Abstract
In Gram-negative bacteria, outer membrane transporters import nutrients by coupling to an inner membrane protein complex called the Ton complex. The Ton complex consists of TonB, ExbB, and ExbD, and uses the proton motive force at the inner membrane to transduce energy to the outer membrane via TonB. Here, we structurally characterize the Ton complex from Escherichia coli using X-ray crystallography, electron microscopy, double electron–electron resonance (DEER) spectroscopy, and crosslinking. Our results reveal a stoichiometry consisting of a pentamer of ExbB, a dimer of ExbD, and at least one TonB. Electrophysiology studies show that the Ton subcomplex forms pH-sensitive cation-selective channels and provide insight into the mechanism by which it may harness the proton motive force to produce energy.
Similar content being viewed by others
Main
Gram-negative bacteria contain no known energy source located in the outer membrane. To overcome this deficiency, bacteria have evolved systems to harness the energy produced from the proton motive force (PMF) generated at the inner membrane and to transduce it for transport at the outer membrane1,2. An example is the Ton system, which mediates the uptake of metals, carbohydrates, iron-siderophore complexes, cobalamin, and many bacteriocins3,4,5,6. The Ton system consists of two elements: an energy-transducing Ton complex located within the inner membrane, and a ligand-specific TonB-dependent transporter within the outer membrane, which interacts with the ligands to be transported2,7,8,9 (Fig. 1a). The Ton complex is formed by three integral polytopic membrane proteins: ExbB, ExbD, and TonB. The energy transfer is mediated by a conserved 5–7 residue TonB-box at the N terminus of all TonB-dependent transporters2,10. Upon ligand binding to the TonB-dependent transporter, the TonB box becomes exposed and the interaction with TonB leads to conformational changes in the TonB-dependent transporter that are coupled to ligand transport across the outer membrane.
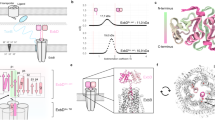
a, Schematic of the Ton system for energy transduction. IM, inner membrane; OM, outer membrane; TBDT, TonB-dependent transporter. b, SEC profiles of the Ton subcomplexes (1, ExbB–ExbD; 2, ExbB–ExbDΔperi; representative purification from 50 or 30 experiments, respectively). c, SDS–PAGE analysis of the Ton subcomplexes purified in b. d, Cartoon representation of the ExbB monomer, consisting of seven α-helices. Peri, periplasm; Cyto, cytoplasm. e, The ExbB pentamer structure shown as cartoon and transparent surface. f, Perpendicular view of the cytoplasmic end of the ExbB pentamer depicting the five-fold symmetry with each edge measuring ~45 Å. g, Perpendicular view of the periplasmic end of the ExbB pentamer. h, The ExbB pentamer was analysed with the programs MOLE 2.039 (spheres representation) and HOLE40 (purple dots). i, Perpendicular view of the cavities shown in h to better illustrate the five fenestrations (vents).
ExbB is predicted to contain three transmembrane spanning helices (TMHs) and a large cytoplasmic domain, whereas ExbD and TonB are each predicted to contain a single N-terminal TMH that anchors a large C-terminal periplasmic domain in the inner membrane11,12,13,14,15 (Fig. 1a). The exact stoichiometry of components of the Ton complex has been a matter of debate for years16,17,18,19. Evidence favouring a dynamic mechanism has been reported in which fluorescence anisotropy studies showed that the presence of TonB within the Ton complex sustains a rotational motion that depends on the PMF at the inner membrane20.
The Ton complex is often compared to the Tol complex, which consists of the analogous components TolQ, TolR, and TolA21,22. The Tol complex is required for cell envelope integrity23,24 and to maintain cellular structure during cell division25. Similar to TonB for the Ton system, TolA has been shown to undergo energy-dependent conformational changes20,26,27. The Ton complex is also evolutionarily related to the Mot complex, which drives bacterial flagellar motion22,28,29,30.
To better understand the role of the Ton complex in energy transduction to the outer membrane, we solved crystal structures of the E. coli Ton subcomplex. We further characterized the assembly of the complex using electron microscopy, crosslinking, and DEER spectroscopy, which reveal that the fully assembled Ton complex consists of a pentamer of ExbB, a dimer of ExbD, and at least one TonB.
Crystal structure of the Ton subcomplex
Constructs of the Ton subcomplex (ExbB–ExbD) were purified using a C-terminal 10×His tag on ExbD (Fig. 1b, c) and crystals grown by vapour diffusion (see Methods). Initial phases were calculated using a 5.2 Å Se-SAD (single-wavelength anomalous diffraction) dataset of ExbB–ExbDΔperi, allowing an initial poly-alanine model to be built (Extended Data Fig. 1). This starting model was then used as a search model to solve the structures at pH 4.5 and 7.0 by molecular replacement (Supplementary Table 1).
The structure of the ExbB–ExbDΔperi complex at pH 7.0 was solved to 2.6 Å resolution. However, only ExbB could be built, owing to insufficient density for ExbDΔperi (Extended Data Fig. 2). The ExbB monomer adopts an extended conformation sitting perpendicular to the membrane, consisting of seven α-helices with α2 and α7 measuring 80–100 Å in length and α5 and α6 forming an extended helix (~100 Å) separated by a kink (Fig. 1d). The transmembrane domain consists of three transmembrane helices (α2, α6, and α7) which extend into the cytoplasm to form a 5-helix bundle with cytoplasmic domain 1 and the C-terminal domain.
The quaternary structure of ExbB is a pentamer in which the five transmembrane domains form a transmembrane pore (α6 and α7), while the cytoplasmic domains form a large enclosed cavity extending as far as ~60 Å into the cytoplasm (Fig. 1e–g). The cytoplasmic domain of ExbB retains five-fold symmetry with each edge measuring around 45 Å, while the periplasmic domain is arranged in pseudo-five-fold symmetry with each edge measuring around 35 Å. ExbB forms a large extended cavity (largest pore radius around 11 Å) along the cytoplasmic and transmembrane domains that is open but constricted at each end (pore radius approximately 2 Å on the cytoplasmic side and 4 Å along the transmembrane side; Fig. 1h). Each monomer has approximately 3,000 Å2 of buried surface area with the two adjacent molecules (about 20% of total surface area), indicating a stable oligomeric state. For the cytoplasmic cavity, five side fenestrations are observed that could allow solvent or ion passage (Fig. 1h, i). Sparse electron density indicated that the transmembrane pore of ExbB is probably filled by the TMH of ExbDΔperi; however, this density was too diffuse to allow a model to be built unambiguously (Extended Data Fig. 2). Two ExbB pentamers were observed per asymmetric unit and alignment of these pentamers revealed some helical shifts, possibly indicating a propensity for movement within the membrane domain (Extended Data Fig. 3).
To verify the presence of the TMH of ExbDΔperi within the transmembrane pore of the ExbB pentamer, we solved the structure of ExbB–ExbDΔperi at pH 4.5 to 3.5 Å resolution and observed a single α-helix (Fig. 2a, b, Supplementary Table 1 and Extended Data Fig. 4). An extended α-helix could be built consisting of residues 22–45, which correlated well with the hydrophobic residues inside the transmembrane pore of ExbB, although it was offset by about 10 Å from the position of the transmembrane domains of ExbB, which are predicted to be embedded into the membrane. The exact position of each residue was less precise owing to the lack of well-defined side-chain density. These results suggest that movements of the TMH of ExbDΔperi may be modulated by changes in pH (Extended Data Fig. 5).
a, The ExbB–ExbDΔperi complex highlighting the transmembrane helix of ExbD (blue) located within the transmembrane pore of the ExbB pentamer (grey). b, Residues from helices α6 and α7 line the transmembrane pore of ExbB (grey) and mediate interactions with the transmembrane helix of ExbD (blue). For clarity, only two monomers of the ExbB pentamer are shown. c, The cytoplasmic domain of ExbB forms a large enclosed cavity that includes 12 arginines, 6 lysines, 11 glutamates, and 7 aspartates from each monomer (acidic residues in red, basic residues in blue). d, Electronegatively charged residues E105 and E109 line the cytoplasmic pore and interact with a single calcium ion (green). Left, expanded view of dashed box on right. e, Electrostatic surface representation of ExbB showing the electropositive ‘belt’ and the electronegative ‘cap’. Blue and red shades indicate electropositivity (blue) or electronegativity (red). f, Cutaway view showing the electrostatic surface properties of the inside cavity.
