Abstract
In mammals, chromatin organization undergoes drastic reprogramming after fertilization1. However, the three-dimensional structure of chromatin and its reprogramming in preimplantation development remain poorly understood. Here, by developing a low-input Hi-C (genome-wide chromosome conformation capture) approach, we examined the reprogramming of chromatin organization during early development in mice. We found that oocytes in metaphase II show homogeneous chromatin folding that lacks detectable topologically associating domains (TADs) and chromatin compartments. Strikingly, chromatin shows greatly diminished higher-order structure after fertilization. Unexpectedly, the subsequent establishment of chromatin organization is a prolonged process that extends through preimplantation development, as characterized by slow consolidation of TADs and segregation of chromatin compartments. The two sets of parental chromosomes are spatially separated from each other and display distinct compartmentalization in zygotes. Such allele separation and allelic compartmentalization can be found as late as the 8-cell stage. Finally, we show that chromatin compaction in preimplantation embryos can partially proceed in the absence of zygotic transcription and is a multi-level hierarchical process. Taken together, our data suggest that chromatin may exist in a markedly relaxed state after fertilization, followed by progressive maturation of higher-order chromatin architecture during early development.
Similar content being viewed by others
Main
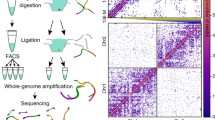
To examine three-dimensional chromatin structure in early development, we developed a low-input Hi-C method based on proximity ligation within nuclei2 and in situ Hi-C3 (Extended Data Fig. 1; see Methods), which we termed small-scale in situ Hi-C (sisHi-C). This method can generate high-quality Hi-C data using only 500 cells that accurately recapitulate chromatin interaction patterns derived from millions of cells4 (Extended Data Fig. 2a–c). We then crossed C57BL/6N female mice and PWK/PhJ male mice, and collected gametes, pronuclear stage 5 (PN5) zygotes, early 2-cell, late 2-cell, 8-cell embryos, and inner cell masses (ICMs) from blastocysts (Fig. 1; see Methods). We conducted sisHi-C for each stage and obtained high coverage data for most stages, including 134–192 million monoclonal read pairs for early embryos from long-distance (more than 20 kb) intra-chromosomal interactions (replicates combined) (Supplementary Table 1). These Hi-C data are highly reproducible among the replicates, and our data in sperm are also consistent with published data5 (Extended Data Fig. 2d–f). We then examined chromatin organization in MII oocytes, which are arrested in the metaphase of meiosis II. Consistent with the segregation of individual chromosomes6, MII oocytes showed a lower percentage of inter-chromosomal read pairs than did early embryos (Extended Data Fig. 3a). Strikingly, MII oocytes lacked typical chromatin higher-order structures, including TADs4,7 and chromatin compartments (represented by plaid chromatin interaction patterns8) (Fig. 1). Instead, these cells show a uniform interaction pattern along the entire chromosomes that appears to be locus-independent, which strongly resembles that of mitotic chromatin9 (Extended Data Fig. 3b). Notably, the interactions decrease abruptly beyond 4 Mb (Extended Data Fig. 3b). This ‘interaction insulation boundary’, which is likely to reflect the interaction unit in a linearly organized array of consecutive chromatin loops9, appears to be shorter than that (10 Mb) of human mitotic chromatin9. These data indicate that the chromatin of MII oocytes is in a uniform folding configuration that lacks both TADs and chromatin compartments.
Heatmaps showing normalized Hi-C interaction frequencies (100-kb bin, chromosome 2) in mouse gametes and preimplantation embryos (pooled data from 2–4 biological replicates). Zoomed-in views (40-kb bin) are also shown.
We then investigated the dynamics of chromatin architecture after fertilization. Strikingly, most chromatin interactions are restricted to local regions in PN5 zygotes, with very weak TADs and sparse distal interactions compared to those of later stage embryos (Fig. 1; Extended Data Fig. 4a). Similarly, we observed weak TADs and distal chromatin interactions in both early and late 2-cell embryos. During the mitotic cycle, TADs are dissolved during mitosis before being rapidly established once cells exit mitosis9. However, the diminished higher-order chromatin structure in early embryos is not simply due to cell cycle, as it was observed in the PN5 zygote (G2), early (G1) and late (S–G2) 2-cell stages. TADs and distal interactions gradually become more evident as development proceeded, indicating increasing chromatin compaction (Extended Data Fig. 4a, b, discussed below). The presumed loss of TADs in mitosis9 raises the possibility that chromatin organization is slowly re-established after each mitosis in early development but with growing kinetics. Together, these data demonstrate unexpectedly relaxed chromatin states after fertilization, with weak TADs and depleted distal chromatin interactions.
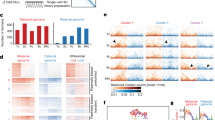
To investigate how TADs are established during early development, we identified TAD boundaries using insulation score10 in ICMs (n = 2,121) where TADs appear to be well established (Extended Data Fig. 4a, Supplementary Table 2). The majority of ICM TAD boundaries (80.7%) were also present in mouse embryonic stem cells (mES cells) (Extended Data Fig. 4c). Using differential interaction heatmap analysis11 at individual loci (Extended Data Fig. 4b) and across the genome (Fig. 2a), we found that these TADs showed increased intra-domain interactions between nearby regions at early stages, and later between distal regions within the domains. We further calculated a ‘consolidation score’, which is defined as the ratio between average interaction frequency within each TAD and that from local background (Extended Data Fig. 4d; see Methods). We confirmed that consolidation scores increased gradually during early development, indicating the maturation of TADs (Extended Data Fig. 4e). Consistently, we also observed progressive insulation around TAD boundaries (Fig. 2a, b). The insulation of boundaries was observed as early as in PN5 zygotes (P = 2.86 × 10−16 compared to a random control, two-tailed t-test, see Methods) (Fig. 2b and Extended Data Fig. 5a), as confirmed using directionality index4 (Extended Data Fig. 5b). We considered TADs at early stages to be in ‘priming’ states (characterized by weak intra-TAD interactions), in contrast to TADs in ‘mature’ states at later stages (showing strong intra-TAD interactions). Together, these data demonstrate step-wise establishment of TADs in early development.
a, Heatmaps showing the normalized average interaction frequencies for all TADs (defined in ICM) as well as their nearby regions (±0.5 TAD length) (top), and differential interactions between consecutive stages (replicates pooled, n = 2–4) (bottom). b, The average insulation scores at TADs (defined in ICM) and nearby regions are shown. Insulation scores generated by a random valid read pair data set (see Methods) are also shown as a control.
