Abstract
Cellular potassium import systems play a fundamental role in osmoregulation, pH homeostasis and membrane potential in all domains of life. In bacteria, the kdp operon encodes a four-subunit potassium pump that maintains intracellular homeostasis, cell shape and turgor under conditions in which potassium is limiting1. This membrane complex, called KdpFABC, has one channel-like subunit (KdpA) belonging to the superfamily of potassium transporters and another pump-like subunit (KdpB) belonging to the superfamily of P-type ATPases. Although there is considerable structural and functional information about members of both superfamilies, the mechanism by which uphill potassium transport through KdpA is coupled with ATP hydrolysis by KdpB remains poorly understood. Here we report the 2.9 Å X-ray structure of the complete Escherichia coli KdpFABC complex with a potassium ion within the selectivity filter of KdpA and a water molecule at a canonical cation site in the transmembrane domain of KdpB. The structure also reveals two structural elements that appear to mediate the coupling between these two subunits. Specifically, a protein-embedded tunnel runs between these potassium and water sites and a helix controlling the cytoplasmic gate of KdpA is linked to the phosphorylation domain of KdpB. On the basis of these observations, we propose a mechanism that repurposes protein channel architecture for active transport across biomembranes.
Similar content being viewed by others
Main
Bacteria use multiple systems to maintain K+ homeostasis2. TrkH and KtrB are gated channels that belong to the superfamily of K+ transporters (SKT)3, which supply K+ to the cell under normal growth conditions. When K+ concentrations fall into the micromolar range, many bacteria use the inducible Kdp system, which produces the four-subunit KdpFABC membrane complex that actively drives K+ into the cell. This complex has high selectivity, binding affinity in the low micromolar range and can maintain cytoplasmic K+ concentrations against up to 104-fold gradients4. Mutagenesis has been used to establish that K+ is transported through KdpA5,6 and that the energy of ATP is harnessed by KdpB7. These subunits are joined by KdpC, which has been proposed to be a catalytic chaperone8, and KdpF9; both have single transmembrane helices and no known homologues outside Kdp.
As a P-type ATPase, KdpB operates according to the Post–Albers scheme, which involves two main conformational states: E1 and E2 (ref. 10). In the E1 state, ATP is bound by the cytoplasmic domains in order to autophosphorylate a conserved aspartate, thus stepping to E1P; this high-energy phosphoenzyme is typically formed in response to cytoplasmic ions binding at a canonical transmembrane site. The energy is used in converting E1P to E2P, where ion binding sites are exposed to the other side of the membrane with lowered affinity. After ions leave, the aspartyl phosphate is hydrolysed to produce E2, which then reverts back to E1 to complete the cycle. As K+ is bound by a different subunit in KdpFABC, it is unclear whether E1P formation in KdpB is associated with ion binding from the periplasm or with release to the cytoplasm by KdpA. Furthermore, it is unclear whether counterions, which generally facilitate E2 formation in other P-type ATPases, are involved in this process.
K+ is expected to move through KdpA, like all members of the SKT family, by way of a selectivity filter descended from that of the bacterial channel KcsA3. The selectivity filter has multiple, tandem binding sites for dehydrated K+ ions that are derived from four repeated M1PM2 motifs, in which two transmembrane helices (M) sandwich a reentrant pore helix (P). Whereas the KcsA channel is a homotetramer, TrkH, KtrB and KdpA are all single polypeptides with four pseudo repeats (D1–D4). Structures of TrkH and KtrB11,12 show a kinked helix in the third repeat (D3M2) with a loop that forms a regulatory gate on the cytoplasmic side of the selectivity filter13.
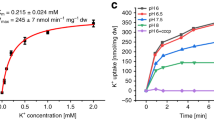
For this work, we used the KdpFABC complex from E. coli carrying the Gln116Arg mutation in KdpA. This mutant exhibits lowered apparent K+ affinity (Michaelis constant (Km) = 6 mM versus 10 μM for wild-type)5 and has been widely used in previous biochemical studies4,7,14. The structure was solved by X-ray crystallography to 2.9 Å resolution using experimental phases from tungsten and mercury with an Rfree of 27.5% (Extended Data Table 1). The large asymmetric unit contains three KdpFABC complexes that adopt identical conformations (Extended Data Fig. 1). KdpA has ten transmembrane helices with four M1PM2 repeats (D1–D4) and a K+ ion bound in the central selectivity filter (Fig. 1). KdpB has seven transmembrane helices (bM1–bM7) and three cytoplasmic domains found in all P-type ATPases: phosphorylation (P) domain, nucleotide-binding (N) domain, and dephosphorylating actuator (A) domain15. KdpC has a single transmembrane helix (cM1), but the topology is inverted relative to previous models (Extended Data Fig. 1d). This topology puts the soluble domain, which appears to have a novel fold, on the periplasmic side of the membrane. KdpF is a single transmembrane helix with a position that is distinct from those of transmembrane helices or accessory elements in other P-type ATPases. KdpF is not present in some species16 and the E. coli complex lacking KdpF is fully functional in the membrane-bound state or after addition of lipids9, suggesting that it plays a structural role in stabilizing the complex.
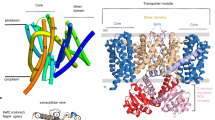
a, ATPase activity using purified, detergent-solubilized KdpFABC demonstrates robust stimulation by K+ with an apparent affinity of 24.8 mM and Vmax of 6.6 μmol mg−1 min−1. There is negligible activity with other ions. Error bars indicate s.e., n = 3. b, KdpA (green) has a K+ ion bound in the middle of the membrane (purple sphere). KdpB has a transmembrane domain (brown) and three cytoplasmic domains responsible for nucleotide binding (N, red), phosphoenzyme formation (P, blue) and dephosphorylation (A, yellow). KdpC (purple) and KdpF (cyan) have a single transmembrane helix each and the soluble domain of KdpC is periplasmic. c, Transmembrane topology diagram, coloured as in b.
KdpA displays pseudo-four-fold symmetry with M1PM2 motifs surrounding a central selectivity filter (Fig. 2a). A large number of mutations that affect apparent K+ affinity5,6 map directly onto the selectivity filter or the associated pore helices (Extended Data Fig. 2). The presence of a K+ ion within the selectivity filter is evidenced by a strong density peak that colocalizes with an anomalous density peak (Extended Data Fig. 1e). In K+ channels, four distinct sites (designated S1–S4) are characterized by ‘rings’ of oxygen ligands above and below each site17 (Extended Data Fig. 3). In KdpA, most of these oxygens contribute to the selectivity filter and the observed K+ ion occupies site S3 (Fig. 2b). Sites S2 and S4 lack ions and are distorted, whereas the S1 site is occupied by the charged side chain of mutated Arg116, which is hydrogen bonded to selectivity filter carbonyls (Gly232, Gly345, Gly468) and to Asn239, all of which produce lowered K+ affinity when mutated5,6. The position of the Arg116 side chain suggests that it is a surrogate for a K+ ion in the S1 site. Thus, this side chain is likely to interfere with K+ ions entering the selectivity filter in the Gln116Arg mutant, whereas this site would assist in recruiting K+ to the selectivity filter of the native protein and could thus contribute to its high apparent affinity.