A striking feature of the ExbB pentamer is the very large cytoplasmic domain and its electrostatic properties, which include a strongly electropositive ‘basic belt’ that sits close to the membrane interface and a strongly electronegative ‘cap’ that sits at the cytoplasmic end of the structure (Fig. 2c–f). For the basic belt, each monomer contributes six lysine residues at positions 44, 52, 56, 81, 108, and 206 and twelve arginine residues at positions 53, 54, 57, 66, 110, 114, 117, 118, 124, 128, 200, and 222. For the cap, each monomer contributes seven aspartates at positions 73, 77, 102, 103, 211, 223, and 225, and 11 glutamate residues at positions 47, 58, 64, 90, 94, 96, 99, 105, 109, 116, and 227. Residues E105 and E109 line the cytoplasmic pore, where we observed a single calcium ion in our structure (Fig. 2c, d).
ExbB is a pentamer in the Ton complex
Negative stain electron microscopy was performed on 2D crystals of the full-length ExbB–ExbD complex (Fig. 3a). The best images were used to generate an averaged 2D projection map of the unit cell, which revealed five domains arranged as a pentamer, each with a diameter of 20–25 Å and with the edges of the pentamer measuring about 45 Å (Fig. 3b and Extended Data Fig. 6).
a, Electron microscopy analysis was performed using 2D crystals (left) of the Ton subcomplex with a power spectrum out to ~30 Å (top right). Five images were analysed, and a representative averaged projection map calculated from 900 sub-images shows that the complex is pentameric (bottom right). b. The electron microscopy studies are consistent with ExbB being a pentamer with edges measuring ~45 Å. c. DEER spectroscopy was performed on the Ton subcomplex labelled with MTSL at position C25 of ExbB. The experimentally measured traces and distance distributions (inset, red lines) agree well with those calculated from the in silico labelled ExbB (black dashed lines). d, DEER analyses of the Ton subcomplex are consistent with ExbB being a pentamer. e, Purification of the fully assembled Ton complex (orange) compared with the subcomplex (blue). f, Comparison of distance distributions of the fully assembled Ton complex (solid orange line) to those of the Ton subcomplex in DDM lacking TonB (dashed red line) showed minimal differences. c, e, and f show data from single experiments.
The Ton subcomplex was also studied using DEER spectroscopy, in which ExbB was labelled at C25 using the spin label S-(1-oxyl-2,2,5,5-tetramethyl-2,5-dihydro-1H-pyrrol-3-yl)methyl methanesulfonothioate (MTSL). Using this method, distance distributions were obtained experimentally and compared to simulations of the in silico labelled crystal structure (Fig. 3c and Extended Data Fig. 7). The experimental results agree well with the simulated distances, with peaks at approximately 35 and 50–60 Å (Fig. 3d). Together with the crystal structure and electron microscopy studies, these results further verify the stoichiometry of ExbB as a pentamer within the Ton subcomplex containing a centralized transmembrane pore (Fig. 3b, d).
To determine the oligomeric state of ExbB in the presence of TonB, the fully assembled Ton complex was expressed and purified, and found to have a larger hydrodynamic radius than the ExbB–ExbD subcomplex (Fig. 3e and Extended Data Fig. 8). We then labelled ExbB at position 25 with MTSL and repeated the DEER spectroscopy analysis. The distance distributions were nearly identical to those of the subcomplex (Fig. 3f and Extended Data Fig. 7), confirming that ExbB is a pentamer in both the absence and the presence of TonB.
ExbD is a dimer in the Ton complex
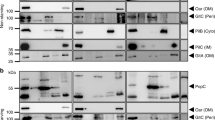
Previous studies have suggested that the Ton complex may contain a dimer of ExbD31. To investigate this possibility, we engineered an ExbBC25S–ExbDE113C construct of the Ton subcomplex. The sample was incubated with the crosslinker 1,8-bismaleimidodiethylenglycol (BM(PEG)2) and then separated by size exclusion chromatography (SEC) and compared to a control sample that was not crosslinked (Fig. 4a). SDS–PAGE analysis confirmed the shift of ExbD from monomer to dimer for the crosslinked sample; however, no shift was induced by SEC, indicating that the ExbD-crosslinked dimer was formed within a single complex (intra) rather than between two different complexes (inter).
a, Crosslinking studies targeting ExbD are consistent with a dimer within the Ton subcomplex, as evidenced by an observed crosslinked dimer (red, lane 2) compared to the non-crosslinked sample (blue, lane 1). b, c, DEER spectroscopy was performed on ExbD labelled at position 78 (purple lines, b) and position 113 (cyan lines, c). The experimentally measured traces and distance distributions (insets, purple and cyan lines) are consistent with those calculated (black dashed lines) from the in silico labelled ExbD dimer model (PDB ID 2PFU), which is based on the reported TolR structure (PDB ID 2JWK). d, The distance measurements within the in silico labelled ExbD dimer model are in agreement with those obtained experimentally at each site using DEER analysis. e, DEER spectroscopy was performed in DDM on the fully assembled Ton complex labelled at position 78 on ExbD. Comparison of distance distributions of the fully assembled Ton complex (solid orange line) to those of the Ton subcomplex in DDM lacking TonB (dashed purple line) show minimal differences. a–c and e show data from single experiments.
DEER spectroscopy was performed on the subcomplex by labelling ExbD at residues 78 and 113 individually, and constructs of ExbBC25S–ExbDN78C and ExbBC25S–ExbDE113C were labelled with MTSL. Distance distributions were detected experimentally and compared to simulations of an in silico labelled model of the ExbD dimer (PDB ID 2PFU)15, which was based on the related TolR dimer structure (PDB ID 2JWK)32 (Fig. 4b, c and Extended Data Fig. 7). According to the dimer model, labelling at residue 78 would yield distances of 32–44 Å, which is consistent with the peaks observed experimentally at 35 and 43 Å (Fig. 4b, d). Furthermore, labelling at residue 113 would yield distances of 15–35 Å, which is also consistent with the peaks observed experimentally at 23 and 34 Å, within the accuracy of the rotamer library approach (Fig. 4c, d and Extended Data Fig. 7).
To determine the oligomeric state of ExbD in the presence of TonB, DEER spectroscopy was performed on the fully assembled Ton complex containing the TonBC18A, ExbBC25S and ExbDN78C mutations and labelled with MTSL. The distance distributions for the labels on ExbD were nearly identical to those of the subcomplex (Fig. 4e and Extended Data Fig. 7), confirming that ExbD is a dimer in both the absence and the presence of TonB.
Ton subcomplex channel properties
To investigate ion conduction by the Ton subcomplex (ExbB–ExbD)27,33, the subcomplex was reconstituted into liposomes that were fused with a preformed planar bilayer membrane34. Single- and multichannel recordings revealed that channels formed by the Ton subcomplex display a conductance of 120 ± 30 pS at neutral pH (Fig. 5a, b), whereas channels formed by the ExbB pentamer are nearly twice as large with a conductance of 220 ± 50 pS (Fig. 5b). This is consistent with our structure, which shows the transmembrane helix of ExbD plugging the transmembrane pore of the ExbB pentamer.
a, Representative spectra for single-channel measurements of the Ton subcomplex (n = 15). b, Representative spectra of multichannel measurements performed on the Ton subcomplex (blue) and ExbB alone (green; n = 15 for each sample). c, Dependence of the macroscopic current amplitude on pH for the Ton subcomplex (blue) and the D25A mutation in the TM helix of ExbD (green) with a holding potential of +50 mV (circles and squares) or −50 mV (triangles and diamonds). d, Dependence of single-channel conductance of the Ton subcomplex on pH measured at +50 mV. Bars show mean ± s.d. with n = 5. e, Structural model of the Ton complex consisting of a pentamer of ExbB, a dimer of ExbD, and at least one TonB. LPS, lipopolysaccharide. f, The ExbB–ExbDΔperi complex showing electric fields for the ExbB pentamer. The modelled location of residue D25 of the transmembrane helix of ExbD (purple) is indicated as a red sphere. g, Proposed mechanisms for how the Ton subcomplex may use the PMF to produce energy.
