Abstract
Metabolic production of acetyl coenzyme A (acetyl-CoA) is linked to histone acetylation and gene regulation, but the precise mechanisms of this process are largely unknown. Here we show that the metabolic enzyme acetyl-CoA synthetase 2 (ACSS2) directly regulates histone acetylation in neurons and spatial memory in mammals. In a neuronal cell culture model, ACSS2 increases in the nuclei of differentiating neurons and localizes to upregulated neuronal genes near sites of elevated histone acetylation. A decrease in ACSS2 lowers nuclear acetyl-CoA levels, histone acetylation, and responsive expression of the cohort of neuronal genes. In adult mice, attenuation of hippocampal ACSS2 expression impairs long-term spatial memory, a cognitive process that relies on histone acetylation. A decrease in ACSS2 in the hippocampus also leads to defective upregulation of memory-related neuronal genes that are pre-bound by ACSS2. These results reveal a connection between cellular metabolism, gene regulation, and neural plasticity and establish a link between acetyl-CoA generation ‘on-site’ at chromatin for histone acetylation and the transcription of key neuronal genes.
Similar content being viewed by others
Main
Memory formation involves synaptic restructuring and requires the coordinated expression of neuronal genes through poorly understood processes that modify chromatin1,2. Histone acetylation is a key regulator of memory storage and restructures chromatin in distinct brain regions that have been implicated in learning and memory, most prominently in the hippocampus3. Hippocampal memory consolidation requires the transcription factor CREB and the coactivator CREB binding protein (CBP), specifically the histone acetyltransferase (HAT) activity of CBP4,5. Furthermore, inhibitors of histone deacetylases enhance memory consolidation3. However, the mechanisms that regulate neuronal histone acetylation in long-term memory remain incompletely understood.
Direct sensing of intermediary metabolites by chromatin-modifying enzymes such as acetyltransferases can dynamically adapt chromatin structure and gene expression6,7. Alteration of pools of intracellular acetyl-CoA manipulates histone acetylation8,9; thus, metabolic enzymes that generate nuclear acetyl-CoA might directly control histone acetylation and gene expression10,11. In mammalian cells, there are two principal enzymes that generate acetyl-CoA for histone acetylation: acetate-dependent acetyl-CoA synthetase 2 (ACSS2) and citrate-dependent ATP-citrate lyase (ACL)11. The relative importance of ACSS2 and ACL for nuclear histone acetylation differs by tissue type, developmental state, and disease9,11, but the roles of these enzymes in post-mitotic neuronal cells are unknown.
The observation that ACSS2 is highly expressed in the mouse hippocampus12 led us to investigate the role of ACSS2 in neuronal histone acetylation and gene expression. Our findings support the hypothesis that neuronal ACSS2 has a critical function in linking acetate metabolism to neuronal gene regulation through direct binding of chromatin by ACSS2, and identify a prominent role of this mechanism in hippocampal memory consolidation.
ACSS2 regulates neuronal gene expression
We investigated the function of ACSS2 in neurons using the Cath.-a-differentiated (CAD) cell line derived from mouse catecholaminergic cells. Upon serum deprivation, CAD cells differentiate to form neuronal processes and become excitable, similar to functional neurons13. Immunofluorescence showed that endogenous ACSS2 was primarily cytoplasmic in undifferentiated CAD cells (Fig. 1a), but shifted primarily to the nucleus upon differentiation (Fig. 1b, Extended Data Fig. 1a). Whole-cell and nuclear levels of ACSS2 increased upon differentiation of CAD cells into neurons, whereas cytoplasmic ACL expression remained constant (Fig. 1c). In primary hippocampal and cortical neurons from mouse brain, even 14 days after isolation, ACSS2 was chiefly nuclear and ACL was primarily cytoplasmic (Extended Data Fig. 1c–f). We conclude that ACSS2, unlike ACL, is localized to nuclei during neuronal differentiation.
a, ACSS2 localizes to the cytoplasm in undifferentiated CAD neurons. ACSS2 was imaged by immunofluorescence microscopy in CAD cells (4′,6-diamidino-2-phenylindole (DAPI) and α-tubulin (α-Tub) immunostaining show nuclei and cytoplasm, respectively). b, ACSS2 localizes to the nucleus in differentiated CAD neurons. c, Western blot analysis of cytoplasmic (CE) and nuclear (NE) extracts from undifferentiated CAD cells (undiff.) and differentiated CAD neurons (diff.) for ACSS2, ACL and histone H3. Nuclear ACSS2 expression is higher in differentiated cells (p.d.u., procedure defined unit; t-test P = 0.002, n = 3, mean ± s.d.). d, ACSS2 knockdown reduces differentiation-linked upregulation of neuronal gene expression program. Scatter plot contrasts the fold-change fragments per kilobase of transcript per million mapped (FPKM) of induced genes (Extended Data Fig. 2c) between wild-type (WT) and ACSS2 knockdown (KD) cells. Marginal distributions show histogram and kernel density estimation. Ordinary least squares linear regression is displayed with 95% confidence interval. e, Western blot of lysates from differentiated CAD neurons that were infected with lentiviral control (WT) or ACSS2 knockdown vector (shACSS2) (quantification shown in Extended Data Fig. 1g; n = 3). f, ACSS2 knockdown greatly reduces gene upregulation. Quintiles of upregulated genes (red dots in Extended Data Fig. 2c) with the greatest fold-change increase in wild-type cells (grey). Corresponding gene quintiles depict fold-change FPKM in ACSS2 knockdown cells (green) (for each quintile, columns represent the mean induction value of genes; mean ± s.e.m.). g, ACSS2i treatment of differentiated CAD neurons results in reduced expression of differentiation-induced genes. All genes are plotted in order of fold-change in wild-type CAD differentiation, and z-scores were computed for ACSS2i treatment and control, representing upregulation as blue and downregulation as red (RNA-seq in 24-h ACSS2i-treated and DMSO-treated control neurons, genes removed with z-score <0.5). Scale bar, 10 μm (a, b).
We investigated the role of ACSS2 in upregulation of canonical neuron-specific protein markers in differentiated CAD neurons. Pre-differentiation knockdown of ACSS2 reduced differentiation-linked expression of nuclear NeuN, activity-regulated Nr4a2, and the cytoplasmic markers synaptophysin, Map2 and Snap25, without an associated decrease in ACL (Extended Data Fig. 1g), indicating that ACSS2 has a key role in neuronal differentiation.
Transcriptome analysis by mRNA sequencing (mRNA-seq) upon CAD neuronal differentiation identified 894 upregulated genes (Extended Data Fig. 3a–c; Supplementary Table 1). Gene ontology analysis revealed that these differentiation-linked genes were neuron-specific; gene ontology terms included neuron differentiation, synaptic transmission, ion transport, and neuron projection morphogenesis (Extended Data Fig. 3e). We built a protein interaction framework that produced a neuronal network centred on activity-dependent signalling and synaptic plasticity: calmodulin 1 (Calm1), glutamate ionotropic receptor NMDA type subunit 1 (Grin1), and inositol 1,4,5-trisphosphate receptor type 1 (Itpr1) (Extended Data Fig. 3d). Calm1 mediates the control of neuronal proteins by Ca2+ during synaptic plasticity, including Ca2+/calmodulin-dependent protein kinase II (CaMKII). Such Ca2+ signalling is regulated by Grin1, an NMDA receptor subtype of glutamate-gated ion channels, and also by the ion channel Itpr1, which mobilizes intracellular Ca2+ stores, an important process in activity-dependent signalling that underlies synaptic plasticity during learning.
The metabolite acetyl-CoA is required by HAT enzymes for histone acetylation. To investigate histone acetylation during differentiation of CAD cells into neurons, we performed chromatin immunoprecipitation with high-throughput DNA sequencing (ChIP–seq) for histone H3 lysine 9 acetylation (H3K9ac), H4K5ac, and H4K12ac (see Methods). All marks were enriched upon differentiation at upregulated neuronal genes (for example, at nudix-type motif 1 (Nudt1)) (Extended Data Fig. 2f). Overall, the 894 upregulated neuronal genes displayed higher acetylation than all other genes (Extended Data Fig. 2g).
