Abstract
Chemokines and their G-protein-coupled receptors play a diverse role in immune defence by controlling the migration, activation and survival of immune cells1. They are also involved in viral entry, tumour growth and metastasis and hence are important drug targets in a wide range of diseases2,3. Despite very significant efforts by the pharmaceutical industry to develop drugs, with over 50 small-molecule drugs directed at the family entering clinical development, only two compounds have reached the market: maraviroc (CCR5) for HIV infection and plerixafor (CXCR4) for stem-cell mobilization4. The high failure rate may in part be due to limited understanding of the mechanism of action of chemokine antagonists and an inability to optimize compounds in the absence of structural information5. CC chemokine receptor type 9 (CCR9) activation by CCL25 plays a key role in leukocyte recruitment to the gut and represents a therapeutic target in inflammatory bowel disease6. The selective CCR9 antagonist vercirnon progressed to phase 3 clinical trials in Crohn’s disease but efficacy was limited, with the need for very high doses to block receptor activation6. Here we report the crystal structure of the CCR9 receptor in complex with vercirnon at 2.8 Å resolution. Remarkably, vercirnon binds to the intracellular side of the receptor, exerting allosteric antagonism and preventing G-protein coupling. This binding site explains the need for relatively lipophilic ligands and describes another example of an allosteric site on G-protein-coupled receptors7 that can be targeted for drug design, not only at CCR9, but potentially extending to other chemokine receptors.
Similar content being viewed by others
Main
To obtain a crystal structure of human CCR9, a thermostabilized receptor (StaR) was generated8,9 containing eight amino-acid substitutions (Extended Data Figs 1 and 2). These modifications did not alter vercirnon binding properties of the receptor compared with wild-type (Extended Data Fig. 3); however, stabilization with the [3H]vercirnon antagonist precludes G-protein coupling of the final StaR (Data not shown). To further facilitate crystallization, amino (N) and carboxy (C) termini were truncated resulting in the construct designated CCR9-StaR(25-340). No fusion partner(s) were used to aid crystallization, and the receptor was crystallized in lipidic cubic phase (LCP) in the presence of the antagonist vercirnon10 (4-tert-butyl-N-{4-chloro-2-[(1-oxidopyridin-4-yl)carbonyl]phenyl}benzenesulfonamide, GSK1605786, CCX282-B). The structure was determined to 2.8 Å resolution with two copies in the asymmetric unit arranged in a parallel fashion with TM4–TM4-mediated interactions (Extended Data Fig. 4). Details of data collection and refinement are in Extended Data Table 1. For discussion purposes, molecule A is used forthwith.
CCR9 exhibits the core canonical arrangement of seven transmembrane helices (TM1–TM7) with continuous density observed for all intracellular loops (ICLs) and helix 8 (Fig. 1a). Only extracellular loop 3 was resolved on the extracellular side of the receptor. Additionally, only residual signal is present for the conserved disulfide bridging the top of TM3 (Cys1193.25) and extracellular loop 2. A second disulfide is present in CCR9 linking the N terminus (Cys38) with the top of TM7 (Cys2897.25) as for the related chemokine receptor structures of CCR5/maraviroc11 and CXCR4/IT1t12. Structural superposition of the 7TM core of CCR9 with both CCR5 and CXCR4 (sequence identity 35%, Extended Data Fig. 5) achieves a Cα root mean square deviation of 1.9 Å and 2.5 Å, respectively, with the main differences across the extracellular halves of the receptors (Fig. 1b–g). Compared with CCR5 and CXCR4, the tops of TM3 and TM6 of CCR9 are moved in towards the central axis of the helical bundle, and TM5 is moved outwards, with the differences being greatest between CCR9 and CXCR4. These changes in transmembrane helix position are possibly a consequence of the lack of a small molecule bound in the extracellular portion of the CCR9 transmembrane bundle.
a, Ribbon representation of CCR9 (cyan) viewed parallel to the membrane. Vercirnon is shown in sphere and stick representation, with carbon, nitrogen, chlorine, sulfur and oxygen atoms coloured magenta, blue, green, yellow and red, respectively. b–d, Superposition of CCR9 with CCR5/maraviroc (orange, maraviroc in green) viewed from the membrane, intracellular and extracellular space, respectively. e–g, Superposition of CCR9 with CXCR4/IT1t (blue, IT1t in yellow) viewed from the membrane, intracellular and extracellular space, respectively. Significant changes in transmembrane positions are denoted by red arrows.
Strong and unambiguous density is found for vercirnon on the intracellular side of the receptor contacting TM1, TM2, TM3, TM6, TM7 and helix 8 in an allosteric pocket within the helix bundle open to the cytoplasm (Fig. 2a and Extended Data Fig. 6). So far, the only other structural examples of small molecules binding towards the intracellular side of a receptor to effect allosteric antagonism are provided by the class B structures of corticotropin-releasing factor receptor type 1 (CRF1R) in complex with the small-molecule antagonist CP-376395 (ref. 13) and the glucagon receptor (GCGR) in complex with MK-0893 (ref. 14). However, while CP-376395 is found in a pocket approximately 18 Å from the centre of the orthosteric cavity of CRF1R, and MK-0893 adopts an extra-helical binding mode towards the bottom of TM6 in GCGR, the position of vercirnon bound to CCR9 is unique in both distance from the orthosteric site (approximately 33 Å) and in occupying a pocket with cytoplasmic access.
a, Electrostatic surface representation of the intracellular surface of CCR9 with vercirnon bound (coloured as in Fig. 1) in the allosteric pocket open to the cytoplasm. b, Ligand interactions in the intracellular allosteric binding pocket; specific interactions are depicted as dashed red lines with distances labelled (inset) Fo − Fc OMIT density contoured at 2.0σ calculated before vercirnon inclusion in the model.