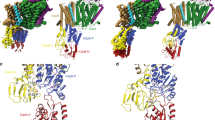
a, KdpA, as seen from the periplasmic side, has four M1PM2 units (D1–D4) that form the selectivity filter (yellow backbone) and bind K+ (purple sphere). KdpB (brown) includes a water molecule (red sphere) at its canonical cation binding site. b, Side view of the KdpA selectivity filter shows Arg116 in the S1 site and main chain carbonyls coordinating K+ in the S3 site. c, Side view shows the gating loop below the selectivity filter with the coupling helix (pink). On the periplasmic side, the soluble domain of KdpC (purple) is firmly held by loops from repeats D2 (dark green) and D3 (light green) of KdpA (Extended Data Fig. 2).
Unlike permeation through channels, the reaction cycle of an ATP-driven transporter involves transient occlusion of ions in order to prevent the leakage of ions and futile ATP hydrolysis10. Such occlusion requires gating elements on both sides of the binding site. Towards the cytoplasm, the selectivity filter of KdpA is blocked by a loop directly below the K+ ion (Fig. 2c). As in TrkH and KtrB, this loop is derived from D3M2, which forms a kinked helix (Fig. 1c, Extended Data Fig. 3). On the periplasmic side, access to the filter is unimpeded, except by Arg116, which is not present in the native sequence. Nevertheless, the soluble domain of KdpC is held nearby by two loops from the D2 and D3 repeats of KdpA (Fig. 2c). The robustness of the hydrogen bond network that mediates this interaction may explain why KdpA and KdpC can be co-purified in the absence of KdpB18. Furthermore, this interaction suggests that relative movements between the repeats of KdpA could move the soluble domain of KdpC into an occluding position in response to the relevant conformational change in KdpB.
KdpB is characterized by seven transmembrane helices, the first six of which are consistent with the ‘core’ of other P-type ATPases19. The middle of bM4 is unwound at the conserved proline motif (IP264TTI) and is very similar to M4 of Ca2+-ATPase (SERCA1a) in the Ca2+-bound E1 state10 (Extended Data Fig. 4). We observe a strong density peak at the unwound part of bM4 (Extended Data Fig. 1f). Neither K+ nor Na+ fit the binding geometry and both introduce positive charge adjacent to Lys586. There is no anomalous signal, and refinement with K+ led to an anomalously high temperature factor for this atom. Therefore, this peak was assigned as water on the basis of the presence of four binding ligands within 2.4–2.5 Å, although the planar geometry of these bonds indicates a strained environment. Given the canonical nature of this cation site in most P-type ATPases, this water could serve as a substrate mimic to assist communication between the cytosolic and transmembrane domains of KdpB. Although the cytosolic domains of KdpB adopt a unique configuration due to the unexpected presence of a phosphoserine in the A domain (Fig. 1b, Extended Data Fig. 5), the general decoupling of the A domain, the juxtaposition of the N and P domains, the lack of phosphorylation at Asp307, and the unwound configuration of bM4 are all consistent with an E1 enzymatic state.
To evaluate the accessibility of the bound water molecule in KdpB, we searched for exit pores and found that the site is blocked from both the cytosol and the periplasm. However, this analysis revealed a tunnel that connects the water site in KdpB with the K+ ion binding site in KdpA (Fig. 3). This tunnel runs for 40–45 Å parallel to the membrane surface and to the coupling helix of D3M2 and is completely encased within the membrane domain of the complex. The KdpA part of the tunnel is reminiscent of fenestrations found in other channels20,21. Unlike these fenestrations, however, the tunnel stays within the membrane and connects functional sites in the two subunits. Whereas the middle is hydrophobic or neutral, the two ends are distinctly electronegative owing to Glu370 and carbonyl oxygens in the gating loop of KdpA, and to Asp583 and side chain oxygens of Ser579 and Thr266 in KdpB. Although no density is visible within the tunnel, its diameter would accommodate water molecules and, like cavities identified in other membrane proteins, it is likely to be filled with water22.
a, A tunnel (>1.4 Å radius, blue surface) connects the water molecule in KdpB to the K+ ion in KdpA. The coupling helix (pink) runs adjacent to the tunnel on the cytosolic side of the membrane. b, Salt bridges attach the coupling helix to the P domain of KdpB (blue). c, Close-up from the periplasmic side shows charged residues at either end of the tunnel: Glu370 and Arg493 in KdpA and Asp583 and Lys586 in KdpB. The tunnel could facilitate charge transfer between these two sites. d, In KdpB, an unwound portion of transmembrane helix bM4 (shown in stick representation) forms a binding site similar to those of other P-type ATPases. The modelled water is coordinated by main chain carbonyls from bM4 and side chain oxygens from Thr265 and Asn624.
Coupling between KdpA and KdpB requires two-way communication. On the one hand, the presence or absence of K+ in KdpA should initiate autophosphorylation of Asp307 in KdpB and, on the other hand, conformational change in KdpB should control gating of ion sites in KdpA. The latter step is likely to involve the kinked helix from the distal part of the D3M2 element of KdpA, which forms a strong interaction with the P domain of KdpB involving a salt bridge between Arg400 from KdpA and Asp300 and Asp302 from KdpB (Fig. 3b). In TrkH and KtrB, this same ‘coupling helix’ interacts with regulatory cytoplasmic subunits that undergo conformational changes in response to ATP binding12,23. In P-type ATPases, the E1P-to-E2P transition is associated with a marked inclination of the P domain24, which in KdpFABC would pull on this coupling helix and open the cytosolic gate. Indeed, this helix is relatively free to move, in part owing to the tunnel that is located directly above it, which ensures minimal interactions with other parts of KdpA. This idea is supported by mutation of Asp300 in KdpB, which has been shown to increase ATPase activity and decrease K+ affinity, as would be expected from an uncoupled complex25. To initiate autophosphorylation (E1-to-E1P transition), P-type ATPases rely on binding of cations to their canonical transmembrane site. The water-filled intramembrane tunnel provides a possible proton wire for introducing charge into that site in response to K+ binding to the selectivity filter of KdpA. The tunnel begins near Asp583 and Lys586 in KdpB, and passes Glu370 before ending next to Arg493 in KdpA (Fig. 3c, d). The functional importance of the tunnel is supported by mutation of Asp583, which uncouples K+ transport and ATP hydrolysis26,27, and mutation of Arg493, which disrupts KdpFABC function27. All of these residues are conserved, except for Glu370, which is a Gln or an Asn in about half of bacterial species (Extended Data Figs 6, 7, 8, 9).
