Abstract
Insulin-dependent diabetes is a complex multifactorial disorder characterized by loss or dysfunction of β-cells. Pancreatic β-cells differ in size, glucose responsiveness, insulin secretion and precursor cell potential1,2,3,4,5; understanding the mechanisms that underlie this functional heterogeneity might make it possible to develop new regenerative approaches. Here we show that Fltp (also known as Flattop and Cfap126), a Wnt/planar cell polarity (PCP) effector and reporter gene6, acts as a marker gene that subdivides endocrine cells into two subpopulations and distinguishes proliferation-competent from mature β-cells with distinct molecular, physiological and ultrastructural features. Genetic lineage tracing revealed that endocrine subpopulations from Fltp-negative and -positive lineages react differently to physiological and pathological changes. The expression of Fltp increases when endocrine cells cluster together to form polarized and mature 3D islet mini-organs7,8,9. We show that 3D architecture and Wnt/PCP ligands are sufficient to trigger β-cell maturation. By contrast, the Wnt/PCP effector Fltp is not necessary for β-cell development, proliferation or maturation. We conclude that 3D architecture and Wnt/PCP signalling underlie functional β-cell heterogeneity and induce β-cell maturation. The identification of Fltp as a marker for endocrine subpopulations sheds light on the molecular underpinnings of islet cell heterogeneity and plasticity and might enable targeting of endocrine subpopulations for the regeneration of functional β-cell mass in diabetic patients.
Similar content being viewed by others
Main
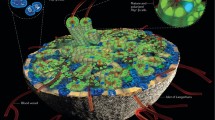
The islet of Langerhans is a complex mini-organ that regulates energy metabolism10. Pancreatic β-cells are organized in polarized rosette-like structures around blood vessels7 and are coupled by gap junctions to allow them to sense glucose and secrete appropriate amounts of insulin into the blood stream11. The conserved Wnt/planar cell polarity (PCP) pathway regulates the orientation of cells and organelles within the plane of a tissue and thus determines the function of cells in an organ12,13. As self-organized 3D tissue architecture is required for organ formation and terminal differentiation14,15, we tested whether acquisition of tissue polarity and architecture could influence β-cell heterogeneity and maturation.
To analyse the establishment of tissue polarity at the molecular level, we used the FltpZV knock-in/knockout mouse, in which the entire open-reading frame of Fltp is replaced by a multicistronic lacZ–Venus reporter cassette6. In this model, Fltp promoter-driven histone 2B (H2B)–Venus reporter (FVR) activity is confined to terminally differentiated cells with established PCP6. We first analysed reporter activity in the pancreas of FltpZV/+ heterozygous embryos at embryonic day (E)18.5, when endocrine cells are still organized in cord-like structures outside the ductal tubules and start to aggregate and form 3D mini-organs (Extended Data Fig. 1). Notably, we detected heterogeneous reporter activity in β-cells that expressed high levels of Nkx6.1 in compacted islet structures, but not in scattered pre-β-cells in cord-like structures that expressed low levels of Nkx6.1. This finding suggests that FVR is activated in endocrine cells during islet compaction and establishment of polarization. Because β-cell maturation occurs in the first two weeks after birth9, we analysed reporter activity in postnatal and adult islets from heterozygous FltpZV/+ mice. At postnatal day (P)1, reporter activity was detectable in less than 50% of Nkx6.1+ β-cells; activity was significantly increased at P11 and reached a maximum of 80% in adult islets (Fig. 1a, b). In adult islets, FVR was expressed in ~50% of glucagon-secreting α-cells, ~80% of insulin-secreting Nkx6.1+ β-cells, ~50% of somatostatin-secreting δ-cells and ~50% of pancreatic polypeptide-secreting PP-cells (Fig. 1c, d). Together, these observations suggest that islet cell heterogeneity develops during islet neogenesis and is likely to be determined by the terminal differentiation and planar polarization status of endocrine cells.
a, b, Laser scanning confocal microscopy (LSM) images (a) and quantification (b) of FVR+ β-cell distribution during postnatal β-cell maturation (P1, P11, P25) and in adulthood (12 weeks (w)) (P1: n (mice) = 5 (14,766 Nkx6.1+ cells), P11: n (mice) = 6 (13,789 Nkx6.1+ cells), P25: n (mice) = 5 (8,733 Nkx6.1+ cells), 12 weeks: n (mice) = 9 (17,369 Nkx6.1+ cells); ****P < 0.0001). c, d, LSM images (c) and quantification (d) of FVR expression in endocrine cell lineages in adulthood (n (mice) = 3, 1,676 PP+ cells, 1,656 Sst+ (δ-)cells and 1,975 Gcg+ (α-)cells); n (mice for β-cells) = 5, β-cells (7,327 Nkx6.1+ cells)). e, f, LSM images (e) and quantification (f) of Ki-67-expressing cells among FVR− and FVR+ β-cell subpopulations at P1, P11, 12 weeks and in pregnant mice (G15.5; arrowheads indicate proliferating cells; red arrowheads, Nkx6.1; green arrowheads, Venus and Nkx6.1; P1: n (mice) = 5 (14,766 Nkx6.1+ cells), P11: n (mice) = 6 (13,789 Nkx6.1+ cells), 12 weeks: n (mice) = 5 (10,042 Nkx6.1+ cells) and pregnant G15.5: n (mice) = 5 (12,691 Nkx6.1+ cells); (P1) **P = 0.0048, (P11) ****P < 0.0001, (G15.5) **P = 0.0039). Scale bars, 50 μm (a, c, e). Two-sided unpaired t-test was performed (b, e). Data represent mean ± s.e.m. Source data for all graphs and quantifications are provided online.
To understand the biological relevance of islet cell heterogeneity in more detail, we specifically focused on β-cells in heterozygous FltpZV/+ mice. We first compared the proliferation rate of FVR− and FVR+ β-cells during homeostasis, postnatal β-cell expansion and physiological insulin resistance. FVR− Nkx6.1+ β-cells showed a significantly increased replication rate during the postnatal expansion phase at P1, at P11 and upon metabolic demand during pregnancy at gestational day (G)15.5, as measured by Ki-67 immunoreactivity (Fig. 1e, f) and EdU incorporation (Extended Data Fig. 2a, b), when compared to FVR+ Nkx6.1+ β-cells; by contrast, FVR+ Nkx6.1+ β-cells showed increased levels of the cell-cycle inhibitor p27 (Extended Data Fig. 2h–j). When we compared the head and tail regions and different-sized islets, we noticed remarkable intra-islet and inter-islet heterogeneity in terms of FVR marker expression and proliferation rate (Extended Data Fig. 2c–g), probably due to different cell-type compositions in the islet niches16. Collectively, our data show that β-cell subpopulations, depending on their niche and environmental conditions, show markedly different FVR expression levels correlating with their proliferative capacity.
The data presented so far suggest that the expression of FVR probably subdivides β-cells into proliferative and more mature β-cells. To test this idea directly, we isolated adult islets from FltpZV/+ mice and purified the FVR endocrine subpopulations for genome-wide mRNA expression profiling (Fig. 2a, b). We found 997 genes that showed more than 1.5-fold differences in expression between FVR− and FVR+ endocrine cells (Fig. 2c and Supplementary Table 1). Strikingly, unbiased gene ontology term (Fig. 2g and Supplementary Table 2) and ingenuity analysis (Fig. 2h and Supplementary Table 3) revealed that the FVR− subpopulation showed significant enrichment of genes associated with G-protein coupled receptor (GPCR), Wnt and MAPK signalling transduction, whereas the FVR+ subpopulation showed significant enrichment of genes that are important for mature β-cell function. This difference is reflected by differential mRNA expression of hormones, β-cell maturation markers, mitochondrial and metabolic genes, receptors and signalling components, as revealed by microarray analysis (Fig. 2d–f) and confirmed by qPCR analysis on highly enriched β-cell subpopulations (Extended Data Fig. 3a–h). Mitochondrial gene expression (Fig. 2e, g, h), ultrastructural (Extended Data Fig. 4) and biochemical analysis (Fig. 2k, l), and mitochondrial network architecture (Extended Data Fig. 5e–k) collectively suggest that the two isolated β-cell subpopulations have similar numbers of mitochondria (Extended Data Fig. 3i) but that the mitochondria differ in architecture and physiology. Measuring the oxygen consumption rate of sorted FVR− and FVR+ β-cells from FltpZV/+ islets using an XF96 extracellular flux analyser revealed that glucose-induced ATP synthesis and glucose responsiveness were not significantly different between the two subpopulations (Extended Data Fig. 5a–d), whereas static glucose-stimulated and arginine-stimulated insulin secretion were significantly enhanced in the sorted and reaggregated FVR+ β-cell enriched subpopulation (Extended Data Fig. 5l). These results are consistent with the differences in the expression of genes involved in glucose sensing and insulin secretion (Fig. 2 and Extended Data Fig. 3g). Finally, the high expression of the proprotein convertase subtilisin/kexin type-1 gene (Pcsk1) in the FVR+ cells (Extended Data Fig. 3g) and the higher number of mature secretory granules in Fltp-lineage+ cells (Fig. 2i, j and see below) further support the idea that FVR+ and Fltp-lineage+ cells are mature β-cells.