Moving to the molecular details of the CCR9–StaR–vercirnon interaction, the sulfone group of vercirnon hydrogen bonds with the backbone amino groups of Glu322, Arg323 and Phe324, acting as a helix cap for the N terminus of helix 8 in CCR9. Favourable interactions are also made with the side chain of Tyr3177.53 (of the conserved NP7.50xxY(x)5,6F motif) from above the sulfone group. Mutation of Tyr3177.53, Phe324 and Gly3217.57 to Ala, three highly conserved residues across all chemokine receptors (Fig. 3a, b and Extended Data Table 2), severely decreases vercirnon binding to CCR9 (Fig. 3c and Extended Data Fig. 7), highlighting the importance of these residues in forming the core scaffold of the intracellular allosteric binding site, with Gly3217.57 contributing the necessary conformational flexibility in the junction of TM7–helix-8 to orient the N terminus of helix 8 for ligand interaction.
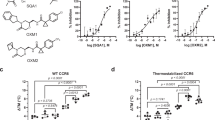
a, CCR9 allosteric site in surface representation with residues in rainbow spectrum according to conservation across chemokine receptors (red = 100%; blue = 0%). b, Two-dimensional schematic of a. c, [3H]vercirnon binding analysis of point mutations in allosteric site. Top: pKd from saturation binding analysis; T81E (P = 0.0027), F324A (P = 0.0116). Bottom: cell surface expression (percentage of wild-type (WT) allophycocyanin (APC) staining); T81E (P = 0.0134), L87F (P = 0.0232), Y317A (P = 0.002). Data shown as mean ± s.e.m. representative of three independent experiments performed in duplicate. Statistical difference, represented with asterisks calculated from unpaired two-tailed t-tests. *P ≤ 0.05; **P ≤ 0.01. †Ambiguous values due to near-complete loss of specific binding.
The ligand pyridine-N-oxide group is oriented towards the intracellular face of the receptor at the cytoplasmic entrance to the ligand binding cavity. The pyridine-N-oxide is surrounded by polar residues located on the intracellular extremities of TM2, TM3 and the TM7–helix-8 hinge region including Thr832.39, Asp842.40, Arg1443.50, Arg323 (on helix 8) Thr81 (on ICL1)—the last two being within hydrogen bonding distance of the pyridine-N-oxide (Fig. 2b). Mutation of Thr81 to glutamic acid reduces vercirnon binding compared with wild type (Fig. 3c), presumably as a result of the glutamic-acid side chain no longer being poised to make a polar contact with the ligand and/or fully engaging with Arg323 on helix 8. Finally, the ketone group of vercirnon is engaged in a hydrogen bond with the side chain of Thr2566.37, resulting in a ligand-mediated polar network linking TM6 across to ICL1, TM7 and the junction with helix 8 (Fig. 2b).
The tert-butylphenyl group anchors vercirnon in a cavity formed by TM1, TM2, TM7 and helix 8 and is characterized by concurrent lipophilic and hydrophilic residues. The lipophilic tert-butyl group faces towards the TM1–TM2 interface and makes hydrophobic interactions with Val691.53, Val721.56, Tyr731.57 in TM1 and Leu872.43 in TM2. Phe324 (on helix 8) and Tyr3177.53 make edge-to-face π–π stacking with the aromatic core of the tert-butylphenyl group (Fig. 2b). Other favourable hydrophobic interactions occur with the aliphatic portion of Arg323 and the side chain of Leu872.43, which is concomitantly engaged with the chlorophenyl part of the ligand.
The chlorophenyl moiety of vercirnon is located in a narrow, apolar cavity surrounded by several hydrophobic residues from TM2, TM3, TM6 and TM7. The chloro group, pointing up towards the central core of the receptor between TM3 and TM6, is located between the residues Leu872.43, Ile1403.46 and Val2596.40. The aromatic part of the chlorophenyl group is located between the hydrophobic surface of Leu872.43 and the main chain of Ala2556.36. Mutation of Leu872.43 to phenylalanine abolishes vercirnon binding to CCR9 (Fig. 3c), probably as a result of filling the cavity between TM2, TM3 and TM6 with a bulky aromatic side chain. The aromatic ring of Tyr3177.53 and the methyl group of Thr832.39 also contribute to favourable interactions with the chlorophenyl ring. The conservative stabilizing mutation V255A is found in the proximity of the chlorophenyl group of vercirnon; however, none of the stabilizing mutations altered vercirnon binding properties of the receptor compared with wild type (see earlier).
Molecular dynamics simulations of vercirnon bound to CCR9 (both StaR and wild type) showed stable interactions between the ligand and the residues in the binding site, with hydrogen bonds anchoring the sulfone group to the backbone of Arg323 and Phe324. However, after removal of ligand, molecular dynamics simulations of the pseudo-apo model showed a reorientation of side chains of Tyr3177.53, Arg323 and Phe324 towards the centre of the transmembrane bundle (Extended Data Fig. 8). Interestingly, the corresponding region in the CCR5 (ref. 11) structure is similar to the CCR9 pseudo-apo model after molecular dynamics.