Our results suggest the following model for transport by KdpFABC (Fig. 4). With KdpB in the E1 conformation, as observed in the structure, K+ ions enter the selectivity filter of KdpA from the periplasmic side of the membrane. Selectivity filters have innate binding affinity for K+ ions at the micromolar level28 and the relatively slow rate of transport by KdpFABC would allow equilibration of these sites and explain the selectivity for K+ over Na+. Permeation of ions into the S4 site would induce a proton charge transfer through the tunnel to the structural water molecule bound to the canonical ion binding site in KdpB. This mimic of cation binding to a P-type ATPase would induce a conformational change in KdpB that would lead to phosphorylation of Asp307. The consequent conformational changes associated with E1P formation in KdpB would lead to outward occlusion of the selectivity filter in KdpA, perhaps by movement of the periplasmic KdpC domain. As in SERCA1a, the transition of KdpB to E2P would induce an inclination of the P domain away from KdpA24,29, which would pull the coupling helix towards KdpB and open the gating loop to allow K+ to escape to the cytoplasm. This could be associated with a reverse charge movement from KdpB to KdpA through the tunnel due to expected changes in the conformation of bM429. A removal of the transmembrane charge in KdpB would then trigger hydrolysis of E2P, returning the system to the E1 state24,30.
According to the Post–Albers scheme, transitions between open and closed states (black boxes) are driven by phosphorylation and dephosphorylation events in the cytosolic domains of KdpB. The cycle is initiated by K+ binding to the E1 state from the periplasm (1). The presence of K+ in the S4 site of the selectivity filter of KdpA leads to charge transfer through the tunnel to the transmembrane domain of KdpB (2). The presence of charge at the canonical site in KdpB triggers phosphorylation through a conserved P-type ATPase mechanism (3). K+ occlusion, which may involve the periplasmic domain of KdpC, leads to the occluded E1P state (4). The transition to the E2P state in P-type ATPases involves inclination of the P domain away from KdpA, which will pull the D3 coupling helix (pink) of KdpA (5). This movement opens the cytoplasmic gate, thereby allowing K+ release to the cytosol (6). The models are derived from our structure of an inhibited E1 state and SERCA1a structures of E1P (1T5T) and E2P (3B9B) states.
Ion transport by channels and by transporters such as P-type ATPases are fundamentally different processes and KdpFABC is, so far, unique in pairing these two disparate functionalities to enable active transport. Although the structure provides the first hints as to how this is accomplished, there are many questions to be answered. This structure provides a template with which we can design experiments to address these questions for a better understanding of this system and of transport mechanisms in general.
Methods
Sample expression
The kdp operon encoding the four subunit KdpFABC complex with a single mutation (Q116R) in the KdpA subunit and an 8 × histidine tag at the C terminus of the KdpC subunit was cloned into plasmid pSD107, which was originally derived by inserting the KdpFABC operon with a C-terminal histidine tag into the pBR322 plasmid, thus providing ampicillin resistance6. This plasmid was transformed into the E. coli KdpFABC knockout strain TK2498, with genotype: F− thi lacZ nagA rha trkA405 trkD1 Δ(KdpFAB)5 Δ(ompT). Both the plasmid and the E. coli strain were obtained from W. Epstein (University of Chicago). Expression was controlled by the native kdp promoter9,31, with KdpD and KdpE being present in the chromosome. Expression was induced by a potassium-free K0-medium (46 mM Na2PO4, 23 mM NaH2PO4, 25 mM (NH4)2SO4, 0.4 mM MgSO4, 6 μM FeSO4, 1 mM sodium citrate, 0.2% glucose, 1 μg ml−1 thiamine, 50 μg ml−1 carbenicillin) supplemented with different amounts of KCl32,33. To start, cells from a glycerol stock were incubated overnight at 37 °C in 10 ml K5-medium (K0-medium supplemented with 5 mM KCl). This culture was transferred to 500 ml K1-medium (K0-medium supplemented with 1 mM KCl) and incubated at 37 °C for 8 h. The 500 ml cell culture was transferred again to 18 l K0.2-medium (K0-medium supplemented with 0.2 mM KCl) and incubated at 31 °C to induce expression of KdpFABC. Cells were harvested when the culture density reached OD600 ~1. For production of protein with selenomethionine (SeMet) substitution, we used high concentrations of isoleucine, leucine, phenylalanine, lysine, and threonine to inhibit the methionine biosynthesis pathway in E. coli34. Specifically, 100 mg/l each of lysine, phenylalanine, and threonine, 50 mg/l each of isoleucine, leucine, and valine, and 60 mg ml−1 SeMet were added to the K0.2 culture medium as the protein expression was induced.
Sample purification
The harvested cells were resuspended in 50 mM Tris pH 7.5, 1.2 M NaCl, 10 mM MgCl2, 10% glycerol, protease inhibitor tablets (Roche), 25 mg ml−1 DNase, and 1 mM DTT, and lysed using an Emulsiflex C3 high-pressure homogenizer (Avestin). After centrifugation at 10,600g for 15 min to remove unbroken cells and debris, the supernatant was centrifuged at 90,140g for 2 h to pellet cell membranes. This membrane fraction was solubilized in 50 mM Tris pH 7.5, 600 mM NaCl, 10 mM MgCl2, 10% glycerol, 1 mM DTT and 1.2% n-decyl-β-maltoside (DM) at 4 °C for at least 2 h, and then centrifuged at 90,140g for 30 min to remove insoluble components. The supernatant containing solubilized protein was loaded onto a 5 ml Ni+-charged HiTrap chelating column (GE Healthcare), which had been pre-equilibrated with buffer A (50 mM Tris pH 7.5, 600 mM NaCl, 10 mM MgCl2, 10% glycerol, 0.15% DM, 20 mM imidazole). A linear gradient of imidazole (20–500 mM based on buffer A) was used to elute the KdpFABC complex using a Biologic LP chromatography system (Bio-Rad). The eluted fractions were pooled together and concentrated using 100 kDa cut-off concentrators to ~10 mg ml−1. This sample was further purified by applying 0.5 ml onto a preparative size exclusion column Superdex 200 (GE Healthcare) equilibrated with OG-DMPC buffer composed of 25 mM Tris pH 7.5, 100 mM KCl, 10% glycerol, 1.1% n-octyl-β-d-glucoside (OG), and 0.5 mg ml−1 1,2-dimyristoyl-sn-glycero-3-phosphocholine (DMPC). The eluted fractions containing KdpFABC complex were combined, concentrated to 15–20 mg ml−1 and stored at −80 °C. An identical protocol was used for purification of SeMet-substituted KdpFABC complex.
