Abstract
Chromosomes in proliferating metazoan cells undergo marked structural metamorphoses every cell cycle, alternating between highly condensed mitotic structures that facilitate chromosome segregation, and decondensed interphase structures that accommodate transcription, gene silencing and DNA replication. Here we use single-cell Hi-C (high-resolution chromosome conformation capture) analysis to study chromosome conformations in thousands of individual cells, and discover a continuum of cis-interaction profiles that finely position individual cells along the cell cycle. We show that chromosomal compartments, topological-associated domains (TADs), contact insulation and long-range loops, all defined by bulk Hi-C maps, are governed by distinct cell-cycle dynamics. In particular, DNA replication correlates with a build-up of compartments and a reduction in TAD insulation, while loops are generally stable from G1 to S and G2 phase. Whole-genome three-dimensional structural models reveal a radial architecture of chromosomal compartments with distinct epigenomic signatures. Our single-cell data therefore allow re-interpretation of chromosome conformation maps through the prism of the cell cycle.
Similar content being viewed by others
Main
As cells progress from mitosis to interphase and back again, their chromosomes undergo notable structural alterations that have intrigued microscopists for more than a century1. In preparation for mitosis and cell division, chromosomes are packaged into rod-like structures that are functionally inert. Upon mitotic exit, chromosomes rapidly decondense into structures that allow regulated access to functional elements, enabling genome functions such as transcription, gene silencing and DNA replication. Although the structural and organizational principles of the interphase genome have been the subject of intense study, in particular using Hi-C2,3,4,5,6,7,8,9,10, genome-wide, high-resolution understanding of chromosome cell-cycle dynamics11 is still limited by current technologies.
Here we scale up single-cell Hi-C12 using flow cytometry sorting, a new library preparation protocol and a new automation scheme, and apply this methodology to thousands of diploid and haploid mouse embryonic stem (ES) cells. Having thousands of single-cell maps has allowed us to develop a strategy for in silico cell-cycle phasing of single nuclei, and use it to study the cell-cycle dynamics of chromosomal structural features. Such features include insulation profiles and looping contacts that together define TADs, and preferential contacts between generalized chromosome pseudo-compartments. We also present whole-genome, restraint-based three-dimensional (3D) models from G1 phase haploid nuclei, following the structural dynamics during the exit from mitosis. Our results reveal a continuum of dynamic chromosomal structural features throughout the cell cycle, creating a new point of reference for interpretation of Hi-C maps.
Improved multiplexed single-cell Hi-C
We redesigned our original single-cell Hi-C approach12 to use a four-cutter restriction enzyme, Tn5-transposase tagmentation, flow cytometry sorting of single cells into 96-well plates and partially automated library preparation (Fig. 1a). We processed a total of 2,924 single F1 hybrid 129 × Castaneus mouse ES cells grown in 2i media using 1.5 million reads per cell on average (Extended Data Fig. 1a–c). Analysis of various quality control metrics (Extended Data Fig. 1d–i) confirmed the robustness of the protocol over different batches, and in particular identified karyotypic imperfections within the single-cell sample (Extended Data Fig. 1j). Overall, 1,992 cells (68.1%) passed stringent quality control filters for further analysis. This provided median coverage of 393,506 restriction fragments, and 127,233 distinct >1 kb contacting pairs per cell (Fig. 1b). We found the incidence of trans-chromosomal contacts in the single-cell maps was low (median 5.87%, Fig. 1c), indicative of high library quality13 and reflecting specific and restricted interfaces of chromosome territories (Extended Data Fig. 2). These data provided us with the opportunity to study variation in chromosomal conformation across a large single-cell pool, on the basis of a high-resolution reference ensemble map (Extended Data Fig. 3).
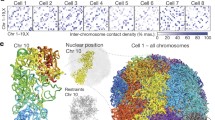
a, Single-cell Hi-C schematic. b, Number of informative contacts retrieved per cell that passed the quality control filter. Median 127,233 (dashed line). c, Percentage of trans-chromosomal contacts per cell that passed the quality control filter. Median 5.87% (dashed line). d, Genome-wide contact map of a representative mitotic cell (1CDX4_242). e, Percentage of short-range (<2 Mb) versus mitotic band (2–12 Mb) contacts per cell (left), and repli-score (right). Cells are grouped by percentage short-range and percentage mitotic contacts and coloured by group. f, Single-cell contact decay profiles ordered by in silico inferred cell-cycle phasing, with approximate cell-cycle phases shown on top. Each column represents a single cell. g, Selected phased and pooled contact maps.
Integrated cell-cycle phasing
Mammalian mitotic chromosomes are rod-shaped, and bulk Hi-C analysis on sorted M phase cells has revealed a characteristic scale of cis-chromosomal contact distances peaking between 2 and 12 Mb11. We examined the cells with the highest fraction of contacts at this mitotic range, and found their overall distribution of contact distances was similar to that previously observed for mitotic cells (Extended Data Fig. 4a). These single-cell mitotic maps were enriched for notable patterns of trans-chromosomal alignments (Fig. 1d, Extended Data Fig. 4b), suggesting that the chromosomes interacted along their entire length (head-to-head and tail-to-tail) (1CDX4_242 in Extended Data Fig. 4b) as though organized by the mitotic spindle in telophase (see also 1CDX_74 and 1CDX_453 in Extended Data Fig. 4b for partial or disordered chromosomal alignments). Comparison of the frequency of contacts in the mitotic range to the frequency of contacts at shorter distances (short-range contacts less than 2 Mb; Fig. 1e) shows a remarkable circular pattern evocative of a gradual chromosome remodelling process as cells enter and exit mitosis. These observations suggest that chromosome conformation may be used to phase single cells at various stages of the cell cycle around mitosis.