To investigate whether the two parental alleles show differential reprogramming of chromatin topological structure, we assigned Hi-C sequencing reads to their parental origins based on single nucleotide polymorphisms (SNPs) (see Methods). We observed few inter-chromosomal read pairs between the two parental genomes in PN5 zygotes, indicating that the parental genomes were spatially segregated despite the fusion of pronuclei (Extended Data Fig. 6a). Such spatial segregation can be found as late as the 8-cell stage (P < 1 × 10−300 compared to cortex (control), two-tailed t-test). Notably, the paternal allele in PN5 zygotes appeared to have fewer distal interactions than the maternal allele, before the two genomes gradually converged at later stages (Fig. 3a and Extended Data Fig. 6b). These data raise the possibility that perhaps the paternal chromatin is more relaxed at the PN5 zygote stage. The sisHi-C analysis of an even earlier stage, the PN3 zygote, also revealed similar results (Extended Data Fig. 6c). Together with the depletion of TADs on both alleles (Extended Data Fig. 6d), these data indicate that the sperm chromatin organization has largely been disassembled by the PN3 stage after protamine-histone exchange1. In sum, our results suggest that the two parental genomes remain partially segregated as late as the 8-cell stage, and show differential chromatin organization at early stages.
a, Heatmaps showing allelic chromatin interaction frequencies (100-kb bin; pooled data from 2–4 biological replicates; chromosome 15). b, Correlation heatmaps showing correlations between any two region pairs along the chromosome for their intra-chromosomal interaction frequency patterns (300-kb bin; chromosome 12). P, paternal; M, maternal. The principal component (PC) 1 values are also shown. c, Boxplots showing the ratios for average interaction frequency between different classes of compartments (AB) compared to those between the same classes of compartments (AA and BB) for each chromosome (X chromosome excluded) of each replicate separately (n = 2–4). P values calculated by Wilcoxon rank-sum test (two-tailed, with Benjamini–Hochberg multiple testing correction) are also shown.
The genome is typically organized into large, self-interacting chromatin compartments A and B, resulting in plaid patterns in chromatin interaction or the derived correlation heatmap8. Despite the weak distal chromatin interactions, our correlation heatmap analysis revealed plaid patterns of chromatin interactions, although less well segregated, at early stages (Fig. 3b). We first identified compartments8 (see Methods) in ICM which we considered to be in a ‘mature’ state, and examined the interactions between compartments (defined in ICM) across stages. We found more inter-compartment interactions (between A–B compartment pairs along the same chromosomes) at early stages than at late stages (Fig. 3c) (for example, P < 9.7 × 10−11 between PN5 zygote and ICM). Notably, the chromatin compartment is clearly visible for the paternal allele (Fig. 3b). By contrast, the maternal genome in PN5 zygotes appears to be poorly segregated (Fig. 3b) with frequent contacts across compartments A and B (P = 2.2 × 10−9) (Fig. 3c), and its correlation matrix showed lower correlation with that of ICM (Extended Data Fig. 7a). This was also true for the earlier stage PN3 zygote (Fig. 3c). These results are further echoed by a global clustering analysis demonstrating that the two parental alleles show differential compartment patterns at early stages but are clustered together at late stages (Extended Data Fig. 7b). The allelic differences in chromatin compartments can be found as late as the 8-cell embryo stage (Fig. 3b, c). We then attempted to identify compartments A and B at non-ICM stages. Despite the relatively poor chromatin compartmentalization at early stages, the positions of chromatin compartments were largely consistent from early to late stages, especially from the late 2-cell stage onward (Extended Data Fig. 7c, Supplementary Table 3). As validations, compartment A, but not compartment B, was correlated with enrichment of accessible chromatin12 and gene expression at these stages (Extended Data Fig. 7c–e). Together, these data suggest that chromatin compartmentalization is observed as early as in zygotes for the paternal genome, followed by further segregation of compartments A and B on both alleles in preimplantation development.
One intriguing question is whether the maturation of chromatin organization in early development requires zygotic transcription. To investigate this, we blocked transcription with alpha-amanitin (see Methods), which also arrested embryos at the late 2-cell stage13. We then collected these embryos for Hi-C analyses when the control group had grown to the late 2-cell (20 h) or 8-cell stage (45 h) (Extended Data Fig. 8a, b). Unexpectedly, we found that TADs continued to consolidate in the presence of alpha-amanitin (Fig. 4a and Extended Data Fig. 8c, d), indicating that the maturation of higher-order chromatin organization can at least partially proceed in the absence of zygotic transcription. These data also suggest that the weak chromatin organization in early development is due to unusually slow establishment rather than early breakdown.
a, Hi-C interaction heatmaps (40-kb bin) showing an example region on chromosome 13 for the establishment of TADs in embryos (pooled data from two biological replicates) with or without alpha-amanitin. b, A schematic model showing the reprogramming of chromatin organization in early mouse development. The MII oocytes are characterized by a uniform chromatin configuration that lacks both TADs and compartments. TADs start to appear in zygotes in the priming state (open red circles) and become mature at the later stage (solid red circles). Long-distance chromatin interactions and chromatin compaction increase as development proceeds (indicated by the shortening horizontal lengths). Weak chromatin compartments (A/B) appear first on the paternal genomes of zygotes, along with much weaker or non-existent compartments on the maternal genome, and become increasingly strong on both alleles at later stages. Open chromatin and closed chromatin are shown as yellow and blue loops, respectively.