We reduced levels of ACSS2 or ACL using short hairpin RNAs (shRNA) before cell differentiation and performed RNA-seq (Extended Data Fig. 2h–k). The induction of neuronal genes was lost in the ACSS2 knockdown cells (Pearson r = 0.15; Fig. 1d, e), whereas the same genes retained a strong correlation in transcriptional fold-change in ACL knockdown cells (Pearson r = 0.53; Extended Data Fig. 1l). We stratified the top 10% upregulated genes in differentiated wild-type cells into quintiles (Fig. 1i), and found that ACSS2 knockdown strongly lowered upregulation across all quintiles (Fig. 1i: green bars; P = 7.2 × 10−252, Wilcoxon rank-sum test). The ACL-knockdown cells showed the same upward trend as wild-type cells, in contrast to the severe defect in ACSS2-knockdown cells (Extended Data Fig. 2m; P = 1.1 × 10−25, Wilcoxon rank-sum test). Notably, ACL-knockdown cells showed lower global transcript levels (P = 1.91 × 10−7, Mann–Whitney U-test), unlike ACSS2-knockdown cells, which showed a less severe genome-wide defect (Extended Data Fig. 2n; P = 0.04, Mann–Whitney U-test). ACL thus has a broad but non-specific effect on gene expression, whereas ACSS2 is required for upregulation of the neuronal gene expression program upon differentiation of CAD cells into neurons.
Further, we tested whether ACSS2 catalytic activity is required for the neuronal gene expression program using a small-molecule specific inhibitor of ACSS2 (ACSS2i)14. RNA-seq showed a reduction in differentiation-induced genes (Fig. 1g), and the genes whose expression was affected by ACSS2-knockdown were also highly sensitive to the ACSS2i (Extended Data Fig. 2o, P = 1.62 × 10−6).
Recruitment of ACSS2 to chromatin
We investigated the direct association of ACSS2 with chromatin using ChIP–seq of differentiated CAD cells (see Methods). Two ACSS2 antibodies showed a high correlation both for model-based analysis of ChIP–seq (MACS) overlapping peaks (Spearman r = 0.82; Extended Data Fig. 3a), and for global enrichment over 1-kb genomic windows (Spearman r = 0.73; Extended Data Fig. 3b). Binding of ACSS2 correlated with increases in histone H3K9ac, H4K5ac, and H4K12ac in differentiated relative to undifferentiated CAD cells, for instance at the promoters of Nudt1 and Tab2 (TAK1-binding protein 2; Extended Data Fig. 3c, d). Both genes have been linked to neurodegenerative disorders; the Nudt1 hydrolase oxidizes purine nucleoside triphosphates to prevent RNA incorporation, and Tab2 regulates signal transduction pathways in neurons15. Gene ontology analysis showed that genes proximal to ACSS2 peaks were linked to neuronal differentiation (Extended Data Fig. 3e). Hence, chromatin-associated, neuronal gene promoter-proximal ACSS2 may provide a local source of acetyl-CoA to HAT enzymes.
We examined ACSS2 binding relative to histone acetylation and found that 80% of ACSS2 peaks upstream of the nearest target gene overlapped an acetylation peak or had an acetylation peak downstream towards the targeted transcription start site (TSS; Extended Data Fig. 3f, g). A substantial number (13–15% of all ACSS2 peaks genome-wide) directly overlapped peaks of H3 and H4 acetylation (Extended Data Fig. 3h). In addition, the height of ACSS2 peaks correlated overall with intersected histone acetylation peaks (Extended Data Fig. 3i–k). This peak height correlation suggests that H4 acetylation might be most responsive to ACSS2-derived acetyl-CoA, in particular H4K12ac, a mark that has been linked to defective memory formation during ageing16. In general, the most enriched ACSS2 peaks displayed the strongest histone acetylation enrichment (Extended Data Fig. 3l–n).
We investigated putative recruitment of ACSS2 by transcription factor binding using de novo motif discovery over ACSS2 ChIP–seq peaks, which revealed binding sequences predicted for neuronal transcription factors. The most enriched motif was Yin Yang1 (YY1) (Extended Data Fig. 3o; P = 1 × 10−599), which recruits the two acetyl-CoA-dependent HAT enzymes CREB-binding protein (CBP) and E1A binding protein (p300)17, consistent with the idea that ACSS2 fuels nearby catalytic HAT activity.
Initial peak analysis did not identify all peaks of ACSS2 or acetylation (Extended Data Fig. 4a, b), so we analysed gene body enrichment and found additional prominent examples, such as Camk2a (Fig. 2a), which encodes the CaMKII alpha chain that is required for hippocampal long-term potentiation (LTP) and spatial learning. ACSS2 and acetylation co-occupancy profiles were similar at Camk2a and Nudt1 (Extended Data Fig. 3c). Meta-gene analysis indicated that the top 5% of ACSS2-enriched genes had levels of acetylation up to threefold higher than the mean across all genes (Extended Data Fig. 5a–d), and genes with the greatest fold-change in differential ACSS2 binding had the highest histone acetylation levels (Fig. 2c; Extended Data Fig. 5e–h). Gene ontology term enrichment showed that the top ACSS2-bound and acetylated genes were neuron-specific (Fig. 2b).
a, Genome browser tracks showing ChIP–seq over the Camk2a locus show that increases in H4K5, H4K12, and H3K9 acetylation co-occur with proximate ACSS2 enrichment upon CAD neuron differentiation (chromosome 18: 60,920,000–60,990,000). b, Gene ontology term enrichment analysis of top 5% genes that become ACSS2-bound during CAD neuron differentiation show neuronal pathways. c, Violin-contour plots show that ChIP–seq enrichment of the indicated histone acetylation occurs with top-ranked ACSS2 enrichment during neuronal differentiation of CAD cells. d, ChIP–seq enrichment of the 299 genes that are reduced upon ACSS2i treatment (see Methods) shows high correlation (P < 2.2 × 10−16 for all) with histone acetylation in the differentiated state (AUC, area under the curve; d, differentiated; u, undifferentiated). e, Analysis of all genes previously linked to neuronal differentiation (ND genes, AmiGO annotation set of 1,315 genes), and the subset of known ND genes that are induced during differentiation of CAD cells (Induced), shows reduced expression in ACSS2i-treated CAD neurons (inh.) compared to DMSO-treated control neurons (con.). Inhibitor-treated versus control, P < 2.2 × 10−16. f, Nuclear acetyl-CoA levels are reduced in response to either knockdown of ACSS2 (shACSS2; mean Δ = −0.19 ± 0.03, **P = 0.003) or application of the ACSS2 inhibitor (mean Δ = −0.25 ± 0.05, **P = 0.006; n = 3, mean ± s.d.). g, Western blot analysis of whole-cell lysates shows that lentiviral shRNA-mediated knockdown of ACSS2 lowers H3K9 and H3K27 acetylation (quantifed in Extended Data Fig. 6a). h, Western blot analysis of immunoprecipitation eluates shows that CBP is co-immunoprecipitated with ACSS2 but not with control Ig. i, Immunofluorescence in primary hippocampal neurons shows nuclear localization of ACSS2 (day 7 of in vitro differentiation culture, isolated from C57BL/6 embryos). Scale bar, 50 μm. j, Western blots of lysates from primary hippocampal neurons (d7) treated for 24 h with ACSS2i and probed with the indicated antibodies (quantified in Extended Data Fig. 6c) show reduction of histone acetylation.
At all induced genes, ACSS2 binding was concomitant with increased histone acetylation (Extended Data Fig. 3p), and the 299 genes that showed reduced expression upon ACSS2i treatment were those with the greatest differentiation-linked increases in histone acetylation (Fig. 2d). In total, about 9% of genes previously linked to neuronal differentiation (ND genes, AmiGO annotation set of 1,315 genes) were induced in differentiated CAD cells, and these induced genes were exceptionally sensitive to ACSS2i treatment (Fig. 2e, ‘Induced’). Moreover, although the entire ND gene class did not change expression in differentiated CAD cells, their expression was markedly reduced by ACSS2i treatment (Fig. 2e, ‘ND genes’). We visualized the interaction between differentiation-linked gene expression changes and ACSS2 recruitment to chromatin using multiple linear regression analysis, and found a remarkable relationship between higher ACSS2 enrichment (red) and increased gene expression (Extended Data Fig. 5i). Overall, the CAD cell genomic data demonstrate dynamic ACSS2 enrichment in differentiated neurons linked to increased histone acetylation and involvement in transcriptional upregulation of neuronal genes.
ACSS2 functions in neuronal histone acetylation
We measured nuclear acetyl-CoA levels in ACSS2 knockdown cells (Fig. 2f; mean Δ = −0.19 ± 0.03, P = 0.003) and in cells treated with ACSS2i (Fig. 2f; mean Δ = −0.25 ± 0.05, P = 0.006) and found that levels of acetyl-CoA were similarly decreased. This finding supports the theory that ACSS2 enzymatic activity supplies nuclear acetyl-CoA. Global histone acetylation levels of transcription-linked H3K27ac and H3K9ac were reduced in ACSS2 knockdown cells (Fig. 2g, Extended Data Fig. 6a), and these marks are key substrates of the transcriptional coactivators CBP and p300 with roles in hippocampal LTP and long-term memory18. ACSS2 co-immunoprecipitated with acetylated chromatin, specifically H3K9ac, H3K27ac, and H4K12ac (Extended Data Fig. 6b), and also with CBP (Fig. 2h), suggesting that ACSS2 binds chromatin at transcriptionally active genes to increase histone acetylation during memory formation in vivo4,5,18.