Early- and late-replicating domains are defined by their copy number dynamics during S phase, and show distinct enrichment for active and inactive epigenetic marks respectively14. Indeed, we found that normalized TAD coverage across the single-cell dataset reflects a strong correlation between domains previously annotated as early- or late-replicating in mouse ES cells15 (Extended Data Fig. 4c, d). We computed a ‘repli-score’ for each cell based on the copy-number ratio of early-replicating regions to total coverage. G1 cells would be expected to have low repli-scores, whereas early S cells would be associated with increasing scores, peaking in mid-S before declining again in late S phase and returning to a low level in G2–M phase. Combining repli-score with the circular pattern exhibited by the frequencies of short-range and mitotic contacts yields exactly that expected trajectory (Fig. 1e). Together, the mitotic signatures and time of replication analysis define two major anchors to support phasing of the entire single-cell Hi-C dataset along the cell cycle.
We studied the distributions of contact distance scales in single cells using nonlinear dimensionality reduction (Extended Data Fig. 4e) and clustering (Extended Data Fig. 5). Both approaches indicate the data can be globally represented by a cyclic, gradual process of coordinated change that links the highly distinct mitotic and replicating states characterized above. To capture this variation precisely, we used a supervised approach that groups cells to phases and then orders cells within phases. We first identified cells approaching mitosis (pre-M regime, orange) and coming out of mitosis (post-M regime, red) by thresholding short-range and mitotic contact frequencies (based on Fig. 1e, see Supplementary Methods). Cells in both regimes can be ordered by the ascending and descending frequency of mitotic contacts for the pre-M and post-M regimes, respectively. Next, we observed that cells with less than 63% short-range contacts do not include the replication group and probably represent a G1 regime (blue). The mean distance of long-range contacts in these cells is correlated to the overall frequency of short-range contacts, allowing ordering of cells within this group (Extended Data Fig. 6a). Cells with over 63% short-range contacts show a continuum of repli-score and short-range contact enrichments (Extended Data Fig. 6b). These were modelled using two additional regimes, one ordered based on a gradual increase in repli-score and short-range contacts (the early-S regime, green), and one ordered based on a gradual decrease in repli-score and further increase in short-range contacts (the late-S/G2 regime, purple). Using this phasing strategy, we observed a smooth, cyclical transition of the distribution of genomic contact distances (Fig. 1f), which is insensitive to batch effects (Extended Data Fig. 6c, d) and robust to subsampling of the data or analysis of specific chromosomes (Extended Data Fig. 6e, f). In summary, we developed an in silico cell-cycle phasing approach that organizes single cells along a smooth trajectory of highly varied chromosome conformations (Fig. 1g, Extended Data Fig. 5) leading from the mitotic state to replication and back into mitosis.
To confirm our Hi-C-based cell-cycle phasing, we analysed 1,171 cells sorted based on levels of the DNA replication marker geminin and DNA content using fluorescence-activated cell sorting (FACS), and found that the ordering of cells based on their contacts is remarkably consistent with their stage as determined by FACS (Fig. 2a, b). We further hypothesize that the refined ordering we impose within each coarse-grained stage is likely to represent cell-cycle progression at high resolution. This was initially demonstrated by correlating Hoechst and geminin FACS indices to the inferred phasing rank of 420 single-cell Hi-C profiles (Fig. 2c, Extended Data Fig. 6g), deriving correlation even within classical sorting gates. To support this idea further, we analysed the distribution of key statistics for single cells ordered according to their inferred cell-cycle stage. For reference, we projected the repli-score on the cell-cycle progression (Fig. 2d), validating that it was low during G1 and G2 phases, showed a consistent gradual increase during early S phase, and decreased during late S phase. The total number of contacts (Fig. 2e), which unlike the repli-score is independent of the inferred phasing, was shown to increase gradually during S phase, peak at G2 phase at an almost twofold higher level than at G1 phase, before dropping sharply after mitosis. The fraction of trans-chromosomal contacts exhibits an unforeseen cell-cycle dynamic, with a smooth curve from its lowest point of around 3% in G2 and mitosis, to approximately 8% in early S phase (Fig. 2f). Next, we quantified the trans-alignment of chromosomes, which is a strong characteristic of mitotic cells. Remarkably, despite being a priori unrelated to the inferred G1 ordering scheme, we found the trans-alignment score declined smoothly from its peak at mitosis to its nadir at the end of the G1 phase, immediately before the start of S phase (Fig. 2g; see Extended Data Fig. 6h for examples). Collectively, these results validate our phasing strategy and open the way to analyse cell-cycle dynamics of Hi-C architectural features at high resolution.
a, FACS sorting scheme. b, Positions of FACS-sorted cells along a circular layout of their in silico phasing (clockwise, M is on top). c, Comparing in silico phasing rank to combined Hoechst and geminin FACS indices for 420 indexed-sorting cells. Black lines demarcate approximate cell-cycle phases. d, Repli-score per cell ordered by cell-cycle phasing (left to right). FACS-sorted cells coloured as in a and b, and remaining cells are grey. Outliers (0.5%) are red. Cyclic average window (n = 100) trend shown in black. e–g, Similar to d, showing additional single-cell statistics. Arrows in g correspond to the heat maps in Extended Data Fig. 6h.