Finally, we attempted to decode the spatiotemporal chromatin packaging in early development from a global view, by examining and comparing how the chromatin contact probability (P(s)) depends on genomic distance (s) among different stages. As the interaction is strongly dominated by short-distance interactions (Extended Data Fig. 9a), we computed the relative interaction probability by normalizing the distance effects against a reference curve (P(s) ~ s−1 in this case, which represents the expected interaction–distance relationship for the fractal globule state8) (Extended Data Fig. 9b). As a validation, this analysis showed relative depletion of local interactions (<0.6 Mb, indicating lack of TADs) and enrichment of distal interactions (~1–7 Mb) for MII oocytes (Extended Data Figs 3b, 9b). By contrast, sperm chromatin showed strong interactions over even longer distances (>15 Mb), indicating tight packaging5. Consistent with possible chromatin relaxation after fertilization, chromatin interactions over 1 Mb were generally reduced in PN5 zygotes (Extended Data Fig. 9b). Distinct interaction patterns were evident between the two alleles in zygotes, and the differences gradually diminished during development (Extended Data Figs 9b, 10a–e). Notably, from the zygote stage onward, we observed chromatin compaction primarily at three levels. First, both alleles showed increasing chromatin interactions within 1 Mb (short-distance chromatin folding) from PN5 zygotes to ICMs, indicating hierarchical consolidation of TADs (Extended Data Figs 4a, 9c). We also observed a second type of strong chromatin interaction at ‘long distances’ (2–20 Mb) from zygotes to early 2-cell embryos (Extended Data Fig. 9b, c). These long-distance interactions started to decrease beyond 15 Mb, showing an interaction insulation boundary (Extended Data Figs 9c, 10b, 10f (arrowheads)). Such long-distance interactions were also observed in late 2-cell and 8-cell embryos to a lesser extent, but were not apparent in ICMs (Fig. 1 and Extended Data Fig. 10c–e). As the long-distance interaction boundary partially resembles that for mitotic chromatin, but with larger distances, these data raise the question of whether such chromatin organization represents a transition state between interphase and mitotic chromatin. Finally, we found increased chromatin interactions for extra-long-distance region pairs (>20 Mb) specifically in ICMs (Extended Data Figs 9b, c, 10e), which may reflect the ultimate compaction of chromatin, allowing interactions between even more distant regions. Therefore, chromatin packaging in early development is likely to occur in a coordinated manner at different distance levels (Fig. 4b).
Chromatin undergoes marked reorganization during early development in mammals. However, the molecular basis of the reprogramming of higher-order chromatin structure in this process remains unclear. Here, using an improved Hi-C approach, we examined 3D chromatin architecture in mouse gametes and preimplantation embryos (Fig. 4b). Unexpectedly, although TADs appear as early as the zygote stage, they are largely in priming states at early stages, being characterized by weak consolidation and boundary insulation. Together with the lack of strong distal interactions and weak chromatin compartmentalization, these data indicate that interphase chromatin is likely to be in a relatively relaxed state after fertilization12,14,15,16. Notably, the maturation of 3D chromatin architecture in early development is partially independent of transcription, a finding that echoes a recent study in fly17. Chromatin organization undergoes stage-specific regulation during the cell cycle9,18 and our Hi-C data cover a wide spectrum of cell cycle stages, including PN3 zygotes (S), PN5 zygotes (G2, with clearly visible pronuclei), early 2-cell (G1), and late 2-cell (S–G2). Cells become more asynchronized in 8-cell embryos and ICMs. As weak TADs and compartments were observed at all early stages from PN3 zygotes to late 2-cell embryos, these data suggest that chromatin organization in early development is likely to be a combinatorial result of generally relaxed architecture and cell cycle stage. We speculate that, in early development, the chromatin architecture is slowly re-established after each mitosis but with increasing kinetics. While our paper was under revision, a separate study reported the chromatin structure of GV-stage mouse oocytes (compared to MII oocytes in our study) and allele-specific compartmentalization in zygotes using single-nucleus Hi-C19. Future studies are needed to identify the key factors and molecular mechanisms that underlie the slow kinetics of chromatin assembly and the establishment of 3D chromatin architecture in early development.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Early embryo, oocyte and sperm collection
Preimplantation embryos were collected from 5–6-week-old C57BL/6N female mice (Vital River) mated with PWK/PhJ males (Jackson Laboratory). To induce ovulation, females were treated with 5 IU human chorionic gonadotropin (hCG) intraperitoneally, 44–48 h after injection of 5 IU pregnant mare’s serum gonadotropin (PMSG) (San-Sheng Pharmaceutical Co. Ltd). Each set of embryos at a particular stage was collected from the reproductive tract at defined time periods after hCG administration: 20 h (MII oocyte), 22 h (PN3 zygote), 27–28 h (PN5 zygote), 30 h (early 2-cell), 43 h (late 2-cell), 68–70 h (8-cell) and 92–94 h (blastocysts) in Hepes-buffered CZB medium. Embryos were selected by cell numbers or morphology with the zona pellucida gently removed by treatment with 10 IU/ml pronase (Sigma P8811) for several minutes. The embryos were then manually picked and prepared for the Hi-C experiments. Blastocysts were incubated in a 1:3 dilution of anti-mouse rabbit serum in DMEM for 20 min, washed in PBS and further incubated for 20 min in a 1:5 dilution of rat serum in DMEM for the complement reaction. The ICM was subsequently cleaned from lysed trophectoderm with a narrow glass pipette. Mature mouse sperm cells were obtained from 8-week-old PWK/PhJ males with a swim-up procedure to avoid somatic contamination20. Sperm were first squeezed out from the cauda epididymis and placed in Hepes-buffered CZB medium for 4 h at 37 °C. Only the top fractions containing motile sperm were collected. Cortex was isolated from 4-week-old PWK/PhJ × C57BL/6N F1 mice. All animal maintenance and experimental procedures were carried out according to the guidelines of the Institutional Animal Care and Use Committee (IACUC) of Tsinghua University, Beijing, China.