a, Experimental design for microarray analysis of endocrine subpopulations from isolated FltpZV/+ islets. b, Fluorescent activated cell sorting (FACS) separation of FVR+ and FVR− endocrine subpopulations. c, Scatter plot of the 997 genes that were differentially regulated in FVR+ and FVR− endocrine cells (1.5-fold regulated, P < 0.01, limma t-test). d–f, Heatmaps depicting regulated genes among FVR subpopulations involved in β-cell function (d), mitochondrial function (e) and receptors and signalling (f) (*validated by qPCR, median centre was applied to each gene of the heatmaps across different samples). g, h, Bar graph of selected, significantly enriched gene ontology terms (g) and ingenuity pathways (h) of regulated genes among FVR subpopulations shown in c. i, j, Transmission electron microscopy images (i; scale bars, 1 nm) and quantification (j) of insulin granules in immunogold-labelled β-cells from FltpT2AiCre/+;Gt(ROSA)26mTmG/+ mice (m, mature; im, immature) (n (independent experiments) = 5, n (β-cells) = 50, n (granules) = 12,254; **P = 0.0015, two sided unpaired t-test). k, l, Western blot (k) and quantification (l) of mitochondria complexes I–V from FACS-sorted FVR+ and FVR− subpopulations (protein expression normalized to γ-tubulin and to the FVR− population (dotted line), n (independent experiments) = 5, whiskers represent minimum and maximum values). Data in j represent mean ± s.e.m. See Supplementary Fig. 1 for gel source data.
The differential expression of signalling components and the distinct physiologies of the two β-cell subpopulations suggested that they might react differently to environmental changes. To test this hypothesis, we used a Cre recombinase/loxP-mediated genetic lineage tracing approach, which allowed us to follow the cell populations over time (Fig. 3a). We isolated FltpT2AiCre/+;Gt(ROSA)26mTmG islets (refs 17, 18) and performed in vitro single cell-tracking experiments. This showed that Fltp-lineage− endocrine cells turned into Fltp-lineage+ cells (Fig. 3b). As FVR− and FVR+ β-cells are molecularly and biologically distinct, these results suggest that FVR− precursor β-cells differentiate into FVR+ mature β-cells. Notably, the FVR− β-cell enriched subpopulation expressed low levels of Slc2a2 (Extended Data Fig. 3g), which has been recently shown to be expressed in insulin-positive pancreatic multipotent precursor cells in mouse and human islets3,4.
a, FltpT2AiCre/+;Gt(ROSA)26mTmG/+ lineage-tracing model. Fltp lineage− cells (mT, red) convert into Fltp lineage+ cells (mG, green) upon Fltp-promoter driven Cre expression via an intermediate (mTmG, yellow) cell state. b, e, Time-lapse imaging of in vitro cultured FltpT2AiCre/+;Gt(ROSA)26mTmG/+ islets (b; n (cells) = 7) and in vivo islets transplanted into the anterior chamber of the eye (AC) using laser scanning confocal microscopy (LSM) (e) shows conversion of endocrine Fltp lineage− cells (mT) into Fltp lineage+ cells (mG) (white dotted line shows converting cells). c, d, FACS analysis (c) and quantification (d) of mT, mTmG and mG Nkx6.1+ β-cells from control (Ctrl) and pregnant mice (G15.5) (n (mice per group) = 4, ***P = 0.0001, ***P = 0.0003, Bonferroni’s multiple comparison test). f, g, LSM images (f) and quantification (g) of a FltpT2AiCre/+;Gt(ROSA)26mTmG/+ islet transplanted into the AC over engraftment time (n (islets) = 15 islets from 4 mice; **P = 0.0086, two sided unpaired t-test). h, i, LSM images of a FltpT2AiCre/+;Gt(ROSA)26mTmG/+ islet in the AC before (Pre-diet; h) and after 4 weeks on a high-fat diet (HFD; i). j, k, Quantification of mT and mG cell cross-sectional areas (j) and compartment volume change among mT and mG endocrine cells (k) of intraocular FltpT2AiCre/+;Gt(ROSA)26mTmG/+ islets before and during HFD (4 w, 8 w) (j, n (islets) = 34, *P = 0.0112, **P = 0.00732; k, n (islets) = 34, *P = 0.026, ****P < 0.0001, Sidak’s multiple comparison test). Scale bars, 10 μm (b); 20 μm (e, f); 50 μm (h, i). Data shown as mean ± s.e.m.
To analyse the in vivo β-cell subpopulation dynamics at the time when compensatory β-cell proliferation reaches peak levels19, we isolated adult islets from non-pregnant female mice and from pregnant female mice at G15.5 (Fig. 3c, d). Quantitative analysis showed that the Fltp-lineage− β-cell subpopulation compensates for the increased metabolic demand in pregnant mice by proliferating. The higher number of mTmG double-positive cells in pregnant mice further suggested that Fltp-lineage− cells differentiate into mature β-cells in vivo. Next, we transplanted FltpT2AiCre/+;Gt(ROSA)26mTmG islets into the anterior chamber of the eye in mice for noninvasive longitudinal in vivo imaging20. After donor islet isolation and transplantation onto the iris, islets first shrank in volume during engraftment, owing to the lack of nutritional supply, before revascularization counteracted islet degeneration (Extended Data Fig. 6d). Concomitant with the revascularization and decrease in islet size we observed a relative increase in the Fltp-lineage− and decrease in the Fltp-lineage+ subpopulation volume (Fig. 3f, g) and number (Extended Data Fig. 6e) 13 days after transplantation, suggesting that the more mature and metabolic active endocrine subpopulation undergoes apoptosis, probably due to hypoxia, until revascularization. Four weeks after transplantation the Fltp-lineage+ subpopulation had recovered, probably owing to the maturation of Fltp-lineage− cells into mature Fltp-lineage+ cells (Fig. 3e). Finally, we investigated the endocrine subpopulation dynamics in a high-fat diet (HFD) mouse model. HFD causes insulin resistance and compensatory β-cell hyperplasia followed by hypertrophy, but the exact pathomechanism of β-cell failure remains unclear21. Over an eight-week period, we observed increased glucose intolerance (Extended Data Fig. 6c) and a steady increase in islet volume accompanied by a significant increase in the cross-sectional area of the Fltp-lineage+ endocrine cell population, indicating that mature endocrine cells are more metabolically active and prone to hypertrophy commonly associated with cytotoxic stress (Fig. 3h–k and Extended Data Fig. 6a, b). This observation might explain why the Fltp-lineage+ subpopulation predominantly decreased after transplantation (Fig. 3g). Together, our results suggest that, in pregnant mice, Fltp− endocrine cells undergo compensatory proliferation, whereas more mature Fltp-lineage+ endocrine cells probably account for islet cell hypertrophy and seem more sensitive to cytotoxic stress.
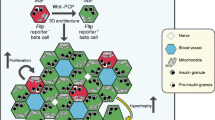
In our molecular analysis we noticed that apical–basal polarity and expression of Wnt/PCP genes were increased in FVR+ β-cells (Fig. 2 and Extended Data Fig. 3g, j). Notably, cellular polarization and compaction in 3D pseudo-islet cultures22,23 strongly induced protein expression of a variety of β-cell maturation markers, including Nkx6.1, MafA and Ucn3, as revealed by quantitative PCR (qPCR) and immunocytochemistry in time-course experiments in Min6 insulinoma cells (Extended Data Fig. 7a–j). Consistent with the induction of the Wnt/PCP effector Fltp and Nkx6.1 during islet neogenesis (Extended Data Fig. 1), we observed that treatment of dispersed and reaggregated P5 islet cells (Fig. 4a–c) and pseudo-islets of Min6 insulinoma cells (Extended Data Fig. 7k–n) with the Wnt/PCP ligand Wnt5a significantly upregulated β-cell maturation makers. Human microislets and human islets show heterogenous expression of WNT4 in endocrine and β-cells, consistent with the differential expression of Wnt4 and Wnt5b in the FVR− and FVR+ subpopulations (Fig. 4d–g and Extended Data Fig. 3g). Treatment of human EndoC-β H1 β-cells with the canonical Wnt ligand WNT3A increased proliferation, whereas treatment with WNT4 or WNT5A did not (Extended Data Fig. 8a–e). Notably, WNT4 induced JNK activation (Extended Data Fig. 8f, g) and increased levels of NKX6.1 and PDX1 protein in human EndoC-β H1 β-cells (Fig. 4h, i) and human islets (Fig. 4j, k), suggesting that WNT4 triggers the non-canonical Wnt/PCP pathway. Finally, WNT5A increased glucose-stimulated insulin secretion (GSIS) in human microislets (Extended Data Fig. 8h). Together, these results suggest that 3D architecture and Wnt/PCP pathway activation induce β-cell maturation and increase GSIS.
a, b, LSM images (a) and quantification (b) of Ucn3 protein in reaggregated islet cells from P5 wild-type mice treated with or without Wnt5a (n (independent experiments) = 3 (DAPI+ cells control = 680, DAPI+ cells Wnt5a = 877), ***P = 0.001, two sided unpaired t-test). c, Wnt5a stimulation of mRNA expression of β-cell maturation markers in P5 reaggregated islet cells (n (independent experiments) = 4, *P < 0.05, two sided unpaired t-test). d–g, LSM images of a human islet (d, e) and a human microislet (f, g) showing heterogeneity of WNT4-expressing NKX6.1+ β-cells (white dotted lines and arrowheads). h, i, Western blot (h) and quantification (i) of NKX6.1 and PDX1 in EndoC-β H1 β-cells after 4 days of treatment with WNT4 and WNT5A (n (independent experiments) = 5; *P = 0.030 for PDX1, *P = 0.0349 for NKX6.1, two-sided paired t-test). j, k, Western blot (j) and quantification (k) of PDX1 expression in human islet after 4 days of treatment with WNT4 (n (independent experiments) = 9, 4 human donors, two-sided paired t-test). l, m, RNA sequencing expression data of FLTP and PDX1 in human pancreatic islets from healthy donors (normal glucose tolerance (NGT), n = 66), pre-diabetic donors (impaired glucose tolerance (IGT), n = 21) and type 2 diabetic donors (T2D, n = 19; one-way ANOVA). Scale bars, 10 μm (a, d–g). Data represent mean ± s.e.m. See Supplementary Fig. 1 for gel source data.