We also determined the ion selectivity of the channels. Channels of the Ton subcomplex have a pronounced cation selectivity with sevenfold greater permeability for K+ than for Cl− (Vrev, 24.7 ± 0.9 mV; pK+/pCl−, 7.0 ± 0.9) (Supplementary Table 2). Channels formed by the ExbB–ExbDΔperi complex are less cation selective (Vrev, 13.7 ± 4.5 mV; pK+/pCl−, 2.6 ± 1.0), which implies that the periplasmic domain of ExbD enhances cation selectivity. However, the ExbB pentamer is anion-selective (Vrev, −12.6 ± 2.8 mV; pK+/pCl−, 0.43 ± 0.09) (Supplementary Table 2), indicating that ExbDΔperi is sufficient to serve as a cation-selective filter. The point mutation D25A in the transmembrane helix of ExbD, which sits in the pore of the ExbB pentamer, markedly decreases the cation selectivity of the Ton subcomplex (Vrev, 17.0 ± 1.5 mV; pK+/pCl−, 3.3 ± 0.5) (Supplementary Table 2), indicating that D25 makes a substantial contribution toward ion selectivity.
The channel activity of the Ton subcomplex has a pronounced pH dependence, showing a marked decrease in transmembrane current upon a decrease in pH from neutral to acidic (Fig. 5c). However, the transmembrane helix of ExbD is not the major contributor to the observed pH dependence, as the D25A mutant shows a nearly identical pH dependence to that of the wild type (Fig. 5c), suggesting that the unique electrostatic properties of the ExbB pentamer may be responsible. The decrease in transmembrane current amplitude in the pH range 4.5–8.0 is explained by a decrease in single-channel conductance from 120 pS at pH 8.0 to 70 pS at pH 4.5 (Fig. 5d). Below pH 4.5, the decrease in transmembrane current is caused by channel closure at both positive and negative potentials (Fig. 5c). The ion channel conductance properties of the Ton subcomplex demonstrate that it is being modulated by pH, possibly through movement of the transmembrane helix of ExbD within the transmembrane pore of the ExbB pentamer, such that at low pH, the transmembrane helix of ExbD is in a more closed/fixed conformation (Fig. 2b).
Model of a fully assembled Ton complex
On the basis of our findings, we propose a model in which the Ton complex consists of a pentamer of ExbB, a dimer of ExbD, and at least one TonB (Fig. 5e and Extended Data Fig. 9). As only a single transmembrane helix can fit within the transmembrane pore of ExbB, and dimerization of ExbD is hypothesized to be mediated by its periplasmic domain, we propose that a second copy of ExbD is located outside the ExbB pentamer. Previous studies have indicated that TonB may exchange for one of the ExbD monomers during energy transduction17. However, our studies show that association of TonB does not notably affect the structure or stoichiometry of ExbB or ExbD within the Ton complex. The interaction of TonB with ExbD leads to a functional Ton complex, triggering energy production and transduction in the form of conformational changes in TonB that lead to ligand uptake by the transporter at the outer membrane35,36.
The Ton complex relies on the PMF for its function27,33 and it has been proposed that the Ton complex acts as a proton-conducting channel that shuttles protons from the periplasm to the cytoplasm and that this powers a mechanical motion within the complex20. Mutagenesis studies have previously identified a number of residues that are necessary for harnessing the PMF, including D25 of ExbD and T148 and T181 of ExbB37,38. These residues all map to the interior of the transmembrane pore of ExbB, where protons would be translocated (Extended Data Fig. 10). Our studies indicate that the transmembrane helix of ExbD is quite dynamic within the transmembrane pore of ExbB, and together with the electrophysiology experiments, show that this dynamic behaviour can be modulated by pH. The electrostatics of the ExbB pentamer may also create an ‘electrostatic funneling’ effect that helps to draw protons from the periplasm and steer them through the transmembrane pore of ExbB into the cytoplasm (Fig. 5f). Therefore, we suggest two plausible mechanistic models for how the Ton complex harnesses the PMF for energy production and transduction (Fig. 5g). The first is the ‘electrostatic piston’ model, in which the transmembrane helix of ExbD moves translationally within the transmembrane pore of ExbB, thereby creating a piston-like motion. The second is the ‘rotational’ model, in which the transmembrane helix of ExbD rotates within the transmembrane pore of ExbB, creating rotational motion. A combination of the two mechanistic models is also plausible. While we observe minor conformational shifts within the transmembrane helices of ExbB in our structures, it is also feasible that the ExbB pentamer cycles through more pronounced conformations to either drive or accommodate the dynamics of the transmembrane helix of ExbD.
Methods
Cloning of E. coli (K-12 strain) ExbB, ExbD, and TonB constructs and mutants
The ExbB construct with and without a C-terminal 6×His tag was subcloned into pET26b (Novagen). ExbD was subcloned into pACYCDuet-1 vector (Novagen) with an N-terminal Strep-tag and a C-terminal 10×His tag. ExbD was also subcloned into a pCDF-1b vector (Novagen) containing a C-terminal TEV protease site followed by a 10×His tag. An ExbDΔperi construct containing a C-terminal TEV protease site (preceded by a Gly-Gly-Gly linker for efficient digestion by TEV protease) followed by a 10×His tag was constructed by deletion of the sequence encoding the periplasmic domain of ExbD (residues 50–141). TonB was cloned into a pACYCDUET-1 vector with an N-terminal 10×His tag followed by a TEV protease site. Mutants of TonB (C18A), ExbD (D25A, N78C and E113C), and ExbB (C25S) were prepared by site-directed mutagenesis (primer sequences for all cloning and mutagenesis experiments are available upon request). The sequences of all plasmid constructs and mutations were verified by sequence analysis (Macrogen USA and Eurofins Genomics GmbH).
Expression and purification of the Ton complex, subcomplexes, and components
Expression of ExbB with a C-terminal 6×His tag was performed by transforming E. coli BL21(DE3) cells (NEB) with the pET26b/ExbB vector. Co-expression was performed by co-transforming E. coli BL21(DE3) cells with the respective ExbB, ExbD, and/or TonB plasmids. For all transformations, cells were plated onto LB agar plates supplemented with appropriate antibiotics. Colonies were then used for a starter culture to inoculate 12 flasks containing either 1 l 2×YT medium (Ton subcomplex) or SelenoMet medium supplemented with l-methionine at 40 mg/l (Molecular Dimensions) (Ton complex), with appropriate antibiotics. Cultures were grown at 37 °C with shaking at 220 r.p.m. until they reached an OD600 of 0.5–1.0, induced with isopropyl β-d-1-thiogalactopyranoside (IPTG) to 0.1 mM final concentration, and then allowed to continue to grow overnight at 28 °C. For selenomethionine-substituted samples for experimental phasing, B834(DE3) cells (NEB) were co-transformed with pET26b/ExbBC25S and pCDF-1b/ExbDΔperi plasmids. Single colonies were used to inoculate 12 flasks containing 1 l SelenoMet medium (Molecular Dimensions) supplemented with 40 mg/ml l-selenomethionine and appropriate antibiotics. Cultures were grown at 37 °C with shaking at 220 r.p.m. until they reached an OD600 of 0.5–1.0, induced with IPTG to 0.1 mM final concentration, and then allowed to continue to grow overnight at 28 °C. Cells were harvested and used immediately or stored at −80 °C.