To inhibit transcription in early embryos, PN3 zygotes were cultured in CZB supplemented with alpha-amanitin (100 μg/ml) for about 20 h or 45 h.
Cell culture
The mouse R1 ES cell line was derived from 129X1/SvJ × 129S1 F1 mice and was a gift from Y.-H. Jiang at Duke University. Mouse ES cells were cultured without irradiated mouse embryonic fibroblasts (MEFs) in DMEM containing 15% FBS, leukaemia inhibiting factor (LIF), penicillin/streptomycin, l-glutamine, β-mercaptoethanol, and non-essential amino acids. These cells were tested and found to be free of mycoplasma contamination.
sisHi-C library generation and sequencing
The procedure for sisHi-C is similar to that for in situ Hi-C3, with further optimization for low input cells achieved by scaling down the reaction volume, reducing experimental procedures and minimizing tube exchanges to avoid sample loss. Briefly, embryos or mouse ES cells were fixed with 1% formaldehyde at room temperature (RT) for 10 min. Formaldehyde was quenched with glycine for 10 min at RT. Embryos or mouse ES cells were then washed twice with 1 × PBS. The exchange of buffers was done by transferring embryos or mouse ES cells with a mouth capillary pipette. Embryos were lysed in 50 μl lysis buffer (10 mM Tris-HCl pH 7.4, 10 mM NaCl, 0.1 mM EDTA, 0.5% NP-40 and proteinase inhibitor) on ice for 50 min. After spinning at 3,000 r.p.m. at 4 °C for 5 min, the supernatant was discarded carefully with a pipette. Chromatin was solubilized in 10 μl 0.5% SDS and incubated at 62 °C for 10 min. SDS was quenched with 5 μl 10% Triton X-100 at 37 °C for 30 min. Then the nuclei were digested with 50 U MboI at 37 °C overnight with rotation with a total volume of 50 μl. MboI was then inactivated at 62 °C for 20 min. To fill in the biotin to the DNA, 1.5 μl 1 mM dATP, 1.5 μl 1 mM dGTP, 1.5 μl 1 mM dTTP, 3.75 μl 0.4 mM biotin-14-dCTP and 10 U Klenow were added to the solution and the reaction was carried out at 37 °C for 1.5 h with rotation. After adding 60 μl ligation mix (38.8 μl water, 12 μl 10 × NEB T4 DNA ligase buffer, 7 μl 10% Triton X-100, 1.2 μl 10mg/ml BSA and 1 μl 400U/μl T4 DNA ligase), the fragments were ligated at RT for 6 h with rotation. This was followed by reversal of crosslinking and DNA purification. DNA was sheared to 300–500 bp with Covaris M220. The biotin-labelled DNA was then pulled down with 10 μl Dynabeads MyOne Streptavidin C1 (Life Technology). Sequencing library preparation was performed on beads, including end repair, dATP tailing and adaptor ligation. DNA was eluted twice by adding 20 μl water to the tube and incubating at 66 °C for 20 min. 12–15 cycles of PCR amplification were performed with Extaq (Takara). Finally, size selection was done with AMPure XP beads and fragments ranging from 200 bp to 1,000 bp were selected. All the libraries were sequenced on Illumina HiSeq2500 or HiSeq XTen according to the manufacturer’s instruction. For sperm samples, the lysis buffer (same as for mouse embryos) was added with 0.05% l-α-lysophosphatidylcholine (Sigma L4129). The cortex was crosslinked with 2% formaldehyde for 20 min at RT and was homogenized before lysis as described previously21.
RNA sequencing library preparation and sequencing
The RNA sequencing (RNA-seq) libraries were generated with alpha-amanitin-treated and control embryos using the Smart-seq2 protocol as described previously with minor modifications22. Cells were lysed in hypotonic lysis buffer (Amresco, M334), and the polyadenylated mRNAs were captured with PolyT primers. After lysis for about 3–10 min at 72 °C, the Smart-seq2 reverse transcription reactions were performed. After pre-amplification and AMPure XP bead purification, cDNAs were sheared with Covaris and were subject to Illumina TruSeq library preparation. All libraries were sequenced on Illumina HiSeq 2500 according to the manufacturer’s instructions.
Hi-C data mapping
Paired-end raw reads of Hi-C libraries were aligned, processed and iteratively corrected using HiCPro (version 2.7.1b) as described23. Briefly, sequencing reads were first independently aligned to the mouse reference genome (mm9) using the bowtie2 end-to-end algorithm and ‘-very-sensitive’ option. To rescue the chimaeric fragments spanning the ligation junction, the ligation site was detected and the 5′ fraction of the reads was aligned back to the reference genome. Unmapped reads, multiple mapped reads and singletons were then discarded. Pairs of aligned reads were then assigned to MboI restriction fragments. Read pairs from uncut DNA, self-circle ligation and PCR artefacts were filtered out and the valid read pairs involving two different restriction fragments were used to build the contact matrix. Valid read pairs were then binned at a specific resolution by dividing the genome into bins of equal size. We chose 100-kb or 300-kb bin size for examination of global interaction patterns of the genome, and 40-kb bin size to show local interactions and to perform TAD calling. The binned interaction matrices were then normalized using the iterative correction method23,24 to correct for biases such as GC content, mappability and effective fragment length in Hi-C data.
To eliminate the possible effects on data analyses of variable sequencing depths, we randomly sampled equal numbers of long range (>20-kb) intra-chromosomal read pairs (n = 115 million) from each stage for most downstream analyses involving comparison analyses among stages.