Small-molecule chemokine receptor antagonists may be split into two broad chemical classes: tertiary amines and non-amines. Tertiary amines represent most compounds identified so far and probably engage a buried acidic residue (E2837.39 in the CCR5–maraviroc complex11) in the now well-understood class A transmembrane ligand-binding site region, explaining the preponderance of these molecules in chemical literature. Non-amines, such as vercirnon, have been less frequently reported and display pharmacological properties inconsistent with typical receptor antagonism. Interestingly, pepducin ATI-2341, a potent agonist of CXC-type receptor 4 (CXCR4) and whose peptide sequence derives from the first ICL of the receptor, suggests modulation of receptor activity by acting at the intracellular receptor surface15. Furthermore, mutagenesis studies have repeatedly suggested that many of the non-amine class of chemokine antagonists bind near the intracellular surface of receptors, for example the highly CCR4 selective pyrazinyl-sulfonamide series16. For the dual CXCR1/2 squaramide antagonist SCH-527123, mutagenesis of CXCR2 suggests an intracellular allosteric pocket17,18 lined by Thr832.39, Asp842.40, Tyr3147.53 and Lys3207.59, correlating with the vercirnon binding site in CCR9 (Extended Data Fig. 5); indeed a similar intracellular interaction mode may also exist for SB-656933 (ref. 19) binding to CXCR2. Additionally, investigation of two CXCR2 antagonists exhibiting 1000-fold selectivity over CXCR1, shows that selectivity can be reversed by swapping the receptor C-terminal tails, specifically mapping to residue Lys/Asn7.59 (correlating to Arg3237.59 in CCR9 which makes a direct contact to the pyridine-N-oxide of vercirnon)20. Pharmacological evidence for an intracellular allosteric binding site in CXCR2 is further provided by the insurmountable inhibition of CXCL8-promoted β-arrestin-2 recruitment by SB-265610 (ref. 21). Triazolylpyridylbenzenesulfonamides (CCR2), indazolesulfonamides (CCR4), repertaxin (CXCR1) and dihydroquinazolines (CXCR3) represent additional examples of non-amine chemokine antagonists5. The chemical similarity of several of these compounds to vercirnon, particularly the CCR2 and CCR4 antagonists that contain an aromatic sulfonamide (found capping helix 8 in CCR9), is highly suggestive of analogous sites on the intracellular face of their respective receptors. Overall, a consideration of the chemical nature of non-amine ligand classes, their pharmacological behaviour and evidence from mutagenesis supports the notion that an intracellular binding site may exist in many chemokine receptors, and that subtype-selective ligands can often be identified. Resolution of the structural details of this site in CCR9 facilitates further studies of non-amine chemokine antagonists using structure-based drug design.
In response to chemokine binding, CCR9 and chemokine receptor signalling in general have been most widely characterized via the heterotrimeric G-protein Gi transducer. However, C-terminal receptor phosphorylation by GRK can mediate β-arrestin binding, desensitization and internalization, alongside activation of, for example, Src, PI3K and MAPK22, with vercirnon inhibiting such signalling10. In structural terms, class A receptor agonist binding elicits a rigid-body movement along TM6, altering the interface to TM5 and causing an outward movement of the intracellular half of TM6 alongside an upward movement of TM3 (refs 23, 24). Superposition of CCR9–vercirnon with the β2–AR–Gs complex structure25 using the core transmembrane bundles provides a structural basis for intracellular allosteric antagonism (Fig. 4a). Assuming that Gi binds analogously, the G-protein clashes with vercirnon and TM6/ICL3 of CCR9, a likely consequence of vercirnon mediating a network of polar contacts (see earlier) from TM6 across to TM7/helix 8 and ICL1, which holds TM6 inwards towards the receptor’s central helical axis. This, alongside acting as a steric wedge within the helical bundle, restricts the required movements of TM6/TM3, thereby abrogating G-protein binding. Superposition with the structure of rhodopsin bound to arrestin26 demonstrates a similar situation where vercirnon–CCR9 interactions specifically occupy two of the major arrestin-receptor interfaces. Additionally, the junction of TM7–helix-8 in rhodopsin and the finger loop of arrestin directly clash with vercirnon (Fig. 4b).
a, Structural superposition of CCR9–vercirnon with the β2–AR–Gs complex structure (Protein Data Bank accession number 3SN6). b, Structural superposition of CCR9–vercirnon with the rhodopsin–arrestin complex structure (Protein Data Bank accession number 4ZWJ). Vercirnon exerts intracellular antagonism by holding the intracellular half of the receptor in a conformation sterically incompatible with G-protein or arrestin binding, both of which clash with the small molecule itself.