Finally, we investigated whether the Wnt/PCP effector Fltp is required for β-cell function beyond its role as a maturation marker. Terminally differentiated cell types in the inner ear and lung depend on Fltp function as a modulator of the actin and microtubule cytoskeleton6 and mature β-cells require cell polarity24 and the cytoskeleton25 for proper insulin secretion. Fltp knockout (FltpZV/ZV) β-cells do not show impaired development, proliferation or maturation (Extended Data Figs 9, 10a, c, d) or altered glucose tolerance or insulin sensitivity (Extended Data Fig. 10b, e). However, when we analysed basal glucose and insulin levels in adult mice we found small but significantly reduced levels in starved FltpZV/ZV mice, suggesting that Fltp knockout causes a minor defect in insulin secretion (Extended Data Fig. 10b, f). To rule out systemic effects on basal insulin levels in FltpZV/ZV animals, we isolated islets from adult mice and performed GSIS. Consistent with the finding that insulin secretion depends on cell polarity and the cytoskeleton24,25 we found reduced GSIS in FltpZV/ZV islets (Extended Data Fig. 10g). To investigate whether the human FLTP orthologue (CFAP126, also known as C1orf192) is associated with metabolic traits, we genotyped seven single nucleotide polymorphisms (SNPs) tagging all the common variation in this gene (minor allele frequency ≥ 0.01) and analysed their associations with insulin secretion, insulin sensitivity and body fat content and distribution in a cohort of 2,228 human prediabetic subjects26. The results showed that the intronic SNP rs75715534 (minor allele frequency 0.037) was significantly associated with increased insulin secretion in lean subjects (body mass index (BMI) < 25), whereas the same SNP was associated with decreased insulin secretion in obese people (BMI ≥ 35) (Extended Data Fig. 8j, k and Supplementary Table 4). CFAP126, SLC2A2 and PDX1 were significantly downregulated in isolated human islets from pre-diabetic (IGT) when compared to non-diabetic (NGT) subjects and further downregulated in islets from type 2 diabetic (T2D) individuals (Fig. 4l, m and Extended Data Fig. 8i), suggesting that Wnt/PCP signalling and β-cell maturation or function are impaired by glucolipotoxicity.
We have demonstrated that proliferative and mature β-cells can be distinguished by expression of the Wnt/PCP effector and reporter gene Fltp. Several studies have indicated that loss of β-cell identity and maturation underlies the pathogenesis of type 2 diabetes27,28,29,30. The availability of a unique marker for mature β-cells opens new ways to explore pathways regulating islet heterogeneity and might allow researchers to target β-cell maturation for cell-replacement and regenerative therapy.
Methods
Animal studies
Animal experiments were carried out in compliance with the German Animal Protection Act and with the approved guidelines of the Society of Laboratory Animals (GV-SOLAS) and of the Federation of Laboratory Animal Science Associations (FELASA). Mouse lines used: FltpZV (C57BL/6J; ref. 6) and FltpT2AiCre/+ (mixed C57BL/6J, CD1 background; ref. 17) crossed with Gt(ROSA)26mTmG (129/SvJ, C57BL/6J mixed background; ref. 18) (FltpT2AiCre/+;Gt(ROSA)26mTmG). Albino B6 mice (B6N-Tyrc/BrdCrCrl; Charles River) crossed with B6 Rag1−/− mice (B6.129S7-Rag1tm1Mom/J; Jackson Laboratories) were used as recipients for FltpT2AiCre/+;Gt(ROSA)26mTmG islet transplants. All experiments were performed using 3–6-month-old mice, unless indicated otherwise. Islet isolation was carried out by collagenase P (Roche) digestion and centrifugation using Optiprep density gradient (Sigma) as described31. Isolated islets were handpicked twice under a microscope. For analysis of islet β-cell proliferation during pregnancy, mice were euthanized at G15.5 and for histological analysis EdU was administrated 24 h before sacrifice. For HFD analysis, fourteen weeks after islet transplantation, mice were divided into two groups and either continued on a normal diet (Ssniff GmbH) or switched to HFD (60 kcal % fat; Research Diets, Inc.) with ad libitum access to water and chow.
For intraperitoneal glucose tolerance tests (IPGTTs) and insulin tolerance test (ITT) mice were fasted for 6 h and injected intraperitoneally with 2 g glucose per kg (BW) or 0.75 U insulin per kg (BW), respectively. Blood glucose was measured at indicated time points after glucose injection using Accu-Chek Aviva glucometer (Roche).
Cell culture and human islets
Min6 (clone 9) murine β-cell line and EndoC-β H1 were cultured in adherence (2D culture) as described previously32,33. For floating 3D (3D) culture Min6 were plated on uncoated ibidi chambers (ibidi) for 3, 6 or 7 days. For 3D matrigel based cultures, Min6 and EndoC-β H1 were cultured in Growth-Factor-Reduced Matrigel Matrix (BD Bioscience, Germany) diluted 1:1 in their respective medium. All the cell lines used were tested for mycoplasma and were negative. Isolated islet cells from P5 pancreata were cultured overnight in RPMI 1640 containing P/S and 5% FCS. For analysis of Wnt5a induced β-cell maturation, samples were stimulated with 400 ng/ml of Wnt5a (R&D systems, Germany) for 12 h, 3 or 6 days in RPMI1640 supplemented with P/S and 3% FCS. Human islets isolated from cadaver and pancreatic microislets were obtained from the JDRF award 31-2008-416 (ECIT Islet for Basic Research program) and from InSphero (Switzerland), respectively. All experiments with human islets were approved by the Ethical commission of the Technical University of Munich (Germany). Human islet donor data: 6 donors, 3 males and 3 females; age mean 53.6 yrs (range: 41–65); BMI, mean 24.25 (range: 20.8–30.1); purity mean 78 (range 89–45%). Human microislet donor data: 3 donors, 2 female and 1 male; age mean 40.10 yrs (range: 37–56); BMI mean 23.19 (range: 22.7–23.56).
Upon arrival, human islets were collected by centrifugation, selected by hand picking and cultured in CMRL1066 (Gibco, Thermo Fisher Scientific) supplemented with 10% human serum (Sigma), 1% P/S and 2 mM l-glutamine. Pancreatic microislets were cultured according to the manufacturer’s instruction. Human islets were treated with 100 ng/ml WNT4 (R&D systems) or 400 ng/ml WNT5A in CMRL 1066 supplemented with 2% human serum albumin (Sigma), 1% P/S, 2 mM l-glutamine, 5.5 mM glucose, 100 ng/ml mouse Noggin (Peprotech) and 10 μM Y-27632 (Santa Cruz) for 4 days (medium was changed every 2 days).
Microislets were treated with 100 ng/ml WNT4 (R&D Systems) or 400 ng/ml WNT5A in CMRL 1066 supplemented with 2% human serum albumin, 1% P/S, 2 mM l-glutamine, 5.5 mM glucose and 100 ng/ml mouse Noggin (Peprotech) for 4 days (medium was changed every 2 days).
EndoC-β H1 cells were treated with 100 ng/ml WNT4 (R&D systems) or 400 ng/ml WNT5A or 200 ng/ml WNT3A (Peprotech) in their culture medium5 supplemented with 100 ng/ml mouse Noggin (Peprotech) and 10 μM Y-27632 (Santa Cruz) for 4 h or 4 days (medium was changed every 2 days).