Cell-cycle chromosomal dynamics
We identified 3,434 TAD borders in the pooled Hi-C map using analysis of contact insulation signatures6 (Extended Data Fig. 3). We created sub-pools of G1, early-S and late-S–G2 cells in silico to compare insulation patterns between the three major cell-cycle phases. We found that the locations of TAD borders are generally unchanged in these pools, but that the average intensity of the borders is dynamic (Extended Data Fig. 7a–c). We calculated the mean insulation at borders for each cell (Fig. 3a) and found that the intensity of insulation varies markedly over the cell cycle. Insulation is not detectable in mitotic maps (Extended Data Fig. 7d, e), and emerges rapidly upon exit from mitosis in early G1. Insulation increases to its maximum during G1 phase, but declines when replication begins, plateauing at mid-S phase at its lowest interphase level and remaining fixed through G2 until mitosis. We note that since insulation is defined by the depletion of short-range contacts crossing TAD borders, it may be affected by changes in the frequency of contacts in the 25 kb–2 Mb distance range. However, this overall frequency only partially predicts single-cell insulation (Extended Data Fig. 7f).
a, Mean contact insulation at TAD borders, phased cells coloured as in Fig. 2. b, A/B compartmentalization score. c, Clustering borders (k-means, k = 4) according to insulation profile over inferred phasing. Mean insulation of the borders in a cluster per cell (blue dots), mean cluster insulation trend (blue line) and global trend (black line). d, Mean time of replication (ToR) ± 200 kb of the Fig. 3c clustered borders. Early-replicating positive, late-replicating negative. e, Convergent CTCF loops contact enrichment. Showing data for 762 early–early loops and 456 late–late loops. f, Aggregated contacts around convergent CTCF loops distanced 0.2–1 Mb from each other, pooling cells by their in silico phasing group and normalizing by the expected profile using the shuffled pooled contact map. g, Mean intra-TAD contact distance stratifying domains by length, and mean time of replication.
Next, we studied the dynamics of compartmentalization of the genome into active (A) and non-active (B) groups of TADs by analysing the cis-chromosomal compartment linkage. For each cell, we estimated the depletion of A–B contacts given the observed A–A and B–B counts (Supplementary Methods, Fig. 3b). Similar to insulation, compartmentalization is weakest during mitosis, as expected from the condensed uniform structure at that stage11. However, in contrast to TAD insulation, compartmentalization is weak during G1 phase and increases upon entering S phase, with a further increase in late S phase, reaching a maximum in G2, just before a rapid and complete loss approaching mitosis. Compartmentalization calculated from trans-chromosomal contacts shows exactly the same trend (Extended Data Fig. 7g). The distinct cell-cycle dynamics of TAD insulation and compartment structure suggest that independent mechanisms contribute to these phenomena.
To study the cell-cycle dynamics of insulation at individual TAD borders and compare it to the global trend above, we grouped single cells into 20 time slots according to their inferred cell-cycle phase, and pooled data from each time slot to compute an insulation time series for each border. We then clustered the borders based on their dynamics in these time slots, and examined the mean insulation in each cluster per cell, along the inferred phasing (Fig. 3c). Borders demonstrate similar qualitative dynamics in all clusters. However, each cluster varies in timing and intensity of deterioration in insulation during S phase. Cluster C4 includes borders showing the strongest G1 insulation and C2 borders show the earliest loss of insulation in S. Notably, borders in both clusters tend to be located between early-replicating TADs (Fig. 3d). Borders in C3 show a gradual increase in insulation well into S phase, followed by abrupt loss of insulation in mid-S phase. This cluster is enriched for borders separating early-replicating from mid/late-replicating TADs (Fig. 3c, d). C1, which has the weakest G1 insulation, shows gradual and delayed loss of insulation, and is enriched for borders between late-replicating domains. These data suggest that loss of insulation in S phase is temporally coupled to the replication process. Specifically, replication through a border or in both adjacent TADs is linked to loss of insulation (C1, C2, C4), whereas replication of only one adjacent TAD is linked to a temporary increase in insulation (C3). In all cases, insulation remains low after replication, not re-emerging until the next cell cycle.
We identified pairs of looping loci by analysis of pooled matrices and shuffled pooled matrices (Extended Data Fig. 7h). We focused on 2,036 top-scoring loops that were associated with convergent CTCF motifs and evidence for CTCF/cohesin binding (Supplementary Methods). We then grouped loops according to the replication time of their anchors (early–early versus late–late) and computed for each group and each single cell the total number of looping contacts in 20 × 20-kb bins around the loop anchors, normalized by the number of contacts in the surrounding 60 × 60-kb bin (Fig. 3e). We found that loop enrichment increases after mitosis, is generally stable through G1 phase, but then decreases slightly in early S phase for early-replicating loops and in late S phase for later-replicating loops. Spatial enrichment analysis shows conserved localized loop enrichment (Fig. 3f). We do not detect loops that are specific to a certain cell-cycle phase (Extended Data Fig. 7i), suggesting that even when decreased in average intensity during replication, loops are markedly observed in the data. However, the resolution of the assay is not sufficient to allow detection of individual loop turnover at single-cell resolution, leaving unresolved regimes for rapid loop dynamics. Notably, loops are still apparent in cells approaching mitosis despite the fast chromosomal condensation—unlike cells after mitosis, which do not have any contact enrichment around loops (Fig. 3f). The notable lack of loop contact enrichment in mitotic maps does not necessarily imply loops are disassembled at this stage, since mitotic condensation may increase the local contact frequency around loop anchors and decrease Hi-C sensitivity.
