Abstract
Mammalian sperm and oocytes have different epigenetic landscapes and are organized in different fashions. After fertilization, the initially distinct parental epigenomes become largely equalized with the exception of certain loci, including imprinting control regions. How parental chromatin becomes equalized and how imprinting control regions escape from this reprogramming is largely unknown. Here we profile parental allele-specific DNase I hypersensitive sites in mouse zygotes and morula embryos, and investigate the epigenetic mechanisms underlying these allelic sites. Integrated analyses of DNA methylome and tri-methylation at lysine 27 of histone H3 (H3K27me3) chromatin immunoprecipitation followed by sequencing identify 76 genes with paternal allele-specific DNase I hypersensitive sites that are devoid of DNA methylation but harbour maternal allele-specific H3K27me3. Interestingly, these genes are paternally expressed in preimplantation embryos, and ectopic removal of H3K27me3 induces maternal allele expression. H3K27me3-dependent imprinting is largely lost in the embryonic cell lineage, but at least five genes maintain their imprinted expression in the extra-embryonic cell lineage. The five genes include all paternally expressed autosomal imprinted genes previously demonstrated to be independent of oocyte DNA methylation. Thus, our study identifies maternal H3K27me3 as a DNA methylation-independent imprinting mechanism.
Similar content being viewed by others
Main
Sperm and oocytes are generated from primordial germ cells through distinct processes. Consequently, their genomes are packaged differently with distinct epigenetic landscapes1. After fertilization, paternal chromatin releases protamines and is repackaged with maternally stored histones that are devoid of most histone modifications, while maternal chromatin harbours various histone modifications inherited from oocytes2,3. The different processes of parental chromatin formation result in parental epigenetic asymmetry in zygotes1, which becomes largely equalized during subsequent development with the exception of certain genomic loci, including imprinting control regions (ICRs)4.
Transcriptional regulatory elements, such as promoters and enhancers, can be mapped by DNase I hyper-sensitivity assay5,6. By using a low-input DNase I-sequencing (liDNase-seq) technique, we recently mapped the transcriptional regulatory landscape of preimplantation embryos; single nucleotide polymorphism (SNP)-based analysis revealed that chromatin accessibility of the two parental alleles is, overall, comparable, except for imprinted gene promoters7. A similar conclusion was also reached using an assay for transposase-accessible chromatin with high-throughput sequencing (ATAC-seq)8. However, the mechanisms underlying parent-of-origin specific chromatin accessibility are unknown.
Allelic DNase I hypersensitive sites in zygotes
To comprehensively profile parental allele-specific DNase I hypersensitive sites (DHSs) in zygotes, we isolated paternal and maternal pronuclei from pronucleus stage 5 (PN5) zygotes and performed liDNase-seq (Fig. 1a and Extended Data Fig. 1a). Using stringent criteria (Extended Data Fig. 1b) and excluding data for sex chromosomes, we identified 3,462, 687, and 169 bi-allelic DHSs, paternal allele-specific DHSs (Ps-DHSs), and maternal allele-specific DHSs (Ms-DHSs), respectively (Fig. 1b, Extended Data Fig. 1c and Supplementary Table 1). The genomic location of allelic DHSs was heavily biased to non-promoter elements compared with bi-allelic DHSs enriched in promoters and CpG islands (Extended Data Fig. 1d, e). Similar to previous findings7, Ps-DHSs included ICRs of known imprinted genes (Extended Data Fig. 1f). Interestingly, both Ps- and Ms-DHSs also included promoters of genes not previously known to be imprinted (Extended Data Fig. 1g, h).
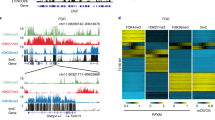
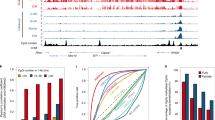
a, Schematic for identifying parental allele-specific DHSs in zygotes. IVF, in vitro fertilization. PN, pronucleus. b, Heat map showing bi-allelic, Ps-, and Ms-DHSs in zygotes. Each row represents liDNase-seq signal intensity at a DHS ± 5 kb. c, Representative androgenesis- and gynogenesis-specific DEGs harbouring allelic promoter DHSs in zygotes. Top, genome browser views of DHSs in paternal and maternal pronuclei with biological duplicates. The DHS signal intensity and the genomic length of each view (in kilobases) are indicated at the top left and the bottom of each panel, respectively. Bottom, gene expression levels in androgenetic (AG), gynogenetic (GG), and α-amanitin-treated (Ama) two-cell embryos. Error bar, s.d. of biological duplicates. Note that gynogenetic-specific expression is evident after subtraction of maternal pool transcripts.
Since promoter DHSs can prime gene expression at the next developmental stage7, we asked whether allelic DHSs in zygotes can prime allelic gene expression at zygotic genome activation (ZGA). RNA sequencing (RNA-seq) analysis of two-cell stage androgenetic and gynogenetic embryos, using α-amanitin treatment as a negative control, identified 107 androgenetic- and 14 gynogenetic-specific differentially expressed genes (DEGs), including 8 known imprinted genes (Extended Data Fig. 2a–d and Supplementary Table 2).
Integrated analysis of allelic ZGA and allelic promoter DHSs in zygotes revealed that most (59% and 79%) of the androgenetic- and gynogenetic-specific DEGs were associated with paternal and maternal allele-biased chromatin accessibility, respectively (Extended Data Fig. 2e, f). Genes showing such a correlation included not only known imprinted genes but also those not known to be imprinted (Fig. 1c). These results suggest that allelic DHSs in zygotes can mark promoters that are primed for allelic ZGA.
DNA methylation and allelic DHSs
To understand how allelic DHSs in zygotes are specified, we first examined whether they were inherited from gametes. We profiled DHSs of fully grown oocytes (Extended Data Fig. 3a) and analysed sperm DHSs7. Although sperm had only 34 reproducible DHSs (Supplementary Table 3), some of them contributed to Ps-DHSs (Extended Data Fig. 3b). However, most Ps-DHSs were absent in sperm and oocytes, indicating that they were generated after fertilization (Extended Data Fig. 3c, d). In contrast, most Ms-DHSs and bi-allelic DHSs were already present in oocytes (Extended Data Fig. 3e–h), indicating that most maternal DHSs are inherited from oocytes.
To determine how the maternal allele at Ps-DHSs remains inaccessible, we first hypothesized that maternal DNA methylation prevents DHS formation. Analysis of a public whole-genome bisulfite sequencing (WGBS) data set of oocytes and sperm9 revealed that only 17% of Ps-DHSs overlapped with oocyte germline differentially methylated regions (gDMRs) (Extended Data Fig. 4a and Supplementary Table 4). Despite extending to a ±100 kilobase (kb) region flanking Ps-DHSs, only an additional 21% were found to be associated with oocyte gDMRs (Extended Data Fig. 4a and Supplementary Table 4). Even when the oocyte DNA methylation level alone was considered, 48% of Ps-DHSs were devoid of oocyte DNA methylation (Extended Data Fig. 4b), indicating the existence of a DNA methylation-independent mechanism that prevents accessibility of the maternal allele.
Maternal allelic protection by H3K27me3
The fact that Polycomb-mediated H3K27me3 can mediate silencing of DNA hypomethylated promoters10 prompted us to postulate that H3K27me3 might be responsible for maternal allele inaccessibility. Analyses of public ChIP–seq (chromatin immunoprecipitation followed by sequencing) data sets11 revealed that the H3K27me3 level in oocytes was much higher than that of sperm at DNA hypomethylated Ps-DHSs, while it was reversed at DNA hypermethylated Ps-DHSs (Extended Data Fig. 4c, left). SNP tracking analysis revealed that the hypomethylated Ps-DHSs maintained maternal allele-specific H3K27me3 in zygotes (Extended Data Fig. 4c, right), indicating that H3K27me3 might be responsible for maternal allele inaccessibility at DNA hypomethylated regions.