Islet transplantation and in vivo imaging of islets engrafted in the anterior chamber of the eye
For islet transplantation into the anterior chamber of the mouse eye, mice were anaesthetized by inhalation of 2% isoflurane in 100% oxygen via a face mask. The mouse head was fixed in a head holder and a 25-gauge needle used to make a small incision into the cornea, close to the corneal limbus. Next, 30–40 islets in PBS were slowly injected into the anterior chamber of the eye using a custom made beveled glass cannula (outer diameter 0.4 mm, inner diameter 0.32 mm; Hilgenberg GmbH). For in vivo imaging mice were intubated (BioLite, Braintree Scientific, Inc.) and anaesthetized by 2% isoflurane in 100% oxygen with 270 μl stroke volume at 250 strokes per min for ≤90 min. In vivo imaging was performed as previously described34. For confocal and two-photon imaging, an upright laser-scanning microscope (LSM780 NLO; Zeiss) with a two-photon laser (Chameleon Vision II; Coherent, Inc.) and W Plan-Apochromat 20×/1.0 DIC M27 75 mm objective (Zeiss) was used. Imaging of intraocular islets was performed during islet engraftment (days 3, 13 and 4 weeks post-transplantation) and repeated four and eight weeks after diet switch. Backscattered laser light was detected at 633 nm. Total islet volume was calculated using surface rendering (Imaris 7.4, Bitplane AG). GFP (mG) and Tomato (mT) were excited by two-photon laser at 930 nm and detected at 500–550 nm and at 575–610 nm, respectively. mG and mT islet fractions were assessed from Gaussian filtered z-stacks (step size of 1.5 μm) of the whole islet for the revascularization or within the top 80 μm of the islet for the HFD time points using surface rendering (Imaris 7.4). Compartment volumes were calculated in relation to total islet volume obtained from backscatter imaging. Vessels were visualized by injecting 0.8 μM Qtracker 705 (Life Technologies) in 100 μl PBS into the tail vein and detected at 690–730 nm. mG and mT cell cross-sectional areas were assessed manually in individual optical planes of the islet z-stacks (Fiji software)35.
Transmission electron microscopy
Immunogold staining was performed on isolated islets from FltpT2AiCre/+;Gt(ROSA)26mTmG/+ mice as follows. After islet isolation the islets were cultured overnight in RPMI1640 supplemented with 10% FCS and P/S, then fixed in 2.5% electron microscopy grade glutaraldehyde in 0.1 M sodium cacodylate buffer pH 7.4 (Science Services), stained with rabbit anti-GFP (Invitrogen, A11122, 1:500) and a gold-conjugated goat anti-rabbit antibody (Aurion, 806.011, 1:40). Isolated FACS-sorted FVR islet cells or immunogold-stained islets were fixed in 2.5% electron microscopy grade glutaraldehyde in 0.1 M sodium cacodylate buffer pH 7.4 (Science Services) as pellets, post-fixed in 2% aqueous osmium tetraoxide36, dehydrated in gradual ethanol (30–100%) and propylene oxide, embedded in Epon (Merck) and cured for 48 h at 60 °C. Semi-thin sections were cut and stained with toluidine blue. Ultrathin sections of 50 nm were collected onto 200 mesh copper grids, stained with uranyl acetate and lead citrate before examination by transmission electron microscopy (Zeiss Libra 120 Plus, Carl Zeiss NTS GmbH). Pictures were acquired using a Slow Scan CCD-camera and iTEM software (Olympus Soft Imaging Solutions). Serial pictures for image analyses were acquired by transmission electron microscopy (Zeiss EM10CR, Carl Zeiss NTS GmbH) with a MegaView III camera system (Olympus Deutschland GmbH). Images were quantified using image analysis software Definiens Developer XD2 (Definiens AG) according to a previously published procedure37. Initially mitochondria were annotated manually. A rule set was developed in order to detect and quantify the length of the inner mitochondrial membrane within these marked mitochondria, based on intensity, morphology and neighbourhood. Additionally the area for each mitochondrion and the number of mitochondria per cell area were calculated. The β-cells were identified as Fltp lineage+ or Fltp lineage− according to their immuno-gold labelling. Insulin granules were counted after their classification as immature or mature granules according to their morphology.
Immunofluorescence staining and western blot
For immunohistochemistry, pancreas samples were fixed in 4% formalin, cryoprotected by incubation in a progressive sucrose gradient and embedded in Optimum Cutting Temperature (OCT). Sorted cells, Min6 and EndoC-β H1 were fixed in 4% formalin before staining. For whole-mount staining, embryos were cleared using BABB solution and stained using a modified previously described protocol6.
Primary antibodies used for immunostaining: goat anti-Nkx6.1 (R&D system, AF5857, 1:400), rabbit anti-Nkx6.1 (Acris/Novus, NBP1-82553, 1:100), rat anti-RFP (Chromotek, ORD003515, 1:1000), chicken anti-GFP (Aves Labs, GFP-1020, 1:800), guinea pig anti-glucagon (Millipore, 4031-01F, 1:500), goat anti-somatostatin (Santa Cruz, sc-7819, 1:300), rabbit anti-insulin (Thermo Scientific, PA-18001, 1:200), guinea pig anti-insulin (Thermo Scientific, PA-26938, 1:300), goat anti-pancreatic polypeptide (PP) (Abcam, ab77192, 1:300), rabbit anti-Ki-67 (Abcam, ab15580, 1:200), rabbit anti-MafA (Bethyl, IHC-00352-1, 1:100), mouse anti-β-catenin (BD, 610154, 1:2000), rabbit anti-Urocortin 3 (Phoenix Pharmaceuticals, H-019-29, 1:300), rabbit anti-p27 (Santa Cruz, sc-528, 1:100), goat anti-Wnt4 (R&D Systems, AF475, 1:100), Alexa Fluor 546 phalloidin (Invitrogen, A22283, 1:200), rat anti-E-cadherin (raised by E. Kremmer (Helmholtz Zentrum München), 1:200), MitoTracker Deep Red FM (Life Technologies, M22426, final concentration in the medium: 200 nM). Secondary antibodies used for indirect fluorescence staining (dilution always 1:800): goat anti-chicken Alexa Fluor 488 (Dianova, 103-545-155), donkey anti-goat Alexa Fluor 488 (Invitrogen, A11055), donkey anti-mouse Alexa Fluor 555 (Invitrogen, A31570), donkey anti-goat Alexa Fluor 555 (Invitrogen, A21432), donkey anti-rabbit Alexa Fluor 555 (Invitrogen, A31572), donkey anti-goat Alexa Fluor 594 (Invitrogen, A11058), donkey anti-mouse Alexa Fluor 594 (Invitrogen, A21203), donkey anti-guinea-pig Alexa Fluor 649 (Dianova, 706-495-148). Nuclear staining was performed with DAPI (Life Technology, 1:800). EdU staining was performed using the Click-iT Staining Kit (Life Technology) according to the manufacturer’s instructions. For analysis of proliferation of EndoC-β H1, cells were pulsed with 10 μM EdU for 2 days. Cryosection imaging was performed using a Leica SP5 confocal microscope. Analysis was performed either manually or by using IMARIS software (Bitplane) or the Leica LAS-AF (Version 2.7.3.9723) software. For western blot analysis of mitochondrial complexes, islet cells were isolated and FACS sorted in RIPA buffer. Cell lysates were resolved by SDS–PAGE, transferred to PVDF membrane (Biorad) and incubated with the following primary antibodies: anti-Rt/Ms Total OxPhos Complex Kit (Life technologies, 458099, 1:250) and γ-tubulin (Abcam, ab11316, 1:10.000). For western blot analysis of Fltp and Nkx6.1 expression, tissues and Min6 were dissociated in RIPA buffer. Cell lysates were resolved by SDS–PAGE, transferred to PVDF membrane (Biorad) and incubated with the following antibodies: rabbit anti-Fltp38 (1:500), goat anti-Nkx6.1 (R&D system, AF5857, 1:2.000) and mouse anti-γ-tubulin (Abcam, ab11316, 1:10.000). For western blot analysis of pJNK, NKX6.1 and PDX1 expression, human islet and EndoC-β H1 were dissociated in RIPA buffer. Cell lysates were resolved by SDS–PAGE, transferred to PVDF membrane (Biorad) and incubated with the following primary antibodies: Rabbit SAPK/JNK (56G8) (New England Biolabs, 9258, 1:1.000), rabbit SAPK/JNK-Phospho (New England Biolabs, 4668, 1:1.000), goat anti-NKX6.1 (R&D Systems, AF5857, 1:2.000), rabbit anti-PDX1 (New England Biolabs, 5679, 1:500) and mouse anti GAPDH (Merck Bioscience, CB 1001, 1:5.000-10.000). Protein bands were visualized using the following horseradish peroxidase (HRP)-conjugated antibody: Goat anti-mouse HRP (Dianova, 115-036-062, 1:10.000), rabbit anti goat HRP (Dianova, 305-035-045, 1:5.000), goat anti rabbit HRP (Dianova, 111-036-045, 1:5.000) and chemiluminescence reagent (Millipore). The bands were quantified using ImageJ.
Fluorescent intensity analysis
Nkx6.1 and MafA fluorescent intensity analysis of Min6 was performed with Imaris software, using the spot function. Spots of the same size were chosen and the mean fluorescence within these spots was calculated. The median fluorescent intensity was used as a threshold to distinguish between Nkx6.1-high and -low cells.
To show the fluorescent intensity profiles a line was drawn and the fluorescence intensity over this line calculated and plotted against the length of the line.
To calculate the Fltp reporter fluorescent intensity of the BABB cleared pancreata Imaris software was used. A surface on the Nkx6.1 positive cells was calculated and the mean Fltp reporter fluorescent intensity was calculated within the described surfaces.