For purification, cells were resuspended in either 1×PBS (Ton subcomplex) or TBS (Ton complex) supplemented with 100 μM 4-(2-aminoethyl)benzenesulfonyl fluoride (AEBSF), 100 μM DNase, and 50 μg/ml lysozyme, and disrupted with two passages through an EmulsiFlex-C3 (Avestin) operating at ~15,000 p.s.i. Membranes were pelleted by ultracentrifugation in a Type 45 Ti Beckman rotor at 200,000g for 1 h at 4 °C. Membranes were then resuspended in 1×PBS or TBS using a dounce homogenizer and solubilized by the addition of Triton X-100 (Ton subcomplex) or DDM (Anatrace) (Ton complex) to a final concentration of 1% by stirring at medium speed for 1 h to overnight at 4 °C. Insoluble material was pelleted by ultracentrifugation in a Type 45 Ti Beckman rotor at 200,000g for 1 h at 4 °C and the supernatant was used immediately.
Immobilized metal affinity chromatography (IMAC) was performed on an AkTA Purifier (GE Healthcare) using a 15-ml Ni-NTA agarose column (Qiagen) equilibrated with 1×PBS or TBS supplemented with 0.1% Triton X-100 or 0.1% DDM. The supernatant was supplemented with 10 mM imidazole and loaded onto the column. The column was washed in three steps with 1×PBS or TBS supplemented with 20, 40 and 60 mM imidazole, respectively, and eluted with 1×PBS or TBS supplemented with 250 mM imidazole in 2-ml fractions. Fractions were analysed by SDS–PAGE and those fractions containing the complex were pooled. To remove the 10×His tag, TEV protease was added to the sample at 0.1 mg/ml final concentration and rocked overnight at 4 °C. For the Ton complex, the sample was then diluted 2–3 times with 25 mM HEPES, pH 7.3, and 0.1% DDM and loaded onto an anion exchange 6-ml ResourceQ column (GE Healthcare). Elution was performed with a 0–1 M NaCl gradient over 5 column volumes. For the Ton subcomplex, the sample was concentrated using an Amicon Ultra-15 Centrifugal Filter Unit with a 50-kDa MW cut-off (Millipore), filtered, and purified by size-exclusion chromatography using a Superdex 200 HL 16/600 column (GE Healthcare) at a flow rate of 0.5–1.0 ml/min. The buffer consisted of 20 mM HEPES-NaOH, pH 7.0, 150 mM NaCl, 0.01% NaN3, and 0.08% C10E5. For the Ton complex, eluted fractions were concentrated using an Amicon Ultra-15 Centrifugal Filter Unit with a 100-kDa MW cut-off (Millipore), and passed over a Superose6HR 10/30 column (GE Healthcare) at a flow rate of 0.5 ml/min using 20 mM HEPES-NaOH, pH 7.0, 150 mM NaCl, and 0.05% DDM.
Densitometry analysis was performed using ImageJ software41.
Circular dichroism
Far-UV circular dichroism (CD) spectra (185–260 nm) were measured in 0.1 M NaPi, pH 7.0, and 0.03% DDM using quartz cuvettes with a 0.02–0.2 mm optical path length. The results were analysed using the DichroWeb package of programs42 and different sets of reference proteins, including the SMP180 set of membrane proteins. The analysis of the thermal stability of the complexes reconstituted into liposomes was measured by the temperature dependence of the CD signal amplitude at 222 nm. Thermal melting was performed in a magnetically stirred 1-cm quartz cuvette containing 10 mM HEPES, pH 7.0, and 100 mM NaCl with a rate of temperature increase of 0.5 °C/min. Melting curves were normalized to the measured value of the molar ellipticity change at 10 °C.
Crystallization and data collection
For crystallization, samples were concentrated to ~10 mg/ml and sparse matrix screening was performed using a TTP Labtech Mosquito crystallization robot using hanging drop vapour diffusion and plates incubated at 15–21 °C. Initially, many lead conditions were observed to produce crystals with hexagonal morphology; however, none diffracted to better than ~7 Å and most suffered from anisotropy. To avoid this packing, we performed reductive methylation of our samples before crystallization using the Reductive Alkylation Kit (Hampton Research), followed by an additional size-exclusion chromatography step. This led to a condition which produced diffraction spots to ~4 Å resolution. Further optimization and screening allowed us to grow crystals in 100 mM Na-acetate, pH 4.5, 100 mM MgCl2, and 25% PEG 400 that routinely diffracted to ~3.5 Å resolution or better. For heavy atom soaking, crystals were transferred to a drop containing 1 mM HgCl2 and incubated overnight at room temperature and then harvested directly from the soaking condition. The best native crystals for the ExbB–ExbDΔperi complex, however, were grown from 100 mM HEPES-NaOH, pH 7.0, 100 mM CaCl2, and 22% PEG MME 550 and diffracted to 2.6 Å resolution; these crystals were also used for heavy atom soaking experiments. Unfortunately, none of the heavy atom soaked crystals (nor the selenomethionine substituted crystals) were useful for phasing owing to crystal pathologies, which we suspected were twinning related. However, selenomethionine substituted crystals of the ExbBC25S–ExbDΔperi complex were obtained using 100 mM MES/imidazole, pH 6.5, 30 mM MgCl2, 30 mM CaCl2, 50% ethylene glycol, and 8% PEG 8000 and diffracted to 5.2 Å resolution with no twinning-related issues. Both native and selenomethionine-substituted crystals were harvested directly from the crystallization drops. Screening for diffraction quality was performed at the GM/CA-CAT and SER-CAT beamlines at the Advanced Photon Source at Argonne National Laboratory and at beamlines 5.0.1 and 8.2.1 at the Advanced Light Source at Lawrence Berkeley National Laboratory. Final datasets were collected at the SER-CAT beamline and all data were processed using either HKL200043 or Xia244. A summary of the data collection statistics can be found in Supplementary Table 1. The presence of both components of the Ton subcomplex within the crystals was confirmed by SDS–PAGE and mass spectrometry analyses of harvested crystals.
Structure determination
For phasing the ExbB–ExbDΔperi complex structure, three datasets were collected on selenomethionine substituted crystals of the ExbBC25S–ExbDΔperi complex at a wavelength of 0.979 Å. The data were processed with Xia244 and, based on non-isomorphism, one dataset was removed. The final two datasets were processed together in space group P43212 to a final resolution of 5.2 Å. Selenium sites (35 total) were located using HKL2MAP45 after 5,000 tries within SHELXD at a resolution range of 20–6 Å. The sites were then fed into AutoSol (PHENIX)46 which removed one site, producing a phase-extended density-modified electron density map into which we could build an initial poly-alanine model. Five-fold symmetry was clearly observed, with each monomer consisting of very elongated α-helices, and directionality was determined on the basis of the predicted topology of ExbB, which contains a single large cytoplasmic domain. This model was then used as a search model to solve the native and Hg-soaked structures by molecular replacement using PHASER/PHENIX46,47 and the sequence docked on the basis of anomalous peaks from the SeSAD dataset. The ExbB–ExbDΔperi complex was solved in space group P21 to 2.6 Å resolution with R/Rfree values of 0.21/0.26 and the Hg-soaked structure in space group P212121 to 3.5 Å resolution with R/Rfree values of 0.25/0.30. All model building was performed using COOT and subsequent refinement done in PHENIX46. r.m.s.d. analysis was performed within PyMOL (Schrödinger). Electrostatic surface properties (calculated using the Linearized Poisson-Boltzman Equation mode with a solvent radius of 1.4), including generation of the electric field lines, were analysed and visualized using the APBS plugin within PyMOL (Schrödinger). Buried surface area was calculated using the PDBePISA server48. Structure-related figures were made with PyMOL (Schrödinger) and Chimera49 and annotated and finalized with Adobe Photoshop and Illustrator.