To test this possibility, we injected messenger RNA (mRNA) encoding an H3K27me3-specific demethylase Kdm6b (Kdm6bWT) with its catalytic mutant (H1390A) (Kdm6bMUT) as a control12 (Fig. 2a). Similarly, we prepared zygotes injected with an H3K9me3-specific demethylase Kdm4d or its catalytic mutant (H189A)13. Both wild-type and mutant Kdm6b and Kdm4d were expressed at a similar level (Extended Data Fig. 4d), and Kdm6bWT and Kdm4dWT, but not their mutants, significantly reduced H3K27me3 and H3K9me3 levels, respectively (Extended Data Fig. 4e, f). liDNase-seq of isolated pronuclei (Extended Data Fig. 4g, h) revealed that 78 and 150 of the 431 most reliable Ps-DHSs became bi-allelic in Kdm6bWT- and Kdm4dWT-injected zygotes, respectively, while their catalytic mutants had little effect (Fig. 2b, c, Extended Data Fig. 4i and Supplementary Table 5). This result indicates that both maternal H3K27me3 and H3K9me3 are involved in maternal allele inaccessibility. Importantly, Kdm6b-affected Ps-DHSs were largely devoid of oocyte DNA methylation, which was markedly different from Kdm4d-affected Ps-DHSs that located at DNA hypermethylated regions (Fig. 2d). Consistently, Ps-DHSs specifically affected by Kdm6b, but not Kdm4d, overlapped maternal allele-specific H3K27me3 (Extended Data Fig. 4j). These results suggest that maternal H3K27me3 and H3K9me3 restrict maternal allele accessibility at regions with hypomethylated and hypermethylated DNA, respectively.
a, Schematic for studying the role of histone methylations in maternal chromatin inaccessibility. b, Heat map showing the allelic bias at Ps-DHSs in Kdm6b- or Kdm4d-injected zygotes. WT, wild-type. Mut, catalytic mutant. c, Genome browser view of representative Ps-DHSs affected by Kdm6bWT. Pat, paternal; Mat, maternal. d, Kdm6b- or Kdm4d-affected Ps-DHSs organized on the basis of their oocyte DNA methylation levels.
H3K27me3-dependent imprinting
To understand to what extent allelic DHSs exist at a later embryonic stage, we generated androgenetic and gynogenetic morula embryos (Fig. 3a) and performed liDNase-seq (Extended Data Fig. 5a). Using the same criteria for allelic DHSs as in zygotes and excluding data for sex chromosomes, we identified 36,569 common DHSs, 247 androgenetic-specific DHSs (AG-DHSs), and 176 gynogenetic-specific DHSs (GG-DHSs) (Fig. 3b and Supplementary Table 6). By SNP tracking analyses of a public DHS profile of hybrid morula embryos7, we confirmed that AG-DHSs, but not GG-DHSs, recapitulated the corresponding parental allele-specific DHSs (Extended Data Fig. 5b), indicating that AG-DHSs are physiological. Interestingly, AG-DHSs included almost all known maternally methylated ICRs (Extended Data Fig. 5c). This finding raised the possibility that AG-DHSs could serve as indicators of genomic imprinting.
a, Schematic for identifying parental allele-specific DHSs in morula embryos. b, Heat map showing AG-DHSs and GG-DHSs in morula embryos. Each row represents liDNase-seq signal intensity at a DHS ± 5 kb. c, Scatterplot showing allelic enrichment of H3K27me3 ChIP-seq signal at AG-DHSs ± 1 kb in ICM of blastocyst embryos. AG-DHSs with reads per million > 0.5, fold change (maternal/paternal) > 2, were considered to harbour maternal allele-biased H3K27me3 (red dots). d, Heat map showing parental allele-specific gene expression of putative H3K27me3-dependent imprinted genes. Genes expressed in androgenetic morula embryos (RPKM > 0.5) are shown. The left column represents the ratio of androgenetic/gynogenetic gene expression. The two right columns represent relative gene expression in hybrid morula embryos. B × C, B6/CAST; C × B, CAST/B6. The four known non-canonical imprinted genes are indicated in bold. White boxes indicate not determined (ND) owing to lack of SNP reads (<20 reads).
Because both DNA methylation and H3K27me3 can contribute to maternal allele inaccessibility (Fig. 2), we determined their respective contribution to the generation of the 247 AG-DHSs. Analyses of the oocyte DNA methylome9 identified 183 (74%) AG-DHSs in DNA hypomethylated regions (Extended Data Fig. 5d). Allelic H3K27me3 enrichment analysis revealed that 112 of the 183 were marked with maternal allele-biased H3K27me3 in inner cell mass (ICM) of blastocysts (Fig. 3c). Of the 112 AG-DHSs, 105 showed maternal allele-specific H3K27me3 enrichment in zygotes (reads per million > 0.5, fold change (maternal/paternal) > 4), suggesting that the maternal allele-biased H3K27me3 is inherited from zygotic maternal chromatin. By associating the 105 H3K27me3-marked AG-DHSs with their nearest genes, we obtained 76 genes (Supplementary Table 7) as putative H3K27me3-dependent imprinted genes.
To examine the allelic gene expression of the 76 genes, we performed RNA-seq analysis for androgenetic and gynogenetic morula embryos (Extended Data Fig. 6a and Supplementary Table 8). After confirming androgenetic- or gynogenetic-specific expression of known imprinted genes (Extended Data Fig. 6b), we calculated the relative androgenetic/gynogenetic expression levels for each candidate. Among the 76 genes, 28 were expressed in either androgenetic or gynogenetic embryos (fragments per kilobase of exon per million mapped fragments (FPKM) > 0.5). Interestingly, 27 of the 28 genes exhibited biased (fold change > 2), and 23 genes exhibited highly biased (fold change > 8), expression in androgenetic embryos (Fig. 3d, left column). Using an RNA-seq data set of hybrid morula embryos14, we confirmed that all 13 SNP-trackable genes exhibit paternal allele-specific expression (Fig. 3d, right columns). Importantly, the 28 genes include Sfmbt2, Gab1, Slc38a4, and Phf17 (also known as Jade1) (Fig. 3d), whose imprinted expression was suggested to be independent of oocyte DNA methylation15,16,17,18. These ‘non-canonical’ imprinted genes are coated with oocyte-specific H3K27me3 domains that are retained even in blastocysts (Extended Data Fig. 6c), which is in contrast to DNA methylation-dependent ‘canonical’ imprinted genes that are devoid of oocyte H3K27me3 (Extended Data Fig. 6d). Collectively, these results demonstrate that maternal H3K27me3 may serve as a DNA methylation-independent imprinting mark.
To determine whether maternal H3K27me3 is responsible for maternal allele repression of the putative H3K27me3-dependent imprinted genes, we injected Kdm6bWT or Kdm6bMUT mRNAs into one-cell-stage parthenogenetic embryos (Fig. 4a). After verifying that the injection did not affect embryo development to the morula stage (Extended Data Fig. 7a), we performed RNA-seq analysis (Extended Data Fig. 7b). Of the 28 putative imprinted genes expressed in androgenetic morula embryos (Fig. 3d), 16 were significantly de-repressed in a catalytic activity-dependent manner, including all four known non-canonical imprinted genes (Fig. 4b and Supplementary Table 9). In contrast, canonical imprinted genes were not affected by Kdm6bWT injection (Extended Data Fig. 7c), demonstrating that H3K27me3 is specifically required for maternal allele repression of the putative H3K27me3-dependent imprinted genes.
a, Schematic for studying the role of H3K27me3 in maternal allele repression. Kdm6bMUT-injected parthenogenetic (PG) embryos were used as a negative control. b, Relative gene expression levels (log scale) of putative H3K27me3-dependent imprinted genes. Shown are genes expressed in androgenetic morula embryos (RPKM > 0.5) and significantly de-repressed by Kdm6bWT. The expression level of gynogenetic (GG) morula embryos was set as 1. The genes are ordered by statistical significance (P values by DEseq) between Kdm6bWT and Kdm6bMUT samples. Arrows indicate known non-canonical imprinted genes. c, Heat map showing parental allele-specific gene expression of putative H3K27me3-dependent imprinted genes in Kdm6bWT- and Kdm6bMUT-injected hybrid morula embryos. Among the 28 genes listed in Fig. 3d, those with >10 SNP reads in both samples are shown. Known non-canonical imprinted genes are indicated in bold. Allelic expression levels of representative canonical imprinted genes are shown at the bottom. d, Heat map showing the levels of chromatin accessibility at AG-DHSs in Kdm6bWT- and Kdm6bMUT-injected morula parthenogenetic embryos. The DHS signal intensity in androgenetic embryos was set as 100%. AG-DHSs are ordered by Δ(Kdm6bWT − Kdm6bMUT). Known imprinted genes are indicated at right, with non-canonical imprinted genes shown in red. e, Genome browser view of gain-of-accessibility at AG-DHSs of putative H3K27me3-dependent imprinted genes.