Nkx6.1, Venus, EdU and p27 fluorescent intensity analysis of pancreatic islet sections were performed with Imaris software, by using the surface function. Surfaces using the same threshold were rendered for the Nkx6.1 positive cells and the mean fluorescence intensity within these surfaces was calculated for Venus, EdU and p27. To discriminate between p27-high and -low cells, the total mean fluorescent intensity of all islets of one mouse was used as a threshold. The threshold for EdU was set by eye.
Mitochondria network analysis and oxygen consumption rate
The mitochondria network was analysed using Imaris. The mitochondrial surface was rendered by thresholding the Mitotracker Deep Red FM (Thermo Fisher Scientific) intensity. The analysis was performed on nine Nkx6.1–Venus double-positive cells and on four Nkx6.1 single-positive cells from two islets cultured in culture medium and on nine Nkx6.1–Venus double-positive cells and on six Nkx6.1 single-positive cells from three islets exposed to 2 mM glucose for 2 h. For the flux analysis experiment, FVR+ and FVR− β-cells were sorted and recovered. Around 30,000 single cells were plated in a 96-well plate for flux analysis using 2 μl Growth-Factor-Reduced Matrigel Matrix (BD Bioscience) diluted 1:2 in medium per well and rested overnight. The cells were starved for 2 h in Hank’s Salt (HBSS) supplemented with 2 mM glucose and 2 mM glutamax (Thermo Fisher Scientific). Oxygen consumption was measured using an XF96 extracellular flux analyser (Seahorse Bioscience) with sequential injections of 13 mM glucose, 4 μM oligomycin (Sigma) and 10 μM antimycin + 4 μM rotenone (Sigma).
FACS analysis, gene profiling and endocrine cell reaggregation
For gene profiling, qPCR, transmission electron microscopy and western blotting, cells from isolated islets were sorted and analysed using FACS-Aria III (BD Bioscience). The percentage and purity of both endocrine populations were controlled by cytospins and quantitative PCR (qPCR) and 90% purity of Nkx6.1+ β-cell subpopulations was achieved using an optimized FACS sorting scheme. For FACS analysis, cells were fixed in 4% PFA and intracellular staining for Nkx6.1 was performed by using goat anti-Nkx6.1 (R&D Systems, AF5857, 1:100) and donkey anti-goat APC (Dianova, 705-136147, 1:1.000). The ratio of Fltp lineage+ and Fltp lineage− populations during pregnancy was calculated over the Nkx6.1+ cells. The results were analysed using FlowJo software. For qPCR on sorted populations, total RNA was extracted from FACS-sorted endocrine cells, using a miRNeasy micro kit (Qiagen) and amplified with the Ovation PicoSL WTA System. For gene profiling, total RNA was extracted from FACS-sorted FVR endocrine cells, using a miRNeasy micro kit (Qiagen) and amplified with the Ovation PicoSL WTA System V2 in combination with the Encore Biotin Module (Nugen). Amplified cDNA was hybridized on Affymetrix Mouse Gene 1.0 ST arrays. Staining and scanning were performed according to the Affymetrix expression protocol including minor modifications as suggested in the Encore Biotion protocol. Expression console (v.1.3.0.187, Affymetrix) was used for quality control and to obtain annotated normalized RMA (robust microarray analysis) gene-level data (standard settings including median polish and sketch-quantile normalization). TaqMan qPCR was assessed using the probe list provided in Supplementary Table 5, according to the manufacturer’s instructions. For quantification of mitochondrial DNA copy number, DNA from sorted cells was extracted using QIamp DNA Blood Mini kit (QiAgen) according to the manufacturer’s instructions. The ratio of mtDNA to nDNA was determined by Syber green qPCR (Dynamo Flash Syber green, Byozym) using the primers listed in Supplementary Table 5. For endocrine cell re-aggregation, FVR+ and FVR−-enriched β-cells were FACS sorted and cultured by shaking in DMEM (containing 4.5 g/l glucose, 5% FCS and 1% v/v penicillin/streptomycin) diluted 1:1 in conditioned medium (DMEM containing 4.5 g/l glucose, 5% FCS and 1% v/v penicillin/streptomycin) from MS1 cells (pancreatic islets endothelial cell line, ATCC CRL-2279, LGC Standards GmbH). Cells were aggregated for 6 days; 10 μM Y-27632 (Santa Cruz) was added on day 1 and left for the first 3 days, at which point the medium was changed. One day before the GSIS assay, the cells were cultured in DMEM (containing 11 mM glucose, 5% FCS and 1% v/v penicillin/streptomycin).
GSIS assays and pancreatic insulin and glucagon concentration
Isolated murine islets were allowed to recover overnight in islet medium (DMEM containing 11 mM glucose, 10% v/v FBS, 1% v/v penicillin/streptomycin). For the static GSIS assay, islets of the same size were handpicked and assayed with different glucose concentrations as previously described39. For static GSIS assays on endocrine aggregates, modified Krebs Ringer phosphate Hepes (KRPH) buffer supplemented with 0.1% BSA was used. The aggregates were incubated for 30 min in KRPH with 2.8 mM glucose and sequentially incubated with different glucose concentrations as indicated. For static GSIS assays on human microislets, sequential incubation with 2.8 mM glucose and 16.8 mM glucose in KRPH buffer was assessed for 2 h each. Assessment of pancreatic insulin and glucagon concentration was performed by acid ethanol extraction. Murine insulin and glucagon concentrations were determined using Ultrasensitive Insulin ELISA kit (Cristal Chem) and Mouse Glucagon ELISA Kit (Cristal Chem), respectively, and normalized over protein content. Human insulin concentrations were determined using an Ultrasensitive Insulin ELISA kit (Mercodia).
Association analysis
The study population consisted of 2,228 white individuals at risk for type 2 diabetes (family history of type 2 diabetes, body mass index (BMI) ≥ 27 kg/m2, impaired fasting glycaemia and/or previous gestational diabetes) recruited from the ongoing Tübingen Family study for type 2 diabetes. All participants underwent assessment of medical history, smoking status and alcohol consumption habits; the subjects furthermore agreed to undergo physical examination, routine blood tests and oral glucose tolerance tests (OGTTs). Only individuals with complete phenotypic and genotypic data sets and documented absence of medication known to influence glucose tolerance, insulin sensitivity or insulin secretion were included. All study participants gave informed written consent to the study and protocols adhered to the Declaration of Helsinki.
The study protocol was approved by the Ethics Committee of the Eberhard Karls University, Tübingen.
OGTT and laboratory measurements
A standardized 75 g OGTT was performed following a 10 h overnight fast. For the determination of plasma glucose, insulin and C-peptide levels, venous blood samples were drawn at baseline and at time-points 30, 60, 90 and 120 min of the OGTT40. Plasma glucose levels (in mmol/l) were measured with a bedside glucose analyser (glucose oxidase method, Yellow Springs Instruments). Plasma insulin and C-peptide levels (in pmol/l) were determined by commercial chemiluminescence assays for ADVIA Centaur (Siemens Medical Solutions). BMI was calculated as weight divided by squared height (in kg/m2). OGTT-derived insulin sensitivity was estimated as proposed earlier41: 10,000/[c(Glc0)*c(Ins0)*c(Glcmean)*c(Insmean)]½ (with c = concentration, Glc = glucose and Ins = insulin). OGTT-derived insulin secretion was estimated as area under the curve (AUC) Cpep0–30/AUC Glc0–30 according to the formula: [c(Cpep0)+c(Cpep30)]/[c(Glc0)+c(Glc30)] (Cpep = C-peptide).
Selection of tagging SNPs and genotyping
Based on publicly available data from the 1000 Genomes Project (http://browser.1000genomes.org/index.html), we analysed in silico a genomic area on human chromosome 1q23.3 spanning the CFAP126 gene (3.143 kb, five exons, four introns, located on the reverse strand) and 2 kb of the gene’s 5′-flanking region. Within the analysed CFAP126 locus, 36 SNPs were found. Using the tagger analysis tool of Haploview (see http://www.broadinstitute.org/scientific-community/science/programs/medical-and-population-genetics/haploview/%20haploview), seven tagging SNPs were identified that cover all the other common SNPs (minor allele frequency ≥ 0.01) with an r2 ≥ 0.8. These SNPs were rs17399583 (C/T), rs11584714 (C/G) and rs57835711 (C/G) in the 5′-flanking region, rs114482063 (A/T) and rs182840301 (C/A) in intron 1 and rs16832872 (G/A) and rs75715534 (C/G) in intron 3. For genotyping, DNA was isolated from whole blood using a commercial kit (NucleoSpin, Macherey & Nagel). The tagging SNPs were genotyped using the mass spectrometry system massARRAY from Sequenom and the manufacturer’s iPLEX software (Sequenom). The call rates were ≥ 96.1%. The mass spectrometric results were validated in 50 randomly selected subjects by bidirectional sequencing and both methods gave 100% identical results.