ATPase assay and dephosphorylation
A coupled enzyme assay was used to measure ATPase activity35. The assay solution contained 50 mM Tris pH 7.5, 5 mM MgCl2, 0.15% DM, 2.4 mM ATP, 0.18 mM NADH, 0.5 mM phosphoenol pyruvate, 4.8 units of pyruvate kinase, and 4.8 units of lactate dehydrogenase in 10 ml. This assay couples the hydrolysis of ATP with the oxidation of NADH, which was followed in real time using the absorbance at 340 nm. For each reaction, 10 mg KdpFABC complex and various concentrations of cations were mixed with 0.5 ml assay buffer at 25 °C and specific activity (μmol Pi min−1 mg−1) was calculated as (OD340 per min) × (1/6.22) × (1/mg protein) × 0.5 ml. To study the effect of Ser162 phosphorylation, a given preparation was incubated in the presence and absence of lambda phosphatase at 30 °C for either 4 or 16 h and then assayed for ATPase activity in the presence of 50 mM KCl and for the level of phosphorylation using mass spectrometry.
Mass spectrometry
The presence of phosphoserine in the sample was confirmed using electrospray ionization with liquid chromatography and tandem mass spectrometry (ESI–LC–MS/MS). KdpB was isolated by SDS–PAGE from various different samples and digested with trypsin within the gel. After elution, the peptides were fractionated using high-performance liquid chromatography integrated with tandem mass spectrometers. The peptide containing the non-phosphorylated or phosphorylated Ser162 was identified by MS1, the sequence of which was further confirmed by MS2. The relative abundance of the non-phosphorylated and phosphorylated Ser162 of KdpB was calculated from the peak heights of the two from MS1.
Crystallization
Purified KdpFABC complex was adjusted to 8 mg ml−1 using OG-DMPC buffer and supplemented with 5 mM Mg-AMPPCP for crystallization. Mg-AMPPCP was required for crystallization, despite not being visible in the crystal structure. We speculate that the Mg-AMPPCP may trap the KdpFABC complexes that did not carry the Ser162 phosphorylation in an unfavourable conformation, thus excluding them from the crystal lattice. Native and SeMet-derived crystals were grown at 18 °C using the hanging-drop vapour diffusion method: 2 μl protein solution was mixed with 2 μl reservoir solution containing 20% (w/v) PEG3350, 0.5–1.0 M NaCl, 0.05 M sodium citrate pH 5.5. Crystals appeared within a week and grew to full size (100 × 300 × 40 μm3) after three weeks. Drops containing suitable crystals were dehydrated against a reservoir containing 36% glycerol for 1 day36 and flash-frozen in liquid nitrogen. To obtain heavy atom derivatives, crystals were soaked with osmium, platinum and tantalum salts; despite strong anomalous signal, however, these derivatives could not be used to solve the structure. Tungsten derivatives were obtained by soaking native crystals in 1 mM Na6(H2W12O40) for 1 day. The mercury derivative was obtained by soaking native crystals in 5 mM Hg(OOCCH3)2 for 2–3 days. Data used for structure solution were collected at the Advanced Photon Source beamline 23-ID-B and 23-ID-D. Additional screening and preliminary data collection was done at the National Synchrotron Light Source X4A and X4C and at Stanford Synchrotron Radiation Lightsource 14-1.
Data processing
Datasets were processed and scaled using XDS37 in space group P21, which suggested the presence of three KdpFABC complexes in the asymmetric unit (~70% solvent content). The positions of tungsten clusters were determined by single-wavelength anomalous dispersion (SAD) in SHELXC/D38 and initial phases were calculated in SHARP39. After solvent flattening with Solomon40 or DM40, the electron density map based on data from the tungsten cluster derivative at 4.0 Å resolution enabled us to identify KdpA and KdpB subunits and the expected 19 transmembrane helices, but the quality of this map was not sufficient to establish the orientation of the pseudo-four-fold symmetric KdpA or to place the soluble domains of KdpC unambiguously. Independent phase information to higher resolution (3.3 Å) was then obtained from the mercury derivatives. Owing to excessive non-isomorphism, no native or derivative datasets could be combined for phase calculation. However, mercury sites could be identified by anomalous difference Fourier maps using the phases from the tungsten cluster derivative. Mercury derivative SAD phases were calculated in SHARP, and were refined and extended using DM to 3.3 Å resolution, exploiting histogram matching, solvent flattening and three-fold non-crystallographic symmetry averaging. The resulting electron density map was of high quality, allowing for a continuous trace of the main chain. The model was built using COOT41 with KtrB12 and Cu+-ATPase42 as guides for building initial models of KdpA and KdpB, respectively. Objective aides in building this model came from 44 selenomethiones and 7 mercury atoms bound to cysteines in each KdpFABC complex, which were identified in anomalous difference Fourier maps from the SeMet and mercury derivatives, respectively (Extended Data Fig. 1b). The anomalous peak from K+ was identified from the native 2.9 Å dataset. It was not possible to use Rb+ as a K+ congener as the mutation Q116K not only lowers K+ affinity but also inhibits Rb+ coupling43 (Fig. 1a). Iterative model building in COOT and refinement using phenix.refine44 gradually improved the model and the fit to the experimental map. At later stages, the model was of sufficient quality to be used for molecular replacement into the native dataset (2.9 Å) using the program PHASER45; afterwards, model building was guided by 2mFo−DFc maps using model phases. Final refinement in phenix.refine exploited three-fold non-crystallographic symmetry with a refinement strategy of individual sites, individual ADP, and group TLS (18 groups), against a maximum likelihood (ML) target with reflections in the 20–2.9 Å range. The final model yielded an Rwork of 24.3% and an Rfree of 27.5% (Extended Data Table 1). MolProbity46 evaluation of the Ramachandran plot gave 95.9% in favoured regions and 0.1% outliers. The KdpA–KdpB tunnel was identified with CAVER47 using default settings and a probe radius of 1.4 Å, which is equivalent to the radius of water. All structural figures were prepared using PyMOL (The PyMOL Molecular Graphics System, Version 1.5.0.4 (Schrödinger LLC, 2012)).