To demonstrate that Kdm6b-mediated maternal allele de-repression occurs in a physiological context, we performed RNA-seq analysis in hybrid morula embryos that had been injected with Kdm6bWT or Kdm6bMUT mRNA at the one-cell stage. Of the 28 putative imprinted genes, 17 had sufficient SNP reads, and 16 of them showed paternal allele-biased expression in Kdm6bMUT-injected embryos (Fig. 4c and Supplementary Table 10). Notably, the extent of the paternal allelic bias of all these genes became milder in Kdm6bWT-injected embryos, whereas that of canonical imprinted genes was not affected (Fig. 4c). These data strongly suggest that paternal allele-specific expression of these genes depends on maternal H3K27me3.
To determine whether maternal allele de-repression couples with gain of maternal chromatin accessibility, we performed liDNase-seq for Kdm6bWT- and Kdm6bMUT-injected parthenogenetic morula embryos (Extended Data Fig. 7d). We found that Kdm6bWT, but not Kdm6bMUT, markedly increased chromatin accessibility in AG-DHSs of putative H3K27me3-dependent imprinted genes, including all four non-canonical imprinted genes (Fig. 4d, e, Extended Data Fig. 7e and Supplementary Table 11). In contrast, ICRs of canonical imprinted genes were not affected (Fig. 4d and Extended Data Fig. 7f, g). These results suggest that maternal H3K27me3 restricts maternal allele accessibility to mediate H3K27me3-dependent genomic imprinting.
Imprinting status in blastocysts
We next analysed the allelic expression of putative H3K27me3-dependent imprinted genes in blastocyst embryos by SNP tracking of recently published data sets14. Of the 28 putative imprinted genes (Fig. 3d), 15 had sufficient SNP reads in both reciprocal crosses (Fig. 5a). Among them, 12 (80%) showed paternal allelic expression in both crosses (Fig. 5a), demonstrating that H3K27me3-dependent imprinting is largely maintained in blastocysts.
a, Heat map showing parental allele-specific gene expression of putative H3K27me3-dependent imprinted genes in hybrid blastocyst embryos. B × C, B6/CAST. C × B, CAST/B6. Known non-canonical imprinted genes are indicated in bold in a–d. The colour scheme in a–d follows Fig. 3d. b, Heat map showing androgenesis/gynogenesis relative expression of putative H3K27me3-dependent imprinted genes in ICM and trophectoderm (TE) of blastocyst embryos. Arrows indicate genes showing a milder level of androgenetic-bias in ICM compared with trophectoderm. White boxes indicate not determined owing to low gene expression levels (RPKM < 0.5). c, Heat map showing parental allele-specific gene expression of putative H3K27me3-dependent imprinted genes in epiblast (EPI), visceral endoderm (VE), and extra-embryonic ectoderm (EXE) of E6.5 embryos. Genes with >20 SNP reads in both reciprocal crosses are shown. B × P, B6/PWK; P × B, PWK/B6. Arrowheads indicate genes showing imprinted expression. d, Heat map showing parental allele-specific gene expression of putative H3K27me3-dependent imprinted genes in pure fetus-derived E9.5 placenta cells. Genes with >20 SNP reads in both reciprocal crosses are shown. Arrowheads indicate genes showing imprinted expression. e, Model illustrating the fate of H3K27me3-dependent genomic imprinting during development.
Since previous studies have indicated that Gab1, Sfmbt2, and Phf17 are imprinted only in extra-embryonic tissues19,20, we examined their imprinting status in trophectoderm and ICM, respectively, by RNA-seq analysis for androgenetic and gynogenetic blastocysts (Supplementary Table 12). Marker gene expression confirmed no cross-contamination (Extended Data Fig. 8a). Of the 28 putative imprinted genes (Fig. 3d), 23 and 24 were expressed in trophectoderm and ICM, respectively (reads per kilobase per million (RPKM) > 0.5). Of these, 18 (78%) in trophectoderm and 16 (67%) in ICM showed androgenetic-biased expression (fold change > 2) (Fig. 5b). Notably, nine genes showed weaker androgenetic-bias in ICM compared with trophectoderm (Fig. 5b, arrows), suggesting that H3K27me3-dependent imprinting might start to diminish in ICM.
Post-implantation imprinting dynamics
To determine the imprinting status in post-implantation embryos, we dissected hybrid embryonic day (E)6.5 embryos into epiblast, visceral endoderm, and extra-embryonic ectoderm, and performed RNA-seq analysis (Extended Data Fig. 8b and Supplementary Table 13). We confirmed their cell identify by analysing cell-lineage-specific marker gene expression21 (Extended Data Fig. 8c) and identified 17 paternally expressed genes (PEGs) and 8 maternally expressed genes (MEGs) in epiblast, and 19 PEGs and 12 MEGs in both visceral endoderm and extra-embryonic ectoderm, including new imprinted genes such as D7Ertd715e (also known as Snhg14), Smoc1, and Mas1 (Extended Data Fig. 8d, e and Supplementary Table 13).
Among the 76 putative H3K27me3-dependent imprinted genes (Supplementary Table 7), 25, 23, and 18 genes had enough SNP reads in both reciprocal crosses in epiblast, visceral endoderm, and extra-embryonic ectoderm, respectively (Fig. 5c). We found that one, three, and five genes were paternally expressed in epiblast, visceral endoderm, and extra-embryonic ectoderm, respectively (Fig. 5c, arrowheads). The genes imprinted in extra-embryonic ectoderm included the four non-canonical imprinted genes, Gab1, Phf17, Sfmbt2, and Slc38a4, and a new imprinted gene, Smoc1 (Fig. 5c). These results suggest that H3K27me3-dependent imprinting is completely erased in the epiblast with the exception of Slc38a4, but that some are maintained in the extra-embryonic cell lineages.
To analyse the imprinting status in E9.5 placentae avoiding possible maternal cell contamination, we purified fetus-derived placental cells from green fluorescent protein (GFP) transgenic embryos by fluorescence-activated cell sorting (FACS) (Extended Data Fig. 9a) and performed RNA-seq analysis (Extended Data Fig. 9b and Supplementary Table 14). After confirming cell purity by demonstrating comparable total SNP reads from parental alleles (Extended Data Fig. 9c), we identified 25 PEGs and 21 MEGs, including new imprinted genes such as D7Ertd715e, Smoc1, Cbx7, and Thbs2 (Extended Data Fig. 10a, b and Supplementary Table 14). Among the 76 putative H3K27me3-dependent imprinted genes, 27 genes had sufficient SNP reads in both reciprocal crosses (Fig. 5d). Among them, Gab1, Sfmbt2, Slc38a4, and Smoc1 were paternally expressed (Fig. 5d). The allelic bias of Phf17 in one cross was weak (fold change = 1.87) (Fig. 5d and Supplementary Table 14), which was consistent with a previous study23. Taken together, our data not only identify Smoc1 as a new H3K27me3-dependent imprinted gene, but also suggest that H3K27me3-dependent imprinting is mostly transient in preimplantation embryos, while some genes remain imprinted in the extra-embryonic cell lineage (Fig. 5e).
Discussion
Since the identification of DNA methylation as a master regulator of genomic imprinting more than 20 years ago22,23,24, it has been the only known mammalian germline imprinting mark4. However, recent studies have identified several imprinted genes capable of maintaining paternal allele-specific expression in the absence of oocyte DNA methylation15,16, suggesting the existence of a DNA methylation-independent imprinting mechanism. Here we have revealed that these non-canonical imprinted genes harbour high levels of oocyte-specific H3K27me3, and that loss of H3K27me3 results in loss of imprinting. Although previous studies have revealed a link between a repressed allele and repressive histone modifications, including H3K27me3, at certain imprinted loci25,26,27,28,29,30,31,32,33,34, the imprinting status of these loci originally depends on neighbouring gDMRs17,26,34,35. Consistently, ectopic removal of H3K27me3 specifically affected non-canonical imprinted genes, indicating that the regulatory mechanism of H3K27me3-dependent imprinting is fundamentally different from that of gDMR-mediated canonical imprinting.