We examined ACSS2 in primary mouse hippocampal neurons, given their capacity for depolarization and expression of key memory-related neuronal genes. ACSS2 was localized to the nucleus (Fig. 2i), and ACSS2i treatment reduced neuronal marker expression and histone acetylation without lowering ACSS2 levels (Fig. 2j, Extended Data Fig. 6c). Expression of ACL did not change (Fig. 2j), indicating that ACL is less important than ACSS2 in the regulation of histone acetylation in hippocampal neurons.
We investigated chromatin association of ACSS2 and H3K9ac in vivo using ChIP–seq in mouse hippocampus. Our hippocampal H3K9ac mapping strongly correlated with ENCODE mouse forebrain H3K9ac ChIP–seq (Spearman coefficient of multiple correlation R = 0.67), with similar peak distribution (Extended Data Fig. 7a). Hippocampal ACSS2 and H3K9ac corresponded genome-wide and over three canonical neuronal genes involved in memory (Arc, Egr2 and Nr2f2; Fig. 3a, b). In addition, ACSS2 promoter binding and H3K9ac correlated with RNA–seq in the hippocampus (Fig. 3c). We found a small number of genes that were ACSS2-bound but not H3K9ac enriched that were silent, similar to genes not enriched for ACSS2 or H3K9ac (Fig. 3d). By contrast, genes enriched for H3K9ac were actively transcribed, but genes enriched for both ACSS2 and H3K9ac were most highly expressed (Fig. 3d).
a, ChIP–seq for ACSS2 and H3K9ac in mouse hippocampus. Track views show ACSS2 and H3K9ac for three neuronal genes involved in memory: Arc, Egr2 and Nr2f2 (chr15:74,496,025–74,506,488; chr10:66,991,018–67,006,804; and chr7:77,488,549–77,516,626, respectively). b, In vivo hippocampal ACSS2 and H3K9ac peaks co-localize with the nearest gene TSS (<1 kb from peak) among all RefSeq transcripts. c, RNA-seq expression in dorsal hippocampus (dHPC) correlates with hippocampal ACSS2 binding and enrichment of H3K9 acetylation. d, Expression profile of genes classified by their ACSS2 and H3K9ac enrichment state. e, Overlap between ACSS2 targeted genes (hippocampus) and CBP and H3K27ac enrichment for entire set of peaks (ENCODE CBP and H3K27ac ChIP–seq in mouse forebrain and cortex). f, Motif analysis at ACSS2 peaks from in vivo ChIP–seq in hippocampus showing top enrichment of NRF1, a neuronal transcription factor.
Physical association of ACSS2 and CBP in differentiated CAD cells (Fig. 2h) correlated with gene colocalization of ACSS2 and CBP in the hippocampus, together with H3K27ac (from public mouse cortex CBP and H3K27ac ChIP–seq data; ACSS2:CBP overlap P = 3.23 × 10−16 by hypergeometric test). Overall, 57% of ACSS2-associated genes were co-targeted by H3K27ac (Fig. 3e), and ACSS2 and CBP co-targeted genes were enriched for gene ontology terms involved in synaptic membrane potential (Extended Data Fig. 7b, c). Motif analysis at hippocampal ACSS2 peaks show that Nrf1—a transcription factor that regulates neurite growth—predicted binding at 45% of ACSS2 sites (Fig. 3f), evoking an ACSS2–CBP recruitment mechanism. Moreover, ACSS2i-sensitive genes (50%: 145 of 289) had proximal ACSS2 within 10 kb of the TSS (hypergeometric analysis, P = 7.6986 × 10−8), consistent with a direct role for chromatin-bound ACSS2 in transcription.
ACSS2 regulates long-term memory storage
Hippocampus-dependent spatial memory occurs through activity-dependent changes in gene expression that are coordinated, in part, through epigenetic modifications, specifically histone acetylation19,20. ACSS2 is expressed throughout the hippocampus12,21 (Extended Data Fig. 8a), and thus could mediate histone acetylation to upregulate neuronal gene expression during memory consolidation19,22. To investigate the role of ACSS2 in the adult hippocampus, we attenuated ACSS2 expression in the dorsal hippocampus by shRNA knockdown using a viral vector (Fig. 4a, b). Compared to control-injected mice, ACSS2 knockdown mice showed similar levels of locomotion, coordination, body weight, and anxiety-related thigmotaxis during open field exploration23 (Extended Data Fig. 8b–d; not significant, n = 10 per group); therefore, ACSS2 knockdown did not cause gross behavioural alterations.
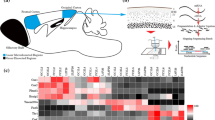
a, Stereotactic surgery was performed to deliver AAV9 knockdown vector into the dorsal hippocampus (AP, −2.0 mm; DV, −1.4 mm; ML, ±1.5 mm from bregma); 4 weeks later, habituated mice were trained in object location memory (OLM; four 5-min training sessions in arena with three different objects). Twenty-four hours later the mice were given a retention test in which one object was moved to a novel location (n = 10 per cohort). b, Western blot analysis of hippocampal tissue removed from mice injected into the dorsal (d) or ventral (v) hippocampus with either eGFP control or ACSS2 knockdown vector shows specific reduction of ACSS2 in dorsal hippocampus. c, ACSS2-knockdown mice are impaired in object location memory. eGFP control and shACSS2 AAV9 mice display no preference for any of three objects (O1–3) during the object location memory training session (TR). In the retention test 24 h later, control mice show a preference for the novel object location (NL), whereas the knockdown mice display no such preference. ***P < 0.001; n = 10, mean ± s.d. d, The spatial memory defect in ACSS2-knockdown mice manifests in a lowered discrimination index (% DI = (t NL – t FL)/(t NL+ t FL)) compared to control mice (ΔDI = −29.5 ± 11.4, *P = 0.02; n = 10, mean ± s.d.). e, Training-induced expression of a cohort of immediate early genes (Extended Data Fig. 8h) is greatly attenuated in ACSS2-knockdown mice (n = 4 mice per group, 2 replicates for each condition, P < 0.0001, paired t-test, mean ± s.d.). f, Model for function of ACSS2 as a chromatin-bound coactivator to provide acetyl-CoA locally to promote histone acetylation and activity-induced upregulation of immediate early genes.
To assess hippocampus-dependent spatial memory, we used an object-location memory paradigm24. Animals explored three different objects during training, and long-term memory was tested by re-exposure 24 h later with one object moved to a different location (Fig. 4a, right). In training, control and knockdown mice showed no difference in exploration (Fig. 4c, left). During memory retrieval, control mice showed increased exploration of the object that had been moved (Fig. 4c). By contrast, ACSS2 knockdown mice were impaired in spatial object memory (Fig. 4c and Extended Data Fig. 8e, mean Δ = −5.01 ± 1.21; P = 8.3 × 10−5), and displayed a lower discrimination index (Fig. 4d; % ΔDI = −29.5 ± 11.4; P = 0.02). ACSS2 knockdown mice showed reduced total object exploration during the test (Fig. 4c), suggesting diminished novelty associated with intact recognition of the objects from the training session (mean Δ = −6.13 ± 2.15; P = 0.02, n = 10 per group).
Previous studies have shown that long-term contextual fear memory is mediated by the ventral hippocampus when manipulations of the dorsal hippocampus occur before training25. Therefore, as a control experiment, mice injected with the ACSS2 knockdown shRNA or eGFP in the dorsal hippocampus were subjected to a contextual fear conditioning paradigm. During the 24-h test session, there was no significant difference in the amount of freezing behaviour between ACSS2-knockdown and eGFP mice (Extended Data Fig. 8f, g) suggesting that the ventral hippocampus successfully mediated context–shock association. Overall, we conclude that ACSS2 has a critical role in dorsal hippocampus-mediated long-term spatial memory.