The structure of CCR9 complexed with vercirnon provides the first detailed view of a small molecule bound on the intracellular surface of a G-protein-coupled receptor, in a pocket within the helical bundle of the receptor but open to the cytoplasm. This novel allosteric pocket may be targeted for the design of selective small-molecule antagonists of CCR9 (or related chemokine receptors). Since the intracellular regions of the receptor that interact with G proteins are overlapping but not identical to those that engage β-arrestin, a unique opportunity may now exist to deploy structure-based drug design techniques in fine-tuning molecules that differentially modulate biased signalling cascades and functional outcomes in the chemokine receptor family.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Preparation of [3H]vercirnon and vercirnon
Pyridine intermediate 1 (4-tert-butyl-N-[4-chloro-2-(pyridin-4-ylcarbonyl)phenyl]benzenesulfonamide) was prepared according to published procedures27. Intermediate 2 and the radioligand [3H]vercirnon 3 were prepared by Quotient Bioresearch (see Supplementary Fig. 1). Briefly, intermediate 2 was prepared by reaction of a solution of intermediate 1 in DCM with tritium gas in the presence of (1,5-cyclooctadiene)(tricyclohexylphosphine)(pyridine)iridium(I) hexafluorophosphate (Crabtree’s catalyst), followed by purification by high-performance liquid chromatography (HPLC). Subsequently, [3H]vercirnon was prepared by N-oxidation according to published procedures for the intermediate 1 (ref. 27) and purified by preparative HPLC. Mass spectrometry of [3H]vercirnon gave a spectrum which was consistent with vercirnon and implied the incorporation of on average between one and two tritium atoms per molecule, which was consistent with the preparation of deuterated intermediate 2 in an analogous catalytic isotope exchange reaction with deuterium gas. The radiochemical was determined to have a purity of 99.9% by HPLC and a specific activity of 35 Ci/mmol. Cold vercirnon was prepared from intermediate 1 by N-oxidation according to published procedures27 and purified by preparative thin-layer chromatography.
StaR generation
Full-length human CCR9 (1–369) was used as background for the generation of the conformationally thermostabilized receptor using a mutagenesis approach described earlier9. Mutants were analysed for thermostability in the presence of the radioligand [3H]vercirnon. The CCR9-StaR is the full-length receptor with eight thermostabilizing mutations.
Cell culture
HEK293T cells were purchased from the American Type Culture Collection and were cultured in DMEM supplemented with 10% (v/v) fetal bovine serum (FBS). Cells were transfected using GeneJuice (Merck Millipore) according to the manufacturer’s instructions and harvested after 48 h.
Thermostability measurement
Transiently transfected HEK293T cells were incubated in 50 mM HEPES–NaOH pH 7.5, 150 mM NaCl, supplemented with cOmplete Protease Inhibitor Cocktail tablets (Roche), with 1% (w/v) n-dodecyl-β-d-maltopyranoside (DDM) or 1% (w/v) n-decyl-β-d-maltopyranoside (DM) at 4 °C for 1 h. All subsequent steps were performed at 4 °C. Samples were incubated with 250 nM [3H]vercirnon for 1 h and crude lysates cleared by centrifugation at 16,000 g for 15 min. Thermostability of the receptor was determined as previously described14. Thermal stability (Tm) is defined as the temperature at which 50% ligand binding is retained.
FACS analysis
HEK293T cells transiently expressing CCR9–enhanced green fluorescent protein (eGFP) constructs and mock-transfected cells were harvested 40 h post-transfection using non-enzymatic cell dissociation solution (Sigma-Aldrich). Cells were washed with FACS buffer (PBS, 0.1% sodium azide, supplemented with cOmplete Protease Inhibitor Cocktail tablets (Roche)) before counting. Half a million cells per staining sample were taken and re-suspended in 200 μl FACS buffer containing 2% BSA and Mouse anti-CCR9 (R&D systems, MAB179) at 5 μg/ml. After incubation for 1 h at room temperature, samples were washed three times with 200 μl FACS buffer, then resuspended in 200 μl FACS buffer containing 2% BSA and APC-conjugated Goat anti-Mouse IgG2A (Southern Biotech, 1080-11S) at 0.5 μg/ml and incubated at room temperature for 1 h in the dark. The cells were washed three times with 200 μl FACS buffer and finally resuspended in 200 μl FACS buffer before FACS analysis using BD FACSCantoII and FACSDiva software. Bound APC was detected using excitation wavelength (λex) = 633 nm and emission wavelength (λem) = 660 nm.
Radioligand binding
For saturation binding experiments HEK293 membranes transiently expressing CCR9 (5 μg per well) or CCR9-StaR(1-369) (2.5 μg per well) were incubated with varying concentrations of [3H]vercirnon (final assay concentration ≈ 0–50 nM) in the presence or absence of 1 μM vercirnon to define non-specific binding (assay buffer: 25 mM HEPES–NaOH pH 7.1, 140 mM NaCl, 1 mM CaCl2, 5 mM MgCl2, 0.2% BSA). Binding assays were incubated for 3 h at 25 °C. The reaction was terminated by rapid filtration through 96-well GF/B filter plates pre-soaked with 0.1% polyethyleneimine (PEI) using a 96-well head harvester (Tomtec, USA) and plates washed with 5 × 0.5 mL phosphate buffered saline. For saturation binding experiments of mutants, HEK293T cells transiently expressing CCR9–eGFP constructs or mock transfected cells were resuspended in buffer (50 mM HEPES–NaOH pH 7.5, 150 mM NaCl, supplemented with cOmplete Protease Inhibitor Cocktail tablets (Roche)) and homogenized using a Tissuemiser. Homogenized cells (5 × 104 cells per well) were incubated with varying concentrations of [3H]vercirnon (final assay concentration ≈ 0–15 nM) for 2.5 h at 25 °C. Non-specific binding was defined using mock transfected cells. The reaction was terminated by rapid filtration through 96-well GF/C filter plates pre-soaked with Milli-Q water using a 96-well head harvester (Tomtec, USA) and plates washed with 5 × 1 mL Milli-Q water. Specific binding was determined by subtracting mock transfected controls. Plates were dried, and bound radioactivity was measured using scintillation spectroscopy on a Microbeta counter (PerkinElmer, UK). Data were analysed using GraphPad Prism version 5 (San Diego, USA). Saturation binding data was globally fitted to one site total and non-specific binding, or one site-specific binding.