The dynamics of H3K27me3-dependent imprinting is strikingly different from DNA methylation-dependent imprinting that is largely maintained in both embryonic and extra-embryonic lineages36. The H3K27me3 imprinting is probably established during oogenesis and maintained in preimplantation embryos (Fig. 5e). While it begins to dilute in ICM and is almost completely lost in the epiblast of E6.5 embryos, it is maintained in some genes at least until E9.5 placentae. Further investigation is warranted to understand why and how these genes are selected to maintain imprinting and how cell-lineage-specific imprinting is achieved. Furthermore, what other organisms may conserve H3K27me3-dependent genomic imprinting is a fascinating question given that flowering plants also adopt this mechanism37,38.
Methods
Isolation of maternal and paternal pronuclei from PN5 stage zygotes
All animal studies were performed in accordance with the guidelines of the Institutional Animal Care and Use Committee at Harvard Medical School. MII-stage oocytes were collected from 8-week-old B6D2F1/J (BDF1) female mice superovulated by injecting 7.5 IU of pregnant mare serum gonadotropin (PMSG; Millipore) and hCG (Millipore). For in vitro fertilization, MII oocytes were inseminated with activated spermatozoa obtained from the caudal epididymis of adult BDF1 male mice in HTF medium supplemented with 10 mg ml−1 bovine serum albumin (BSA; Sigma-Aldrich). Spermatozoa capacitation was attained by 1 h incubation in the HTF medium. Zygotes were cultured in a humidified atmosphere with 5% CO2/95% air at 37.8 °C. At 10 h post-fertilization (h.p.f.), zygotes were transferred into M2 media containing 10 μg ml−1 cytochalasin B (Sigma-Aldrich). Zona pellucidae were cut by a Piezo impact-driven micromanipulator (Prime Tech, Ibaraki, Japan) and the pronuclei were isolated from the zygotes. At 12 h.p.f. (PN5 stage), isolated pronuclei were washed with 0.2% BSA/PBS, transferred into Eppendorf LoBind 1.5 ml tubes, and placed on ice until DNase I treatment. For each experiment, 150–200 pronuclei were collected. The parental pronuclei were distinguished by (1) the distance from the second polar body and (2) the size of the pronucleus.
Preparation of androgenetic and gynogenetic embryos
MII oocytes were collected from 8-week-old superovulated BDF1 females and inseminated with BDF1 sperm. At 7 h.p.f., zygotes were transferred into M2 media containing 5 μg ml−1 cytochalasin B, and parental pronuclei were exchanged by using a Piezo impact-driven micromanipulator. The sendai virus (HVJ, Cosmo-bio) was used for fusing karyoplasts with cytoplasms as previously described39. After reconstruction, embryos were cultured in KSOM.
When collecting embryos for RNA-seq or/and liDNase-seq, we removed zona pellucidae by a brief exposure to Acidic Tyrode’s Solution (Sigma-Aldrich), then the embryos were washed with M2 media, and then 0.2% BSA/PBS. For liDNase-seq, ten morula embryos were transferred into an Eppendorf LoBind 1.5 ml tube, and placed on ice until DNase I treatment. For RNA-seq, seven to ten embryos were transferred into thin-walled RNase-free PCR tubes (Ambion). The two-cell and morula embryos were collected at 30 and 78 h.p.f., respectively. When preparing α-amanitin-treated two-cell embryos, 5 h.p.f. zygotes were transferred into KSOM containing 25 μg ml−1 α-amanitin (Sigma-Aldrich) and cultured in the presence of α-amanitin until collection (30 h.p.f.).
ICM and trophectoderm were isolated as described previously40 with slight modifications. Briefly, androgenetic and gynogenetic embryos at 120 h.p.f. were treated with Acidic Tyrode’s Solution to remove zona pellucidae. After being washed in M2 media, the embryos were incubated in KSOM containing rabbit anti-mouse lymphocyte serum (Cedarlane, 1:8 dilution) for 45 min at 37 °C. After being washed in M2 media, they were transferred into KSOM containing guinea-pig complement (MP Biomedicals, 1:3.3 dilution). After incubation for 30 min at 37 °C, lysed trophectoderm cells were removed by pipetting with a glass capillary. The remaining ICM clumps were incubated in 0.25% trypsin/EDTA (Thermo Fisher, 25200) for 10 min at 37 °C, and then dissociated into single cells to avoid contamination of lysed trophectoderm cells. One hundred to 200 cells were collected for RNA-seq.
Isolation of germinal vesicle nuclei from fully grown oocytes
Fully grown germinal-vesicle-stage oocytes were obtained from 3-week-old BDF1 mice 44–48 h after injection with 5 IU PMSG. The ovaries were transferred to M2 media. The ovarian follicles were punctured with a 30-gauge needle, and the cumulus cells were gently removed from the cumulus–oocyte complexes using a narrow-bore glass pipette. The oocytes were then transferred into α-MEM (Life Technologies, 12571-063) supplemented with 5% fetal bovine serum (FBS) (Sigma-Aldrich, F0926), 10 ng ml−1 epidermal growth factor (Sigma-Aldrich, E4127), and 0.2 mM 3-isobutyl-1-methylxanthine (IBMX; Sigma-Aldrich). One hour after collection, germinal vesicle oocytes exhibiting visible perivitelline spaces, which had the surrounding-nucleolus-type chromatin, were culled41. They were then incubated in M2 media containing 10 μg ml−1 cytochalasin B, 0.1 μg ml−1 colcemid (Sigma-Aldrich), and 0.2 mM IBMX for 15 min. Then, germinal vesicle nuclei were isolated by using a Piezo-driven micromanipulator. After washing with 0.2% BSA/PBS, the germinal vesicle nuclei were transferred into an Eppendorf LoBind 1.5 ml tube. For each experiment, 115–150 germinal vesicle nuclei were collected.
Dissection of E6.5 embryos and FACS sorting of GFP-positive E9.5 placental cells
To obtain E6.5 C57BL6(B6)/PWK hybrid embryos, we used a natural mating scheme. To obtain E6.5 PWK/B6 hybrid embryos, we used in vitro fertilization of PWK oocytes with B6 sperm, and the two-cell embryos were transferred into surrogate ICR strain mothers. Dissection of E6.5 embryos into epiblast, visceral endoderm, and extra-embryonic ectoderm was performed as described previously42. To collect E9.5 placental cells, we purchased the B6GFP mice from The Jackson Laboratory (C57BL/6-Tg(CAG-EGFP)131Osb/LeySopJ, stock number 006567). MII oocytes and sperms were collected from superovulated 8-week-old B6GFP or PWK mice. After in vitro fertilization, the two-cell embryos were transferred into surrogate ICR strain mothers. At E9.5, placentae were harvested, cut into ~0.5 mm pieces, transferred into 50 ml tubes, and treated with 2 ml of 0.25% Trypsin-EDTA (Thermo Fisher Scientific, 25200) at 30 °C for 15 min in a shaker at 200 r.p.m. to dissociate placental cells. Trypsin treatment was stopped by the addition of 2 ml DMEM containing 10% FBS. After pipetting, the tubes were centrifuged and the pelleted cells were washed with 0.2% BSA/PBS three times. DAPI (4′,6-diamidino-2-phenylindole) was added at the final concentration of 1 μM in the final cell suspension. The GFP-positive cells were sorted using a BD FACSaria machine (BD Biosciences) with DAPI-positive cells excluded as dead cells. Approximately 10,000–20,000 GFP-positive cells were collected from each placenta, which corresponded to 40–60% of total placental cells.