ACSS2 regulates upregulation of immediate early genes
Long-term spatial memory requires a rapid increase in histone acetylation and immediate early gene transcription to occur in a sensitive time window to enable memory consolidation19,26; during memory retrieval, gene transcription also occurs for memory reconsolidation, and this can be prevented by inhibiting mRNA synthesis during the sensitive post-retrieval period27. We tested whether ACSS2 was involved in dynamic gene upregulation for hippocampal memory consolidation and reconsolidation by performing mRNA-seq on the dorsal hippocampus. We first identified global gene expression changes that were induced by spatial object training, which has not previously been investigated genome-wide to our knowledge. Dorsal hippocampi from control and shACSS2-knockdown mice were removed during the sensitive period of memory consolidation following spatial object training. To control for circadian oscillation, injected animals were included that had been handled but not trained.
Genes that were differentially expressed following training were identified by transcriptome comparison of trained control-injected mice to untrained circadian control-injected mice using Cuffdiff. A small number of genes were induced immediately after training, including immediate early genes such as Egr2, Fos, Nr2f2, Sgk1 and Arc (Fig. 4e). Importantly, baseline expression of these memory-associated genes in untrained control and ACSS2-knockdown mice was remarkably similar (Extended Data Fig. 8h). By contrast, dynamic upregulation of these immediate early genes by training was greatly reduced by ACSS2 knockdown (Fig. 4e).
We further tested whether ACSS2 also regulates immediate-early gene expression in the dorsal hippocampus during memory reconsolidation. Therefore, our hippocampal mRNA-seq analysis was focused on previously identified and validated genes that become upregulated following memory retrieval26,28. Retrieval-associated induction of immediate early genes during the sensitive reconsolidation period was blocked by ACSS2 knockdown, whereas retrieval-linked downregulation during the same period was not (Extended Data Fig. 9a–f).
Discussion
Metabolic state can regulate chromatin structure, in particular by inducing histone modifications10. Here, we establish a connection between cellular metabolism and neuronal plasticity, and reveal a neuronal function of ACSS2 as a chromatin-bound transcriptional coactivator that stimulates histone acetylation and gene expression.
Acetyl-CoA metabolism is cell- and tissue-specific, and is frequently disregulated in malignant transformation14,29. In adipose cells, ACSS2 partially localizes to the nucleus and contributes to histone acetylation under conditions of low glucose9,30, but the principal metabolic determinant of histone acetylation is ACL9. By contrast, we show that post-mitotic neurons rely on chromatin-recruited ACSS2 to supply acetyl-CoA for histone acetylation. Notably, fasting lowers acetyl-CoA and protein acetylation in most tissues, but acetyl-CoA levels remain unchanged in the brain31, indicating that neuronal ACSS2 has an important role in the fasted state when acetyl-CoA production by citrate-dependent ACL is reduced.
Optimal acetyltransferase activity requires an increased local acetyl-CoA to CoA ratio, which determines the catalytic activity and substrate specificity of HAT enzymes8,9,32. This finding suggests that histone acetylation can be controlled by changing levels of nuclear acetyl-CoA. Thus, chromatin-bound ACSS2 could provide acetyl-CoA to fuel HAT activity locally, instantaneously recycling CoA and could also recapture acetate from deacetylation reactions. Here, we show specific chromatin binding by ACSS2 at neuronal genes and link localization to upregulation of histone acetylation and gene transcription in spatial memory (Fig. 4f), which requires increased histone acetylation3,20. We demonstrate a crucial role for ACSS2 in upregulation of immediate early genes with key functions in neuronal plasticity and memory, leading to a critical role in long-term memory consolidation.
Epigenetic mechanisms continue to be revealed as important regulators of neural function and behaviour, and have been implicated in neuropsychiatric diseases, including anxiety disorders and depression1,3,33. In establishing ACSS2 as a key regulator at the interface of metabolic environment and neuronal chromatin, our study provides a previously unrecognized enzymatic target for the development of therapies to treat neurological and cognitive disorders.
Methods
Mouse experiments
Animal use, surgical procedures, and all experiments performed were approved by the Institutional Animal Care and Use Committee (IACUC, protocol 804849). All personnel involved have been adequately trained and are qualified according to the Animal Welfare Act (AWA) and the Public Health Service (PHS) policy. No statistical methods were used to predetermine sample size; prior experiments using the relevant behavioural assays with pharmacological or genetic manipulations determined that effects are achieved when group sizes are at least 7–9 animals. The experiments were randomized and the investigators were blinded to allocation during experiments and outcome assessment.
Cell culture
CAD cells (Cath.-a-differentiated; donated by the Holzbaur laboratory at the University of Pennsylvania and intermittently tested for mycoplasma contamination) were grown in Dulbecco’s modified Eagle’s medium (DMEM):Ham’s F12 (1:1), supplemented with 2 mM glutamine, 1% penicillin/streptomycin, and 10% fetal bovine serum (FBS). To induce neuronal differentiation, sub-confluent CAD cell cultures (50–60%) were transferred to serum-free medium (DMEM:Ham’s F12 (1:1) supplemented with 2 mM glutamine) and maintained in 15-cm2 culture dishes for 5 days. Upon differentiation, CAD neurons exhibit morphological changes that are characteristic of neurons. For knockdown experiments, CAD cells were infected with lentiviral hairpin constructs (TRC collection) designed against ACL (#TRCN0000055217) or ACSS2 (#TRCN0000076124, #TRCN0000076125) in medium containing 8 mg ml-1 polybrene and 10% FBS for 24 h. Cells then underwent selection in culture medium supplemented with 0.5 mg ml-1 puromycin for 5 days to obtain a stably infected population. Cell treatment with ACSS2i (diluted in 1-(2,3-di(thiophen-2-yl)quinoxalin-6-yl)-3-(2-methoxyethyl)urea (DMSO), ChemBridge) was carried out for 24 h at a final concentration of 20 μm (treatment with DMSO alone served as control).
RNA-seq
To generate libraries for RNA-seq, poly(A)+ RNA was extracted using the Dynabeads mRNA Direct kit (Ambion) according to the manufacturer’s instructions. RNA-seq libraries for scrambled control (referred to as wild-type), shACL, and shACSS2 were made using a ScriptSeq v2 RNA-seq Library Preparation Kit from Epicentre (now Illumina). The quantity and quality of the libraries were assessed by BioAnalyzer (Agilent) and qPCR (Kapa Biosystems). The multiplexed libraries were pooled and sequenced on a single lane on the Illumina NextSeq 500 platform (50 bp, single-end reads). All RNA-seq data were prepared for analysis as follows: NextSeq sequencing data was demultiplexed using bcl2fastq2-v02.14.01.07. Demultiplexed FASTQs were aligned by RNA-STAR 2.3.0.e using the genome index mm10 generated from iGenome UCSC mm10 FASTQ genome sequence. The aligned reads were mapped to genomic features using cufflinks-2.2.1, (-G parameter to quantify only known features), and iGenomes mm10 UCSC genomic transcript loci. The rRNA, mRNA, and tRNA of the mouse genome were downloaded from the goldenPath UCSC FTP and were masked from the transcript quantification. After quantification, all data processing was done using python pandas library v.0.14.0. Differential expression in CAD neurons was defined as the top 10% of genes by fold-change, corresponding roughly to 1.6-fold upregulation or higher. Differential expression in the inhibitor and the hippocampal ACSS2 knockdown in vivo were defined using Cuffdiff. The relationship between CAD cell differentiation and inhibitor function (Fig. 1g) was inferred by assessing standardized scores over two RNA-seq replicates each of untreated and ACSS2i-treated cells. These were averaged and genes with |z| < 0.5 in either condition were dropped. Scores for remaining genes were plotted in order of increasing CAD differentiation fold-change. The statistical significance of the trend and reproducibility were assessed by taking the top 20% of genes by loss of expression in the knockdown and comparing the expression of these genes in inhibitor-treated cells to a random sample of genes outside this set (Mann–Whitney test).