Allele assignment of sequencing reads
Allelic interaction frequency matrices were generated with HiCPro23 using the SNPs between two mouse strains (C57BL/6N and PWK/PhJ). Briefly, for allele-specific analysis, the paired end reads were first aligned to a modified mm9 genome where all polymorphic sites were N-masked. Then the polymorphic sites were identified on the aligned reads. Reads without SNP information or containing conflicting allelic polymorphic sites were classified as unassigned. All the read pairs for which both reads were assigned to the same parental allele or for which one read was assigned to one parental allele and the other was unassigned were classifying as allelic reads for downstream analyses.
RNA-seq data processing
All RNA-seq data were mapped to the mouse reference genome (mm9) by TopHat (version 2.0.11). The gene expression level was calculated by Cufflinks (version 2.0.2)25 using the refFlat database from the UCSC genome browser.
Validation of sisHi-C data
The correlation between sisHi-C and conventional Hi-C and between sisHi-C replicates was calculated as following: the interaction frequency was generated for each pair of 100-kb bins. As the interaction matrix was highly skewed towards proximal interactions, we restricted the analysis to a maximum distance of 50 bins (5 Mb) as previously described4. Interaction frequencies were compared between different samples and Pearson correlation coefficients were calculated.
Hi-C interaction heatmap, differential interaction heatmap, and correlation heatmap
All the Hi-C interaction frequency heatmaps of whole chromosomes and the zoom-in views were generated using HiCPlotter version 0.6.05.compare, a Hi-C data visualization tool26. The ‘triangle’ interaction heatmaps to show TADs were generated with 3D Genome Browser (http://www.3dgenome.org). The interactions between loci were shown on 2D heatmaps along a colour scale using the normalized contact matrices.
To demonstrate the establishment of TADs, differential Hi-C interaction heatmaps were calculated as previously described27 with some modifications. In brief, sequencing-depth normalized interaction matrices (40-kb bin) of one stage were subtracted from the next stage. In the differential matrices, positive values indicate that the interaction frequency at the second stage is higher than the first stage, and vice versa.
To generate the correlation heatmap, the total or allelic correlation matrices for each stage were generated as previously described8. To plot the correlation heatmap of two alleles, we first combined the correlation matrix of each allele into one matrix divided by the diagonal. The heatmap was then generated with Java TreeView to ensure that the two alleles were presented with the same parameters.
Comparison of interaction frequencies between developmental stages
To compare interaction frequencies between developmental stages, we first combined the Hi-C data of replicates of each stage. The interaction matrices were then normalized for sequencing depth as previously described27 by making the sum of all interaction frequencies for a given chromosome at each stage equal to that of an arbitrarily selected ‘standard stage’ (ICM in this case).
Analysis of inter-chromosomal read pairs
To examine the possible segregation of the two parental genomes in early development, inter-chromosomal read pairs between the two parental genomes (maternal–paternal, MP) and those between the same parental genomes (maternal–maternal, MM, or paternal–paternal, PP) were counted. For each pair of different chromosomes (chromosomes 1–2, 1–3, 1–4 and so on), the number ratios between read pairs from differential parental genomes and read pairs from the same parental genomes were calculated. Boxplots were used to show the distribution of the ratios for all pairs of chromosomes.
Analysis of TADs
TAD boundaries were identified by calculating the insulation score for each bin using the 40-kb resolution Hi-C data as previously described with minor modifications10. In brief, the insulation score was calculated by sliding a 1 Mb × 1 Mb square along the diagonal of the interaction matrix for every chromosome. A 200-kb window was used for calculation of the delta vector and all boundaries with a ‘boundary strength’ <0.25 were removed. Insulation scores were plotted around all ICM TADs as well as their nearby regions (± 0.5 TAD length). Directionality index (DI) scores were calculated using a previously described pipeline4. The heatmaps were binned at 40-kb resolution and a 2-Mb window was used. DI scores were plotted around boundaries from 500 kb upstream to 500 kb downstream. The random control data set was generated by shifting all PN5 zygote valid pairs to random loci in the same chromosome without altering the distances between the pairs. Interaction frequency, insulation scores and DI values were then computed using this random read pair data set. The insulation scores at ICM TAD boundaries of PN5 zygote and a random control were used for a two-tailed t-test.
Average interaction heatmap of TADs
We used all TADs in ICMs as representatives of mature TADs, and plotted the composite interaction frequency by averaging all TADs along early development. The resulted matrices were then normalized by the average levels of the matrix values to make the sum of matrices for different stages equal. To generate the differential heatmap for average interactions of TADs, interaction matrices of one stage were subtracted from the second stage. In the differential matrices, positive values indicate that the interaction frequency at the second stage is higher than at the first stage, and vice versa.
Consolidation score
To statistically compare the consolidation levels of TADs between different samples, we developed a TAD consolidation score to quantify the states of TADs, which is defined as the ratio of average interaction frequency within each TAD (excluding short-distance interactions <400 kb) and the local background interaction frequency from nearby non-TAD regions. High scores indicate strong consolidation of TADs.
Hierarchical clustering analysis
The hierarchical clustering analysis based on the interaction correlation matrix at various stages for two parental alleles was conducted using an R package (ape) based on the Pearson correlation as indicated between each pair of data sets. The distance between two data sets was calculated by (1 − correlation).
Identification of chromatin compartments
A and B compartments were identified as described previously11 with some modifications. The expected interaction matrices were calculated after removing the bins that had no interactions with any other bins, most of which were unmappable regions in the genome. For normalized 100-kb interaction matrices, observed/expected matrices were generated using a sliding window approach11 with a bin size of 400 kb and a step size of 100 kb. For normalized 300-kb interaction matrices, a bin size of 600 kb and a step size of 300 kb were used. Finally, principal component analysis was performed on the correlation matrices generated from the observed/expected matrices. The first principal component of the correlation matrix coupled with gene density was used to identify A/B compartments. Identification of A and B compartments on each allele was performed similarly. Chromosome 14, which consistently showed incorrect compartment calling (probably owing to a mapping issue), and chromosome X were excluded from the downstream analysis.