Plasmid construction and mRNA preparation
To generate the Kdm6bWT construct, the complementary DNA (cDNA) encoding the carboxyl-terminal part containing the catalytic domain (amino acids 1025 to end) was amplified12. The PCR amplicon was cloned between a Flag tag and poly(A) of the pcDNA3.1–Flag–poly(A)83 plasmid2. The H1390A Kdm6bMUT construct43 were generated by using PrimeSTAR mutagenesis (TAKARA). Primers used for the mutagenesis were 5′-CCAGGCgctCAAGAGAATAACAATTTCTGCTCAGTCAACATCAAC-3′ and 5′-CTCTTGagcGCCTGGCGTTCGGCTGCCAGGGACCTTCATG-3′. All constructs were verified by DNA sequencing. The plasmids for wild-type and H189A mutant Kdm4d were previously described13.
After linearization by a restriction enzyme, the construct was purified with phenol–chloroform extraction. mRNA was synthesized by in vitro transcription using an mMESSAGE mMACHINE T7 Ultra Kit (Life Technologies) according to the manufacturer’s instructions. The synthesized mRNA was purified by lithium chloride precipitation and diluted with nuclease-free water. mRNA aliquots were stored at −80 °C until use.
mRNA injection
MII oocytes were collected from superovulated 8-week-old BDF1 females and inseminated with BDF1 sperm. At 2.5 h.p.f., fertilized oocytes were transferred into M2 media and mRNA was injected using a Piezo impact-driven micromanipulator. mRNA injection was completed by 4 h.p.f. The mRNA concentrations of Kdm6bWT and Kdm6bMUT were 1.8 μg μl−1, and those of Kdm4dWT and Kdm4dMUT were 1.5 μg μl−1. When preparing Kdm6b-injected parthenogenetic embryos, MII oocytes were chemically activated by treating them with 3 mM SrCl2 in Ca2+-free KSOM containing 5 μg ml−1 cytochalasin B. At 4 h post-activation, the embryos were washed with KSOM. At 5 h post-activation, they were injected with mRNA.
Whole mount immunostaining
Zygotes were fixed in 3.7% paraformaldehyde in PBS containing 0.2% Triton X-100 for 20 min. After four washes with PBS containing 10 mg ml−1 BSA (PBS/BSA), zygotes were treated with primary antibodies at 4 °C overnight. The primary antibodies used in this study were mouse-anti-H3K27me3 (1/500, Active Motif, 61017), rabbit anti-H3K9me3 (1/500, Millipore, 07-442), and rabbit anti-Flag (1/2,000, Sigma-Aldrich, F7524). After three washes with PBS/BSA, samples were incubated with a 1:250 dilution of fluorescein-isothiocyanate-conjugated anti-mouse IgG (Jackson Immuno-Research) or Alexa Fluor 568 donkey anti-rabbit IgG (Life Technologies) for 1 h. The zygotes were then mounted on a glass slide in Vectashield anti-bleaching solution with DAPI (Vector Laboratories, Burlingame, California, USA). Fluorescence was detected under a laser-scanning confocal microscope with a spinning disc (CSU-10, Yokogawa) and an EM-CCD (electron multiplier charge-coupled device) camera (ImagEM, Hamamatsu) or Zeiss LSM800.
All images were acquired and analysed using Axiovision software (Carl Zeiss). The fluorescent signal intensity was quantified with the Axiovision software. Briefly, the signal intensity within the maternal pronuclei was determined, and the cytoplasmic signal was subtracted as background. Then, the averaged signal intensity of the no-injection control zygotes was set as 1.0.
Low-input DNase-seq
Low-input DNase-seq libraries were prepared as previously described with minor modifications7. Embryos or nuclei collected in 1.5 ml tubes were resuspended in 36 μl lysis buffer (10 mM Tris-HCl, pH 7.5, 10 mM NaCl, 3 mM MgCl2, 0.1% Triton X-100) and incubated on ice for 5 min. DNase I (10,000 U ml−1, Roche, 04716728001) was added to the final concentration of 80 U ml−1 (for the germinal vesicle nucleus sample) or 40 U ml−1 (for all the other samples) and incubated at 37 °C for exactly 5 min. The reaction was stopped by adding 80 μl Stop Buffer (10 mM Tris-HCl, pH 7.5, 10 mM NaCl, 0.15% SDS, 10 mM EDTA) containing 2 μl Proteinase K (20 mg ml−1, Life Technologies). Then 20 ng of a circular carrier DNA (a pure plasmid DNA without any mammalian genes purified with 0.5× Beckman SPRIselect beads (Beckman Coulter) to remove small DNA fragments) was added. The mixture was incubated at 50 °C for 1 h, then DNA was purified by extraction with phenol–chloroform and precipitated by ethanol in the presence of linear acrylamide (Life Technologies) overnight at −20 °C. Precipitated DNA was resuspended in 50 μl elution buffer (2.5 mM Tris, pH 7.6, 0.05 mM EDTA), and the entire volume was used for sequencing library construction.
Sequencing libraries were prepared using a NEBNext Ultra II DNA Library Prep Kit for Illumina (New England Biolabs) according to the manufacturer’s instructions, with the exception that the adaptor ligation was performed with 0.03 μM adaptor in the ligation reaction for 30 min at 20 °C and that PCR amplification was performed using Kapa Hifi HotStart ReadyMix (Kapa Biosystems) for eight cycles. The PCR products were purified with ×1.3 volume of SPRIselect beads (Beckman Coulter) and then size-selected with ×0.65 volume followed by ×0.7 volume of SPRIselect beads. The sample was eluted in 24 μl elution buffer. The number of cycles needed for the second PCR amplification was determined by quantitative PCR using 1 μl of the 1:1,000 diluted samples. The remaining 23 μl of the samples was then amplified with Kapa Hifi HotStart ReadyMix (we used seven cycles for all samples in this study). The PCR product was purified with ×1.3 volume of SPRIselect beads and then size-selected with ×0.65 volume followed by ×0.7 volume of SPRIselect beads. The DNA was eluted in 30 μl of elution buffer and quantified by a Qubit dsDNA HS assay kit (Thermo Fisher Scientific, Q32854) and Agilent High Sensitivity assay kit (Agilent Technologies). The libraries were sequenced on a Hiseq2500 with single-end 100 bp reads (Illumina).
RNA-seq
RNA-seq libraries were prepared as previously described13. Briefly, reverse transcription and cDNA amplification were performed using whole-embryo lysates with a SMARTer Ultra Low Input RNA cDNA preparation kit (Clontech, 634890). When processing two-cell androgenetic, gynogenetic, and α-amanitin-treated in vitro fertilization embryo samples, 1 μl of 1:40,000 diluted ERCC (External RNA Controls Consortium) standard RNA (Life Technologies) was added to each of the tubes at the step of cell lysis. cDNAs were then fragmented using a Covaris M220 sonicator (Covaris) with microTUBE-50 (Covaris) into average of 150–160 bp fragments. The fragmented cDNAs were end-repaired, adaptor-ligated and amplified using a NEBNext Ultra DNA Library Prep Kit for Illumina according to the manufacturer’s instructions (New England Biolabs). Single-end 100 bp sequencing was performed on a HiSeq2500 sequencer (Illumina).
liDNase-seq data analysis
Reads of liDNase-seq data were first trimmed of low quality and adaptor with trim_galore, and then mapped to the mouse genome (mm9) using Bowtie version 0.12.9. The ‘-m 1’ parameter was used to keep unique mapping hits. The reads with mapping quality (MAPQ) ≤ 10 or redundant reads that mapped to the same location with the same orientation were removed with SAMtools44. The DHS peaks in the liDNase-seq data were identified by the Hotspot program with FDR < 0.01 (ref. 45). The DHS peaks from all 33 libraries were merged using ‘bedtools merge’ from bedtools46. The number of reads in each DHS for each library was calculated using ‘multiBamSummary’ from deepTools47 and normalized to the total number of mapped reads and to the length of DHS (the possibility of a tag located on a position per 1 kb per million mapped reads). Reads of sex chromosomes were removed because the number of sex chromosomes is different between the parental pronuclei and between androgenetic and gynogenetic embryos. The Pearson correlation coefficient (r) of tag densities at genome-wide DHSs was calculated to measure the correlation between replicates. For identification of parental allele-specific DHSs in zygotes and morula embryos, we used a stringent cutoff (RPKM mean > 2, RPKM > 1 in all replicates in a biased allele, and mean value of fold change larger than 4 between the two alleles). The 431 most reliable Ps-DHSs were identified by applying an additional criterion, ‘RPKM > 1 in all replicates of paternal PNs of microinjected zygotes’ to the 687 Ps-DHSs. The RefSeq gene assembly (mm9) from the University of California, Santa Cruz (UCSC) Genome Browser database and CpG islands previously defined9 were used as genomic feature distribution analysis in Extended Data Fig. 1d, e.