Data availability
Coordinates and structure factors for the ExbB/ExbD complexes have been deposited into the Protein Data Bank (PDB accession codes 5SV0 and 5SV1).
2D crystallization
For 2D crystallization experiments, the Ton subcomplex (ExbB–ExbD) was extracted and purified by IMAC as previously described. The sample was passed over a Superose 12 HR 10/30 column using 20 mM Tris-HCl, pH 7, 150 mM NaCl, 0.01% NaN3, and 0.035% Triton X-100. The purified complex was then mixed with a solution stock of E. coli polar lipid (Avanti Polar Lipids, Inc.) at 10 mg/ml in 2% Triton X-100, to reach final concentrations of 0.5–1.0 mg/ml protein and 0.1–0.4 mg/ml lipid. The lipid-protein-detergent samples solutions were placed into Mini Slide-A-Lyser dialysis devices (Pierce) with a 20-kDa MW cutoff, and dialysed in 1 l of 25 mM Tris-HCl, pH 7.0, 150 mM NaCl, and 0.01% NaN3 at 4 °C. Aliquots of dialysed samples were observed periodically by electron microscopy to monitor the formation of 2D crystals.
Electron microscopy
Sample preparation for electron microscopy was carried out by applying a 5-μl drop of protein-lipid material on a glow discharged carbon-coated electron microscopy grid. Staining was performed by addition of 1% (w/v) uranyl acetate and incubation for 1 min. Grids were then imaged on a Tecnai G2 200 LaB6 electron microscope operating at 200 kV at the Institut de Microbiologie de la Méditerranée. Images were recorded with a 2K Eagle CCD camera.
The best 2D crystals were selected through observation of the power spectrum of the images using ImageJ software41. Selected images were processed using the IPLT Correlation Averaging suite program50. A filtered image was generated by optical filtering of the low resolution spots, and padded to contain only 4–6 unit cells. The padded image was cross-correlated with the original large image. The positions of the cross-correlation peaks were determined and used to extract sub-images that were summed to generate an average image of the 2D unit cell.
DEER spectroscopy
Site-directed spin labelling was used to covalently attach the spin label (1-oxyl-2,2,5,5-tetramethyl-∆3-pyrroline-3-methyl) methanethiosulfonate (MTSL) (Toronto Research Chemicals) to Cys25 on ExbB and to cysteines engineered at positions 78 and 113 on ExbD (N78C, E113C; ExbD constructs were in the pACYC vector containing an N-terminal strep-tag and a C-terminal 10×His tag for the Ton subcomplex, and in the pCDF-1b vector for the Ton complex). For labelling with MTSL, samples were first incubated with 2–10 mM dithiothreitol (DTT) for 1–2 h and the DTT then removed by passage over a HiTrap desalting column (GE Healthcare) or during anion exchange (Ton complex). Samples were then incubated with a 10× molar excess of MTSL overnight at 4 °C and then passed over a Superose 6HR 10/30 gel filtration column (GE Healthcare) using 20 mM HEPES-NaOH, pH 7.5, 200 mM NaCl, 0.08% C10E5 or 0.03% DDM (Ton subcomplex); or 20 mM HEPES-NaOH, pH 7.0, 150 mM NaCl, and 0.05% DDM (Ton complex).
For DEER measurements, the samples were diluted with D2O to a final concentration of 30% and cryoprotected with 10% v/v D8-glycerol before being flash frozen in liquid nitrogen. Continuous wave (CW) electron paramagnetic resonance (EPR) experiments were carried out at room temperature on a bench-top X-band MiniScope MS 400 (Magnettech by Freiberg Instrument) at 9.5 GHz (X-band) with 2.5 mW microwave power, 15 mT sweep width and 0.15 mT modulation amplitude. Spin labelling efficiency was calculated from the second integral of the derivative spectra compared to a standard spin concentration of 100 μM (Tempol in water). The ExbB native cysteine C25 was labelled with a 50% efficiency, while the ExbD mutants were labelled with efficiencies >80%. DEER measurements were initially performed at ETH Zurich on a commercial Bruker ELEXSYS-II E580 Q-band spectrometer (34–35 GHz) and later on a Bruker ELEXSYS E580Q-AWG dedicated pulse Q-band spectrometer operating at 34–35 GHz. Both spectrometers were equipped with a TWT amplifier (150 W) and a home-made rectangular resonator (from ETH Zurich) enabling the insertion of 30–40 μl sample volume in quartz tubes with 3 mm outer diameter51. Dipolar time evolution data were acquired using the four-pulse DEER experiment at 50 K. All pulses were set to be rectangular with 12 ns length, with the pump frequency at the maximum of the echo-detected field swept spectrum, 100 MHz higher than the observer frequency. Deuterium nuclear modulations were averaged by increasing the first interpulse delay by 16 ns for 8 steps as previously described51. The background of the normalized DEER primary data (V(t)/V(0)) was fitted with optimized dimensions from 2.5 to 3.2 and the resulting normalized secondary data (F(t)/F(0)) were converted by model-free Tikhonov regularization to distance distributions with the software DeerAnalysis201552,53. The simulation of the possible spin label rotamers populated at selected positions in the protein was performed using the Matlab program package MMM2015.1 using the MTSL ambient temperature library54.
Crosslinking
The ExbBC25S–ExbDE113C complex (ExbDE113C was in the pACYC vector containing an N-terminal strep-tag and a C-terminal 6×HIS tag) was expressed and purified as described earlier. To prepare the sample for crosslinking, the sample was incubated at 4 °C with 5 mM DTT for at least 1 h. The DTT was then removed using a desalting column in 20 mM HEPES, pH 7.0, 150 mM NaCl, and 0.1% DDM. The crosslinker 1,8-bismaleimidodiethylenglycol (BM(PEG)2) (Pierce) was added at a final concentration of 0.2 mM and the reaction was incubated at 4 °C overnight. The sample was concentrated and passed over a Superose 6HR 10/30 gel filtration column using 20 mM HEPES-NaOH, pH 7.0, 150 mM NaCl, and 0.035% DDM on an AkTA Purifier system (GE Healthcare). The results were visualized by SDS–PAGE analysis.
Reconstitution in liposomes
Protein complexes were reconstituted into liposomes by dialysis of the protein–lipid–detergent mixture. Lipids (DOPG, DOPC and DOPE) dissolved in chloroform were mixed in a molar ratio of 2:3:5. Chloroform was removed by vortexing in a stream of nitrogen gas in a glass tube followed by drying in vacuum for 2–3 h. The lipid film was hydrated in 1 ml TN buffer (10 mM Tris-HCl, pH 7.5, 50 mM NaCl), followed by five cycles of freeze–thaw and sonication using a water bath sonicator until the suspension of lipids became clear (10–15 min). For proteoliposome preparation, small unilamellar vesicles (SUVs) were mixed with octylglucoside (final concentration, 2%) and then proteins added to achieve a molar ratio of total lipid to protein ∼500–2,000 mol/mol. After 1 h incubation in ice, the lipid–protein–detergent mixture was dialysed into 10 mM Tris-HCl, pH 7.5, 0.3 M sucrose, and 50 mM KCl for 30–40 h using a dialysis membrane with a MW cut-off pore size of 10 kDa.