ChIP–seq
CAD cells were fixed in 1% formaldehyde for 10 min and fixation was quenched with the addition of glycine to 125 mM for an additional 5 min. Cells were harvested by scraping from plates, and washed twice in 1× PBS before storage at −80 °C. ChIP was performed as previously described34), except that chromatin was sheared to an average size of <500 bp using the Covaris S220 Ultrasonicator. Equal aliquots of sonicated chromatin from undifferentiated and differentiated CAD neurons were used for each immunoprecipitation reaction, and 10% of the amount was saved as input. ACSS2 ChIPs were performed using 2,000 μg extract and 4 μg antibody per sample; all other ChIPs were performed using 500 μg extract and 4 μg antibody per sample. Immunoprecipitation was performed using protein A Dynabeads (Life Technologies). Sequencing libraries were prepared using NEBNext Ultra library preparation procedure, and then assessed for quality and quantity by BioAnalyzer (Agilent) and qPCR (Kapa Biosystems). Sequencing was performed on the Illumina NextSeq 500 platform. All ChIP–seq data were prepared for analysis as follows: NextSeq sequencing data was demultiplexed using bcl2fastq. All reads were aligned to the mm9 or the mm10 referenence genome using bowtie2.2.1. One alignment was allowed per read and one mismatch was allowed in the seed region (-N1 -k1). Reads were tabulated in fixed windows or to genes provided in the iGenome mm10 UCSC annotations using featureCounts from the subread 1.4.6 software package. CAD cell ACSS2 ChIP–seq data were normalized to input controls, while all histone acetylation ChIP–seq data were H3-subtracted. The plot in Extended Data Fig. 5i is the result of performing a multiple linear regression to determine the relationship between expression in undifferentiated and differentiated CAD cells (regressors) and enrichment of ACSS2 in differentiated CAD cells (target). The relationship was used to colour the negative space in the plot by propensity for ACSS2 binding. For the in vivo ChIP, hippocampal tissue pooled from two animals was finely minced and cross-linked with formaldehyde (1% final concentration) for 15 min at room temperature, followed by glycine quenching for an additional 10 min at 4 °C. To create a single-cell suspension, samples were washed once with ice-cold PBS and homogenized by passing through a 22G needle 10 times. Subsequent steps were performed in the same way as described for the in vitro ChIP. In vivo ChIP peaks (ACSS2 (T), H3K9ac, CBP—GSM1629373, and H3K27ac—GSM1629397) were called using MACS v2.1.0 with the false discover rate (FDR) controlled at 1%. Peak scores were assessed by adjusting to millions of aligned tags and subtracting background. Tracks were similarly normalized and are visualized using the UCSC genome browser with a maximum value windowing function and smoothing at 5 pixels (in vitro ChIP–seq) or using default parameters (in vivo ChIP–seq). The Venn diagram in Fig. 3b shows overlap of genes (official gene symbol) that feature ChIP–seq peaks within 1 kb of their nearest TSS. Venn diagrams in Fig. 3e and Extended Data Fig. 7b display overlapping target gene sets (RefSeq transcripts) that were assigned to the entire set of ChIP–seq peaks, because CBP binds to few genes within 1 kb of the TSS. The in vivo mRNA expression versus ChIP analysis (Fig. 3c) was created by sorting genes on log2-transformed expression values (DESeq library-size adjusted, replicate-averaged values) in home cage control mice then displaying ChIP–seq AUCs (RPM-adjusted ChIP minus RPM-adjusted background, length adjusted) for each gene for the peak closest to its TSS within a distance of 1 kb (all genes with more distant peaks were rendered as having a score of zero). Gene targets were inferred by the presence of peaks proximal to the TSS (within 1 kb). To identify enriched motifs (Fig. 3f), in vivo ACSS2 peaks targeting genes that were upregulated in differentiated CAD cells (without respect for distance to the nearest TSS) were compared to a background set of equal-sized regions selected from gene-rich regions using HOMER (peak sizes were fixed at 300 bp). Discovered motifs were filtered for those present at one-third or more of the targeting peaks and with a tenfold or higher enrichment over the gene-rich background. To assess the overlap of ACSS2 and histone acetylation (Extended Data Fig. 3f, g), ACSS2 peaks were filtered to include only those upstream of their nearest target genes. Downstream acetylation was assessed for similarly filtered peaks of H3K9ac, H4K5ac, and H4K12ac from the same cells, as well as cortical H3K27ac. For the in vivo analysis (Fig. 3b), gene targets of ACSS2 or H3K9ac peaks within 1 kb of their nearest TSS were examined for overlap. The acetylation pattern due to differentiation at induced or inhibitor-sensitive genes (Fig. 3d, Extended Data Fig. 3p) was assessed by taking a 20-kb window around the TSS and measuring the input-adjusted ChIP–seq signal. H3K9ac data were validated (Extended Data Fig. 7a) by comparing to ENCODE’s common-replicate peaks for H3K9ac in mouse forebrain (accession ENCSR369RBO) using CEAS (default parameters, with a 1–10 kb window around the TSS and TES). Additional comparisons were made to H3K9ac (NCBI GEO: GSE82643) and H3K27ac (GSE82428), contrasting to input (GSE82659) to control for sonication efficiency. Cortical H3K27ac (NCBI GEO: GSM1629397) and CBP (GSM1629373) were aligned along with the corresponding input (GSM1629381) using bowtie2 (parameters -k 1 -N 1–local) and peaks were called using MACS2 (input control, FDR controlled at 1%) (Fig. 3e, Extended Data Fig. 7b). The combined effect of ACSS2 and histone acetylation targeting on gene expression in vivo (Fig. 3d) was demonstrated by box-plotting expression in home cage control mice at genes targeted by ACSS2 by itself, H3K9ac by itself, ACSS2 + H3K9ac, or neither. Only genes bound at the promoter (1 kb distance) by ACSS2 were considered.
Acetyl-CoA quantification
To extract and quantify acetyl-CoA from differentiated CAD neurons, 4 × 106 cells were washed and incubated in lysis buffer for 30 min (10 mM Tris pH 8, 1 mM KCl, 1.5 mM MgCl2, 1 mM DTT). The nuclei were pelleted at 3,000g for 5 min, and immediately re-suspended in Acetyl CoA Assay Buffer provided in the PicoProbe Acetyl CoA Assay Kit (Abcam, ab87546). The acetyl-CoA assay, including the deproteinization step, was prepared according to the manufacturer’s instructions. The PicoProbe assay was performed in 96-well clear-bottom plates, and the resulting fluorescence was quantified using the Synergy HTX Multi-Mode Microplate Reader (BioTek Instruments).
Western blots
Cells were lysed in buffer containing 50 mM Tris pH 8.0, 0.5 mM EDTA, 150 mM NaCl, 1% NP40, 1% SDS, supplemented with protease inhibitor cocktail (Life Technologies, number 78446). For subcellular fractionation experiments, the cells were processed using the subcellular fractionation kit for cultured cells (Thermo Scientific, number 78840) according to the manufacturer’s instructions. Protein concentration was determined by BCA protein assay (Life Technologies, number 23227), and equal amounts of protein were used in co-immunoprecipitation experiments or directly loaded onto polyacrylamide gels. The endogenous co-immunoprecipitation experiments were performed using antibody-conjugated protein A Dynabeads (Life Technologies) in buffer containing: 20 mM Tris, pH 8.0, 137 mM NaCl, 1 mM MgCl2, 1 mM CaCl2, 1% NP-40, 10% glycerol, with protease and phosphatase inhibitors, and 12.5 U ml−1 benzonase (Novagen, 70746). Proteins or co-immunoprecipitation eluates were loaded and separated on 4–12% Bis-Tris polyacrylamide gels (NuPAGE). After transfer to nitrocellulose membrane, 3% BSA in TBS supplemented with 0.1% Tween 20 (TBST) was used to block the membrane at room temperature for 1 h. Primary antibodies were diluted in TBST, and incubated at 4 °C overnight. Primary antibodies are listed below. The membrane was washed three times with TBST, each for 10 min, followed by incubation with HRP-conjugated secondary antibodies at room temperature for 1 h, in TBST. The membrane was washed again three times, and imaged with a Fujifilm LAS-4000 imager. Original gel blots are provided as Supplementary Fig. 1.
Immunofluorescence
Cells were fixed in 4% PFA in PBS for 20 min at room temperature. Cells were washed twice with PBS and permeabilized with 0.5% Triton X-100 in PBS for 10 min. After being washed twice, cells were blocked in 10% BSA in PBS for 1 h at room temperature. Cells were incubated with primary antibodies in 5% BSA in PBS supplemented with 0.1% Tween 20 (PBST) overnight at 4 °C. Antibodies are listed below. Then cells were washed four times with PBST, each for 10 min, followed by incubation with fluorophore-conjugated secondary antibody in 5% BSA in PBST for 1 h at room temperature. F-actin was labelled using Alexa Fluor 488 Phalloidin (Thermo A12379). Cells were then washed three times in PBST, once with PBS, and incubated with 1 μg ml−1 DAPI for 5 min. The cells were then washed twice with PBS and mounted with ProLong Gold (Invitrogen). The slides were observed and imaged using a Nikon Eclipse microscope. Microscopy settings were unchanged between samples.