Sequence alignment
Sequence conservation of KdpA and KdpB were initially evaluated by applying Clustal Omega alignment48 to a list of genes obtained from the Divblast server49. The Divblast server ensures that the list represents the diversity across all KdpFABC homologues, which are otherwise dominated by a large number of E. coli strains. Alignment of KdpA and KdpB with distantly related members of the SKT and P-Type ATPase families was done with the Promals3d server50. The ability of Promals3d to consider structural features as well as sequence produced a reasonably accurate alignment that nevertheless required minor manual adjustments based on comparison of tertiary structures of the respective atomic models.
Data availability
Atomic coordinates and structure factors have been deposited in the Protein Data Bank (PDB) with accession code 5MRW. All other data are available from the corresponding authors upon reasonable request.
Accession codes
References
Greie, J. C. The KdpFABC complex from Escherichia coli: a chimeric K+ transporter merging ion pumps with ion channels. Eur. J. Cell Biol. 90, 705–710 (2011)
Epstein, W. The roles and regulation of potassium in bacteria. Prog. Nucleic Acid Res. Mol. Biol. 75, 293–320 (2003)
Diskowski, M., Mikusevic, V., Stock, C. & Hänelt, I. Functional diversity of the superfamily of K+ transporters to meet various requirements. Biol. Chem. 396, 1003–1014 (2015)
Epstein, W., Whitelaw, V. & Hesse, J. A K+ transport ATPase in Escherichia coli. J. Biol. Chem. 253, 6666–6668 (1978)
Buurman, E. T., Kim, K. T. & Epstein, W. Genetic evidence for two sequentially occupied K+ binding sites in the Kdp transport ATPase. J. Biol. Chem. 270, 6678–6685 (1995)
Dorus, S., Mimura, H. & Epstein, W. Substrate-binding clusters of the K+-transporting Kdp ATPase of Escherichia coli investigated by amber suppression scanning mutagenesis. J. Biol. Chem. 276, 9590–9598 (2001)
Siebers, A. & Altendorf, K. Characterization of the phosphorylated intermediate of the K+-translocating Kdp-ATPase from Escherichia coli. J. Biol. Chem. 264, 5831–5838 (1989)
Irzik, K. et al. The KdpC subunit of the Escherichia coli K+-transporting KdpB P-type ATPase acts as a catalytic chaperone. FEBS J. 278, 3041–3053 (2011)
Gassel, M., Möllenkamp, T., Puppe, W. & Altendorf, K. The KdpF subunit is part of the K+-translocating Kdp complex of Escherichia coli and is responsible for stabilization of the complex in vitro. J. Biol. Chem. 274, 37901–37907 (1999)
Møller, J. V., Olesen, C., Winther, A. M. & Nissen, P. The sarcoplasmic Ca2+-ATPase: design of a perfect chemi-osmotic pump. Q. Rev. Biophys. 43, 501–566 (2010)
Cao, Y. et al. Crystal structure of a potassium ion transporter, TrkH. Nature 471, 336–340 (2011)
Vieira-Pires, R. S., Szollosi, A. & Morais-Cabral, J. H. The structure of the KtrAB potassium transporter. Nature 496, 323–328 (2013)
Hänelt, I. et al. Gain of function mutations in membrane region M2C2 of KtrB open a gate controlling K+ transport by the KtrAB system from Vibrio alginolyticus. J. Biol. Chem. 285, 10318–10327 (2010)
Fendler, K., Dröse, S., Epstein, W., Bamberg, E. & Altendorf, K. The Kdp-ATPase of Escherichia coli mediates an ATP-dependent, K+-independent electrogenic partial reaction. Biochemistry 38, 1850–1856 (1999)
Toyoshima, C., Nakasako, M., Nomura, H. & Ogawa, H. Crystal structure of the calcium pump of sarcoplasmic reticulum at 2.6 Å resolution. Nature 405, 647–655 (2000)
Bramkamp, M., Altendorf, K. & Greie, J. C. Common patterns and unique features of P-type ATPases: a comparative view on the KdpFABC complex from Escherichia coli (Review). Mol. Membr. Biol. 24, 375–386 (2007)
Zhou, Y., Morais-Cabral, J. H., Kaufman, A. & MacKinnon, R. Chemistry of ion coordination and hydration revealed by a K+ channel-Fab complex at 2.0 Å resolution. Nature 414, 43–48 (2001)
Gassel, M., Siebers, A., Epstein, W. & Altendorf, K. Assembly of the Kdp complex, the multi-subunit K+-transport ATPase of Escherichia coli. Biochim. Biophys. Acta 1415, 77–84 (1998)
Sitsel, O. et al. Structure and function of Cu(I)- and Zn(II)-ATPases. Biochemistry 54, 5673–5683 (2015)
Miller, A. N. & Long, S. B. Crystal structure of the human two-pore domain potassium channel K2P1. Science 335, 432–436 (2012)
Payandeh, J., Gamal El-Din, T. M., Scheuer, T., Zheng, N. & Catterall, W. A. Crystal structure of a voltage-gated sodium channel in two potentially inactivated states. Nature 486, 135–139 (2012)
Rasaiah, J. C., Garde, S. & Hummer, G. Water in nonpolar confinement: from nanotubes to proteins and beyond. Annu. Rev. Phys. Chem. 59, 713–740 (2008)
Cao, Y. et al. Gating of the TrkH ion channel by its associated RCK protein TrkA. Nature 496, 317–322 (2013)
Olesen, C., Sørensen, T. L., Nielsen, R. C., Møller, J. V. & Nissen, P. Dephosphorylation of the calcium pump coupled to counterion occlusion. Science 306, 2251–2255 (2004)
Puppe, W., Siebers, A. & Altendorf, K. The phosphorylation site of the Kdp-ATPase of Escherichia coli: site-directed mutagenesis of the aspartic acid residues 300 and 307 of the KdpB subunit. Mol. Microbiol. 6, 3511–3520 (1992)
Bramkamp, M. & Altendorf, K. Single amino acid substitution in the putative transmembrane helix V in KdpB of the KdpFABC complex of Escherichia coli uncouples ATPase activity and ion transport. Biochemistry 44, 8260–8266 (2005)
Becker, D., Fendler, K., Altendorf, K. & Greie, J. C. The conserved dipole in transmembrane helix 5 of KdpB in the Escherichia coli KdpFABC P-type ATPase is crucial for coupling and the electrogenic K+-translocation step. Biochemistry 46, 13920–13928 (2007)