Planar-lipid bilayer measurement of ion-conduction
Mueller-Rudin type planar bilayer membranes were formed on a 0.2-mm diameter aperture in a partition that separates two 1-ml compartments, using a mixture of lipids, DOPG, DOPC and DOPE, at a molar ratio of 2:3:5 (10 mg/ml) in n-decane, applied by a brush technique55. The aqueous solution in both compartments consisted of 2 mM KPi, pH 7.0, and 0.1 M and 0.4 M KCl in the cis- and trans-compartments, respectively. To study the pH dependence of channel activity, bathing solutions were buffered with 2 mM Na-acetate (pK 4.8), Na-cacodylate (pK 6.2), and Tris (pK 8.3). The pH of the bathing solution was changed by adding 10–20 μl 0.1 M HCl or KOH. The cis-side of the planar bilayer is defined as that to which the electrical potential is applied. Proteoliposomes, 0.1–2 μl, were added to the trans-compartment, and the solutions were stirred until the transmembrane current appeared. A large concentration of an osmolyte inside of the liposomes and the transmembrane KCl concentration gradient caused proteoliposome fusion with the pre-formed planar lipid membrane bilayer. The transmembrane current was measured in voltage-clamp mode with Ag/AgCl electrodes and agar bridges, using a BC-525C amplifier (Warner Instruments). The single-channel conductance of the ExbB–ExbD complexes was measured in symmetrical salt conditions: 0.1 M KCl solution, pH 7.5, at a holding potential of +50 or −50 mV. For ion selectivity experiments, zero-current potential (Vrev) was determined from volt-ampere characteristics measured in asymmetric salt conditions. Relative cation/anion permeability was calculated using the Goldman-Hodgkin-Katz equation56.
Change history
05 October 2016
A minor change was made to the reported labelling at residue 113 distance, and to the DEER Spectroscopy section in the Methods.
References
Krewulak, K. D. & Vogel, H. J. TonB or not TonB: is that the question? Biochem. Cell Biol. 89, 87–97 (2011)
Noinaj, N., Guillier, M., Barnard, T. J. & Buchanan, S. K. TonB-dependent transporters: regulation, structure, and function. Annu. Rev. Microbiol. 64, 43–60 (2010)
Bassford, P. J., Jr, Bradbeer, C., Kadner, R. J. & Schnaitman, C. A. Transport of vitamin B12 in tonB mutants of Escherichia coli. J. Bacteriol. 128, 242–247 (1976)
Lohmiller, S., Hantke, K., Patzer, S. I. & Braun, V. TonB-dependent maltose transport by Caulobacter crescentus. Microbiology 154, 1748–1754 (2008)
Schauer, K., Rodionov, D. A. & de Reuse, H. New substrates for TonB-dependent transport: do we only see the ‘tip of the iceberg’? Trends Biochem. Sci. 33, 330–338 (2008)
Cascales, E. et al. Colicin biology. Microbiol. Mol. Biol. Rev. 71, 158–229 (2007)
Postle, K. & Kadner, R. J. Touch and go: tying TonB to transport. Mol. Microbiol. 49, 869–882 (2003)
Clément, E., Mesini, P. J., Pattus, F. & Schalk, I. J. The binding mechanism of pyoverdin with the outer membrane receptor FpvA in Pseudomonas aeruginosa is dependent on its iron-loaded status. Biochemistry 43, 7954–7965 (2004)
Cadieux, N., Barekzi, N. & Bradbeer, C. Observations on the calcium dependence and reversibility of cobalamin transport across the outer membrane of Escherichia coli. J. Biol. Chem. 282, 34921–34928 (2007)
Schramm, E., Mende, J., Braun, V. & Kamp, R. M. Nucleotide sequence of the colicin B activity gene cba: consensus pentapeptide among TonB-dependent colicins and receptors. J. Bacteriol. 169, 3350–3357 (1987)
Brewer, S. et al. Structure and function of X-Pro dipeptide repeats in the TonB proteins of Salmonella typhimurium and Escherichia coli. J. Mol. Biol. 216, 883–895 (1990)
Chang, C., Mooser, A., Plückthun, A. & Wlodawer, A. Crystal structure of the dimeric C-terminal domain of TonB reveals a novel fold. J. Biol. Chem. 276, 27535–27540 (2001)
Witty, M. et al. Structure of the periplasmic domain of Pseudomonas aeruginosa TolA: evidence for an evolutionary relationship with the TonB transporter protein. EMBO J. 21, 4207–4218 (2002)
Ködding, J. et al. Crystal structure of a 92-residue C-terminal fragment of TonB from Escherichia coli reveals significant conformational changes compared to structures of smaller TonB fragments. J. Biol. Chem. 280, 3022–3028 (2005)
Garcia-Herrero, A., Peacock, R. S., Howard, S. P. & Vogel, H. J. The solution structure of the periplasmic domain of the TonB system ExbD protein reveals an unexpected structural homology with siderophore-binding proteins. Mol. Microbiol. 66, 872–889 (2007)
Higgs, P. I., Larsen, R. A. & Postle, K. Quantification of known components of the Escherichia coli TonB energy transduction system: TonB, ExbB, ExbD and FepA. Mol. Microbiol. 44, 271–281 (2002)
Sverzhinsky, A. et al. Membrane protein complex ExbB4-ExbD1-TonB1 from Escherichia coli demonstrates conformational plasticity. J. Bacteriol. 197, 1873–1885 (2015)
Sverzhinsky, A. et al. Coordinated rearrangements between cytoplasmic and periplasmic domains of the membrane protein complex ExbB-ExbD of Escherichia coli. Structure 22, 791–797 (2014)
Pramanik, A. et al. Oligomeric structure of ExbB and ExbB-ExbD isolated from Escherichia coli as revealed by LILBID mass spectrometry. Biochemistry 50, 8950–8956 (2011)
Jordan, L. D. et al. Energy-dependent motion of TonB in the Gram-negative bacterial inner membrane. Proc. Natl Acad. Sci. USA 110, 11553–11558 (2013)
Braun, V. The structurally related exbB and tolQ genes are interchangeable in conferring tonB-dependent colicin, bacteriophage, and albomycin sensitivity. J. Bacteriol. 171, 6387–6390 (1989)
Cascales, E., Lloubès, R. & Sturgis, J. N. The TolQ-TolR proteins energize TolA and share homologies with the flagellar motor proteins MotA-MotB. Mol. Microbiol. 42, 795–807 (2001)
Lloubès, R. et al. The Tol-Pal proteins of the Escherichia coli cell envelope: an energized system required for outer membrane integrity? Res. Microbiol. 152, 523–529 (2001)
Yeh, Y. C., Comolli, L. R., Downing, K. H., Shapiro, L. & McAdams, H. H. The caulobacter Tol-Pal complex is essential for outer membrane integrity and the positioning of a polar localization factor. J. Bacteriol. 192, 4847–4858 (2010)
Gray, A. N. et al. Coordination of peptidoglycan synthesis and outer membrane constriction during Escherichia coli cell division. eLife 4, 07118 (2015)
Germon, P., Ray, M. C., Vianney, A. & Lazzaroni, J. C. Energy-dependent conformational change in the TolA protein of Escherichia coli involves its N-terminal domain, TolQ, and TolR. J. Bacteriol. 183, 4110–4114 (2001)
Larsen, R. A., Thomas, M. G. & Postle, K. Protonmotive force, ExbB and ligand-bound FepA drive conformational changes in TonB. Mol. Microbiol. 31, 1809–1824 (1999)
Zhai, Y. F., Heijne, W. & Saier, M. H., Jr. Molecular modeling of the bacterial outer membrane receptor energizer, ExbBD/TonB, based on homology with the flagellar motor, MotAB. Biochim. Biophys. Acta 1614, 201–210 (2003)
Kojima, S. & Blair, D. F. Conformational change in the stator of the bacterial flagellar motor. Biochemistry 40, 13041–13050 (2001)
Saier, M. H., Jr. Tracing pathways of transport protein evolution. Mol. Microbiol. 48, 1145–1156 (2003)
Ollis, A. A., Manning, M., Held, K. G. & Postle, K. Cytoplasmic membrane protonmotive force energizes periplasmic interactions between ExbD and TonB. Mol. Microbiol. 73, 466–481 (2009)
Parsons, L. M., Grishaev, A. & Bax, A. The periplasmic domain of TolR from Haemophilus influenzae forms a dimer with a large hydrophobic groove: NMR solution structure and comparison to SAXS data. Biochemistry 47, 3131–3142 (2008)