Given the relative stability of looping contacts that demarcate TAD structures and the dynamic cell-cycle rise and fall in TAD insulation, we next asked whether chromatin structure within TADs changes during the cell cycle, or whether TADs constitute stable chromatin blocks that are re-organized dynamically. We computed average intra-TAD contact distances and used these as a measure of TAD condensation per cell, reasoning that condensed TADs will be characterized by higher mean contact distance than de-condensed TADs3,16. We compared condensation for groups of TADs while controlling for size and time of replication (Fig. 3g). In G1 phase, early-replicating TADs are de-condensed compared to mid- or late-replicating TADs, consistent with open chromatin in active regions. Early-replicating TADs are further de-condensed in early S phase, before showing a remarkable increase in condensation at mid-S phase, probably after their replication is complete. Following that, and during late S and G2 phases, early-replicating TADs increase their mean condensation to levels comparable to those of late-replicating TADs in G1 phase. Condensation of mid-early replicating TADs follows a similar trend, with some delay and damped amplitude. Mid-late and late-replicating TADs, on the other hand, show constitutively high condensation in G1 and throughout S phase, with further marked increases observed for mid-late TADs in G2 phase. We note that the potential involvement of trans sister chromatid contacts in driving the observed increase in mean contact distance cannot be determined unambiguously. However, the delay between replication of early domains at the start of S phase and their increased compaction in mid/late S phase suggests sister-chromatid contacts cannot fully account for the dynamics observed.
We next generated 1,247 quality control positive single-cell Hi-C maps from haploid mouse ES cells, to allow systematic analysis of conformation in single-copy chromosomes. We confirmed haploid ES cell maps show comparable quality control metrics to diploid cells, with less than half of the contacts per cell and some specific patterns of chromosome loss that we filtered in silico (Extended Data Fig. 8a–c). We derived a pooled ensemble haploid Hi-C map and computed TAD structures, insulation and A/B compartments (compared to diploid in Extended Data Fig. 3f). We then used the same approach for cell-cycle phasing that was developed for diploid cells (Fig. 4a). Remarkably, we observe a similar cyclical trend, and analogous dynamics of insulation, loop enrichment and TAD condensation (Extended Data Fig. 8d–g). The universality of these processes shows that haploid ES cells are comparable to normal diploid cells, and a good model for studying chromosome conformation and dynamics, while providing the opportunity for unambiguous 3D modelling of single-copy chromosomes for G1 cells.
a, Similar to Fig. 1g, but analysing haploid cells. b, Chromosome idiogram depicting inferred A-score of each TAD. c, The mean time of replication versus trans A-score for each TAD. d, The mean cis A-association score versus trans A-score for each TAD. e, Standard deviation versus mean A-association scores per TAD across cells. Colours mark groups of TADs stratified by mean A-association. f, Standard deviation of A-association score across TADs stratified as in e and averaged separately over cells in each of the three main inferred cell-cycle phases: G1, early S and late S–G2. *P < 0.05, **P < 0.01, ***P < 0.001, one-sided Mann–Whitney U-test. Outliers not shown. g, Per-TAD distributions of A-association scores in the three phased cell-cycle populations. h, i, Long-range (>2 Mb) cis-chromosomal (h) and trans-chromosomal (i) contact frequency between TAD groups according to pseudo-compartments.
A spectrum of epigenetic compartments
We defined A-scores for each TAD as the fraction of trans-chromosomal contacts made with the A compartment, and found a spectrum that generalizes the original discrete partitioning of the genome into two compartments (Fig. 4b). A-scores correlate with time of replication; high A-score TADs are early-replicating and low A-score TADs are late-replicating (Fig. 4c). These scores are conserved between diploid and haploid cells and growth conditions (Extended Data Fig. 9a), suggesting that a continuum of compartment affinities is a robust property of chromosomal domains. Such a continuum may be created if individual TADs switch compartments in individual cells, thereby creating a gradient of A-compartment affinities. Alternatively, it is possible that the A-score gradient is conserved at the single-cell level, representing a structural continuum, rather than a two-compartment structure that is heterogeneous across a cell population. To explore these questions at single-cell resolution, we needed sufficient TAD coverage in each cell, so we computed a distinct single-cell A-association score for each TAD, using the mean A-score of its long-range (>2 Mb) cis-contacting TADs. The cis A-association scores and trans A-scores are highly consistent on average (Fig. 4d), but the cis A-association score is defined by more contacts, while being constrained by the chromosomal larger-scale domain architecture. The standard deviation of A-association scores for individual TADs across the entire haploid single-cell cohort was less than 0.06 (Fig. 4e), suggesting limited variability of A-association score between cells. Further analysis (Fig. 4f) showed that A-association score variation decreases in early and late S phase, consistent with the overall increase in A/B compartment strength above. Both sorting TADs by their mean A-association scores (Fig. 4g) and grouping these distributions by unsupervised clustering (Extended Data Fig. 9b) showed that TADs vary quantitatively in their A-association, but no significant group of TADs shows a bimodal compartment association distribution. The same analysis indicated that compartment association polarizes in late S/G2 phase, with many TADs shifting from broadly spread association in G1 or early S, to preferential contacts with TADs in a narrower range of A-association scores following replication.
To explore this effect further, we discretized the A-association score continuum, deriving 20 groups of TADs that corresponded to putative pseudo-compartments (Supplementary Methods). We tested how such pseudo-compartments contact one another trans- and cis-chromosomally, and observed significant cis-chromosomal interaction between TADs within the same and neighbouring pseudo-compartments (Fig. 4h and Extended Data Fig. 9e, top). In trans we observed a more binary contact structure in G1 cells, transitioning towards a gradient of pseudo-compartment preferences following replication (Fig. 4i). A-association analysis of diploid cells shows similar behaviour (Extended Data Fig. 9c, d, e (bottom), f). Analysis of RNA sequencing, histone marks15 and lamin association17 suggested pseudo-compartments are characterized by a gradient of epigenetic signatures (Extended Data Fig. 9g, h). Overall, the ordering of TADs according to A-association score is significantly correlated with a linear model weighting H3K4me1 and H3K4me3 scores (Pearson’s r = 0.79, P < 0.001). Together, these data suggest pseudo-compartments capture large-scale trans- and cis-chromosomal conformation at the population and single-cell level, generalizing the binary A/B compartment scheme.