Liu, S. & Lockless, S. W. Equilibrium selectivity alone does not create K+-selective ion conduction in K+ channels. Nat. Commun. 4, 2746 (2013)
Toyoshima, C., Nomura, H. & Tsuda, T. Lumenal gating mechanism revealed in calcium pump crystal structures with phosphate analogues. Nature 432, 361–368 (2004)
Møller, J. V., Lenoir, G., Le Maire, M., Juul, B. S. & Champeil, P. Proteolytic studies on the transduction mechanism of sarcoplasmic reticulum Ca2+-ATPase: common features with other P-type ATPases. Ann. NY Acad. Sci. 986, 82–89 (2003)
Walderhaug, M. O. et al. KdpD and KdpE, proteins that control expression of the kdpABC operon, are members of the two-component sensor-effector class of regulators. J. Bacteriol. 174, 2152–2159 (1992)
Siebers, A. & Altendorf, K. The K+-translocating Kdp-ATPase from Escherichia coli. Purification, enzymatic properties and production of complex- and subunit-specific antisera. Eur. J. Biochem. 178, 131–140 (1988)
Laimins, L. A., Rhoads, D. B. & Epstein, W. Osmotic control of kdp operon expression in Escherichia coli. Proc. Natl Acad. Sci. USA 78, 464–468 (1981)
Doublié, S. Production of selenomethionyl proteins in prokaryotic and eukaryotic expression systems. Methods Mol. Biol. 363, 91–108 (2007)
Warren, G. B., Toon, P. A., Birdsall, N. J. M., Lee, A. G. & Metcalfe, J. C. Reversible lipid titrations of the activity of pure adenosine triphosphatase-lipid complexes. Biochemistry 13, 5501–5507 (1974)
Wheeler, M. J., Russi, S., Bowler, M. G. & Bowler, M. W. Measurement of the equilibrium relative humidity for common precipitant concentrations: facilitating controlled dehydration experiments. Acta Crystallogr. F Struct. Biol. Cryst. Commun. 68, 111–114 (2012)
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010)
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D Biol. Crystallogr. 66, 479–485 (2010)
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D Biol. Crystallogr. 59, 2023–2030 (2003)
Abrahams, J. P. & Leslie, A. G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D Biol. Crystallogr. 52, 30–42 (1996)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010)
Gourdon, P. et al. Crystal structure of a copper-transporting PIB-type ATPase. Nature 475, 59–64 (2011)
van der Laan, M., Gassel, M. & Altendorf, K. Characterization of amino acid substitutions in KdpA, the K+-binding and -translocating subunit of the KdpFABC complex of Escherichia coli. J. Bacteriol. 184, 5491–5494 (2002)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010)
Petrek, M. et al. CAVER: a new tool to explore routes from protein clefts, pockets and cavities. BMC Bioinformatics 7, 316 (2006)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011)
Eser, E., Can, T. & Ferhatosmanog˘lu, H. Div-BLAST: diversification of sequence search results. PLoS ONE 9, e115445 (2014)
Pei, J., Kim, B. H. & Grishin, N. V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008)
Hu, G. B., Rice, W. J., Dröse, S., Altendorf, K. & Stokes, D. L. Three-dimensional structure of the KdpFABC complex of Escherichia coli by electron tomography of two-dimensional crystals. J. Struct. Biol. 161, 411–418 (2008)
Heitkamp, T., Böttcher, B. & Greie, J. C. Solution structure of the KdpFABC P-type ATPase from Escherichia coli by electron microscopic single particle analysis. J. Struct. Biol. 166, 295–302 (2009)
Heitkamp, T. et al. K+-translocating KdpFABC P-type ATPase from Escherichia coli acts as a functional and structural dimer. Biochemistry 47, 3564–3575 (2008)
Schrader, M. et al. Replacement of glycine 232 by aspartic acid in the KdpA subunit broadens the ion specificity of the K(+)-translocating KdpFABC complex. Biophys. J. 79, 802–813 (2000)
Bertrand, J., Altendorf, K. & Bramkamp, M. Amino acid substitutions in putative selectivity filter regions III and IV in KdpA alter ion selectivity of the KdpFABC complex from Escherichia coli. J. Bacteriol. 186, 5519–5522 (2004)
Sørensen, T. L., Møller, J. V. & Nissen, P. Phosphoryl transfer and calcium ion occlusion in the calcium pump. Science 304, 1672–1675 (2004)
Jensen, A. M., Sørensen, T. L., Olesen, C., Møller, J. V. & Nissen, P. Modulatory and catalytic modes of ATP binding by the calcium pump. EMBO J. 25, 2305–2314 (2006)
Toyoshima, C. & Mizutani, T. Crystal structure of the calcium pump with a bound ATP analogue. Nature 430, 529–535 (2004)
Bond, C. S. & Schüttelkopf, A. W. ALINE: a WYSIWYG protein-sequence alignment editor for publication-quality alignments. Acta Crystallogr. D Biol. Crystallogr. 65, 510–512 (2009)
Pedersen, B. P., Buch-Pedersen, M. J., Morth, J. P., Palmgren, M. G. & Nissen, P. Crystal structure of the plasma membrane proton pump. Nature 450, 1111–1114 (2007)
Andersson, M. et al. Copper-transporting P-type ATPases use a unique ion-release pathway. Nat. Struct. Mol. Biol. 21, 43–48 (2014)
Laursen, M., Yatime, L., Nissen, P. & Fedosova, N. U. Crystal structure of the high-affinity Na+K+-ATPase–ouabain complex with Mg2+ bound in the cation binding site. Proc. Natl Acad. Sci. USA 110, 10958–10963 (2013)
Acknowledgements
The authors acknowledge beamline 23IDB/IDD at the Advanced Photon Source—Argonne National Laboratory, where all the X-ray data were collected, as well as the National Synchrotron Light Source and the Stanford Synchrotron Radiation Light Source. We also thank W. Epstein and D. Sauer for discussions, W. Epstein for the plasmid and bacterial strain for expression, and J. Deng and T. Neubert for mass spectrometry. This work was supported by funding from NIH R01 GM108043 and NIH T32 GM088118 to D.L.S., the European Research Council (grant agreement No. 637372), the Danish Council for Independent Research (grant agreement no. DFF-4002-00052) and an AIAS fellowship to B.P.P.