RNA-seq data analysis
We constructed a custom reference sequence combining mouse genome (mm9) with the ERCC control. Reads of RNA-seq were mapped to the reference genome with TopHat version 2.0.6 (ref. 48) or STAR (https://github.com/alexdobin/STAR). All programs were run with default parameters unless otherwise specified. Uniquely mapped reads were subsequently assembled into transcripts guided by the reference annotation (UCSC gene models) with featureCounts from subread version 1.5.1 (ref. 49). For all two-cell RNA-seq libraries, library size factors were estimated with the ‘estimateSizeFactors’ function from R package DESeq50 only using ERCC read counts. After the library size was normalized, the expression level of each gene was quantified with normalized FPKM. The Pearson correlation coefficient (r) of gene expression level was calculated to indicate the correlation between duplicates. For identification of newly synthesized transcripts at the two-cell stage, we first filtered out statistically non-significant genes between androgenetic or gynogenetic and α-amanitin-treated two-cell embryos. To do this, the adjusted P value was calculated with the ‘nbinomTest’ function from the R package DESeq using a negative binomial model, and only genes with FDR < 0.05 were selected. We then applied additional cutoffs (mean FPKM (androgenetic or gynogenetic) > 2 and fold change (androgenetic/α-amanitin-treated or gynogenetic/α-amanitin-treated) > 2). As a result, 4,381 and 3,916 genes were identified as newly synthesized in androgenetic and gynogenetic two-cell embryos, respectively. For identifying androgenetic- and gynogenetic-specific DEGs in two-cell embryos, the gene expression level (FPKM) of each gene in α-amanitin-treated two-cell embryos was subtracted from that of androgenetic and gynogenetic embryos. Genes showing fold change (androgenetic/gynogenetic or gynogenetic/androgenetic) > 10 were identified as DEGs.
WGBS and H3K27me3 ChIP-seq data analysis
The DNA methylation level at DHSs was calculated using methpipe version 3.4.2 (ref. 51). When calculating the DNA methylation level at each DHS, we extended each DHS 2 kb both upstream and downstream to include enough CpG sites. The oocyte-methylated gDMR was defined by >80% methylation in oocytes and <20% in sperm9. For Extended Data Fig. 4a, ‘bedtools makewindows’ was used to generate a set of non-overlapped 1 kb bins for the ±100 kb flanking region of Ps-DHSs. For H3K27me3 ChIP-seq analysis, Bed files were downloaded from ref. 11 and converted to the bigWig format using ‘bedClip’ and ‘bedGraphToBigWig’ from the UCSC Genome Browser database. ‘multiBigwigSummary’ from deepTools was used to compute the H3K27me3 signal over the DHS and surrounding region.
Statistical analyses and data visualization
Statistical analyses were implemented with R (http://www.r-project.org/). Pearson’s r coefficient was calculated using the ‘cor’ function with default parameters. Figures 2b and 4d were generated with the R function ‘heatmap.2’. Figures 3d, 4c and 5a–d were generated with the R function ‘pheatmap’. Figures 1b and 3b were generated using the ‘computeMatrix’ and ‘plotHeatmap’ functions in deepTools47. Position-wise coverage of the genome by sequencing reads was determined by normalizing to the total unique mapped reads in the library using macs2 version 2.1.0 (ref. 52) and visualized as custom tracks in the Integrative Genomics Viewer genome browser.
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Known imprinting gene information
Known imprinting information was downloaded from http://www.geneimprint.com/site/genes-by-species.Mus+musculus.
Code availability
A customized pipeline was used to split the hybrid RNA-seq data to their parental origin on the basis of SNP information. The code can be found at https://github.com/lanjiangboston/UniversalSNPsplit.
Data availability
All the liDNase-seq and RNA-seq data sets generated in this study are summarized in Supplementary Table 15 and have been deposited in the Gene Expression Omnibus under accession number GSE92605. Sperm liDNase-seq data sets were from a previous publication (GSE76642)7. WGBS data sets for sperm and germinal vesicle oocytes were downloaded from http://www.nodai-genome.org/mouse.html?lang=en (ref. 9). H3K27me3 ChIP-seq data sets of sperm, MII oocytes, and SNP-tracked maternal and paternal alleles of one-cell embryos and ICM were downloaded from a previous publication (GSE76687)11.
Accession codes
Change history
26 July 2017
The label for the colour scale in Fig. 3d was corrected.
References
Burton, A. & Torres-Padilla, M. E. Chromatin dynamics in the regulation of cell fate allocation during early embryogenesis. Nat. Rev. Mol. Cell Biol. 15, 723–735 (2014)
Inoue, A. & Zhang, Y. Nucleosome assembly is required for nuclear pore complex assembly in mouse zygotes. Nat. Struct. Mol. Biol. 21, 609–616 (2014)
Zhou, L. Q. & Dean, J. Reprogramming the genome to totipotency in mouse embryos. Trends Cell Biol. 25, 82–91 (2015)
Ferguson-Smith, A. C. Genomic imprinting: the emergence of an epigenetic paradigm. Nat. Rev. Genet. 12, 565–575 (2011)
Boyle, A. P. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008)
Stergachis, A. B. et al. Developmental fate and cellular maturity encoded in human regulatory DNA landscapes. Cell 154, 888–903 (2013)
Lu, F. et al. Establishing chromatin regulatory landscape during mouse preimplantation development. Cell 165, 1375–1388 (2016)
Wu, J. et al. The landscape of accessible chromatin in mammalian preimplantation embryos. Nature 534, 652–657 (2016)
Kobayashi, H. et al. Contribution of intragenic DNA methylation in mouse gametic DNA methylomes to establish oocyte-specific heritable marks. PLoS Genet. 8, e1002440 (2012)
Deaton, A. M. & Bird, A. CpG islands and the regulation of transcription. Genes Dev. 25, 1010–1022 (2011)
Zheng, H. et al. Resetting epigenetic memory by reprogramming of histone modifications in mammals. Mol. Cell 63, 1066–1079 (2016)
Agger, K. et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449, 731–734 (2007)
Matoba, S. et al. Embryonic development following somatic cell nuclear transfer impeded by persisting histone methylation. Cell 159, 884–895 (2014)
Borensztein, M. et al. Xist-dependent imprinted X inactivation and the early developmental consequences of its failure. Nat. Struct. Mol. Biol. 24, 226–233 (2017)
Okae, H. et al. Re-investigation and RNA sequencing-based identification of genes with placenta-specific imprinted expression. Hum. Mol. Genet. 21, 548–558 (2012)
Okae, H. et al. RNA sequencing-based identification of aberrant imprinting in cloned mice. Hum. Mol. Genet. 23, 992–1001 (2014)
Varmuza, S. & Miri, K. What does genetics tell us about imprinting and the placenta connection? Cell. Mol. Life Sci. 72, 51–72 (2015)
Wang, X., Soloway, P. D. & Clark, A. G. A survey for novel imprinted genes in the mouse placenta by mRNA-seq. Genetics 189, 109–122 (2011)
Babak, T. et al. Genetic conflict reflected in tissue-specific maps of genomic imprinting in human and mouse. Nat. Genet. 47, 544–549 (2015)
Kuzmin, A. et al. The PcG gene Sfmbt2 is paternally expressed in extraembryonic tissues. Gene Expr. Patterns 8, 107–116 (2008)
Blakeley, P. et al. Defining the three cell lineages of the human blastocyst by single-cell RNA-seq. Development 142, 3151–3165 (2015)
Bartolomei, M. S., Webber, A. L., Brunkow, M. E. & Tilghman, S. M. Epigenetic mechanisms underlying the imprinting of the mouse H19 gene. Genes Dev. 7, 1663–1673 (1993)
Ferguson-Smith, A. C., Sasaki, H., Cattanach, B. M. & Surani, M. A. Parental-origin-specific epigenetic modification of the mouse H19 gene. Nature 362, 751–755 (1993)