3D whole-genome modelling and analysis
Using G1 haploid cell data, we developed a structure sampling approach for inferring whole-genome 3D conformation. We applied stringent contact filtering to eliminate contacts that are more likely to be technical artefacts (Supplementary Methods), removing a median of 0.89% of the contacts from each cell (Extended Data Fig. 10a). We applied a simulated annealing protocol12 to sample over 30,000 whole-genome 3D models for 190 single-cell datasets, using contacts as distance restraints at 0.1, 0.5 and 1 Mb resolutions. The resulting 3D models satisfied nearly all observed cis- and trans-chromosomal contacts, with a median of fewer than 0.012% violated constraints per cell (Extended Data Fig. 10b, c). Subsampling 10,000 contacts per cell produced consistent models (Extended Data Fig. 10d, e), while cross-validation tests showed that structural modelling is sufficiently robust to re-discover trans-chromosomal distances between a pair of contacting chromosomes, even when all contacts between those chromosomes are discarded (Extended Data Fig. 10f).
To study changes in chromosomal structure as cells emerge from mitosis into G1 phase, we analysed over 20,000 models from 126 cells with more than 10,000 contacts per cell and fewer than 0.5% constraint violations. Analysis of large-scale cis-chromosomal shapes shows that individual chromosomes rapidly de-condense from a rod-like mitotic conformation to a more spherical structure as G1 progresses (Fig. 5a, b, Extended Data Fig. 10g, h). We monitored the expansion of chromosomes by measuring the distance between adjacent 500 kb chromatin segments in A or B compartments in each cell (Fig. 5c). A and B compartments are similarly compact upon mitotic exit (P > 0.05, two-sided Mann–Whitney U-test), but as cells progress through G1, A compartments expand more rapidly and to a larger extent than B compartments (P < 0.005, one-sided Mann–Whitney U-test), consistent with decondensed euchromatin and compact heterochromatin. Remarkably, the 20 high-resolution pseudo-compartments form a gradient that generalizes this observation, with the strongest B-like pseudo-compartments most compact and the strongest A-like pseudo-compartments the most expanded (Fig. 5d).
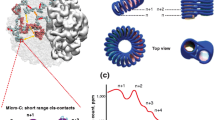
a, Whole-genome conformations, and separated individual chromosomes. Models computed at 100-kb resolution. b, Top, chromosome 1 examples along inferred G1 phase with fitted ellipsoid of inertia. Chromosomes coloured by genomic position pericentromeric (red), telomeric (blue). Bottom, chromosome de-condensation illustrated by distribution of longest-to-shortest (a/c) semi-axis ratios of all chromosomes. c, Decompaction of chromatin in A and B compartments, defined as average distance between neighbouring 500-kb segments of same compartment. d, Same as c for the 20 pseudo-compartments. e, Average radial position of A and B compartments within the nucleus, defined as the volume fraction enclosed by a sphere of that radius. ‘0’ denotes nuclear centre; ‘1’ denotes periphery. f, Same as e for the 20 pseudo-compartments. g, Long-range (>2 Mb) cis-contact enrichment between the 20 pseudo-compartments. h, Trans-chromosomal contact enrichment between the 20 pseudo-compartments.
We then computed the mean nuclear radial positioning of A and B compartments as cells exit mitosis. We observed B compartments occupying more peripheral locations and A compartments tending towards the nuclear interior for cells 9–16 and onward (P < 0.001, one-sided Mann–Whitney U-test, Fig. 5e). Pseudo-compartments generalize this trend, showing progressively more differentiated radial positioning through G1 phase (Fig. 5f). The structural models also confirm and refine the preferential contacts among pseudo-compartments (as shown in Fig. 4h, i), and show that long-range cis-contacts are re-established very early in G1 phase, while trans-compartmentalization is not re-established until late G1 phase (Fig. 5g, h), consistent with 4C observations18. Finally, distributions of chromosomal conformations also suggest preferential contacts of nucleolar organizer regions19 (P < 0.01 one-sided Mann–Whitney U-test; Extended Data Fig. 10i).
Discussion
Using new high-throughput single-cell Hi-C technology, we generated and analysed single-cell contact maps from thousands of cells, and established a strategy for cell-cycle phasing that decouples TADs, compartments and loops that were previously superimposed in bulk maps. We confirmed that genome folding is highly heterogeneous at the single-cell level, but variation is far from random, reflecting a combination of deterministic dynamics and stochastic effects. Our data suggest that the most notable source of deterministic dynamics in the mouse ES cell is the cell cycle. Consistent with previous reports at lower temporal resolution11, we show chromosome condensation in preparation for mitosis and rapid expansion in early G1 phase, while also finding extensive remodelling during replication. Importantly, the effects of these major transitions extend throughout the entire cell cycle, such that the genome is not stably folded at any point in interphase.
Our methodology and other techniques for high-resolution or high-throughput single-cell Hi-C20,21,22 now allow high-resolution analysis of other factors driving chromosome folding and genome regulation. It will be important to interpret future single-cell studies and bulk Hi-C experiments through the cell-cycle prism introduced here. An emerging challenge is therefore the derivation of comprehensive models, combining cell-cycle chromosomal dynamics with gene regulatory processes during differentiation22 or response to stimuli. Using such models, it may become possible to understand how nuclei are capable of supporting the remarkably deterministic and pronounced chromosomal dynamics of mitosis and replication while implementing robust and flexible gene regulatory program—a question that is rapidly becoming one of the key challenges in epigenetics.
Data Availability
All data reported in this study are available at the Gene Expression Omnibus (GEO) under accession GSE94489. All other data are available from the corresponding authors upon reasonable request.
