (2009)
Chapnik, E. et al. miR-142 orchestrates a network of actin cytoskeleton regulators during megakaryopoiesis. eLife 3, e01964 (2014)
Liu, J. et al. MiR-142-3p attenuates the migration of CD4+ T cells through regulating actin cytoskeleton via RAC1 and ROCK2 in arteriosclerosis obliterans. PLoS One 9, e95514 (2014)
Bettencourt, P. et al. Actin-binding protein regulation by microRNAs as a novel microbial strategy to modulate phagocytosis by host cells: the case of N-Wasp and miR-142-3p. Front. Cell. Infect. Microbiol. 3, 19 (2013)
Acknowledgements
We thank A. Whitehouse, G. Smith, F. Wussow and K. Tischer for reagents; T. Toptan, H. Fickenscher, P. Medveczky and A. Bosque for technical advice; J. Marvin and C. Leukel from the Flow Cytometry Core Facility at the University of Utah for help with cell sorting and data analysis; and J. Steitz, J. Cáceres and J. Martínez for critical commentary. This work was supported by a grant from the National Institutes of Health (R01-GM118829).
Author information
Authors and Affiliations
Contributions
C.G. designed and performed most experiments, performed data interpretation, and co-wrote the manuscript. T.M. performed bioinformatic analyses. D.C. conceived and oversaw the project, designed and performed experiments, performed data interpretation, and co-wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks B. Cullen and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 HSUR2 binds host miRNA miR-142-3p and miR-16.
a, Nucleotide sequence of HSUR2. Predicted base-pairing between HSUR2 and miR-142-3p, as well as experimentally confirmed interaction between HSUR2 and miR-167, are shown with miRNA seed sequences in yellow and HSUR2 residues involved in interactions with miRNAs in blue. ARE-like sequences, red boxes; Sm-binding site, black box. b, Western blot analysis of miR-142-3p validated targets GRLF137, Rac138, Rock238, and N-WASP39 in marmoset T cells transformed by wild-type HVS (cj319-WT) or by a mutant version of HVS lacking HSUR1 and HSUR2 (cj319-Δ2a). α-Tubulin provides a loading control. Data shown are representative of three independent experiments.
Extended Data Figure 2
AMT-mediated crosslinking and purification of HSUR2 for RICC–seq. a, Northern blot analyses of purification of HSUR2 for RICC–seq. Cj319-WT cells were transiently transfected with control ASO, and UV crosslinked in the presence of AMT. Total RNA was prepared, polyA+ was selected, and HSUR2 was captured using a specific biotinylated ASO. RNA was isolated from different fractions, crosslinks were reversed, and analysed by northern blot for HSUR2 or U1 snRNA as a crosslinking and purification control. FT, flow-through; αH2, biotinylated HSUR2 ASO. b, Northern blot analyses of HSUR2 and HSUR7 (loading control) on total RNA prepared from cj319-WT cells transiently transfected with either control ASO or HSUR2 ASO. RNA samples prepared from these cells were subjected to the RICC–seq protocol as described in a.
Extended Data Figure 3 HSUR2 regulates the abundance of target mRNAs.
qRT–PCR analyses of HSUR2 target mRNAs in cj319-Δ2a cells, which do not express HSUR1 and HSUR210,21, after transfection with control or HSUR2 ASO (n = 3; two-sided t-test). Dots represent mean values of independent experiments with error bars representing s.d. See Supplementary Table 7 for full description of statistics.
Extended Data Figure 4 HSUR3, HSUR4, HSUR6 and HSUR7 are not required for HSUR2-mediated mRNA repression.
a, Plasmid constructs used for transient transfection of HSURs. GFP (green arrow) expression is driven by the cytomegalovirus promoter (blue box). HSUR encoding regions are represented by red arrows. HVS miRNAs29 are not depicted. b, Plasmid constructs used for transient expression of HSUR2. HSUR2 expression signals (DSE, distal sequence element; PSE, proximal sequence element) are represented by magenta boxes, and the 3′-end processing signal (3′-box) is represented by a purple box. c, Northern blot analysis of GSML cells transiently transfected with pBS-GFP (lane 1), pBS-GFP-H1-5 (lane 2), pBS-GFP-H3-7 (lane 3), and both pBS-GFP-H1-5 and pBS-GFP-H3-7 (lane 4). U2 snRNA provides a loading control. d, GSML cells were transiently transfected with the plasmid constructs described in a, FACS-sorted for GFP expression and analysed as described in Fig. 2e (two-sided t-test in all cases, except for NGDN and HSURs3–7 versus empty vector, which used a two-sided Wilcoxon’s signed rank test). Sample size (n) is indicated below the chart in each case. ****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; see Supplementary Table 7 for full description of statistics.
Extended Data Figure 5 HSUR2 represses target mRNAs in the human B cell line BJAB.
BJAB cells were transiently transfected with plasmids pBS-GFP (empty vector, white bars), pBS-GFP-HSUR2 (HSUR2), or pBS-GFP-ΔHSUR2 (ΔHSUR2), FACS-sorted for GFP expression, and analysed as described in Fig. 2e (two-sided t-test). Dots represent mean values of independent experiments with error bars representing s.d. Sample size (n) is indicated below the chart in each case. ***P < 0.001; **P < 0.01; *P < 0.05; see Supplementary Table 7 for full description of statistics.