RNA sequencing expression data analysis of human pancreatic islets
RNA sequencing expression data on human pancreatic islets from n = 119 donors including n = 14 diagnosed type 2 diabetes (T2D) donors were obtained from the Human Tissue Laboratory (HTL) of Lund University Diabetes Centre (LUDC)42. Gencode v14 RefSeq was used for gene annotation. Across-samples normalization was performed using the TMM normalization method43. HbA1c information available for 106 human pancreatic islets donors was used for classification into a) healthy/non-diabetic donors (normal glucose tolerance, NGT, n = 66, HbA1c < 6), b) pre-diabetic donors (impaired glucose tolerance, IGT, n = 21, 6 < HbA1c < 6.5) and c) diabetic donors (T2D, n = 19, diagnosed + HbA1c ≥ 6.5). Associations of gene expression in human pancreatic islets with blood glucose level (NGT+IGT+T2D status) was performed with ANOVA for linear testing and using the Kruskal–Wallis method for nonlinear and non-parametric testing. Correcting for multiple testing was performed with FDR procedure and 5% significance threshold was selected for the analysis. All calculations were done using R language for statistical computations44. The upper whisker of the boxplots represents min (max (x), Q_3 + 1.5 * IQR) and the lower whisker max (min(x), Q_1 − 1.5 * IQR). IQR represents the box length Q_3 − Q_1.
Statistical analysis
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment. Statistical analysis was performed using GraphPad Prism 6 Software (GraphPad Software, USA). Values were compared using unpaired or paired t-tests or ANOVA as indicated for each experiment. P values of <0.05 were considered statistically significant. Data are expressed as means ± s.e.m. unless otherwise specified. Statistical analysis of microarray profile was performed using the statistical programming environment R (R Development Core Team) implemented in CARMAweb. The P value of the limma t-test was used as a criterion for significance (P < 0.01) and in addition filtered for fold-changes >1.5 to potentially increase biological relevance. Multiple testing corrections were not applied, because with only two replicates this approach would be too stringent (the most significant gene after Benjamini-Hochberg correction had a false discovery rate <22%). Heatmaps were generated with CARMAweb45 and cluster dendrograms with the R script hclust. GO term enrichments were performed for 1.5-fold regulated genes with a P-value <0.005 using the GePS module in the Genomatix Software Suite v3.1 (Genomatix). The pathway analyses (P < 0.05) were generated through the use of QIAGEN’s Ingenuity Pathway Analysis (IPA, QIAGEN Redwood City, http://www.qiagen.com/ingenuity). For association analysis, continuous variables with non-normal distribution were loge-transformed before statistical analysis. Multiple linear regression analysis was performed using the least-squares method. In the regression models, insulin secretion was chosen as an outcome variable, the SNP genotype (in the dominant inheritance model) as an independent variable and gender, age, BMI and insulin sensitivity as confounding variables. SNP–BMI interaction effects on insulin secretion were tested by ANCOVA with gender, age and insulin sensitivity as confounding variables. When testing all seven tagging SNPs in parallel, a Bonferroni-corrected P-value <0.0073 was considered statistically significant. For all analyses, the statistical software JMP 8.0 (SAS Institute) was used.
References
Salomon, D. & Meda, P. Heterogeneity and contact-dependent regulation of hormone secretion by individual B cells. Exp. Cell Res. 162, 507–520 (1986)
Heimberg, H. et al. Heterogeneity in glucose sensitivity among pancreatic β-cells is correlated to differences in glucose phosphorylation rather than glucose transport. EMBO J. 12, 2873–2879 (1993)
Smukler, S. R. et al. The adult mouse and human pancreas contain rare multipotent stem cells that express insulin. Cell Stem Cell 8, 281–293 (2011)
Razavi, R. et al. Diabetes enhances the proliferation of adult pancreatic multipotent progenitor cells and biases their differentiation to more β-cell production. Diabetes 64, 1311–1323 (2015)
Katsuta, H. et al. Single pancreatic β cells co-express multiple islet hormone genes in mice. Diabetologia 53, 128–138 (2010)
Gegg, M. et al. Flattop regulates basal body docking and positioning in mono- and multiciliated cells. eLife 3, e03842 (2014)
Bonner-Weir, S. Morphological evidence for pancreatic polarity of β-cells within islets of Langerhans. Diabetes 37, 616–621 (1988)
Gu, C. et al. Pancreatic beta cells require NeuroD to achieve and maintain functional maturity. Cell Metab. 11, 298–310 (2010)
Blum, B. et al. Functional β-cell maturation is marked by an increased glucose threshold and by expression of urocortin 3. Nat. Biotechnol. 30, 261–264 (2012)
In’t Veld, P. & Marichal, M. Microscopic anatomy of the human islet of Langerhans. Adv. Exp. Med. Biol. 654, 1–19 (2010)
Meda, P., Perrelet, A. & Orci, L. Increase of gap junctions between pancreatic B-cells during stimulation of insulin secretion. J. Cell Biol. 82, 441–448 (1979)
Wang, Y. & Nathans, J. Tissue/planar cell polarity in vertebrates: new insights and new questions. Development 134, 647–658 (2007)
Wallingford, J. B. Planar cell polarity and the developmental control of cell behavior in vertebrate embryos. Annu. Rev. Cell Dev. Biol. 28, 627–653 (2012)
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51–56 (2011)
McCracken, K. W. et al. Modelling human development and disease in pluripotent stem-cell-derived gastric organoids. Nature 516, 400–404 (2014)
Orci, L. et al. A morphological basis for intercellular communication between α- and β-cells in the endocrine pancreas. J. Clin. Invest. 56, 1066–1070 (1975)
Lange, A. et al. Fltp(T2AiCre): a new knock-in mouse line for conditional gene targeting in distinct mono- and multiciliated tissues. Differentiation 83, S105–S113 (2012)
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007)
Rieck, S. & Kaestner, K. H. Expansion of β-cell mass in response to pregnancy. Trends Endocrinol. Metab. 21, 151–158 (2010)
Speier, S. et al. Noninvasive in vivo imaging of pancreatic islet cell biology. Nat. Med. 14, 574–578 (2008)
Golson, M. L., Misfeldt, A. A., Kopsombut, U. G., Petersen, C. P. & Gannon, M. High fat diet regulation of β-cell proliferation and β-cell mass. Open Endocrinol. J. 4, (2010). 10.2174/1874216501004010066
Kitsou-Mylona, I., Burns, C. J., Squires, P. E., Persaud, S. J. & Jones, P. M. A role for the extracellular calcium-sensing receptor in cell-cell communication in pancreatic islets of Langerhans. Cell. Physiol. Biochem. 22, 557–566 (2008)
Schulze, T., Morsi, M., Bruning, D., Schumacher, K. & Rustenbeck, I. Different responses of mouse islets and MIN6 pseudo-islets to metabolic stimulation: a note of caution. Endocrine 51, 440–447 (2016)
Granot, Z. et al. LKB1 regulates pancreatic β cell size, polarity, and function. Cell Metab. 10, 296–308 (2009)
Kalwat, M. A. & Thurmond, D. C. Signaling mechanisms of glucose-induced F-actin remodeling in pancreatic islet β cells. Exp. Mol. Med. 45, e37 (2013)
Staiger, H. et al. Common genetic variation in the human FNDC5 locus, encoding the novel muscle-derived ‘browning’ factor irisin, determines insulin sensitivity. PLoS One 8, e61903 (2013)
Gao, T. et al. Pdx1 maintains β cell identity and function by repressing an α cell program. Cell Metab. 19, 259–271 (2014)
Wang, Z., York, N. W., Nichols, C. G. & Remedi, M. S. Pancreatic β cell dedifferentiation in diabetes and redifferentiation following insulin therapy. Cell Metab. 19, 872–882 (2014)
Talchai, C., Xuan, S., Lin, H. V., Sussel, L. & Accili, D. Pancreatic β cell dedifferentiation as a mechanism of diabetic β cell failure. Cell 150, 1223–1234 (2012)
Jonas, J. C. et al. Chronic hyperglycemia triggers loss of pancreatic β cell differentiation in an animal model of diabetes. J. Biol. Chem. 274, 14112–14121 (1999)
Szot, G. L., Koudria, P. & Bluestone, J. A. Murine pancreatic islet isolation. J. Vis. Exp. 2007, 255 (2007)
Ravassard, P. et al. A genetically engineered human pancreatic β cell line exhibiting glucose-inducible insulin secretion. J. Clin. Invest. 121, 3589–3597 (2011)
Miyazaki, J. et al. Establishment of a pancreatic β cell line that retains glucose-inducible insulin secretion: special reference to expression of glucose transporter isoforms. Endocrinology 127, 126–132 (1990)
Chmelova, H. et al. Distinct roles of β-cell mass and function during type 1 diabetes onset and remission. Diabetes 64, 2148–2160 (2015)
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012)
Dalton, A. J. A chrome-osmium fixative for electron microscopy. Anat. Rec. 121, 281 (1955)
Feuchtinger, A. et al. Image analysis of immunohistochemistry is superior to visual scoring as shown for patient outcome of esophageal adenocarcinoma. Histochem. Cell Biol. 143, 1–9 (2015)
Gegg, M. et al. Flattop regulates basal body docking and positioning in mono- and multiciliated cells. Elife 3, e03842 (2014)
Wu, Y. et al. Growth hormone receptor regulates β cell hyperplasia and glucose-stimulated insulin secretion in obese mice. J. Clin. Invest. 121, 2422–2426 (2011)
Stefan, N. et al. Polymorphisms in the gene encoding adiponectin receptor 1 are associated with insulin resistance and high liver fat. Diabetologia 48, 2282–2291 (2005)
Matsuda, M. & DeFronzo, R. A. Insulin sensitivity indices obtained from oral glucose tolerance testing: comparison with the euglycemic insulin clamp. Diabetes Care 22, 1462–1470 (1999)
Fadista, J. et al. Global genomic and transcriptomic analysis of human pancreatic islets reveals novel genes influencing glucose metabolism. Proc. Natl Acad. Sci. USA 111, 13924–13929 (2014)
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010)
R Computing Team. R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, Vienna, Austria, 2014)
Rainer, J., Sanchez-Cabo, F., Stocker, G., Sturn, A. & Trajanoski, Z. CARMAweb: comprehensive R- and bioconductor-based web service for microarray data analysis. Nucleic Acids Res. 34, W498–W503 (2006)
Acknowledgements
We thank A. Raducanu, A. Böttcher and E. Schlüssel for comments and discussions; A. Theis, B. Vogel and K. Diemer for technical support; the Human Tissue Laboratory (HTL) of Lund University Diabetes Centre (LUDC) for high-quality RNA sequencing expression data from human pancreatic islets donors; to InSphero for human microislets; the European Consortium for Islet Transplantation (ECIT) for human islets; and R. Scharfmann for the EndoC-β H1 cell line. The human islet research was supported by a Strategic Research Grant from the Swedish Research Council (2009-1039). A.M. was funded by the Helmholtz post-doctoral fellowship program. This work was supported by an Emmy-Noether Fellowship, the European Union (ERC starting grant Ciliary Disease) and the HumEn project from the European Union's Seventh Framework Programme for Research, Technological Development and Demonstration under grant agreement No. 602587 (http://www.hum-en.eu/). This work was funded (in part) for H.L. and J.B. by the Helmholtz Alliance ICEMED – Imaging and Curing Environmental Metabolic Diseases, through the Initiative and Networking Fund of the Helmholtz Association. We thank the Helmholtz Society, Helmholtz Portfolio Theme 'Metabolic Dysfunction and Common Disease, German Research Foundation and German Center for Diabetes Research (DZD e.V.) for financial support. This work was supported with funds for S.S. from the Emmy-Noether Program, the Center for Regenerative Therapies Dresden-DFG Research Center for Regenerative Therapies Dresden, Cluster of Excellence (CRTD), the DFG-Collaborative Research Center/Transregio 127 and the German Ministry for Education and Research to the German Centre for Diabetes Research and to the Network of Competence for Diabetes.