Identification of gene dense regions
The genome was split into 1-Mb bins and genes located in each bin were counted. Those bins with more than 10 genes were identified as gene-dense regions.
Analysis of inter-compartment interactions
To compute the inter-compartment interactions between the same classes or different classes, we first removed local interactions that were shorter than 2 Mb, which mainly reflect interactions within TADs rather than long-distance interactions between compartments. The remaining interactions were assigned to two categories: interactions between two bins located in the same class of compartments (including A–A interactions and B–B interactions) and interactions between two bins located in different classes of compartments (A–B interactions). Compartments defined in ICMs were considered as mature compartments and their positions were used for all stages in this analysis. For each stage, the average interaction frequency between a pair of bins was calculated for each of the two categories for each chromosome. Then the ratios between the average interaction frequency per pair of A–B interactions and per pair of A–A or B–B interactions were calculated for each chromosome. Boxplots were used to show the ranges of such ratios for all chromosomes (chr14 and X chromosome excluded) and to measure the degree of compartment segregation for each stage.
Analysis of chromatin accessibility in compartments A/B
To examine the relationship of chromatin accessibility and chromatin compartments in general, ATAC-seq enrichment was calculated as reads per kilobase of transcript per million mapped reads (RPKM) (100-bp bin) for the entire genome. For each compartment bin (300 kb), the average ATAC-seq enrichment was computed and ATAC-seq signals in all 300-kb bins assigned as compartments A or B are shown in boxplots.
Analysis of gene expression in compartments A/B
The ZGA-only genes were selected by requiring FPKM <0.5 in MII oocytes and FPKM >1 at any of the developmental stages including zygotes, early 2-cell, 2-cell, 4-cell and 8-cell embryos, and ICMs. The expression levels of the ZGA genes in compartment A or B are shown as a boxplot.
P(s) analysis
P(s) was calculated with normalized interaction matrices at 100-kb resolution as previously described9. We first divided distances into logarithmically spaced bins with increasing factor 1.15: (100 kb, 100 kb × 1.15, 100 kb × 1.152…). Then, for each bin, we counted the number of interactions at corresponding distances. To obtain the probability P(s), we divided the number of interactions in each bin by the total number of possible region pairs. P(s) was further normalized so that the sum over the range of the distances was 1. As chromatin interactions are strongly correlated with genomic distance, we normalized the P(s) values of each stage to the P(s) ~ s−1 values. The resulting matrices were used to generate the normalized P(s) heatmap.
Data availability
All sequencing data that support the findings of this study have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (GEO) under accession number GSE82185. All other relevant data are available from the corresponding author on request.
Accession codes
References
Burton, A. & Torres-Padilla, M. E. Chromatin dynamics in the regulation of cell fate allocation during early embryogenesis. Nat. Rev. Mol. Cell Biol. 15, 723–734 (2014)
Gavrilov, A. A. et al. Disclosure of a structural milieu for the proximity ligation reveals the elusive nature of an active chromatin hub. Nucleic Acids Res. 41, 3563–3575 (2013)
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014)
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012)
Battulin, N. et al. Comparison of the three-dimensional organization of sperm and fibroblast genomes using the Hi-C approach. Genome Biol. 16, 77 (2015)
Sasaki, H. & Matsui, Y. Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nat. Rev. Genet. 9, 129–140 (2008)
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012)
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009)
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013)
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015)
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015)
Wu, J. et al. The landscape of accessible chromatin in mammalian preimplantation embryos. Nature 534, 652–657 (2016)
Qiu, J. J. et al. Delay of ZGA initiation occurred in 2-cell blocked mouse embryos. Cell Res. 13, 179–185 (2003)
Aoki, F., Worrad, D. M. & Schultz, R. M. Regulation of transcriptional activity during the first and second cell cycles in the preimplantation mouse embryo. Dev. Biol. 181, 296–307 (1997)
Schultz, R. M. The molecular foundations of the maternal to zygotic transition in the preimplantation embryo. Hum. Reprod. Update 8, 323–331 (2002)
Abe, K. et al. The first murine zygotic transcription is promiscuous and uncoupled from splicing and 3′ processing. EMBO J. 34, 1523–1537 (2015)
Hug, C. B., Grimaldi, A. G., Kruse, K. & Vaquerizas, J. M. Chromatin architecture emerges during zygotic genome activation independent of transcription. Cell 169, 216–228 (2017)
Nagano, T. et al. Cell cycle dynamics of chromosomal organisation at single-cell resolution. Preprint available at http://biorxiv.org/content/early/2016/12/15/094466 (2016)
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017)
Brykczynska, U. et al. Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nat. Struct. Mol. Biol. 17, 679–687 (2010)
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012)
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171–181 (2014)
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015)
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012)
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protocols 7, 562–578 (2012)
Akdemir, K. C. & Chin, L. HiCPlotter integrates genomic data with interaction matrices. Genome Biol. 16, 198 (2015)
Zuin, J. et al. Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc. Natl Acad. Sci. USA 111, 996–1001 (2014)
Zhang, B. et al. Allelic reprogramming of the histone modification H3K4me3 in early mammalian development. Nature 537, 553–557 (2016)
Dahl, J. A. et al. Broad histone H3K4me3 domains in mouse oocytes modulate maternal-to-zygotic transition. Nature 537, 548–552 (2016)
Fan, X. et al. Single-cell RNA-seq transcriptome analysis of linear and circular RNAs in mouse preimplantation embryos. Genome Biol. 16, 148 (2015)
Acknowledgements
We thank B. Ren, C. Li, Y. Chen, F. Yue and L. An for discussions, and members of the Xie laboratory for comments. This work was supported by the National Key R&D Program of China (2016YFC0900301 to W.X.), the National Basic Research Program of China (2015CB856201 to W.X.; 2012CB316503 to M.Q.Z.), the National Natural Science Foundation of China (31422031 to W.X.; 61472205 to J.Z.; 31371341 to X.W.; 31361163004 and 31671383 to J.G.; 91519326, 31671384, 31361163004 and 91329000 to M.Q.Z.), the THU-PKU Center for Life Sciences (W.X.), the Youth Thousand Scholar Program of China (W.X., J.Z.), State Key Research Development Program of China (2016YFC1200303 to J.G.), TNLIST Cross-discipline Foundation (M.Q.Z.), the NIH (MH102616 to M.Q.Z.) and Beijing Advanced Innovation Center for Structural Biology (W.X. and J.Z.).