Truncation constructs
A panel of N- and C-terminal truncation variants of CCR9 was designed on the basis of multiple sequence alignment of all human chemokine receptors and secondary structure prediction28,29. Truncated receptors were expressed in HEK293T cells as C-terminal fusions with eGFP followed by a deca-histidine tag. Receptors were solubilized in 50 mM HEPES–NaOH pH 7.5, 150 mM NaCl, and 1% (w/v) n-dodecyl-β-d-maltopyranoside (DDM) and 0.05% (w/v) cholesteryl hemisuccinate (CHS) and their expression levels and stability was assayed by whole-cell fluorescence, western-blotting and fluorescence-detection size-exclusion chromatography (fSEC) as described30. The most suitable construct emerging from this screen comprised residues 25–340. Removal of post-translational modifications (glycosylation at Asn32 and putative palmitoylation at Cys337) was achieved by mutating residues Thr34—part of the glycosylation recognition sequence NXS/T—to Glu and Cys337 to Ala. Inclusion of an N-terminal GP64 signal sequence increased expression levels.
Expression, membrane preparation and protein purification
The truncated CCR9-StaR(25-340) construct was expressed with a C-terminal deca-histidine tag in Spodoptera frugiperda Sf21 cells (Oxford Expression Technologies) using ESF 921 medium (Expression Systems) supplemented with 10% (v/v) fetal bovine serum (Sigma-Aldrich) and 1% (v/v) penicillin/streptomycin (PAA Laboratories) with a Bac to Bac Expression System (Invitrogen). Cells were infected at a density of 2 × 106 to 3 × 106cells per millilitre with baculovirus at an approximate multiplicity of infection of 1. Cultures were grown at 27 °C with constant shaking and harvested by centrifugation 72 h after infection.
All subsequent steps were performed at 4 °C unless otherwise stated. Membranes were prepared by resuspension of cells in PBS supplemented with cOmplete Protease Inhibitor Cocktail tablets (Roche), 10 mM magnesium chloride and 5 μg/ml DNaseI (Roche) followed by disruption using a microfluidizer at 60,000 pounds per square inch (M-110L Pneumatic, Microfluidics). Membranes were collected by ultracentrifugation at 204,700g, resuspended in 50 mM HEPES–NaOH pH 7.5, 250 mM NaCl with cOmplete Protease Inhibitor Cocktail tablets (Roche), and stored at −80 °C until use.
To purify the receptor, membranes were thawed at room temperature and incubated with 10 μM vercirnon for 30 min before solubilization with 1.5% (w/v) n-decyl-β-d-maltopyranoside (DM) for 1 h. Insoluble material was removed by ultracentrifugation at 204,700g and the receptors were immobilized by batch binding to 2.5 ml of NiNTA resin (Qiagen). The resin was packed into an Omnifit column (Kinesis) and washed with ten column volumes of 20 mM HEPES–NaOH pH 7.5, 250 mM NaCl, 0.15% (w/v) n-decyl-β-d-maltopyranoside DM, and 10 μM vercirnon then for ten column volumes with the same buffer supplemented with 64 mM imidazole before bound material was eluted in buffer containing 400 mM imidazole. The protein was then concentrated using an Amicon Ultra-15 centrifugal concentrator (MerckMillipore), MWCO 50 kDa, and subjected to preparative SEC in 20 mM HEPES–NaOH pH 7.5, 150 mM NaCl, 0.15% (w/v) n-decyl-β-d-maltopyranoside (DM), and 10 μM vercirnon on a Superdex 200 10/300 Increase column (GE Healthcare). Receptor purity was analysed by SDS–polyacrylamide gel electrophoresis and liquid chromatography–mass spectrometry, and receptor monodispersity was assayed by analytical SEC. Fractions containing the pure, monomeric receptor were concentrated to 10–20 mg/ml in a Vivaspin 500 centrifugal concentrator (Sartorius). Protein concentration was determined using the receptor’s calculated extinction coefficient at 280 nm (ε280,calc = 56,225 M−1 cm−1) and confirmed by quantitative amino-acid analysis.
Crystallization
CCR9-StaR(25-340) was crystallized in LCP at 20 °C. The protein was concentrated to ~16 mg/ml and mixed with monoolein (Nu-Check) supplemented with 10% (w/w) cholesterol (Sigma Aldrich) and 10 μM vercirnon using the twin-syringe method31. The final protein:lipid ratio was 40:60 (w/w). Boli (70 nl) were dispensed on 96-well glass bases and overlaid with 800 nl precipitant solution using a Mosquito LCP from TTPLabtech. Rod-shaped crystals (40-80 μm) of CCR9-StaR(25-340) were grown in 100 mM 2-(Bis(2-hydroxyethyl)amino)acetic acid (BICINE) at a pH range of 7.9–8.0, 200 mM sodium malonate, 28–43% (v/v) polyethylene glycol 400, 10 mM ammonium formate/ammonium nitrate/magnesium formate and 10 μM vercirnon. Single crystals were mounted for data collection and cryo-cooled in liquid nitrogen without the addition of further cryoprotectant. A complete dataset to 2.8 Å was obtained by merging diffraction data from ten crystals belonging to the triclinic space group P1.