Author information
Authors and Affiliations
Contributions
E.B., A.M. and M.G. conceived and performed the experiments and wrote the manuscript. M.B. established the 3D in vitro cultures. S.S.R. and E.B. helped to perform human islet experiments. N.M., C.L., H.Z., M.J., M.T. and J.G. conceived and performed mitochondria analysis. M.I. and J.B. performed the microarray analysis. M.A. and A.F. analysed mitochondria by TEM. F.M., H.S. and H-U.H. performed the SNP analysis of FLTP in humans. O.K. provided human islets and N.O. provided RNA sequencing data. R.W-S. supervised and trained E.B. in human epidemiology and genetics. H.C., J.A.C. and S.S. performed the islet transplantation and analyzed the data set. H.L. conceived the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
H.L., M.G., E.B. and A.M. are inventors of a European Patent (PCT/EP2015/056664) based on this work.
Additional information
Reviewer Information Nature thanks C. Wollheim and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 FVR expression is induced in compacted islet structures and correlates with high Nkx6.1 expression.
a, Overview of a cleared whole-mount pancreas from an FltpZV/+ mouse at E18.5, stained with anti-Nkx6.1 and anti-GFP antibodies and scanned with a laser confocal microscope. b–e, In compacted islets (dotted line), high FVR expression correlates with high Nkx6.1 expression. f–i, In cord-like structures of endocrine cells that left the ductal epithelium (dotted line), low Nkx6.1 expression correlates with low FVR expression. j, Quantification of the mean fluorescent intensity of Venus (FVR)+ and Nkx6.1+ cells in compacted and cord-like structures (n (mice) = 3, 5 cord-like structures, 6 compacted islets; ***P = 0.0005, two sided unpaired t-test). k, Quantification of Nkx6.1 (red) and Venus (green) single-positive cells as well as all Nkx6.1–Venus double-positive cells (yellow) in compacted and cord-like structures (n (mice) = 3, cord-like structures 570 cells, compacted islets 1,827 cells; **P = 0.0064 for Venus, **P = 0.0041 for Nkx6.1, *P = 0.0182 for Venus–Nkx6.1, two-sided unpaired t-test). Scale bars, 100 μm (a), 20 μm (b–i). Data represent mean ± s.e.m.
Extended Data Figure 2 FVR− and FVR+ β-cell subpopulations exhibit differential proliferative capacity and inter-islet heterogeneity in the head and tail of the pancreas.
a, b, LSM images showing FVR, Nkx6.1 and EdU (a) and quantification (b) of EdU+ in FVR− and FVR+ β-cell subpopulations in control and pregnant (G15.5) mice (green/red arrow heads indicate proliferating FVR+/FVR− β-cells. n (control) = 4 (8,500 Nkx6.1+ cells), n (pregnant) = 4 (10,938 Nkx6.1+ cells); **P (Ctrl) = 0.0049, **P (pregnant) = 0.0059, two-sided unpaired t-test). c, Scatter plot of FVR+ β-cell distribution relative to number of β-cells per islet in FltpZV/+ mice (n (mice) = 10, n (islets) = 512). d, Quantification of islet size distribution in head and tail of FltpZV/+ pancreas sections (n (mice) = 4, n (islets) = 299; *P (20–50) = 0.0419, *P (>100) = 0.0238, two-sided unpaired t-test). e, f, LSM images (e) and quantification (f) of FVR+ β-cell distribution in the pancreas head and tail (n (mice) = 4, n (islets) = 299, n (Nkx6.1+ cells) = 19,054; ****P < 0.0001, two-sided unpaired t-test). g, Quantification of EdU incorporation among FVR− and FVR+ β-cell subpopulations relative to number of β-cells per islet in pregnant (G15.5) FltpZV/+ mice (n (mice) = 4, n (islets) = 196, n (Nkx6.1+ cells) = 10,938). h, LSM images showing Nkx6.1-, p27- and FVR-expressing cells in islets of FltpZV/+ pregnant (G15.5) mice. i, Quantification of p27 and Venus fluorescent intensity shows that p27-high cells have a higher Venus fluorescent intensity than p27-low cells in pregnant (G15.5) mice (n (mice) = 3, n (p27 and Venus positive cells) = 4,204). j, Quantification of Venus and EdU fluorescent intensity shows that EdU-positive cells have lower Venus fluorescent intensity than EdU-negative cells in pregnant (G15.5) mice (n (mice) = 3, n (p27+, EdU+ and Nkx6.1+ cells) = 75, n (p27+ EdU− Nkx6.1+ cells) = 4,845). Scale bars, 50 μm (a, e, h). Data represent mean ± s.e.m.
Extended Data Figure 3 Validation of the sorting scheme and molecular and cellular analysis of FVR− and FVR+ endocrine subpopulations.
a, b, Quantification (a) and LSM images (b) of FACS-sorted FVR populations for the microarray analysis show equal numbers of endocrine cell lineages among the subpopulations (n (DAPI+ cells) = 789). c, Detection and purification of four distinct endocrine subpopulations (I–IV) on the basis of granularity and Venus expression. d, e, Quantification (d) and LSM images (e) of sorted FVR subpopulations I and II confirms high enrichment of Nkx6.1+ β-cells whereas III and IV exhibit high enrichment of glucagon+ α-cells (n (DAPI+ cells) = 3,275). f, Expression analysis of endocrine hormones and the exocrine marker Amy2a3 reveals minimal contamination with other endocrine cell types (n (independent experiments) = 6 (FVR+) and 5 (FVR−)). g, h, Validation of the microarray analysis by real-time qPCR of β-cell-enriched subpopulations (I, II) for β-cell maturation, mitochondria and metabolic signalling pathway genes (Atp5b, Hadh, Wnt4, Grb10, Wnt5b, Ins1, GFP, Fltp, Pcsk1, Gipr, Mafa: n (independent experiments) = 6; Bace2, Alpk1, Npy: n (independent experiments) = 5, Slc2a2: n (independent experiments) = 3; *P < 0.05, **P < 0.01, ***P < 0.001, two-sided unpaired t-test). i, Real-time qPCR of mtDNA (S12) versus nDNA (HBB) from β-cell enriched populations (I, II) (n (independent experiments) = 5, expression in the FVR+ subpopulation normalized on each FVR− subpopulation, error bars represent c.i.). j, Scatter plot of regulated polarity genes among FVR endocrine cells in microarray analysis (thin lines mark 1.5-fold difference in expression, P < 0.05, limma t-test). Scale bars, 100 μm (b, e). Data represent mean ± s.e.m. except where stated.