Author information
Authors and Affiliations
Contributions
Z.D. and W.X. conceived the project. Z.D. developed and conducted the sisHi-C experiments. B.H., Z.D., Y.X., and J.M. performed the mouse embryo experiments. Z.D., H.Z., R.M. and J.W. performed bioinformatics analysis. Xi.Z., J.H. and Y.W. helped with the Hi-C analysis pipeline setup. Y.L. and Q.W. performed NGS sequencing. Xu.Z. and K.Z. helped with various experiments. M.Q.Z., J.G., J.R.D., X.W., J.Z. and W.X. supervised experiments or data analyses. Z.D., H.Z. and W.X. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Z.D. and W.X. are co-inventors on a filed patent application for sisHi-C.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Comparison of sisHi-C and in situ Hi-C3.
sisHi-C procedures are shown by the red boxes and arrows on the left, and in situ Hi-C procedures are represented by blue boxes and arrows on the right. The black boxes in the middle show procedures shared by sisHi-C and in situ Hi-C.
Extended Data Figure 2 Validation of sisHi-C data.
a, Hi-C interaction frequency heatmaps (100-kb bin) of chromosome 19 in mouse ES cells as determined by conventional Hi-C4 or sisHi-C using 500 cells (2 replicates). Zoomed-in views (40-kb bin) are also shown. b, Scatter plots showing the comparison of chromatin interaction frequencies between sisHi-C and conventional Hi-C data in mouse ES cells4, or between the two replicates of 500 mouse ES cell sisHi-C samples. Pearson correlation coefficients are also shown. c, Chromatin compartments represented by the first principal component (PC1) values from PCA analyses are shown for the sisHi-C (2 replicates) and conventional Hi-C data in mouse ES cells4. Positive PC1 values represent compartment A regions (yellow), and negative values represent compartment B regions (blue). The gene-dense regions are also shown. d, Bar charts showing the Pearson correlation between the replicates (n = 2–4) of Hi-C data for mouse gametes and early embryos using chromatin interaction frequency. The error bars denote the s.d. of correlation values among different pairs of replicates. e, Heatmaps showing normalized Hi-C interaction frequency (100-kb bin, chromosome 19) in mouse sperm from this study or a previous study5. Zoomed-in views (40-kb bin) are also shown. f, Scatter plots showing the comparison of chromatin interaction frequencies in mouse sperm from this study and a previous study5, with Pearson correlation coefficient shown.
Extended Data Figure 3 Higher-order chromatin structure of MII oocytes.
a, Percentages of inter-chromosomal read pairs among all valid read pairs for gametes and early embryos (replicates pooled, n = 2–4). The error bars denote the s.e.m. among different replicates. b, Hi-C interaction frequency heatmaps comparing the chromatin interaction frequency between human mitotic chromatin (HeLaS3 cells)9 (top right) and mouse MII oocytes (middle right) at 100-kb resolution. The interaction heatmaps for asynchronized human ES cells4 (top left) and mouse ES cells (middle left) are shown as controls. The chromatin contact probabilities relative to genomic distance (P(s) curves) are also shown for each cell type (bottom). P(s) ~ s−0.5 and ~ s−1 represent the predicted mitotic and fractal globule states, respectively. The estimated turning points of chromatin interactions on the P(s) curves for MII oocytes (solid blue) and mitotic HeLaS3 cells (dashed blue) are shown by arrowheads.
Extended Data Figure 4 Establishment of TADs during mouse early development.
a, Hi-C interaction heatmaps (40-kb bin) showing dynamics of local interactions and TADs in mouse preimplantation development (pooled data from 2–4 biological replicates). b, Differential interaction heatmaps showing the changes in interaction frequency between consecutive developmental stages. Regions with increased and decreased interactions from early to late stages (pooled data from 2–4 biological replicates) are shown in yellow and blue, respectively. Hi-C interaction heatmap and identified TADs in ICMs are also shown to indicate the positions of TADs. A pair of subdomains within a TAD that show increased interactions from the 8-cell to ICM stage are indicated by black arrows. c, Overlap of TAD boundaries identified in mouse ES cells (this study) and ICMs (pooled data from three biological replicates). d, Schematic shows the method to compute consolidation score for a TAD. Consolidation score is the ratio between the average intra-TAD interaction frequencies (red) and the average local non-TAD background interaction frequencies in the nearby regions (green). Short-distance interactions (<400 kb) (orange) are excluded. iTF, intra-TAD interaction frequencies; nTF, non-TAD background interaction frequencies. e, Boxplots showing consolidation scores at different developmental stages (pooled data from 2–4 biological replicates). The significance of differences for consolidation scores between two consecutive stages was evaluated by Wilcoxon rank-sum test (two-tailed, with Benjamini–Hochberg multiple testing correction) as shown.
Extended Data Figure 5 TAD boundaries emerge as early as the zygote stage.
a, Hi-C interaction heatmaps showing the chromatin interaction frequencies for TADs in PN5 zygotes (pooled data, n = 4), ICMs (n = 3) and mouse ES cells (n = 2). ICM TADs identified by insulation scores are also shown. Blue boxes show example ICM TAD boundaries. b, The average DI scores of early embryos and mouse ES cells at ICM TAD boundaries (± 500 kb) are shown. DI scores generated by a random valid pair data set are also shown as a control (see Methods).