Code Availability
Additional analysis files and scripts are available from our website: http://compgenomics.weizmann.ac.il/tanay/?page_id=742 or upon request.
Accession codes
References
Paweletz, N. Walther Flemming: pioneer of mitosis research. Nat. Rev. Mol. Cell Biol. 2, 72–75 (2001)
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009)
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012)
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012)
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012)
Sofueva, S. et al. Cohesin-mediated interactions organize chromosomal domain architecture. EMBO J. 32, 3119–3129 (2013)
Rao, S. S. P. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014)
Zuin, J. et al. Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc. Natl Acad. Sci. USA 111, 996–1001 (2014)
de Wit, E. et al. CTCF binding polarity determines chromatin looping. Mol. Cell 60, 676–684 (2015)
Flavahan, W. A. et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529, 110–114 (2016)
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013)
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013)
Nagano, T. et al. Comparison of Hi-C results using in-solution versus in-nucleus ligation. Genome Biol. 16, 175 (2015)
Pope, B. D. et al. Topologically associating domains are stable units of replication-timing regulation. Nature 515, 402–405 (2014)
Mouse ENCODE Consortium et al. An encyclopedia of mouse DNA elements (Mouse ENCODE). Genome Biol. 13, 418 (2012)
Boettiger, A. N. et al. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 529, 418–422 (2016)
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010)
Dileep, V. et al. Topologically associating domains and their long-range contacts are established during early G1 coincident with the establishment of the replication-timing program. Genome Res. 25, 1104–1113 (2015)
Britton-Davidian, J., Cazaux, B. & Catalan, J. Chromosomal dynamics of nucleolar organizer regions (NORs) in the house mouse: micro-evolutionary insights. Heredity 108, 68–74 (2012)
Ramani, V. et al. Massively multiplex single-cell Hi-C. Nat. Methods 14, 263–266 (2017)
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59–64 (2017)
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017)
Olivares-Chauvet, P. et al. Capturing pairwise and multi-way chromosomal conformations using chromosomal walks. Nature 540, 296–300 (2016)
Acknowledgements
The authors thank M. Leeb, A. Wutz and J. Gribnau for haploid and diploid ES cells, and the Babraham Flow Cytometry and Sequencing Facilities for assistance. This work was supported by grants from the National Institutes of Health 4D Nucleome 1U01HL129971-01, European Research Council 340152 and 309706, and the Flight Attendant Medical Research Institute.
Author information
Authors and Affiliations
Contributions
T.N., Y.L., P.F. and A.T. conceived the project and designed the experiments. T.N. devised new single-cell Hi-C protocol. T.N. and W.L. carried out experiments. Y.L., C.V., C.D., Y.B., N.M.C., S.W. and A.T. analysed data. T.N., Y.L., C.V., C.D., P.F. and A.T. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Technical quality controls.
Data for all the diploid cells analysed (as specified in i). a, Testing library saturation. Showing the fraction of segment chains supported by a single read in each batch, batch colours match their sorting criteria, see i for details. b, Number of unique molecules captured per cell against the number of sequenced reads. c, Number of reads per unique molecule against number of sequenced reads. d, Observed (red) and expected (by binomial reshuffling of the observed contacts) number of cells each trans-chromosomal contact appears in. e, Correlation of the fraction of trans-chromosomal contacts and close cis-chromosomal contacts (<1 kb, non-digested) with the fraction of contacts in different distance bins. f, Stratification of contacts by the orientation of contacting fragments against contact distance. g, Distribution of the logarithmic decay bin with the most contacts per cell, dashed line at 15.5 mark quality control threshold below which cells are discarded. h, Quality control metrics of single cells by batch ID shown in the same colour as i. Vertical lines mark experimental batches. Shown from top to bottom: total number of contacts (coverage); fraction of inter-chromosomal contacts (%trans); fraction of very close (<1 kb) intra-chromosomal contacts (%no dig); early-replicating coverage enrichment (repli-score); fraction of fragment ends (fends) covered more than once (%dup fend). Horizontal dashed lines mark thresholds used to filter good cells. i, Details on the experimental batches, showing the number of cells in each batch, number of cells passed quality control, mean number of reads per cell (MRPC, in million reads), FACS sorting criteria and the medium used to grow the cells. Batches are coloured by FACS sorting criteria: ‘2n DNA’ and ‘2n < DNA ≤ 4n’ by Hoechst staining (H), ‘G1’, ‘early-S’, ‘mid-S’ and ‘late-S/G2’ by Hoechst and geminin (H + G) staining. j, Coverage enrichment per cell (column) and chromosome (rows), in which the expected coverage is calculated from the frequency in the pooled cells and the total number of contacts in each cell. We discard cells that have any aberrantly covered chromosome (at least twofold enrichment or depletion, shown on the right ‘bad’ panel). Left panel shows all cells ordered by batch ID, with lower panel coloured by FACS sorting criteria (as in h and i).
Extended Data Figure 2 Trans-chromosomal contacts.
a, Examples of inter-chromosomal contact maps for several pairs of chromosomes. Showing contact maps of single cells (blue, each point is a contact) and the pooled contact map on the same chromosomes, using 500-kb square bins. b, Distribution of the number of contacts between selected pairs of chromosomes. Showing the number of contacts per square megabase; red dashed line marks the cutoff for interacting chromosomes. c, Fraction of cells in which each pair of chromosomes was interacting (the pair of chromosomes had normalized interaction above the cutoff shown in b).