Antibodies
The antibodies used were anti-H3 (Abcam ab1791), anti-H3K9ac (Abcam ab4441), anti-H3K27ac (Abcam ab4729), anti-H3K122ac (Abcam ab33308), anti-H4 (Millipore 05-858), anti-H4K5ac (Millipore 39-584), anti-H4K12ac (Abcam ab1761), anti-ACSS2 (T) (Thermo MA5-145810), anti-ACSS2 (CS) (Cell Signaling 3658), anti-ACL (Proteintech 15421-1-AP), anti-α-tubulin (Sigma T8328), anti-GAPDH (Fitzgerald Industries 10R-G109A), anti-KAT3A/CBP (Abcam ab2832), anti-SNAP25 (Abcam ab5666), anti-synaptophysin (Millipore MAB368), anti-MAP2 C/D (Cell Signaling 8707), anti-NR4A2 (Santa Cruz sc-991) and anti-NeuN (Millipore ABN78).
Intracranial injection of viral vector
Adult mice (8+ weeks of age) were anaesthetized with isoflurane gas (1–5% to maintain surgical plane) and placed in a sterile field within a stereotaxic device. Animals received an injection of bupivacaine (2.5 mg kg−1) for local anaesthesia before the skin was disinfected with betadine solution and the skull exposed with a short incision using sterile surgical equipment. Artificial tears were applied to eyes to ensure sufficient lubrication. A small hole (about 0.5 mm) was drilled in the skull over the target area using a stereotax and a stereotactic drill. A micro-syringe filled with viral vector was inserted into the dorsal hippocampus and slowly removed following injection (AP, −2.0 mm; DV, −1.4 mm; ML, ±1.5 mm from bregma). ACSS2 knockdown vector, AAV2/9.U6.shACSS2.CMV.EGFP; eGFP control vector, AAV2/9.CMV.EGFP.polyA. All animals received a single dose of subcutaneous meloxicam (5 mg kg) as analgesia at induction and one dose per day for two days postoperatively as needed.
Object location memory task
The object location memory procedure is used to test spatial memory. The procedure consists of a training phase and a testing phase. Prior to training, each mouse was handled for 3 min a day for 3 days. On the training day, mice are placed in an arena (approx. 1 square foot) containing three different objects. The objects used were a glass bottle, a metal tower (h × w × l, 5 × 2 × 2 inches), and a plastic cylinder. Mice were habituated to an empty arena with a black and white striped spatial cue on the wall, followed by object exposure in three 6-min trials with an interval of 3 min. The arena and objects were cleaned with 70% EtOH between trials. To diminish biases, the memory test was performed on control and knockdown mice on the same day in the same arena, using every combination of object location (n = 10 mice per study group). After 24 h, the individual mice were placed back in the arena used in the testing phase. For testing, one of the objects was moved to different location in the arena. Mice were allowed to explore freely for 5 min. Each session was recorded using a video camera and time spent exploring (approaches and sniffing) each object was assessed offline. All animals were randomized and preassigned to arena and object the day before testing to ensure that every treatment group explored every object configuration.
Contextual fear conditioning
The mouse was placed in the conditioning chamber (Med Associates) for 5 min before the onset of the unconditioned stimulus (US), a 1.5-mA continuous foot shock. A mild 2-s, 1.5-mA foot shock is used as an aversive stimulus; this does not injure the mice but provides the transient, yet startling and aversive, stimulus that is necessary for conditioning. After an additional 30 s in the chamber, the mouse was returned to its home cage. Twenty-four hours later, the mouse was tested for a freezing response to the chamber (contextual) where training occurred. Time spent freezing in the chamber (motionless except for respiratory movements) was assessed for 5 consecutive minutes.
Data availability
The ChIP–seq and RNA-seq data have been made available at the Gene Expression Omnibus (GEO) repository under the SuperSeries accession code GSE76854. All other data are available from the authors upon reasonable request.
Accession codes
References
Kandel, E. R., Dudai, Y. & Mayford, M. R. The molecular and systems biology of memory. Cell. 157, 163–186 (2014)
Zovkic, I. B., Guzman-Karlsson, M. C. & Sweatt, J. D. Epigenetic regulation of memory formation and maintenance. Learn. Mem. 20, 61–74 (2013)
Gräff, J. & Tsai, L.-H. Histone acetylation: molecular mnemonics on the chromatin. Nat. Rev. Neurosci. 14, 97–111 (2013)
Wood, M. A. et al. Transgenic mice expressing a truncated form of CREB-binding protein (CBP) exhibit deficits in hippocampal synaptic plasticity and memory storage. Learn. Mem. 12, 111–119 (2005)
Korzus, E., Rosenfeld, M. G. & Mayford, M. CBP histone acetyltransferase activity is a critical component of memory consolidation. Neuron. 42, 961–972 (2004)
Kaelin, W. G. Jr & McKnight, S. L. Influence of metabolism on epigenetics and disease. Cell. 153, 56–69 (2013)
Katada, S., Imhof, A. & Sassone-Corsi, P. Connecting threads: epigenetics and metabolism. Cell. 148, 24–28 (2012)
Cai, L., Sutter, B. M., Li, B. & Tu, B. P. Acetyl-CoA induces cell growth and proliferation by promoting the acetylation of histones at growth genes. Mol. Cell. 42, 426–437 (2011)
Wellen, K. E. et al. ATP-citrate lyase links cellular metabolism to histone acetylation. Science. 324, 1076–1080 (2009)
Gut, P. & Verdin, E. The nexus of chromatin regulation and intermediary metabolism. Nature. 502, 489–498 (2013)
Pietrocola, F., Galluzzi, L., Bravo-San Pedro, J. M., Madeo, F. & Kroemer, G. Acetyl coenzyme A: a central metabolite and second messenger. Cell Metab. 21, 805–821 (2015)
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature. 445, 168–176 (2007)
Qi, Y., Wang, J. K., McMillian, M. & Chikaraishi, D. M. Characterization of a CNS cell line, CAD, in which morphological differentiation is initiated by serum deprivation. J. Neurosci. 17, 1217–1225 (1997)
Comerford, S. A. et al. Acetate dependence of tumors. Cell. 159, 1591–1602 (2014)
Sardi, S. P., Murtie, J., Koirala, S., Patten, B. A. & Corfas, G. Presenilin-dependent ErbB4 nuclear signaling regulates the timing of astrogenesis in the developing brain. Cell. 127, 185–197 (2006)
Peleg, S. et al. Altered histone acetylation is associated with age-dependent memory impairment in mice. Science. 328, 753–756 (2010)
Aoyama, T. et al. Histone modifiers, YY1 and p300, regulate the expression of cartilage-specific gene, chondromodulin-I, in mesenchymal stem cells. J. Biol. Chem. 285, 29842–29850 (2010)
Vecsey, C. G. et al. Histone deacetylase inhibitors enhance memory and synaptic plasticity via CREB:CBP-dependent transcriptional activation. J. Neurosci. 27, 6128–6140 (2007)
Barrett, R. M. et al. Hippocampal focal knockout of CBP affects specific histone modifications, long-term potentiation, and long-term memory. Neuropsychopharmacology. 36, 1545–1556 (2011)
Gräff, J., Woldemichael, B. T., Berchtold, D., Dewarrat, G. & Mansuy, I. M. Dynamic histone marks in the hippocampus and cortex facilitate memory consolidation. Nat. Commun. 3, 991 (2012)
Ariyannur, P. S. et al. Nuclear-cytoplasmic localization of acetyl coenzyme a synthetase-1 in the rat brain. J. Comp. Neurol. 518, 2952–2977 (2010)
Maren, S. & Holt, W. The hippocampus and contextual memory retrieval in Pavlovian conditioning. Behav. Brain Res. 110, 97–108 (2000)
Stanford, S. C. The Open Field Test: reinventing the wheel. J. Psychopharmacol. 21, 134–135 (2007)
Balderas, I. et al. The consolidation of object and context recognition memory involve different regions of the temporal lobe. Learn. Mem. 15, 618–624 (2008)
Rogers, J. L., Hunsaker, M. R. & Kesner, R. P. Effects of ventral and dorsal CA1 subregional lesions on trace fear conditioning. Neurobiol. Learn. Mem. 86, 72–81 (2006)
Peixoto, L. L. et al. Memory acquisition and retrieval impact different epigenetic processes that regulate gene expression. BMC Genomics. 16, S5 (2015)
Mamiya, N. et al. Brain region-specific gene expression activation required for reconsolidation and extinction of contextual fear memory. J. Neurosci. 29, 402–413 (2009)
Poplawski, S. G. et al. Object-location training elicits an overlapping but temporally distinct transcriptional profile from contextual fear conditioning. Neurobiol. Learn. Mem. 116, 90–95 (2014)
Mashimo, T. et al. Acetate is a bioenergetic substrate for human glioblastoma and brain metastases. Cell. 159, 1603–1614 (2014)
Gao, X. et al. Acetate functions as an epigenetic metabolite to promote lipid synthesis under hypoxia. Nat. Commun. 7, 11960 (2016)
Mariño, G. et al. Regulation of autophagy by cytosolic acetyl-coenzyme A. Mol. Cell. 53, 710–725 (2014)
Takahashi, H., McCaffery, J. M., Irizarry, R. A. & Boeke, J. D. Nucleocytosolic acetyl-coenzyme a synthetase is required for histone acetylation and global transcription. Mol. Cell. 23, 207–217 (2006)
Walker, D. M., Cates, H. M., Heller, E. A. & Nestler, E. J. Regulation of chromatin states by drugs of abuse. Curr. Opin. Neurobiol. 30, 112–121 (2015)
Shah, P. P. et al. Lamin B1 depletion in senescent cells triggers large-scale changes in gene expression and the chromatin landscape. Genes Dev. 27, 1787–1799 (2013)
Acknowledgements
We thank the NeuronsRUs core of the Mahoney Institute for Neurological Sciences for preparations of primary hippocampal neurons. T.A. is supported by RO1 MH 087463. P.M. and S.L.B. are supported by NIH P01AG031862.