Stöger, R. et al. Maternal-specific methylation of the imprinted mouse Igf2r locus identifies the expressed locus as carrying the imprinting signal. Cell 73, 61–71 (1993)
Mager, J., Montgomery, N. D., de Villena, F. P.-M. & Magnuson, T. Genome imprinting regulated by the mouse Polycomb group protein Eed. Nat. Genet. 33, 502–507 (2003)
Lewis, A. et al. Imprinting on distal chromosome 7 in the placenta involves repressive histone methylation independent of DNA methylation. Nat. Genet. 36, 1291–1295 (2004)
Umlauf, D. et al. Imprinting along the Kcnq1 domain on mouse chromosome 7 involves repressive histone methylation and recruitment of Polycomb group complexes. Nat. Genet. 36, 1296–1300 (2004)
Terranova, R. et al. Polycomb group proteins Ezh2 and Rnf2 direct genomic contraction and imprinted repression in early mouse embryos. Dev. Cell 15, 668–679 (2008)
Pandey, R. R. et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 32, 232–246 (2008)
Yamasaki-Ishizaki, Y. et al. Role of DNA methylation and histone H3 lysine 27 methylation in tissue-specific imprinting of mouse Grb10. Mol. Cell. Biol. 27, 732–742 (2007)
Sanz, L. A. et al. A mono-allelic bivalent chromatin domain controls tissue-specific imprinting at Grb10. EMBO J. 27, 2523–2532 (2008)
Duffié, R. et al. The Gpr1/Zdbf2 locus provides new paradigms for transient and dynamic genomic imprinting in mammals. Genes Dev. 28, 463–478 (2014)
Monk, D. et al. Comparative analysis of human chromosome 7q21 and mouse proximal chromosome 6 reveals a placental-specific imprinted gene, TFPI2/Tfpi2, which requires EHMT2 and EED for allelic-silencing. Genome Res. 18, 1270–1281 (2008)
Henckel, A. et al. Histone methylation is mechanistically linked to DNA methylation at imprinting control regions in mammals. Hum. Mol. Genet. 18, 3375–3383 (2009)
Kobayashi, H. et al. Imprinted DNA methylation reprogramming during early mouse embryogenesis at the Gpr1-Zdbf2 locus is linked to long cis-intergenic transcription. FEBS Lett. 586, 827–833 (2012)
Saitou, M., Kagiwada, S. & Kurimoto, K. Epigenetic reprogramming in mouse pre-implantation development and primordial germ cells. Development 139, 15–31 (2012)
Pires, N. D. & Grossniklaus, U. Different yet similar: evolution of imprinting in flowering plants and mammals. F1000Prime Rep. 6, 63 (2014)
Moreno-Romero, J., Jiang, H., Santos-González, J. & Köhler, C. Parental epigenetic asymmetry of PRC2-mediated histone modifications in the Arabidopsis endosperm. EMBO J. 35, 1298–1311 (2016)
Inoue, A., Nakajima, R., Nagata, M. & Aoki, F. Contribution of the oocyte nucleus and cytoplasm to the determination of meiotic and developmental competence in mice. Hum. Reprod. 23, 1377–1384 (2008)
Ohnishi, Y. et al. Small RNA class transition from siRNA/piRNA to miRNA during pre-implantation mouse development. Nucleic Acids Res. 38, 5141–5151 (2010)
Inoue, A., Akiyama, T., Nagata, M. & Aoki, F. The perivitelline space-forming capacity of mouse oocytes is associated with meiotic competence. J. Reprod. Dev. 53, 1043–1052 (2007)
Sugimoto, M. et al. A simple and robust method for establishing homogeneous mouse epiblast stem cell lines by Wnt inhibition. Stem Cell Reports 4, 744–757 (2015)
Xiang, Y. et al. JMJD3 is a histone H3K27 demethylase. Cell Res. 17, 850–857 (2007)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
John, S. et al. Chromatin accessibility pre-determines glucocorticoid receptor binding patterns. Nat. Genet. 43, 264–268 (2011)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44 (W1), W160–W165 (2016)
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013)
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Song, Q. et al. A reference methylome database and analysis pipeline to facilitate integrative and comparative epigenomics. PLoS ONE 8, e81148 (2013)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Acknowledgements
We thank S. Matoba for technical advice and L. Tuesta for reading the manuscript. This project was supported by the Howard Hughes Medical Institute. F.L. is supported by a Charles A. King Trust Postdoctoral Research Fellowship. Y.Z. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
A.I. and Y.Z. conceived the project and designed the experiments. A.I. performed the experiments; L.J. analysed sequencing datasets. F.L. performed liDNase-seq and RNA-seq; T.S. assisted in embryo manipulation experiments; A.I., L.J., and Y.Z. interpreted the data. A.I. and Y.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks R. Feil and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Identification of parental allelic DHSs.
Related to Fig. 1. a, Correlation of DHSs between three biological replicates in paternal and maternal pronuclei (PN). b, Bi-allelic DHSs (grey), Ps-DHSs (blue), and Ms-DHSs (red). The cutoffs used to define these DHS groups are indicated. c, Averaged DHS signals of Ps-DHSs and Ms-DHSs within ±5 kb around DHSs. d, Genomic distribution of DHSs. Promoters represent ±1 kb around transcription start sites. ‘Random’ indicates the percentages of each genomic element of the mouse genome. e, Percentages of DHSs located at CpG islands (CGIs). The genomic locations of CGIs are defined previously9. f, Genome browser view of Ps-DHSs at ICRs of representative imprinted genes. The genomic locations of ICRs were referred in ref. 9. g, List of genes harbouring promoter Ps-DHSs or Ms-DHSs in zygotes. h, Genome browser view of representative allelic promoter DHSs at genes not previously known to be imprinted.
Extended Data Figure 2 Correlation between allelic ZGA in two-cell embryos and allelic DHSs in zygotes.
Related to Fig. 1. a, Schematic for identifying parental allele-specific gene expression at ZGA. Androgenetic embryos and gynogenetic embryos were produced by pronuclear transfer. Androgenetic two-cell embryos contained paternally expressed nascent transcripts and maternally stored transcripts. Gynogenetic two-cell embryos contained maternally expressed nascent transcripts and maternally stored transcripts. α-Amanitin-treated (Ama) two-cell embryos contained maternally stored transcripts only. b, Correlation between biological duplicate of two-cell RNA-seq samples. c, Flowchart for avoiding maternally stored transcripts and identifying nascent allelic transcripts at ZGA. d, Nascent transcripts in androgenetic and gynogenetic two-cell embryos. For each gene, the FPKM value in α-amanitin-treated embryos was subtracted from that in androgenetic and gynogenetic embryos, respectively. Androgenetic- and gynogenetic-specific DEGs (fold change > 10) are indicated in blue and red, respectively. Known imprinted genes are indicated in green. e, f, DHS allelic bias at promoters (±0.5 kb at transcription start sites) of androgenesis- (e) and gynogenesis- (f) specific DEGs. Fold change > 2 was considered as ‘bias’ (blue or red).
Extended Data Figure 3 Zygotic Ms-DHSs are inherited from oocyte DHSs.
Related to Fig. 2. a, Correlation between three biological replicates of liDNase-seq for germinal vesicle nuclei isolated from fully grown oocytes. b, Genome browser view of sperm DHSs passed on to paternal pronuclei of zygotes. The nearest gene names are indicated at the top of each panel. c, Heat map showing Ps-DHSs. Each row represents the liDNase-seq signal intensity at DHS ± 5 kb. Note that Ps-DHSs are largely absent in both sperm and oocytes. d, Genome browser view of representative Ps-DHSs. e, Heat map showing Ms-DHSs. Note that Ms-DHSs are mostly already present in oocytes. f, Genome browser view of representative Ms-DHSs. g, Heat map showing bi-allelic DHSs. h, Genome browser view of representative bi-allelic DHSs.
Extended Data Figure 4 Distinct epigenetic features of Kdm6b- and Kdm4d-affected Ps-DHSs.