Extended Data Figure 6 Allelic chromatin organization in mouse early development.
a, Boxplots showing the ratios of inter-chromosomal read pairs between maternal and paternal (MP) genomes and between the same parental genomes (MM + PP) (see Methods). The ratios were computed for each pair of different chromosomes (chromosomes 1–2, 1–3 and so on) separately. Boxplots show the variations across all such pairs of chromosomes. b, Percentages of distal intra-chromosomal read pairs (>2 Mb) assigned to the paternal or maternal genome. The significance of allelic imbalance was assessed by comparison with that of the cortex (control) using a two-tailed t-test; **P < 0.001; *P < 0.05; ns, not significant. c, Heatmaps showing the allelic chromatin interaction frequencies (300-kb bin, chromosome 15) of sperm (replicates pooled, n = 3), MII oocyte (n = 2), PN3 (n = 2) and PN5 (n = 2) zygotes. d, Heatmaps showing the normalized average interaction frequencies for all ICM TADs as well as their nearby regions for sperm, MII oocyte and each allele of PN3 zygotes, PN5 zygotes and ICMs.
Extended Data Figure 7 Establishment of chromatin compartments during mouse early development.
a, Pearson correlation coefficients of correlation matrices between each developmental stage (replicates pooled, n = 2–4) and ICM (n = 3) on each allele. b, Hierarchical clustering based on correlation matrix for the two parental alleles. c, UCSC genome browser view showing the enrichment of ATAC-seq and chromatin compartments (PC1 values) for two parental alleles. Gene density is also shown. d, Allelic expression levels of ZGA-only genes (FPKM <0.5 in oocyte and FPKM >1 at any preimplantation stage) that fall into either compartment A or B in early development. The significance of differences between compartments A and B for each allele of each stage was assessed using Wilcoxon rank-sum test (P < 0.02 in all cases, two-tailed, with Benjamini–Hochberg multiple testing correction). e, ATAC-seq enrichment for compartments A and B of two parental alleles in early development. The significance of differences between compartments A and B for each allele of each stage was assessed using two-tailed t-test (P < 5.4 × 10−45 in all cases, with Benjamini–Hochberg multiple testing correction).
Extended Data Figure 8 Chromatin organization establishment in early development can partially proceed in the absence of zygotic genome activation.
a, Gene expression levels (log2[FPKM+1]) show the effects of alpha-amanitin on genes activated during ZGA, genes expressed at all stages or genes repressed at all stages as defined previously28. P values calculated with two-tailed Wilcoxon rank-sum test are shown. Note that genes expressed at all stages presumably have both oocyte-inherited transcripts and zygotically transcribed transcripts. Their decreasing values from 20 h to 45 h in the presence of alpha-amanitin are consistent with continual maternal RNA degradation. Previous RNA-seq data sets29,30 were similarly analysed as controls. b, Heatmaps showing the chromatin interaction frequency (300-kb bin, chromosome 13) for embryos treated with or without alpha-amanitin (replicates pooled, n = 2) (see Methods). The PN3 zygotes were treated with alpha-amanitin and collected for Hi-C analyses at 20 h or 45 h. In both cases the embryos were arrested at the late 2-cell stage. The control groups developed to the late 2-cell and 8-cell stage, respectively. c, Heatmaps showing the normalized average interaction frequencies for all ICM TADs as well as their nearby regions (± 0.5 TAD length) for embryos treated with or without alpha-amanitin (replicates pooled, n = 2) (bottom). Differential interaction heatmap between embryos treated for 20 h and 45 h (top) is also shown. d, Boxplots showing the consolidation scores for embryos treated with alpha-amanitin for 20 h or 45 h for each of the two replicates. P values calculated with two-tailed Wilcoxon rank-sum test are shown.
Extended Data Figure 9 Hierarchical packaging of chromatin in early development.
a, Heatmaps showing the intra-chromosomal interaction probability relative to genomic distance between interacting regions (P(s)) across the whole genome. The P(s) ~ s−1 values are also shown as a reference. b, Heatmaps showing normalized P(s) distributions generated by normalizing P(s) values for each stage to the P(s) ~ s−1 values. c, Hi-C interaction heatmaps showing normalized Hi-C interaction frequency (100-kb bin, chromosome 5). Dashed boxes indicate various regions where chromatin compaction occurs.
Extended Data Figure 10 Hierarchical packaging of chromatin in early development.
a, The contact probability as a function of genomic distance (P(s) analysis) (averaged across all chromosomes) is shown for gametes and zygotes on each allele as previously described9. P(s) ~ s−0.5 and P(s) ~ s−1 are shown to represent the predicted mitotic and fractal globule states, respectively9. b–e, P(s) curves (averaged across all chromosomes) for every two consecutive developmental stages calculated for each allele. P(s) ~ s−1 is shown to represent the predicted fractal globule state9. Arrowheads indicate estimated turning points on the P(s) curves where chromatin interactions drop abruptly (interaction insulation boundary). f, P(s) analyses for individual chromosomes (1, 4, 14 and 19) at early 2-cell stage. P(s) ~ s−1 is shown to represent the predicted fractal globule state. The arrowhead indicates the interaction insulation boundary.
Supplementary information
Supplementary Table
This table contains details of the sequencing information of sisHi-C libraries.
Rights and permissions
About this article
Cite this article
Du, Z., Zheng, H., Huang, B. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547, 232–235 (2017). https://doi.org/10.1038/nature23263
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23263
- Springer Nature Limited
This article is cited by
-
Sex-specific DNA-replication in the early mammalian embryo
Nature Communications (2024)
-
3C methods in cancer research: recent advances and future prospects
Experimental & Molecular Medicine (2024)
-
Antagonism between H3K27me3 and genome–lamina association drives atypical spatial genome organization in the totipotent embryo
Nature Genetics (2024)
-
The impact of DNA methylation on CTCF-mediated 3D genome organization
Nature Structural & Molecular Biology (2024)
-
Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Nature Communications (2024)