Larsen, R. A. & Postle, K. Conserved residues Ser(16) and His(20) and their relative positioning are essential for TonB activity, cross-linking of TonB with ExbB, and the ability of TonB to respond to proton motive force. J. Biol. Chem. 276, 8111–8117 (2001)
Labarca, P. & Latorre, R. Insertion of ion channels into planar lipid bilayers by vesicle fusion. Methods Enzymol. 207, 447–463 (1992)
Ollis, A. A., Kumar, A. & Postle, K. The ExbD periplasmic domain contains distinct functional regions for two stages in TonB energization. J. Bacteriol. 194, 3069–3077 (2012)
Ollis, A. A. & Postle, K. ExbD mutants define initial stages in TonB energization. J. Mol. Biol. 415, 237–247 (2012)
Braun, V. et al. Energy-coupled transport across the outer membrane of Escherichia coli: ExbB binds ExbD and TonB in vitro, and leucine 132 in the periplasmic region and aspartate 25 in the transmembrane region are important for ExbD activity. J. Bacteriol. 178, 2836–2845 (1996)
Braun, V. & Herrmann, C. Point mutations in transmembrane helices 2 and 3 of ExbB and TolQ affect their activities in Escherichia coli K-12. J. Bacteriol. 186, 4402–4406 (2004)
Sehnal, D. et al. MOLE 2.0: advanced approach for analysis of biomacromolecular channels. J. Cheminform. 5, 39 (2013)
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360, 376 (1996)
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012)
Whitmore, L. & Wallace, B. A. Protein secondary structure analyses from circular dichroism spectroscopy: methods and reference databases. Biopolymers 89, 392–400 (2008)
Minor, Z. O. W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010)
Schneider, T. P. T. R. HKL2MAP: a graphical user interface for phasing with SHELX programs. J. Appl. Crystallogr. 37, 843–844 (2004)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007)
Pettersen, E. F. et al. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Schenk, A. D., Philippsen, A., Engel, A. & Walz, T. A pipeline for comprehensive and automated processing of electron diffraction data in IPLT. J. Struct. Biol. 182, 173–185 (2013)
Polyhach, Y. et al. High sensitivity and versatility of the DEER experiment on nitroxide radical pairs at Q-band frequencies. Phys. Chem. Chem. Phys. 14, 10762–10773 (2012)
Jeschke, G. et al. DeerAnalysis2006—a comprehensive software package for analyzing pulsed ELDOR data. Appl. Magn. Reson. 30, 473–498 (2006)
von Hagens, T., Polyhach, Y., Sajid, M., Godt, A. & Jeschke, G. Suppression of ghost distances in multiple-spin double electron-electron resonance. Phys. Chem. Chem. Phys. 15, 5854–5866 (2013)
Polyhach, Y., Bordignon, E. & Jeschke, G. Rotamer libraries of spin labelled cysteines for protein studies. Phys. Chem. Chem. Phys. 13, 2356–2366 (2011)
Mueller, P., Rudin, D. O., Tien, H. T. & Wescott, W. C. Reconstitution of cell membrane structure in vitro and its transformation into an excitable system. Nature 194, 979–980 (1962)
Hille, B. Ion Channels of Excitable Membranes 3rd edn (Sinauer, 2001)
Acknowledgements
We thank the staff at the SER-CAT and GM/CA-CAT beamlines at the Advanced Photon Source (APS), Argonne National Laboratory (use of the APS is supported by the US DOE, Office of Science, Office of Basic Energy Sciences, under Contract No. W-31-109-Eng-38, and by the US DOE, Basic Energy Sciences, Office of Science, under contract No. DE-AC02-06CH11357); the staff at beamlines 5.0.1 and 8.2.1, Advance Light Source at Lawrence Berkeley National Laboratory for their assistance during crystal screening (the Advanced Light Source is supported by the Director, Office of Science, Office of Basic Energy Sciences, of the US DOE under Contract No. DE-AC02-05CH11231); G. Jeschke (ETH Zurich) for providing the Q-band resonator; T. Assafa for the reproducibility measurements; the Deutsche Forschungsgemeinschaft for funding the AWG E580 Q-band spectrometer (INST 130/972-1 FUGG); Y. Li at the NINDS/NIH Protein/Peptide Sequencing Facility for performing mass spectrometry analysis; and members of the Lloubes team, E. Cascales, J. Sturgis and J. P. Duneau for discussions. N.N. is supported by the Department of Biological Sciences, Purdue University and by the National Institute of Allergy and Infectious Diseases (1K22AI113078-01). E.B. is supported by the Cluster of Excellence RESOLV (EXC 1069) funded by the Deutsche Forschungsgemeinschaft. W.A.C. is supported by the NIH (NIH GM 038323) and the Henry Koffler Professorship at Purdue University. H.C. and R.L. are funded by the Centre National de la Recherche Scientifique, the Aix-Marseille Université and grants from the Agence National de la Recherche (BACMOLMOT [ANR-14-CE09-0023]) and from Projets internationaux de coopération scientifique (PICS05853). H.C., T.J.B., I.B. and S.K.B. are supported by the Intramural Research Program of the NIH, NIDDK.
Author information
Authors and Affiliations
Contributions
M.S. prepared the ExbB wild-type construct. H.C., N.N., T.J.B., and R.L. prepared ExbD constructs and mutants of ExbB and ExbD. T.J.B. prepared the TonB constructs. H.C. performed all expression and purification for assays, electron microscopy, DEER spectroscopy, electrophysiology, and crystallization. H.C. and N.N. performed crystallization and H.C., N.N., and I.B. did screening and data collection. N.N. solved the initial crystal structure using crystals grown by H.C. H.C. performed the electron microscopy experiments. E.B. performed the DEER spectroscopy experiments. S.D.Z. and W.A.C. performed the electrophysiology experiments. All authors analysed and discussed the data. R.L., W.A.C., and S.K.B. conceived the original projects. H.C., N.N., S.D.Z., W.A.C., R.L. and S.K.B. contributed to writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks H. Mchaourab, T. Walz and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Structure determination for the Ton subcomplex (ExbB–ExbDΔperi) using Se-SAD at 5.2 Å resolution.
a, The initial structure of the Ton subcomplex was solved by Se-SAD using anisotropic data extending to 5.2 Å resolution. The data from two crystals were processed with Xia2 and the initial sites found using HKL2MAP v0.3, which found a single solution every ~10,000 tries; resolution limits were also important for finding a solution. b, The sites were then input into AUTOSOL/PHENIX for site refinement and density modification, producing density maps (blue mesh) which clearly showed five-fold symmetry and allowed an initial model of a monomer to be built, consisting almost entirely of α-helices. This model was then used as a search model for molecular replacement to solve the 2.6 Å native structure (data obtained from a single crystal). c, Anomalous different map (orange mesh) showing density for the selenium sites in the 5.2 Å Se-incorporated structure.
Extended Data Figure 2 Representative electron density for the native Ton subcomplex (ExbB–ExbDΔperi) solved to 2.6 Å resolution.
a, Representative electron density map (2Fo−Fc contoured at 1.0σ, grey mesh; 2Fo−Fc omit map (omitting residues 113–124) contoured at 1.0σ, magenta mesh) along residues 113–124 within helix α5. b, Cutaway view of the transmembrane pore of ExbB (grey ribbon) from the native structure at pH 7.0 showing ring-like difference density (green isosurface, Fo−Fc map contoured at 2.5σ) along the conserved residues T148 and T181 (grey and red spheres). c, d, Tilted view (c) and an orthogonal view (d) (relative to a) of the ring-like density. Structures were determined using data obtained from a single crystal in each case.