Author information
Authors and Affiliations
Contributions
P.M. and S.L.B. conceived the project. P.M. performed most of the experiments. A.M.D. analysed the CAD RNA-seq datasets. P.M. and G.D. performed and analysed CAD ACSS2i and hippocampal RNA-seq experiments. A.M.D. and G.D. analysed ChIP–seq datasets. P.M. and V.L. performed in vivo ACSS2 knockdown and behavioural characterization. P.M. and S.L.B. wrote the manuscript. All authors reviewed the manuscript and discussed the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 ACSS2 localizes to the nucleus of neurons.
a, Percentage of cells with nuclear staining in ACSS2 immunofluorescence experiments (undiff., undifferentiated CAD cells; diff., differentiated CAD neurons; hippocampal, primary hippocampal neurons day 7; a minimum of 50 cells were examined in three replicate immunofluorescence experiments; t-test undiff. vs diff. P < 0.0001, undiff. vs hippocampal P < 0.0001; error bars, s.e.m.). b, Western blots of cytoplasmic (CE) and nuclear (NE) extracts from undifferentiated CAD cells and differentiated CAD neurons were probed with the indicated antibodies. c, d, Immunofluorescence in primary cortical neurons isolated from C57BL/6 embryos, at day 7 (c) and day 14 (d) of in vitro differentiation culture. ACSS2 locates predominantly to nuclei in differentiated primary cortical neurons. All scale bars, 25 μm. e, Immunofluorescence in primary hippocampal neurons isolated from C57BL/6 embryos at day 14 of in vitro differentiation culture. ACSS2 locates predominantly to nuclei in differentiated primary neurons. f, Immunofluorescence in primary hippocampal neurons at day 7 shows that ACL is chiefly localized to the cytoplasm. g, Neuronal differentiation markers decrease in ACSS2 knockdown cells. CAD cells were infected with lentiviral control (WT) or knockdown vector (shACSS2). Western blots of lysates from stably infected differentiated cells were probed with the indicated antibodies and quantified using ImageJ (n = 3; error bars, s.e.m.).
Extended Data Figure 2 ACSS2 regulates neuronal gene expression.
a, b, Correlation plots of replicate RNA-seq in undifferentiated CAD cells (a) and differentiated CAD neurons (b) for scramble control. c, Transcriptome analysis via RNA-seq, done in two highly correlated biological replicates, identified 894 genes that became upregulated in differentiated CAD neurons (red dots depict genes with >1.6-fold increase). d, Pathway analysis of the 894 upregulated genes (red dots in Fig. 2a) using StringDB. The protein–protein interaction graph depicts a network of binding partners that centres on key players in activity-dependent signalling and synaptic plasticity: Itpr1, Grin1, Nefh, Dync1h1 and Calm1. e, Gene ontology enrichment analysis shows upregulation of neuronal pathways. Gene ontology analysis was used on the 894 genes that become upregulated in differentiated CAD neurons (Extended Data Fig. 2c; identified by RNA-seq, fold-enrichment (FE) > 3.5, FDR < 0.005). f, Genome browser view of Nudt from RNA-seq and ChIP–seq (H4K12ac, H4K5ac, and H3K9ac: mm10 chr5: 140,327,500–140,339,000). g, Relative gene enrichment of H3K9ac, H4K5ac, and H4K12ac at genes that are upregulated during CAD neuron differentiation (>1.6-fold, grey bars) versus all other genes (black bars). h, i, Correlation plots of replicate RNA-seq in undifferentiated CAD cells for ACL knockdown (h), and ACSS2 knockdown (i). j, k, Correlation plots of replicate RNA-seq in differentiated CAD neurons for ACL knockdown (j) and ACSS2 knockdown (k). l, ACL knockdown has a much smaller effect on differentiation-linked upregulation of neuronal gene expression (compare to Fig. 1d). Scatter plot contrasts the fold-change FPKM of induced genes (Extended Data Fig. 2c) between wild-type and ACL knockdown cells. Marginal distributions show histogram and kernel density estimation. Ordinary least squares linear regression is displayed with 95% confidence interval. m, The corresponding quintiles of upregulated genes (red dots in Extended Data Fig. 2c) with the greatest fold-change FPKM increase in wild-type cells. The ACL knockdown showed the same upward trend as the wild-type cells (red bars, compared to black bars in Fig. 1f), contrasting with the severe defect in ACSS2-knockdown cells (green bars; for each quintile, columns represent the mean induction value of genes and error bars represent s.e.m.). n, Box plot of global mRNA transcript levels in undifferentiated and differentiated CAD neurons from RNA-seq in wild-type (scramble control knockdown; grey), ACSS2-knockdown (shACSS2 #25 knockdown; green), and ACL-knockdown (shACL #17 knockdown; red) cells. Genome-wide transcript levels are reduced in differentiated ACL-knockdown cells when compared to differentiated wild-type cells (error bars, s.e.m.), whereas the global reduction in differentiated ACSS2-knockdown cells is less significant when compared to differentiated wild-type cells (error bars, s.d.). o, Genes sensitive to ACSS2 knockdown (top 20%) are also sensitive to ACSS2i treatment, which lowers their expression compared to all genes.
Extended Data Figure 3 ACSS2 is chromatin-bound in differentiated CAD neurons.
a, ChIP–seq in differentiated CAD neurons was performed in replicate with two different antibodies against ACSS2. Correlation plot displays relative enrichment over corresponding MACS peaks (default parameters with input as control, 1,598 peaks). b, Correlation plot displays relative genome-wide ChIP–seq enrichment. c, UCSC Genome Browser views of ChIP–seq tracks show that, upon CAD neuron differentiation, increases in H4K5, H4K12, and H3K9 acetylation over the Nudt1 locus co-occur with ACSS2 enrichment (chr5: 140,326,845–140,339,655). d, UCSC Genome Browser view of indicated ChIP–seq tracks in undifferentiated CAD cells and differentiated CAD neurons over Tab2 locus (chr10: 7,875,000–8,004,000). e, Gene ontology enrichment analysis of the genes most proximate to ACSS2 peaks demonstrates that neuron-specific genes are enriched. f, Frequency of ACSS2 peaks (T antibody) located upstream of their target gene associated with histone acetylation. g, Frequency of ACSS2 peaks (CS antibody) located upstream of their target gene associated with histone acetylation. h, Table shows per cent direct overlap of ACSS2 peaks with H3K9ac, H4K5ac, and H4K12ac broad MACS peaks. i–k, Decile plots depict enrichment of H3K9ac (i), H4K5ac (j), and H4K12ac (k) over ranked deciles of ACSS2 peak enrichment (zeroes removed). l–n, Differentiation-induced co-enrichment of ACSS2 and acetyl broad peaks (MACS). Peak enrichment correlation indicated for H3K9ac (l), H4K5ac (m), and H4K12ac (n). o, Discovered de novo motifs for transcription factor binding sites predicted by HOMER from all ACSS2 ChIP–seq peaks called by MACS in differentiated CAD neurons. p, ChIP–seq enrichment of differentiation-induced genes as a group shows correlation with histone acetylation in differentiated CAD neurons.
Extended Data Figure 4 ACSS2 enrichment co-occurs with histone acetylation at neuronal genes in differentiating CAD neurons.
a, UCSC Genome Browser views of ChIP–seq tracks demonstrate that increases in H4K5, H4K12, and H3K9 acetylation co-occur with ACSS2 enrichment over the Idua (α-l-iduronidase) locus upon CAD neuron differentiation (chr5: 108,649,457–108,687,261). b, At the Slc19A1 (solute carrier family 19 member 1) gene, elevated histone H4K5, H4K12, and H3K9 acetylation levels correspond with increasing ACSS2 enrichment in differentiated CAD neurons (chr10: 76,761,141–77,170,455).