Extended Data Figure 3 Pooled contact map and bulk data.
a, Normalized chromosomal contact maps of the pooled diploid 2i single cells (showing chromosomes 10 and 11). b, A chromosome idiogram showing the division of domains to the A and B compartments by k-means clustering of domains trans-chromosomal contact profiles. c, A chromosome idiogram showing the inferred A-score of each domain. d, Comparing A and B domains by their lengths. e, Comparing A and B domains by their mean time of replication (ToR); percentage of H3K4me3 peaks; mean H3K27me3; mean laminB1 values; mean RNA-seq. f, Genome-wide comparison of insulation scores across reference bulk Hi-C dataset (from ref. 23) and different pools of single cells. Insulation is calculated every 10 kb using a scale of 300 kb; Pearson’s correlation coefficient is shown.
Extended Data Figure 4 Mitotic cells and dynamics of coverage by replication time.
a, Contact decay profile of our pool of diploid 2i singles (pool), a reference mouse ES cell bulk Hi-C dataset (Bulk mES, from ref. 23), human K562 mitotic population Hi-C (K562 M, from ref. 11) and our pooled 8 most mitotic-like cells (top M). b, Genome-wide contact maps of 8 putative mitotic diploid cells (cells with the highest fraction of contacts at 2–12 Mb distances). c, Correlation matrix of domain coverage across cells. Ordered by the mean correlation to the top 50 earliest replicating domains minus the mean correlation with the latest 50 domains. Time of replication of each domain is shown at the bottom. d, Comparing the fraction of contacts associated with the latest- and earliest-replicating fends (ToR less than −0.5 and more than 0.5, respectively). Cells are coloured as in Fig. 1e. e, Projection of diploid 2i cells onto a two-dimensional plane using their contact decay profile (>1 kb) by a nonlinear dimensionality reduction algorithm (see Supplementary Methods). Cells are coloured by their repli-score, which was not used to position them on the plane.
Extended Data Figure 5 Clustering cells by contact decay and phasing over the cell cycle.
a, k-means (k = 12) clusters of single-cell contact decay profiles. b, Chromosomal contact maps (chromosome 6) of pooled cells per cluster defined in a and the mean decay trend of each cluster (red) compared to the rest of clusters (grey). c, Mean far contact distances and repli-score against the fraction of short-range contacts per clusters in a (red, cells in cluster; grey, all cells), ordering clusters by their mean percentage short-range.
Extended Data Figure 6 Phasing cells over the cell cycle and quality controls.
a, Mean contact distance among the long-range (4.5–225 Mb) contacts against the fraction of short-range (23 kb–2 Mb) contacts. Dashed red lines mark the cutoffs used to divide cells into the main three groups (G1, early-S and late-S–G2). b, Similar to a but showing repli-score against the fraction of short-range contacts. c, Batch of origin of each phased cell using all diploid 2i and serum cells, coloured by the batch FACS sorting criteria (see Extended Data Fig. 1h for batch information). d, Similar to c but for haploid 2i and serum cells (see Extended Data Fig. 8b for batch information). e, Testing the stability of our phasing, we compare the positions of cells by phasing using only half the chromosomes (first half: 2, 3, 5, 6, 9, 10, 12, 15, 17, 19; second half: X, 1, 4, 7, 8, 11, 13, 14, 16, 18). Position of only 38 cells (<2%) differ in more than 10% of the number of cells (outliers marked in red, 10% margin in dashed grey). Phasing groups marked in black lines. f, Chromosomal comparison of the contact decay metrics used to phase the cells in each group: mean far contact distance for the G1 cells (left) and percentage short-range for G1, S and G2 cells (right). Showing smoothed (n = 51) trends per chromosome on top, chromosomes coloured by length (light grey, chromosome 19; black, chromosome 1). Data for specific chromosomes: chromosome 1 (middle) and chromosome 11 (bottom) as red dots. g, Hoechst and geminin FACS indices of cells in batches for cells in batches 27–35 (Ext Data Fig. 1i) against the cells’ inferred phasing position. h, Genome-wide contact maps of representative cells along the inferred G1 phase, their position marked at the bottom of Fig. 2g.
Extended Data Figure 7 Insulation and loops by cell cycle phase.
a, Insulation profiles of the main three phased groups (G1, early-S and late-S–G2) over 6 Mb regions in chromosomes 1–5. b, Comparing border insulation of phased groups. Dashed lines show the mean insulation value per group. c, Showing mean insulation per cell on mouse ES cell TAD borders taken from ref. 5 (using the centre point of borders smaller than 80 kb) compared to the mean cell insulation over our inferred borders. d, Insulation profile over a 6 Mb region in chromosome 1 (same as a), showing insulation of pooled cells (black), pooled mitotic cells (orange) and a shuffled pool of mitotic cells (red). e, Genome-wide distribution of insulation values for pooled-, pooled-mitotic- and shuffled-pooled-mitotic cells, same colours as in d. f, Mean border insulation per cell against the fraction of short-range (<2 Mb) contacts. g, A/B compartment score on trans-chromosomal contacts, single-cell mean values in dots coloured by FACS sorting, mean trend (black) and mean trend of cis-chromosomal A/B compartment in dashed black (same trend as in Fig. 3b). h, Normalized contact maps of the regions in a, circling the loops detected in distances 200 kb to 1 Mb (marked in dashed black line). i, Comparing loop foci enrichment (log2) calculated per phased group, showing the Pearson’s correlation coefficient.