Author information
Authors and Affiliations
Contributions
C.-S.H. produced protein, grew crystals, collected data, ran crystallographic analysis programs to solve the structure, interpreted the structure and wrote the manuscript. B.P.P. devised data collection strategies, analysed crystallographic data, interpreted the structure and wrote the manuscript. D.L.S. collected data, analysed sequences, interpreted the structure and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Structure determination.
a, Crystals had plate-like morphology with a substantial level of birefringence. b, Anomalous density maps from Hg (red mesh at 5σ) and seleno-methionine (blue mesh at 3σ) were used to identify positions of cysteine and methionine residues, respectively. This image shows one of the three KdpFABC complexes in the asymmetric unit with the locations of seven Hg atoms and 44 Se atoms superimposed on the ribbon model of the KdpFABC complex (colours as in Fig. 1). Considering all three complexes in the asymmetric unit, we were able to identify a total of 21 Hg sites and 132 Se sites, which were a powerful constraint for building the model before refinement. c, Elution profiles from size-exclusion chromatography using multiple detectors to quantify the size of the complex. Traces are shown for multiple angle laser light scattering (LS), refractive index (RI) and absorption at 280 nm (UV). On the basis of these data, we determined the sizes of the protein-detergent complex (PDC), the KdpFABC complex and the detergent micelle. Given the expected Mr of 160 kD, these data are consistent with a monomeric KdpFABC complex. d, 2mFo−DFc map densities at 1.5σ are superimposed with the refined model at 2.9 Å resolution to demonstrate the quality of the fit. The green chain corresponds to the N terminus of KdpA (upper right corner) and helices from the second M1PM2 repeat. The purple chain corresponds to the transmembrane domain of KdpC (far left) and part of the periplasmic domain (top). A series of bulky aromatic residues towards the periplasmic side of the KdpC helix unambiguously establishes its orientation in the membrane. e, mFo−DFc difference density (purple at 6.5σ) was present in the S3 site of the KdpA selectivity filter. The anomalous signal from K (cyan at 3.8σ) is consistent with the presence of a K+ ion at this site. f, mFo−DFc difference density (purple at 4.5σ) next to the unwound portion of M4 in KdpB was consistent with the presence of a water molecule at this canonical ion binding site in P-type ATPases. No anomalous signal from K was seen at this site. g, Three KdpFABC complexes compose the asymmetric unit and are coloured in shades of red, green and blue. Intermolecular contacts can be seen between the periplasmic domain of KdpC and the cytoplasmic face of KdpA. At the current resolution, the three complexes have identical structures. Although previous results suggest that KdpFABC forms a dimer51,52,53, we did not observe dimer formation either during purification or within our crystal lattice. This result suggests that dimerization may depend on specific conditions, such as the choice of detergent or the presence of a lipid bilayer. h, The three complexes have been coloured according to the temperature factor using a standard colour palette moving from blue (lowest B factor) to red (highest B factor). KdpA appears to be the best ordered and peripheral regions of the N- and A-domains display the most flexibility. i, In addition to the four protein subunits, two detergent molecules (OG) and two lipid molecules (DMPC) were modelled into densities at the periphery of the structure for each complex.
Extended Data Figure 2 Mutations that lower the apparent K+ affinity of KdpFABC.
a, b, Two orthogonal views of KdpA showing the locations of mutations within the selectivity filter that lower the apparent affinity of the pump for K+. The sites of the mutants are indicated by a red sphere at the Cα position; the selectivity filter is coloured yellow and the coupling helix (D3M2) is coloured pink. Although Lys substitutions generally dominate the reported mutations5,6,43,54,55, the selectivity filter has a range of other substitutions that are additionally shown to have an effect. c, d, Additional sites of K+ affinity mutants populate the pore helix in MPM2–4, which presumably support the architecture of the selectivity filter. These are predominantly Lys substitutions, suggesting that the addition of positive charge in this vicinity also plays a role in lowering the affinity5,6.
Extended Data Figure 3 Comparison of selectivity filters from a K+ channel and SKT transporters.
a, KcsA is an archetypal K+ channel and the 2.0 Å resolution structure (1K4C)17 shows four K+ sites within the selectivity filter at sites designated S1–S4. These sites are presumed to be alternatively occupied by K+ and water molecules, but differences have been masked by crystallographic averaging. Additional, partially hydrated K+ ions are seen in cytoplasmic and periplasmic vestibules at either end of the filter. b, TrkH is more closely related to KdpA and displays the gating loop at the cytoplasmic end of the selectivity filter (pink). The crystal structure (3PJZ)11 shows a single K+ ion in the S3 position and the filter is considerably more disordered towards its periplasmic end, perhaps accounting for the lack of further ions. c, KdpA has intermediate order in the selectivity filter. A K+ ion occupies the S3 site and the side chain of Arg116, which was substituted for Gln in our construct, appears to occupy the S1 site. Like TrkH, a gating loop is evident from the kinked helix (pink) in the third M1PM2 repeat. d, The cage of oxygen atoms that coordinate the K+ ions in the selectivity filter of KcsA. The strict fourfold symmetry of this channel produces a very regular geometry. e, The analogous oxygen atoms in the TrkH structure show considerably more distortion of this cage and complete absence of coordination at the S1 sites. f, The coordination cage for KdpA has intermediate order with reasonable evidence for sites at S2–S4.
Extended Data Figure 4 Comparison of KdpB and SERCA1a.
a, In the P domain of KdpB (blue), there is no evidence for phosphorylation of the catalytic Asp307, which is in a binding network with Lys499, Asp522 and Thr309. The location of the nucleotide binding pocket is illustrated by some of the conserved residues (Glu348, Phe377 and Thr429) expected to interact with ATP in the N domain (red). b, Overlay of the N domains of KdpB (red) and SERCA1a (cyan, 1T5T)56 with bound nucleotide (ADP-AlF4). This overlay illustrates how, if the nucleotide were bound to the corresponding site in KdpB, the phosphate groups would clash with the conserved catalytic loop (DKTGTLT in green). The TGES162 loop of KdpB is coloured pink. Taken together, the Asp307 binding network and the positional decoupling of the A domain from the rest of the structure indicate that KdpB is in an E1 state in which Asp307 remains unphosphorylated and the N domain nucleotide binding site is empty15,56,57,58. c, The binding site for Ca(II) (green sphere)15 in the transmembrane domain of SERCA1a (1T5T) consists of main chain carbonyls from the unwound M4 helix near the canonical Pro308 as well as side chain densities from the neighbouring M6 helix (Asn796 and Asp800). The second Ca2+ ion in the Ca(I) site is not shown, but involves additional ligands from the M5 helix. d, The water site (red sphere) in the transmembrane domain of KdpB involves residues homologous to SERCA1a in a very similar configuration: namely, the main chain carbonyls on M4 near Pro264 and the side chain of Asn624. The side chains of Asp583 and Lys586 from M5 are also nearby, which align with Asn768 and Glu771 in SERCA1a that bind Ca(I).