Extended Data Figure 5 Genic ACSS2 enrichment upon CAD neuronal differentiation corresponds to increased histone acetylation.
a–d, Meta-gene enrichment analysis shows ChIP occupancy for ACSS2 (a), H3K9ac (b), H4K5ac (c) and H4K12ac (d) across the top 5% of genes enriched for ACSS2 in differentiated CAD neurons (Top 5% DE; red). The bottom 80% of genes (Bot 80% DE) is shown in blue, and the average signal across all genes (All genes DE) is shown in green. e–h, Meta-gene enrichment analysis shows ChIP occupancy for ACSS2 (e), H3K9ac (f), H4K5ac (g) and H4K12ac (h) at the top 5% of genes that become dynamically bound by ACSS2 upon neuronal differentiation (Top 5% DE; red). The bottom 80% of genes (Bot 80% DE) is shown in blue, and the average signal across all genes (All genes DE) is shown in green. i, Multiple linear regression analysis was implemented to model the interaction between genic ACSS2 enrichment and wild-type gene expression changes, and to visualize the interaction between differentiation-linked gene expression changes and ACSS2 recruitment to chromatin. The contour plot of this fitted regression model displays high levels of ACSS2 enrichment in red and low levels in blue, and is overlaid with the scatter plot of the independent gene expression variables. The visualized model demonstrates that high ACSS2 enrichment corresponds to increased gene expression in differentiated CAD neurons.
Extended Data Figure 6 ACSS2 functions in neuronal histone acetylation.
a, Western blot analysis of whole-cell lysates shows that lentiviral shRNA-mediated knockdown of ACSS2 lowers H3K9 and H3K27 acetylation (compare to Fig. 2g), quantified using ImageJ (n = 3, error bars show s.e.m.). b, Western blot analysis of eluates and supernatants of IgG control and ACSS2 co-immunoprecipitation experiments indicates that ACSS2 binds to acetylated chromatin. c, Western blots of lysates from primary hippocampal neurons (d7) treated for 24 h with the ACSS2i, probed with the indicated antibodies (compare to Fig. 2j), and quantified using ImageJ (n = 3, error bars show s.e.m.).
Extended Data Figure 7 ACSS2 chromatin association and H3K9ac in dorsal hippocampus corresponds to H3K27ac and CBP enrichment in neuronal tissue.
a, Genome-wide compartment analysis of in vivo hippocampal ChIP–seq of H3K9ac and mouse forebrain H3K9ac ChIP–seq from ENCODE, showing a similar peak distribution genome-wide: although they originate in different brain regions, the in vivo H3K9ac ChIP data are in strong agreement (Spearman R = 0.67). b, Overlap of RefSeq transcripts targeted by the indicated enzyme or modification (peaks for CBP (GSM1629373) and H3K27ac (GSM1629397) in mouse cortical neurons were called using MACS2 (narrow peaks, FDR 0.1%) with an input sonication efficiency control (GSM1629381); peaks were associated to the nearest TSS among all RefSeq transcripts). c, Gene Ontology enrichment analysis performed on common CBP–ACSS2 targets, indicating that these enzymes co-target genes that modulate synapse biology and synaptic membrane potential.
Extended Data Figure 8 Attenuation of ACSS2 expression in the dorsal hippocampus impairs object location memory.
a, ACSS2 RNA in situ hybridization on ACSS2 in sagittal section of hippocampal region CA1 (left, reference atlas adapted from Allen Mouse Brain Atlas12; right, in situ hybridization; HPC, hippocampus proper). b, Weight of eGFP-AAV9 control and shACSS2-AAV9 knockdown mice before intracranial surgery, and following recovery before object location memory (OLM) training (NS, n = 10 per group, error bars show s.d.). c, d, ACSS2 knockdown mice showed no defect in locomotion or thigmotaxis (tendency to remain close to vertical surfaces in an open field, a measure of anxiety), as quantified over 5 min in the open field test; c shows example heat map of tracking data (NS, n = 10 per group, error bars show s.d.). e, Exploration times by eGFP-AAV9 control and shACSS2-AAV9 knockdown mice recorded for the three objects (O1–3) during the first OLM training session (TR) and the 24-h retention test (NL, object in novel location; FL, objects in former location). f, Compared to the control eGFP-AVV9 mice, ACSS2-knockdown mice showed no defect in contextual fear memory. Freezing in chamber on day of contextual fear conditioning was recorded and quantified pre-shock (FC Training; NS, n = 10 per cohort, error bars show s.d.). Fear memory was measured as the freezing response after re-exposure to the context 1 day after contextual fear conditioning (aversive stimulus: 1.5 mA electrical shock; NS, n = 10 per cohort, error bars show s.d.). g, RNA-seq was performed on the dorsal hippocampus of eGFP control and shACSS2-knockdown animals. Global transcript levels were not affected by ACSS2 knockdown (dHPC, dorsal hippocampus; four animals per group, two replicates for each condition; NS, paired t-test, error bars show s.d.). h, Baseline expression of immediate-early genes in untrained animals was unaltered in ACSS2-knockdown mice. RNA-seq was performed on the dorsal hippocampus of eGFP control and shACSS2-knockdown mice (r = 0.82, P < 0.0001; HCC, homecage circadian control).
Extended Data Figure 9 ACSS2 regulates retrieval-induced upregulation of immediate-early genes in vivo.
a, Genome-wide RNA-seq was performed on the dorsal hippocampus of eGFP control and shACSS2-knockdown mice. The analysis was focused on the set of previously identified and validated genes that become upregulated during the sensitive period following memory retrieval. The baseline expression of immediate early genes in untrained animals was not changed in shACSS2-AAV9 mice when compared to eGFP-AAV9 control mice (CC, circadian control). b, During the sensitive period following contextual memory retrieval (RT, 30 min post-exposure to conditioning chamber 24 h after fear conditioning), immediate early genes were upregulated in the dorsal hippocampus of control injected mice. By contrast, the dynamic retrieval-induced expression of these early response genes is absent in ACSS2-knockdown mice (P = 0.001, paired t-test). c, Induction defect of immediate early genes in shACSS2-AAV9 injected animals (RT/CC). d, The baseline expression of genes that were downregulated after contextual memory retrieval is not altered in ACSS2-knockdown mice. e, Downregulation of retrieval-responsive genes occurs in both eGFP control and ACSS2-knockdown mice, except for Cldn5. f, Retrieval-induced downregulation of retrieval-responsive genes in the dorsal hippocampus in eGFP control versus shACSS2-knockdown mice (RT/CC).
Supplementary information
Supplementary Figure 1
This file was replaced on 7 June 2017 to remove a duplicate page. Shown are the original western blots with size marker indications. Outlined in red are the cropped blot data presented in Figures 1c, 1e, 2g, 2h, 2j, 4b and Extended Data Figures 1b and 6b. (PDF 424 kb)
Supplementary Table 1
A list of genes upregulated 1.6-fold or higher upon CAD neuronal differentiation, corresponding to the top 10% of upregulated genes by fold-change. NextSeq mRNA sequencing data were aligned by RNA-STAR 2.3.0.e to the mm10 reference genome, and mapped to genomic features using cufflinks-2.2.1 and mm10 UCSC genomic transcript loci. The rRNA, mRNA, and tRNA of the mouse genome were downloaded from the goldenPath UCSC FTP and were masked from the transcriptome analysis. (TXT 17 kb)
Rights and permissions
About this article
Cite this article
Mews, P., Donahue, G., Drake, A. et al. Acetyl-CoA synthetase regulates histone acetylation and hippocampal memory. Nature 546, 381–386 (2017). https://doi.org/10.1038/nature22405
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22405
- Springer Nature Limited
This article is cited by
-
Short-chain fatty acid valerate reduces voluntary alcohol intake in male mice
Microbiome (2024)
-
Histone acetylation in an Alzheimer’s disease cell model promotes homeostatic amyloid-reducing pathways
Acta Neuropathologica Communications (2024)
-
Depletion of SAM leading to loss of heterochromatin drives muscle stem cell ageing
Nature Metabolism (2024)
-
Nuclei facing the tissue surface get fuel for development
Nature (2024)
-
Chemotherapy-induced acetylation of ACLY by NAT10 promotes its nuclear accumulation and acetyl-CoA production to drive chemoresistance in hepatocellular carcinoma
Cell Death & Disease (2024)