Diffraction data collection and processing
X-ray diffraction data were measured on a Pilatus3 6M detector at Diamond Light Source beamline I24 using a beam size of 6 × 8 μm diameter. Crystals displayed diffraction initially out to 2.7 Å after exposure to a beam attenuated down to 60% for 0.12 s per degree of oscillation. It was possible to collect approximately 25° of useful data from each crystal before radiation damage became severe. Further attenuation down to 30% of beam allowed collection of about 60° of useful data. Data from individual crystals were integrated using XDS32. Data merging and scaling was performed using the program AIMLESS from the CCP4 suite33,34. Data collection statistics are reported in Extended Data Table 1.
Structure solution and refinement
The structure of CCR9-StaR(25-340) was solved by molecular replacement with the program Phaser35 using truncated CCR5 (Protein Data Bank accession number 4MBS) as the search model looking for two copies. Here the fusion protein rubredoxin was removed from the CCR5 structure. Manual model building was performed in COOT36 using sigma-A-weighted 2m|Fo| − |DFc|, m|Fo| − D|Fc| maps together with simulated-annealing and simple composite omit maps calculated using Phenix37. Initial refinement was performed with REFMAC5 (ref. 38) using maximum-likelihood restrained refinement in combination with the jelly-body protocol. Further and final stages of refinement were performed with Phenix.refine39 with positional, individual isotropic B-factor refinement and TLS. The final refinement statistics are presented in Extended Data Table 1. Coordinates and structure factors have been deposited in the Protein Data Bank under accession number 5LWE.
Structure analysis
Structures were superposed and aligned for comparison purposes using the program COOT36 to generate global structural superpositions. Figures were prepared using PyMOL (Schrödinger, New York).
The CCR9/vercirnon structure was prepared with the Protein Preparation Wizard method in Maestro version 10.6 (Schrödinger, New York). Hydrogen atoms were energy minimized using the OPLS3 force field. The wild-type molecular model was created in Maestro by changing the StaR mutations to the correspondent wild-type residues. The system was embedded in an equilibrated POPC (1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine) bilayer and parameterized using the OPLS3 force field using the System Builder in Maestro. After the Relax protocol, the system was equilibrated for 100 ns molecular dynamics simulation using Desmond 4.6 (Desmond Molecular Dynamics System, D. E. Shaw Research, New York). The molecular dynamics was performed at 300K/1atm in the NPT ensemble using a Nose-Hoover thermostat and a Martyna–Tobias–Klein barostat40 with a 2.0 ps relaxation time. Coulomb interactions were evaluated using a 9 Å short-range cut-off and smooth particle mesh Ewald as long-range method (Ewald tolerance = 10−9). The resulting molecular dynamics trajectories were analysed with the simulation interactions diagram method in Maestro.
Data availability statement
Coordinates and structure factors have been deposited in the Protein Data Bank under the accession code 5LWE. All other data are available from the corresponding author upon reasonable request.
References
Pease, J. E. Targeting chemokine receptors in allergic disease. Biochem. J. 434, 11–24 (2011)
Wilkin, T. J. & Gulick, R. M. CCR5 antagonism in HIV infection: current concepts and future opportunities. Annu. Rev. Med. 63, 81–93 (2012)
Vela, M., Aris, M., Llorente, M., Garcia-Sanz, J. A. & Kremer, L. Chemokine receptor-specific antibodies in cancer immunotherapy: achievements and challenges. Front. Immunol. 6, 12 (2015)
Solari, R., Pease, J. E. & Begg, M. “Chemokine receptors as therapeutic targets: why aren’t there more drugs?”. Eur. J. Pharmacol. 746, 363–367 (2015)
Pease, J. & Horuk, R. Chemokine receptor antagonists. J. Med. Chem. 55, 9363–9392 (2012)
Wendt, E. & Keshav, S. CCR9 antagonism: potential in the treatment of inflammatory bowel disease. Clin. Exp. Gastroenterol. 8, 119–130 (2015)
Changeux, J. P. & Christopoulos, A. Allosteric modulation as a unifying mechanism for receptor function and regulation. Cell 166, 1084–1102 (2016)
Serrano-Vega, M. J., Magnani, F., Shibata, Y. & Tate, C. G. Conformational thermostabilization of the β1-adrenergic receptor in a detergent-resistant form. Proc. Natl Acad. Sci. USA 105, 877–882 (2008)
Robertson, N. et al. The properties of thermostabilised G protein-coupled receptors (StaRs) and their use in drug discovery. Neuropharmacology 60, 36–44 (2011)
Walters, M. J. et al. Characterization of CCX282-B, an orally bioavailable antagonist of the CCR9 chemokine receptor, for treatment of inflammatory bowel disease. J. Pharmacol. Exp. Ther. 335, 61–69 (2010)
Tan, Q. et al. Structure of the CCR5 chemokine receptor-HIV entry inhibitor maraviroc complex. Science 341, 1387–1390 (2013)
Wu, B. et al. Structures of the CXCR4 chemokine GPCR with small-molecule and cyclic peptide antagonists. Science 330, 1066–1071 (2010)
Hollenstein, K. et al. Structure of class B GPCR corticotropin-releasing factor receptor 1. Nature 499, 438–443 (2013)