Extended Data Figure 6 Construction of cj137-WT and cj137-ΔHSUR2 cell lines.
a, Schematic representation of the construction of HVS-BAC-GFP-WT and HVS-BAC-GFP-ΔHSUR2 bacmids. HVS-BAC-GFP was originally constructed using the HVS-A11-S4 strain31,33 with a cassette containing BAC elements, GFP gene, and chloramphenicol and hygromycin resistance genes placed in the ORF 15 gene30. PCR fragments containing the kanamycin resistance gene and I-SceI restriction site, flanked by a 50-bp duplication of HVS genomic sequence (brown boxes), used for the first recombination are depicted, with the numbers above the ends (in parentheses) indicating the corresponding positions in the HVS A11 genome. After a second recombination event, scarless removal of the kanamycin cassette is achieved, leaving the original 50 bp of genomic sequence (brown box). b, Restriction fragment patterns of recombinant BAC clones. Bacmids HVS-BAC-GFP, HVS-BAC-GFP-WT and HVS-BAC-GFP-ΔHSUR2 were digested with SacI, separated in a 0.7% agarose gel and stained with ethidium bromide. Arrows indicate alterations caused by the addition of recombined fragments. Left, sizes of molecular weight markers in kb. c, Northern blot analysis for HSUR expression on total RNA isolated from cj137-WT and cj137-ΔHSUR2 cell lines. U6 snRNA provides a loading control.
Extended Data Figure 7 Ablation of miR-142-3p and miR-16 expression does not affect HSUR2 expression or its ability to base-pair with mRNAs.
a, Northern blot analysis of total RNA isolated from GSML cells treated as described in Fig. 3a. U6 snRNA provides a loading control. b, AMT-mediated RNA-RNA crosslinking and polyA+ selection performed as in Fig. 1a, on HVS-transformed marmoset T cells transiently transfected with the indicated LNA inhibitors.
Extended Data Figure 8 Interaction of HSUR2 with miR-142-3p and miR-16 is required for HSUR2-mediated mRNA repression.
a, Northern blot analysis of total RNA isolated from GSML cells that were treated as described in Fig. 3c. U6 snRNA provides a loading control. b, As in Fig. 3c, for the other targets of HSUR2. Analysed by two-sided t-test in all cases, except for STAG1, RBM7, RASA2, DDX20, and DDX5 (HSUR2Δ142-3p/16 versus empty vector in all cases), for which a two-sided Wilcoxon signed rank test was used. Dots represent mean values of independent experiments with error bars representing s.d. Sample size (n) is indicated below the chart in each case. ****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05; see Supplementary Table 7 for full description of statistics.
Extended Data Figure 9 Mutant versions of miR-142-3p and miR-16 restore HSUR2-mediated mRNA repression.
a, b, As in Fig. 3d, e, for the other targets of HSUR2 (n = 3; two-sided t-test). Dots represent mean values of independent experiments with error bars representing s.d. Sample size (n) is indicated below the chart in each case. ***P < 0.001; **P < 0.01; *P < 0.05; see Supplementary Table 7 for full description of statistics.
Extended Data Figure 10 HSUR2 confers resistance to apoptosis.
a, b, As in Fig. 4c, d, but cells were treated with TNFα (n = 3; unpaired two-sided t-test). Dots represent mean values of independent experiments with error bars representing s.d. *P < 0.05; see Supplementary Table 7 for full description of statistics.
Supplementary information
Supplementary Figures
This file contains Supplementary Figure 1, the uncropped blots and Supplementary Figure 2, examples of the gating for isolation of cells used in this study. (PDF 4978 kb)
Supplementary Table 1
This file contains a list of HSUR 2 targets identified by RICC-seq. Gene name, genomic coordinates, and ratios of HSUR 2-positive pull down versus HSUR 2-negative pull down and in the HSUR 2-positive pull down versus HSUR 2-positive input are indicated. (XLSX 43 kb)
Supplementary Table 2
This file contains all results from our Ingenuity Pathway Analysis. (XLSX 10 kb)
Supplementary Table 3
This file lists the abundance (in molecules/cell) of miR-142-3p, miR-16, HSUR 2, and ten HSUR 2 target mRNAs. (XLSX 57 kb)
Supplementary Table 4
This file contains the sequences of all oligonucleotides used. (XLSX 41 kb)
Supplementary Table 5
This file contains source data for main figures. (XLSX 247 kb)
Supplementary Table 6
This file contains source data Extended Data figures. (XLSX 300 kb)
Supplementary Table 7
This file contains detailed description of number of replicates, statistical tests, and p-values for main as well as Extended Data figures. (XLSX 39 kb)
Rights and permissions
About this article
Cite this article
Gorbea, C., Mosbruger, T. & Cazalla, D. A viral Sm-class RNA base-pairs with mRNAs and recruits microRNAs to inhibit apoptosis. Nature 550, 275–279 (2017). https://doi.org/10.1038/nature24034
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24034
- Springer Nature Limited
This article is cited by
-
circEIF3I facilitates the recruitment of SMAD3 to early endosomes to promote TGF-β signalling pathway-mediated activation of MMPs in pancreatic cancer
Molecular Cancer (2023)
-
MicroRNA-mediated reprogramming of glucose, fatty acid and amino acid metabolism in cancer
Genome Instability & Disease (2022)
-
Learning noncoding RNA biology from viruses
Mammalian Genome (2022)
-
Developing therapeutic approaches for twenty-first-century emerging infectious viral diseases
Nature Medicine (2021)
-
A structured RNA motif locks Argonaute2:miR-122 onto the 5’ end of the HCV genome
Nature Communications (2021)