Extended Data Figure 5 Serine phosphorylation of KdpFABC.
a, Evidence for the phosphoserine comes from an extra density seen both in the 2mFo−DFc map (grey at 1.2σ) and the mFo−DFc difference density (purple at 4σ) from a refined model that lacks the phosphate group. b, To confirm the presence of the phosphoserine, ESI–LC–MS/MS was performed on KdpB isolated by SDS–PAGE from a crystal. Peptides harbouring phosphorylated and unphosphorylated Ser162 were separated by FPLC and their relative abundance is represented by these elution profiles for peptides with m/z = 1,010.26 (unphosphorylated) or m/z = 1,030.25 (phosphorylated) (z = 4). The sequence of these peptides, in which the cysteine was modified by iodoacetamide, is shown above, as determined by MS2. c, The cytoplasmic domains of KdpB adopt a unique position in our structure compared to other P-type ATPases owing to a salt bridge between the phosphorylated Ser162 in the A domain (yellow) and residues Lys357 and Arg363 of the N domain (red). This interaction is likely to prevent N domain movements, and the close proximity of the P domain to the N domain prevents nucleotide binding, suggesting that the current structure represents an autoinhibited state. d, The level of Ser162 phosphorylation inversely correlated with the ATPase activity of KdpFABC. The absolute level of phosphoenzyme was not determined owing to a lack of a fully dephosphorylated or fully phosphorylated enzyme preparation for calibration of the mass spectrometer detection system; nevertheless, the inverse correlation between the relative levels of Ser162 phosphorylation and ATPase activity supports the inhibitory nature of this modification. Although this serine phosphorylation was observed in samples isolated directly from the bacteria, its physiological relevance is uncertain, especially given that Lys357 and Arg363 are poorly conserved amongst KdpFABC systems (Extended Data Fig. 9). Footnotes for this table: 1apparent percentage of phosphopeptide from mass spectrometry does not reflect lower detection efficiency of the phosphopeptide; 2ATPase assay was determined in the presence of 50 mM KCl; 3200 units of protein phosphatase from bacteriophage lambda (LPP) were incubated with 20 μg KdpFABC under the conditions indicated.
Extended Data Figure 6 Sequence alignment of KdpA with other SKT transporters TrkH and KtrB.
This structure-based alignment was done by the promals3d server50 using PDB depositions 4J7C for KtrB12 and 3PJZ for TrkH11. Sequence conservation is shown by colour with red being most conserved and green least conserved. Secondary structure based on KdpA is shown with the colouring scheme used in Fig. 1c. The four M1PM2 motifs are denoted D1–D4, with S indicating the selectivity filter. The red star indicates the Q116R mutation and red triangles indicate residues implicated in coupling with KdpB, namely Glu370 and Arg493, which reside at the cytoplasmic side of the selectivity filter (Fig. 2c), and Arg400, which forms a salt bridge with the P domain (Fig. 3b). This figure and Extended Data Figs 7, 8, 9 were made with ALINE59.
Extended Data Figure 7 Sequence alignment of KdpB with other P-type ATPases.
This structure-based alignment was done by the promals3d server using PDB depositions 1SU4 for the calcium pump (SERCA)15, 5KSD for the plasma membrane proton pump (AHA2)60, 4BYG for the copper pump (CopA)61 and 4HYT for the Na,K-ATPase62. Sequence conservation is shown by the colour scheme with red being most conserved and green least conserved. Secondary structure elements correspond to KdpB with the same colour scheme as in Fig. 1. The red stars indicate the catalytic Asp307 and other residues involved in the hydrogen bonding network shown in Extended Data Fig. 4a (Thr309, Lys399 and Asp522). The red diamonds indicate residues at the water site along M4 and M6 shown in Extended Data Fig. 4d (Pro264 and Asn624). Red triangles indicate residues implicated in coupling with KdpA, namely Asp583 and Lys586 (Extended Data Fig. 4d) in the transmembrane domain as well as Asp300 and Asp302, which interact with cytoplasmic loops from KdpA (Fig. 3b). The red circles indicate phosphorylated Ser162 and Lys357/Arg363 from the N domain, which form salt bridges to the phosphate (Extended Data Fig. 5c).
Extended Data Figure 8 Sequence alignment of KdpA from a diverse group of bacteria.
Selection of the sequences was done with the divblast server49 and alignment using clustal omega48. Secondary structure corresponds to that observed for KdpA (Fig. 1). Pink stars indicate residues reported in ref. 5 and pink diamonds indicate residues reported in ref. 6 to increase the apparent Kd for K+ to 0.3 mM or higher. Red triangles indicate residues implicated in coupling with KdpB, namely Glu370 and Arg493 at the cytoplasmic end of the selectivity filter (Fig. 2c) and Arg400, which forms a salt bridge to the P domain of KdpB (Fig. 3b). Blue triangles indicate residues forming hydrogen bonds to the periplasmic domain of KdpC.
Extended Data Figure 9 Sequence alignment of KdpB from a diverse group of bacteria.
Selection of sequences was done with the divblast server49 and alignment using clustal omega48. Secondary structure corresponds to that observed for KdpB (Fig. 1). Red stars indicate residues that interact with the catalytic Asp307 in our structure. Red triangles indicate residues that interact with cytoplasmic loops from KdpA (Asp300 and Asp302). Red circles indicate Ser162, which is phosphorylated in our structure, and the residues in the N domain that engage this phosphate in a salt bridge (Lys357 and Arg363).
Rights and permissions
About this article
Cite this article
Huang, CS., Pedersen, B. & Stokes, D. Crystal structure of the potassium-importing KdpFABC membrane complex. Nature 546, 681–685 (2017). https://doi.org/10.1038/nature22970
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22970
- Springer Nature Limited
This article is cited by
-
KdpD is a tandem serine histidine kinase that controls K+ pump KdpFABC transcriptionally and post-translationally
Nature Communications (2024)
-
Enhancing coevolutionary signals in protein–protein interaction prediction through clade-wise alignment integration
Scientific Reports (2024)
-
Potassium Solubilizing Microorganisms as Potential Biofertilizer: A Sustainable Climate-Resilient Approach to Improve Soil Fertility and Crop Production in Agriculture
Journal of Plant Growth Regulation (2024)
-
Wie Bakterien die Aufnahme von Kaliumionen regulieren
BIOspektrum (2022)
-
What do we know about osmoadaptation of Yersinia pestis?
Archives of Microbiology (2022)