Jazayeri, A. et al. Extra-helical binding site of a glucagon receptor antagonist. Nature 533, 274–277 (2016)
Tchernychev, B. et al. Discovery of a CXCR4 agonist pepducin that mobilizes bone marrow hematopoietic cells. Proc. Natl Acad. Sci. USA 107, 22255–22259 (2010)
Andrews, G., Jones, C. & Wreggett, K. A. An intracellular allosteric site for a specific class of antagonists of the CC chemokine G protein-coupled receptors CCR4 and CCR5. Mol. Pharmacol. 73, 855–867 (2008)
Gonsiorek, W. et al. Pharmacological characterization of Sch527123, a potent allosteric CXCR1/CXCR2 antagonist. J. Pharmacol. Exp. Ther. 322, 477–485 (2007)
Salchow, K. et al. A common intracellular allosteric binding site for antagonists of the CXCR2 receptor. Br. J. Pharmacol. 159, 1429–1439 (2010)
Lazaar, A. L. et al. SB-656933, a novel CXCR2 selective antagonist, inhibits ex vivo neutrophil activation and ozone-induced airway inflammation in humans. Br. J. Clin. Pharmacol. 72, 282–293 (2011)
Nicholls, D. J. et al. Identification of a putative intracellular allosteric antagonist binding-site in the CXC chemokine receptors 1 and 2. Mol. Pharmacol. 74, 1193–1202 (2008)
de Kruijf, P. et al. Nonpeptidergic allosteric antagonists differentially bind to the CXCR2 chemokine receptor. J. Pharmacol. Exp. Ther. 329, 783–790 (2009)
Thelen, M. Dancing to the tune of chemokines. Nature Immunol. 2, 129–134 (2001)
Deupi, X. & Standfuss, J. Structural insights into agonist-induced activation of G-protein-coupled receptors. Curr. Opin. Struct. Biol. 21, 541–551 (2011)
Tehan, B. G., Bortolato, A., Blaney, F. E., Weir, M. P. & Mason, J. S. Unifying family A GPCR theories of activation. Pharmacol. Ther. 143, 51–60 (2014)
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011)
Kang, Y. et al. Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser. Nature 523, 561–567 (2015)
Ungashe, S. et al. Aryl sulphonamides. US patent 2006/0111351A1 (2006)
Nugent, T. & Jones, D. T. Membrane protein orientation and refinement using a knowledge-based statistical potential. BMC Bioinformatics 14, 276–285 (2013)
Alva, V. et al. The MPI bioinformatics Toolkit as an integrative platform for advanced protein sequence and structure analysis. Nucleic Acids Res. 44 (Suppl. W1), W410–W415 (2016)
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006)
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nature Protocols 4, 706–731 (2009)
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D 66, 133–144 (2010)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D 69, 1204–1214 (2013)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011)
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012)
Martyna, G. J., Tobias, D. J. & Klein, M. L. Constant pressure molecular dynamics algorithms. J. Chem. Phys. 101, 4177–4189 (1994)
Diederichs, K. & Karplus, P. A. Better models by discarding data? Acta Crystallogr. D 69, 1215–1222 (2013)
Acknowledgements
We thank D. Axford, R. Owen and D. Sherrell at I24, Diamond Light Source, Oxford, UK, for technical support. We thank colleagues at Heptares Therapeutics for suggestions and comments, and G. Brown and S. Bucknell for assistance in radioligand preparation.
Author information
Authors and Affiliations
Contributions
J.K. and A.J. devised and performed the conformational thermostabilization and mutagenesis of the receptor, characterized expression constructs and performed radioligand binding analysis of mutants. Computational analysis of the structure and modelling was performed by F.D. and J.S.M. A.S.D. established the platform/protocols for LCP crystallization and solved the structure. J.C.E. supported expression and scouted purification of the final StaR. M.R. designed and characterized all constructs, collected and processed X-ray diffraction data and solved the structure. C.O. optimized purification, performed LCP crystallization, harvested crystals, collected and processed X-ray diffraction data, and solved and refined the structure. K.B. performed and analysed the pharmacology data. J.A.C. and M.C. identified and sourced the chemical compound(s) used in the study. Project management was performed by J.A.C., R.M.C. and F.H.M. The manuscript was prepared by A.S.D., C.O., F.D., M.C. and F.H.M. All authors contributed to the final editing and approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors are employees of Heptares Therapeutics and are shareholders of Sosei Group, the parent company of Heptares. Heptares is a drug discovery and development company working in the field of G-protein-coupled receptor structure-based drug design.
Extended data figures and tables
Extended Data Figure 1 CCR9 crystallization construct StaR(25-340) in schematic representation.
Thermostabilizing mutations (green) are Thr77Ala, Val79Ala, Met82Ala, Ser141Cys, Thr216Ala, Val255Ala, Asn294Ala, Thr304Ala. Further mutations to remove sites of post-translational modifications (light blue) are Cys337Ala and Thr34Glu. Residues forming the allosteric pocket are pink. Disordered residues in the structure are grey. The disulfide bonds between (Cys1193.25) and extracellular loop 2 and linking the N terminus (Cys38) with the top of TM7 (Cys2897.25) are denoted by dashed yellow lines.