Extended Data Figure 8 Haploid cells technical quality controls.
a, Quality control metrics of single cells. Vertical lines separate experimental batches shown at bottom in the same colour as b. Shown from top to bottom: total number of contacts (coverage); fraction of inter-chromosomal contacts (%trans); fraction of very close (<1 kb) intra-chromosomal contacts (%no dig); early-replicating coverage enrichment (repli-score); fraction of fragment ends (fends) covered more than once (%dup fend). Horizontal dashed lines mark thresholds used to filter good cells. b, Details on the experimental batches, showing the number of cells in each batch, the number of cells that passed quality control, mean number of reads per cell (in million reads), the medium used to grow the cells and FACS sorting criteria: G1 (H) and G1/S (H) by Hoechst staining. c, Coverage enrichment per cell (column) and chromosome (rows), with expected coverage calculated from the pooled cells. We discard cells that have any aberrantly covered chromosome (at least twofold enrichment or depletion, shown on the left ‘bad’ panel). The right panel shows all cells. d, Similar to Fig. 2d–g but produced from the haploid cells, outliers coloured in black. e, Similar to Fig. 3a, b but produced from the haploid cells, outliers coloured in black. f, Similar to Fig. 3e but produced from the haploid cells. g, Similar to Fig. 3g but produced from the haploid cells, outliers coloured in black.
Extended Data Figure 9 Haploid cells clustering, diploid cells pseudo-compartment analysis.
a, Comparing trans A-score of a reference bulk Hi-C dataset (from ref. 23) and different pools of single cells, using the set of domains inferred from the pool of diploid 2i cells (A domains in red and B in black). Showing on top of each panel the correlation of the domains’ A-score in each dataset. b, Per-domain distributions of A-association score across haploid cells in each of the three main cell cycle inferred phases (from left to right: G1, early S, late S to G2). Domains were clustered (k-means, k = 30) by their distribution in all groups. Inter- and intra-cluster ordering by mean A-association score at late-S–G2 phase. c, Similar to Fig. 4g but for diploid 2i cells. d, Contingency table of haploid and diploid inferred pseudo-compartments, considering all overlapping intervals between the haploid and diploid sets of domains. e, Long-range (>2 Mb) cis-chromosomal contact enrichments between TAD groups according to pseudo-compartments for haploid (top) and 2i diploid (bottom) cells. Showing the expected contact frequency by contacts in the shuffled pooled contact maps on the right. f, Long-range (>2 Mb) cis-chromosomal (top) and trans-chromosomal (bottom) 2i diploid contact enrichments between TAD groups according to pseudo-compartments. Showing the expected contact frequency by the marginal pseudo-compartments coverage on the right. g, TADs are sorted by their mean A-association score in late S–G2 phase (x axis, left, strongly B; right, strongly A), and their mean RNA, H3K4me1, laminB1 and H3K4me3 levels are shown (y axis).
Extended Data Figure 10 Whole-genome structure modelling, quality control and clustering of peri-centromeric regions.
a, The distribution of the fraction of unsupported contacts in the pre-filtered contact set in all 190 haploid cells in 2i inferred as being in G1. b, The distribution of the fraction of violated constraints in the 190 haploid G1 cells. c, The fraction of violated constraints in the 3D models of 190 haploid G1 cells at 500 kb versus 100 kb resolutions (Pearson’s r = 0.96). Black dots represent cells that have at least 10,000 contacts and no more than 0 violations at 1 Mb, 0.5% violations at 500 kb and 0.1% violations at 100 kb. Cells with no violations at either 500 kb or 100 kb resolution are not shown. Red dots, cells with low quality models that were filtered out. d, Mean root mean squared deviation (r.m.s.d.) values between all models of the same cell with at least 10,000 (black) and fewer than 10,000 (red) filtered contacts, for cells at 1 Mb (dots), 500 kb (open diamonds) and 100 kb (triangles) resolutions. e, Model reproducibility across cells. The r.m.s.d. distribution between all models for the same cell (red) compared to the models for different cells (blue), for the 126 cells with the highest quality models. The images show aligned structures of 106 × 1 Mb models (top), 80 × 500 kb models (centre) and 5 × 100 kb models (bottom) for NXT-1091 (38th of 126 ordered G1 cells) with a mean r.m.s.d. at the peak of the red curves. f, Cross-validation test. Red curve: average trans-chromosomal 3D distance distribution of the five most strongly contacting chromosome pairs, measured at 200 randomly chosen loci, using all supported contacts. Blue curve: same as the red curve, except after removal of the trans-contacts between one of the five chromosome pairs. Green curve: trans-chromosomal distance distribution control from models of all cells. Cyan curve: trans-distance distribution of loci involved in unsupported contacts. b0 is the bond length parameter. g, Nucleus and chromosome shape properties in the 126 cells with high-quality models: the nuclear radius (in arbitrary units), the longest-to-shortest (a/c) semi-axis ratio of the ellipsoid fitted to the whole nucleus model, and the longest-to-shortest (a/c) and middle-to-shortest (b/c) semi-axis ratios of the ellipsoids fitted to each chromosome of the 3D models. For the nucleus size and a/c ratio, values in each model are shown, for the chromosome fitted ellipsoid semi-axis ratios the model-averaged value is shown for each cell. h, The 7 cell groups in the inferred G1 time order for the 126 cells with the highest-quality models. i, Distances of the top third shortest distances between nucleolar organizer region (NOR) chromosomes or non-NOR chromosomes in 7 cell groups in h. Distances are normalized by the nucleus diameter.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Tables and additional references. (PDF 1239 kb)
Rights and permissions
About this article
Cite this article
Nagano, T., Lubling, Y., Várnai, C. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017). https://doi.org/10.1038/nature23001
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23001
- Springer Nature Limited
This article is cited by
-
Deep neural network models for cell type prediction based on single-cell Hi-C data
BMC Genomics (2024)
-
Single-cell multiplex chromatin and RNA interactions in ageing human brain
Nature (2024)
-
scGHOST: identifying single-cell 3D genome subcompartments
Nature Methods (2024)
-
scHiCyclePred: a deep learning framework for predicting cell cycle phases from single-cell Hi-C data using multi-scale interaction information
Communications Biology (2024)
-
Computational methods for analysing multiscale 3D genome organization
Nature Reviews Genetics (2024)