Related to Fig. 2. a, Percentages of Ps-DHSs that overlap (black) or are associated (grey) with oocyte gDMRs within ±100 kb. Oocyte gDMR was defined by >80% methylation in oocytes and <20% methylation in sperm. b, Percentages of Ps-DHSs organized on the basis of their oocyte DNA methylation levels. c, H3K27me3 signal levels at Ps-DHSs ± 1 kb in gametes (left) and zygotes (right). Ps-DHSs were divided into oocyte DNA hypomethylated (0–20%, n = 296) and hypermethylated groups (80–100%, n = 305). Middle lines in the boxes represent the medians. Box edges and whiskers indicate the 25th/75th and 2.5th/97.5th percentiles, respectively. d, Representative images of Kdm6b- or Kdm4d-injected zygotes stained with anti-Flag antibody, using non-injected zygotes as negative controls. e, Representative images of zygotes stained with anti-H3K27me3 antibody. M, maternal pronucleus; P, paternal pronucleus. The bar graph on the right represents relative immunostaining signal intensity of maternal pronuclei. The averaged signal of non-injected zygotes was set as 1.0. The total numbers of embryos examined were 8 (No injection), 13 (Kdm6bWT), and 10 (Kdm6bMUT). Error bars, s.d. ***P < 0.001 (two-tailed Student’s t-test). NS, statistically not significant. f, Representative images of zygotes stained with anti-H3K9me3 antibody. The bar graph on right represents relative immunostaining signal intensity in the maternal pronuclei. The averaged signal of non-injected zygotes was set as 1.0. The total numbers of embryos examined were 5 (No injection), 5 (Kdm4dWT), and 7 (Kdm4dMUT). Error bars, s.d. ***P < 0.001 (two-tailed Student’s t-test). NS, statistically not significant. g, Correlation between biological duplicates of liDNase-seq for maternal (Mat) and paternal pronuclei (Pat) of Kdm6bWT- and Kdm6bMUT-injected zygotes. h, Correlation between biological duplicates of liDNase-seq for maternal and paternal pronuclei of Kdm4dWT- and Kdm4dMUT-injected zygotes. i, Genome browser view of representative Ps-DHSs affected by Kdm4dWT. j, H3K27me3 signals at Kdm6b- or Kdm4d-affected Ps-DHSs ± 1 kb in gametes (left) and zygotes (right). Middle lines in the boxes indicate the medians. Box edges and whiskers indicate the 25th/75th and 2.5th/97.5th percentiles, respectively.
Extended Data Figure 5 Androgenetic- and gynogenetic-specific DHSs in morula embryos.
Related to Fig. 3. a, Correlation between biological duplicates of liDNase-seq for androgenetic and gynogenetic morula embryos. b, Averaged SNP-tracked liDNase-seq signal intensity of paternal and maternal alleles in hybrid morula embryos. The data were obtained from morula embryos of a BDF1 and JF1 cross7. Plots from the biological duplicates are shown. Note that paternal (JF1), but not maternal (BDF1), SNP reads are enriched in AG-DHSs (left), while neither SNP read is enriched in GG-DHSs (right). c, Genome browser view of DHSs at known ICRs. The genomic locations of ICRs were defined previously9. d, AG-DHSs grouped on the basis of their oocyte DNA methylation levels.
Extended Data Figure 6 Allelic gene expression in morula embryos and allelic H3K27me3 at non-canonical and canonical imprinted genes.
Related to Fig. 3. a, Correlation between biological duplicates of RNA-seq samples. b, Gene expression levels in androgenetic and gynogenetic morula embryos. Androgenetic- and gynogenetic-specific DEGs (fold change > 10) are indicated in blue and red, respectively. Paternally and maternally expressed known imprinted genes are indicated in green and orange, respectively. c, Genome browser views of allelic H3K27me3 levels in non-canonical imprinted genes. Sp; sperm. Oo; MII-stage oocyte. Paternal (Pat) and maternal (Mat) allele signals in one-cell and ICM were based on SNP analyses. d, Genome browser views of allelic H3K27me3 levels in representative canonical imprinted genes. Known ICRs are indicated at the bottom of each imprinted gene.
Extended Data Figure 7 The effect of Kdm6b mRNA injection on maternal allele expression and accessibility.
Related to Fig. 4. a, Developmental ratio of Kdm6bWT- and Kdm6bMUT-injected parthenogenetic (PG) embryos. The total embryo numbers examined were 60 (WT) and 58 (MUT). b, Correlation between biological duplicates of RNA-seq for Kdm6bWT- and Kdm6bMUT-injected parthenogenetic embryos. c, Relative gene expression levels of canonical imprinted genes expressed in androgenetic (AG) morula embryos (RPKM > 0.5). Note that none are de-repressed by Kdm6bWT injection. d, Correlation between biological duplicates of liDNase-seq for Kdm6bWT- and Kdm6bMUT-injected parthenogenetic embryos. e, f, Wide genome browser views of non-canonical (e) and canonical (f) imprinted genes. Arrowheads indicate AG-DHSs at which chromatin accessibility is gained in Kdm6bWT-injected parthenogenetic embryos (shown in Fig. 4e). Known ICRs are indicated above each panel of canonical imprinted genes (f). g, Genome browser view of AG-DHSs at ICRs of representative canonical imprinted genes.
Extended Data Figure 8 Genomic imprinting in E6.5 embryos.
Related to Fig. 5. a, Expression levels of marker genes for trophectoderm (Cdx2) and ICM (Sox2) in the samples. b, Correlation between biological duplicates of the E6.5 epiblast (EPI), visceral endoderm (VE), and extra-embryonic ectoderm (EXE) RNA-seq samples from both B6 × PWK and PWK × B6 crosses. c, Expression levels of marker genes for epiblast (Pou5f1 and Nanog), visceral endoderm genes (Gata6 and Gata4) and extra-embryonic ectoderm (Elf5 and Gata3) in the samples. d, Heat map showing PEGs and MEGs in epiblast, visceral endoderm, and extra-embryonic ectoderm of E6.5 embryos. All genes exhibiting parental allele-specific expression (fold change > 2 in both B6/PWK (B × P) and PWK/B6 (P × B)) in each sample are shown. Genes not previously known to be imprinted are indicated in bold. e, Genome browser view of RNA-seq data of newly identified imprinted genes. D7Ertd715e and Smoc1 are paternally expressed, and Mas1 is maternally expressed.
Extended Data Figure 9 Sample preparation and quality verification.
Related to Fig. 5. a, Experimental scheme of placenta cell purification. Sperm or oocytes were collected from B6GFP mice, and in vitro fertilized with the counterparts collected from the PWK strain. Embryos were transplanted into surrogate mothers. The placentae were harvested at E9.5 and dissociated into single cells by trypsin treatment before FACS of GFP-positive cells. b, Correlation between biological duplicates of RNA-seq samples. c, Total numbers of the paternal and maternal SNP reads in the purified placental cells.
Extended Data Figure 10 Genomic imprinting in E9.5 placentae.
Related to Fig. 5. a, Heat map showing PEGs and MEGs in E9.5 placentae. All genes exhibiting parental allele-specific expression (fold change > 2 in both B6/PWK and PWK/B6) are shown. Genes not previously known to be imprinted are indicated in bold type. b, Genome browser view of RNA-seq data of newly identified imprinted genes. D7Ertd715e and Smoc1 are paternally expressed, and Cbx7 and Thbs2 are maternally expressed.
Supplementary information
Supplementary Data
This zipped file contains Supplementary Tables 1-15 and a Supplementary Table guide.
Rights and permissions
About this article
Cite this article
Inoue, A., Jiang, L., Lu, F. et al. Maternal H3K27me3 controls DNA methylation-independent imprinting. Nature 547, 419–424 (2017). https://doi.org/10.1038/nature23262
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23262
- Springer Nature Limited
This article is cited by
-
Selfish conflict underlies RNA-mediated parent-of-origin effects
Nature (2024)
-
TCF3, TCF12 and distinct enhancers regulate oocyte transcription
Nature Cell Biology (2024)
-
Polycomb function in early mouse development
Cell Death & Differentiation (2024)
-
Inheritance of H3K9 methylation regulates genome architecture in Drosophila early embryos
The EMBO Journal (2024)
-
DNA methylation in mammalian development and disease
Nature Reviews Genetics (2024)