Extended Data Figure 2 Comparison of wild-type and thermostabilized CCR9 in radioligand binding of [3H]vercirnon.
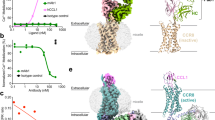
The thermal stability of wild-type CCR9 (filled circles) and CCR9-StaR(1-369) (open circles) analysed in decyl-maltoside are shown. Error bars are derived from standard deviations and calculated from duplicate temperature points (n = 2) within a single experiment. Data shown are representative of three independent experiments. CCR9-StaR(1-369) produced a mean Tm of 39.5 °C. The Tm of wild-type CCR9 was not determinable under these conditions; however, binding is observed and a Tm can be calculated in dodecyl-maltoside (data not shown).
Extended Data Figure 3 Pharmacology of WT CCR9 and CCR9-StaR.
Saturation binding experiments performed in membranes from HEK293 cells transiently expressing (a) human CCR9 or (b) CCR9-StaR(1-369). Non-specific binding was determined by addition of 1 μM cold vercirnon. Data shown as mean ± s.e.m. are representative of three independent experiments performed in duplicate. Data were fitted globally to a one-site saturation isotherm. Affinity and expression level (Bmax) values are given below the graphs for both WT CCR9 and CCR9-StaR(1-369). There was no difference in the affinity of [3H]vercirnon at WT CCR9 or CCR9-StaR(1-369) (unpaired, two-tailed t-test = 0.51). CCR9-StaR(1-369) showed significantly higher expression levels (Bmax) than WT CCR9 (unpaired, two-tailed t-test = 0.0007).
Extended Data Figure 4 Crystal packing in the CCR9-StaR(25-340) triclinic system.
a, Typical CCR9-StaR(25-340) non-fusion crystals grown in LCP complexed with vercirnon. b, The two copies of CCR9-StaR(25-340) in the triclinic asymmetric unit assemble in a parallel fashion with contacts mediated by TM4 – CCR9-StaR(25-340) shown in chainbow colouration (blue to red equals N to C terminus). c, View as in b rotated by 90° with the two copies of CCR9-StaR(25-340) now coloured yellow and cyan. d–f, Views of CCR9-StaR(25-340) packing in the triclinic crystal system along the a, b and c axes respectively, molecules coloured as in c.
Extended Data Figure 6 Electron density around the vercirnon binding site.
a, Cross-eye stereoscopic view of 2Fo − Fc density contoured at 1.6σ covering vercirnon and surrounding residues as viewed from intracellular space. Vercirnon in stick representation, with carbon, nitrogen, chlorine, sulfur and oxygen atoms coloured magenta, blue, green, yellow and red, respectively; CCR9 in stick representation with carbon, nitrogen, oxygen and sulfur atoms coloured white, blue, red and yellow respectively. b, View as in a rotated by 180°.
Extended Data Figure 7 Saturation binding analysis of mutants with [3H]vercirnon.
a–f, Saturation binding of [3H]vercirnon to homogenized cell lysates containing indicated mutant variants of CCR9. Data are representative of three independent experiments performed in duplicate ± s.d. Kd values (inset) are mean of three independent experiments with s.d. in parentheses. The datasets for L87F, Y317A and G321A could not be analysed unambiguously owing to near-complete loss of specific binding.
Extended Data Figure 8 Molecular dynamics analysis of the CCR9–vercirnon complex.
a, Stability of the CCR9–vercirnon complex during 100 ns molecular dynamics. Structural alignment of the wild-type CCR9–vercirnon complex at 0 (blue), 25 (green), 50 (yellow), 75 (orange) and 100 ns (red) molecular dynamics. Proteins are shown as ribbon with TM7 partly hidden for clarity; vercirnon is represented in sticks. b, Two-dimensional representation of the ligand–protein contacts. c, d, Induced-fit binding of vercirnon to CCR9. Superposition of the CCR9–vercirnon complex (cyan) and the pseudo-apo state of CCR9 receptor at 100 ns molecular dynamics (magenta). Vercirnon is shown as sticks with carbons coloured in grey. Arg323, Phe324 and Tyr317 are shown as sticks with carbons coloured in cyan for the crystallographic structure and stick (c) or spheres (d) with carbons coloured in magenta for the molecular dynamics output.
Supplementary information
Supplementary Figure
This file contains Supplementary Figure 1, preparation of [3H]vercirnon and vercirnon. (PDF 39 kb)
Rights and permissions
About this article
Cite this article
Oswald, C., Rappas, M., Kean, J. et al. Intracellular allosteric antagonism of the CCR9 receptor. Nature 540, 462–465 (2016). https://doi.org/10.1038/nature20606
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20606
- Springer Nature Limited
This article is cited by
-
Class B1 GPCR activation by an intracellular agonist
Nature (2023)
-
Allosteric modulation of dopamine D2L receptor in complex with Gi1 and Gi2 proteins: the effect of subtle structural and stereochemical ligand modifications
Pharmacological Reports (2022)
-
Activation pathway of a G protein-coupled receptor uncovers conformational intermediates as targets for allosteric drug design
Nature Communications (2021)
-
G protein-coupled receptors: structure- and function-based drug discovery
Signal Transduction and Targeted Therapy (2021)
-
Ligand-binding and -scavenging of the chemerin receptor GPR1
Cellular and Molecular Life Sciences (2021)