Extended Data Figure 3 Helical shifts and overall flexibility in the ExbB pentamer.
a, Two pentamers were observed per asymmetric unit within the crystal structure. Shown here is pentamer 1 (green) aligned with pentamer 2 (magenta), illustrating slight shifts in a number of the helices (cylinders) between the two pentamers, with the largest shifts indicated by black arrows. The loops connecting α6 and α7 also show variability between monomers and pentamers. b, The TonB subcomplex (ExbB–ExbDΔperi) showing a B-factor putty representation with values ranging from the most ordered in blue to the most disordered in red.
Extended Data Figure 4 Electron density for the transmembrane helix of ExbD.
a, Omit map (2Fo−Fc, contoured at 1.0σ) along the transmembrane pore of ExbB. The density corresponding to the ExbB pentamer is shown in blue mesh, while the density corresponding to the transmembrane helix of ExbD is shown in green mesh. b, Stereoimage showing the density (2Fo−Fc, contoured at 0.8σ, grey mesh; 2Fo−Fc omit map (omitting the transmembrane helix of ExbD), contoured at 0.8σ, green mesh) for the transmembrane helix of ExbD after building and refinement.
Extended Data Figure 5 Comparison of observed density for crystal structures of ExbB–ExbDΔperi solved at pH 7.0 versus pH 4.5.
The presence of electron density for the transmembrane helix of ExbD (magenta ribbon) was dependent on the pH at which the crystals were grown. At pH 7.0, we observed little density (orange mesh) inside the transmembrane pore of the ExbB (grey ribbon) pentamer (see also Extended Data Fig. 3). However, for the structures solved at pH 4.5, we observed clear density (blue mesh) for the transmembrane helix of ExbD, albeit to varying degrees. Density maps (2Fo−Fc) are contoured at 1.0σ.
Extended Data Figure 6 Packing similarities of the 2D and 3D crystals used for electron microscopy and X-ray crystallography.
a, Averaged projection map from the electron microscopy analysis on 2D crystals. Five images were analysed, and a representative averaged projection map was calculated from 900 sub-images. The averaged map shows two different populations of the pentamer that are similar in size but differ in level intensity owing to opposite orientations of the complex within the crystal; a similar packing arrangement was also observed in our crystal structures. ExbD was not detected in our electron microscopy studies, probably owing to disorder of the globular domain, which is anchored to the membrane by a long unstructured linker15. b, Packing of the complex in the X-ray crystal structure from 3D crystals. The right side indicates an orthogonal view highlighting a single row of molecules from the lattice (black dashed box). c, Fitting the row of molecules from the 3D lattice (X-ray) from b onto the averaged projection map from the 2D crystals (electron microscopy) to highlight the consistency observed in packing.
Extended Data Figure 7 DEER traces and analysis.
Ton subcomplex (ExbBC25–ExbD, ExbBC25S–ExbDN78C, and ExbBC25S–ExbDE113C) in 0.08% C10E5 (a) and in 0.03% DDM (b), and the fully assembled Ton complex (TonBC18A–ExbBC25–ExbD and TonBC18A–ExbBC25S–ExbDN78C) in 0.05% DDM (c). Upper panels, experimental Q-band DEER primary data V(t)/V(0) (coloured lines, cyan ExbD113MTSL; violet ExbD78MTSL; red and orange, ExbB25MTSL) and simulated background functions (dotted line). Middle panels, DEER traces after background correction (coloured lines) and fit with DeerAnalysis2015 (dotted lines) with Tikhonov regularization parameters from 10 to 100 adjusted via L-curve analysis and data validation. Lower panels, obtained distance distributions. For the pentameric ExbB sample (50% labelling efficiency), a modulation depth >0.45 was obtained, indicating the presence of a multi-spin system. For the sample solubilized in DDM, longer DEER traces were obtained (4 μs) to better characterize the long distance peak of 5–6 nm in ExbB25MTSL. Additionally, for all panels, another DEER trace was measured after decreasing the microwave power of the 12-ns pump pulse to 25% (orange line) to suppress ghost peaks arising from the presence of more than two spins in the system. The resulting distance distribution (orange) was found to be very similar to that obtained with 100% microwave power (red), showing that no ghost peak artefacts were present. The lower modulation depth observed for the ExbD samples labelled at position 113 with respect to those labelled at position 78 (both labelling efficiency >80%) may be due to the presence of distances <1.5 nm (predicted by the simulations), which are outside of the sensitivity range of the technique, or to destabilization of the ExbD dimer induced by the label. The bottom of c shows a comparison of the Ton subcomplex in DDM (dashed lines from b) to the fully assembled Ton complex (solid lines). All panels show data from single experiments.
Extended Data Figure 8 Densitometry of the purified fully assembled Ton complex.
a, SDS–PAGE gel of the Ton complex (+TonB) and the Ton subcomplex (−TonB) at increasing concentrations. b, Bar graph showing the comparison of the ExbB–ExbD ratio within the Ton complex (+TonB) and the Ton subcomplex (−TonB) indicating that association of TonB with the Ton subcomplex does not change the stoichiometric ratio of the components. While we see a slight difference in the ExbB–ExbD ratio values in the presence or absence of TonB, the observed difference is too small to suggest an altered stoichiometry between ExbB and ExbD. Three representative lanes for each sample are shown in a; however, five lanes were used for all calculations. Densitometry analysis was performed with ImageJ and mean values and standard errors calculated using Microsoft Excel. For purifications of the Ton complex (+TonB), five purification experiments were performed and one representative is shown. For purifications of the Ton subcomplex (−TonB), ~50 purifications were performed and one representative is shown.
Extended Data Figure 9 Circular dichroism analysis of secondary structure and thermal stability of the Ton subcomplex.
Far-UV circular dichroism spectrum (185–260 nm) of the Ton subcomplex (ExbB–ExbD) with the calculated percentage of secondary structure shown. Contents of regular and distorted α-helical structures, 47 and 21%, respectively, were combined during the calculation of secondary structure contributions. Inset, comparison of the thermal stability of the Ton subcomplex (blue) versus ExbB alone (red) measured through the temperature dependence of the circular dichroism signal amplitude at 222 nm. Both panels show data from a single experiment.
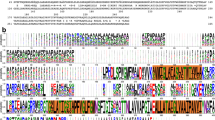
Extended Data Figure 10 Sequence conservation of ExbB orthologues mapped onto the crystal structure.
a, Clustal W alignment of ExbB sequences from: E.coli K12 (P0ABU7), Neisseria meningitidis (P64100), Neisseria gonorrhoeae (Q5F711), Haemophilus ducreyi (O51808), Vibrio harveyi (D0XEN5), Yersinia pestis (D1TTA4), Methanothermobacter thermautotrophicus (O27101), Pseudomonas aeruginosa (G3XCW0), ExbB1 of Vibrio cholerae (O52043) and ExbB2 of Vibrio cholerae (AAC69454). b, Conservation mapped onto the ExbB structure with Chimera. The most conserved residues are in blue and found in α6 (TM2) and α7 (TM3) of the ExbB structure. An extensive alignment that also includes sequences from the Tol and Mot systems shows similar results22. c, Cutaway molecular surface of ExbB pentamer with the most conserved residues mapped onto the surface.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-2. Table 1 contains a summary of data collection and refinement statistics for the Ton subcomplex crystal structures; Table 2 shows how zero-current potential (Vrev) was determined from volt-ampere characteristics measured in asymmetric salt conditions. Relative cation/anion permeability was calculated using the Goldman-Hodgkin-Katz equation. (XLSX 12 kb)
Overall structure of the Ton subcomplex
This video gives a structural insight into the role of the Ton complex in energy transduction. (MOV 29929 kb)
Rights and permissions
About this article
Cite this article
Celia, H., Noinaj, N., Zakharov, S. et al. Structural insight into the role of the Ton complex in energy transduction. Nature 538, 60–65 (2016). https://doi.org/10.1038/nature19757
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19757
- Springer Nature Limited