Extended Data Figure 4 Transmission electron microscopy (TEM) analysis of Fltp− and Fltp+ β-cell subpopulations and quantification of mitochondria membrane length.
a, b, TEM images of immunogold-labelled β-cells of FltpT2AiCre/+;Gt(ROSA)26mTmG/+ mice reveal no structural differences in Golgi complex (a) or endoplasmic reticulum (b). c, d, TEM images and detected areas of the algorithm used for mitochondria analysis (c) and analysis of inner mitochondria membrane length normalized on the mitochondria area in FVR+ and FVR− β-cells (d) (n (independent experiments) = 2, n (FVR+ cells) = 14, n (FVR− cells) = 16, n (mitochondria) = 544). e, TEM images of mitochondria in FVR+ compared to mitochondria in FVR− β-cells. Scale bars, 500 nm (a (Golgi complex and gold staining), c, e), 200 nm (a (insulin granules), b). Data represent mean.
Extended Data Figure 5 Analysis of oxygen consumption rate (OCR), mitochondria network and GSIS in FVR+ and FVR− β-cells.
a, OCR of FACS-sorted β-cell-enriched FVR+ and FVR− subpopulations after Antimycin A subtraction. b, Quantification of absolute OCR rates of FACS-sorted β-cell-enriched FVR+ and FVR− subpopulations after Antimycin A subtraction. FVR+ n (technical replicates) = 9, FVR− n (technical replicates) = 4, for each technical replicate 3 × 104 cells were plated; endocrine cells were isolated from 9 mice; two-sided unpaired t-test. c, OCR of FACS-sorted β-cell-enriched FVR+ and FVR− subpopulations after Antimycin A subtraction and baseline normalization. d, Quantification of relative OCR of FACS-sorted β-cell-enriched FVR+ and FVR− subpopulations after Antimycin A subtraction and baseline normalization. FVR+ n (technical replicates) = 9, FVR− n (technical replicates) = 4, two-sided unpaired t-test. e, h, LSM pictures of immunohistochemistry of isolated FltpZV/+ mouse islets cultured under high (16.5 mM) and low (2 mM) glucose concentrations with single-cell reconstructions (Imaris) showing the reconstructed membrane (light brown), the nuclei (white, green and blue) and the mitochondria (red). f, g, i, j, Enlargement of a single Nkx6.1+ cell under high (f) and low glucose (i) and an Nkx6.1+ FVR+ cell under high (g) and low glucose (j) with reconstruction of the nucleus, the membrane and the mitochondria. k, Quantification of the mitochondrial morphology of Nkx6.1–FVR double-positive cells (yellow) and Nkx6.1 single-positive cells (white) under high and low glucose stimulation (n (Nkx6.1+ FVR+) = 9, n (Nkx6.1+ FVR− in high glucose) = 4, n (Nkx6.1+ FVR− in low glucose) = 6, from three islets for low glucose and two islets for high glucose). l, FACS-sorted and re-aggregated β-cell-enriched subpopulations reveal differences in GSIS and arginine depolarization (n (independent experiments) = 3, n (aggregates FVR+) = 16, n (aggregates FVR−) = 12; *P = 0.0206, **P = 0.007, two-sided unpaired t-test). Scale bars, 10 μm (e, h), 2 μm (f, j), 3 μm (g, i). Data represent mean ± s.e.m.
Extended Data Figure 6 Fltp genetic lineage tracing of endocrine subpopulation dynamics upon islet transplantation and HFD.
a, Maximum intensity projection of laser backscatter to determine the volume of the same transplanted FltpT2AiCre/+;Gt(ROSA)26mTmG/+ islet before (Pre-diet) and after 8 weeks of HFD. b, Quantification of the total islet volume (transplanted FltpT2AiCre/+;Gt(ROSA)26mTmG/+) in mice on normal diet and HFD (n (islets) = 25 (from 4 mice on normal diet), n (islets) = 34 (from 5 mice on HFD); ****P < 0.0001, Sidak’s multiple comparison test). c, Quantification of glucose levels during intraperitoneal GTT by AUC quantification (n (mice on normal diet) = 4, n (mice on HFD) = 5; *P = 0.0223, **P = 0.0064, Sidak’s multiple comparison test). d, Longitudinal quantification of the relative volumes of islet size and vessel volume during engraftment time (n (islets) = 15 (from 4 mice); ****P < 0.0001, ***P = 0.0027, two-sided unpaired t-test). e, Quantification of mG and mT endocrine cell numbers every 10 μm in a confocal z-stack during engraftment (n (islets) = 15 (from 4 mice); ****P < 0.0001, two-sided unpaired t-test). Scale bars, 50 μm (a). Data represent mean ± s.e.m.
Extended Data Figure 7 3D architecture and Wnt/PCP induce β-cell maturation in Min6 cells.
a, c–e, g, h, LSM images (a, e) and quantification (c, g (3 days culture), d, h, (7 days culture)) of Nkx6.1 and MafA mean fluorescent intensity in Min6 cells in 2D and 3D culture (c, n (independent experiments) = 3, ***P = 0.0002; d, n (independent experiments) = 4, *P = 0.0264; g, n (independent experiments) = 2; h, n (independent experiments) = 3; Sidak’s multiple comparison test). b, f, Fluorescent intensity of Nkx6.1 (b) and MafA (f) in a single Min6 cell in 2D and 3D culture. White lines in a and e indicate measured regions. i, j, Gene expression analysis of Min6 cells cultured in 2D and 3D culture by qPCR (n (independent experiments) = 4; *P < 0.05, **P < 0.01, ***P < 0.001, two-sided unpaired t-test). k, l, LSM images (k) and quantification (l) of Nkx6.1+ Min6 cells in floating 3D culture treated with or without Wnt5a (n (independent experiments) = 3; n (cells) = 5,419 (control), 2,419 (Wnt5a); **P = 0.0036, Sidak’s multiple comparison test). m, n, Western blot (m) and quantification (n) of Nkx6.1 and Fltp protein in Min6 after treatment for 6 days with or without Wnt5a in 2D or 3D culture (western blots were normalized on γ-tubulin expression and standardized on 2D control, n (independent experiments) = 5). Scale bars, 20 μm (a, e, k). Data represent mean ± s.e.m.
Extended Data Figure 8 Modulation of maturation and proliferation through WNT signalling in human microislets and EndoC-β H1 human β-cell line and SNP association with insulin secretion defects in the FLTP gene.
a–e, LSM images (a–d; proliferating cells marked by arrow heads) and quantification (e) of NKX6.1–EdU double-positive EndoC-β H1 cells treated with or without WNT3A, WNT4 or WNT5A (n (independent experiments) = 3, 1,617 cells counted; *P = 0.019, two-sided unpaired t-test). f, g, Western blot (f) and quantification (g) of pJNK in EndoC-β H1 treated for 4 h with or without WNT4 (n (independent experiments) = 4; *P = 0.027, two-sided paired t-test). h, Fold change in insulin secretion in response to high glucose in human microislets treated with or without WNT4 or WNT5A (3 human donors; Ctrl: n (microislets) = 23, WNT4: n (microislets) = 25, WNT5A: n (microislets) = 26; *P = 0.0423, two-sided unpaired t-test). i, RNA sequencing expression data for SLC2A2 in human pancreatic islets from healthy donors (NGT, n = 66), pre-diabetic donors (IGT, n = 21) and type 2 diabetic donors (T2D, n = 19; one-way ANOVA). j, k, The minor allele of FLTP rs75715534 was significantly associated with increased insulin secretion in lean subjects (P = 0.0035), but not in obese subjects (P = 0.07). Adjustment of the insulin secretion index AUC Cpep0–30/AUC Glc0–30 (n = 2,228) was achieved by multiple linear regression modelling with gender, age and OGTT-derived insulin sensitivity as confounding variables. Scale bars, 25 μm (a, d). Data represent mean ± s.e.m. (e, g, h); mean diamonds represent mean ± 95% c.i. (j, k).
Extended Data Figure 9 Knockout of Fltp has no effect on β-cell maturation or proliferation.
a–d, Single plane LSM images of cryosections from adult FltpZV/+ (a, c) and FltpZV/ZV (b, d) mice. Some Venus (green), Nkx6.1 (red) and MafA (white; a, b, white arrowheads) or Ucn3 (white; c, d, dashed lines) triple-positive cells are highlighted. e, Quantification of Venus, Nkx6.1 and Ucn3 (white) or MafA (black) triple-positive cells shows no difference in maturation status of β-cells in FltpZV/+ and FltpZV/ZV mice (FltpZV/+: n (mice) = 3, MafA 2,349 cells, Ucn3 1,212 cells; FltpZV/ZV: n (mice) = 3, MafA 867 cells, Ucn3 1,825 cells). f–i, LSM images of immunohistochemistry on cryosections from P11 and adult Fltp+/+ (f, P11; h, adult) and FltpZV/ZV (g, P11; i, adult) mice showing the proliferation of islet cells. j, k, Quantification of proliferating β- and non-β-cells in FltpZV/ZV and Fltp+/+ mice at P11 (j) (Fltp+/+: n (mice) = 4, 6,556 β-cells and 4,325 non-β-cells; FltpZV/ZV: n (mice) = 4, 6,709 β-cells and 5,868 non-β-cells) and in adults (k) (Fltp+/+: n (mice) = 7, 9,803 β-cells and 4,338 non-β-cells; FltpZV/ZV: n (mice) = 9, 14,957 β-cells and 7,955 non-β-cells). Scale bar, 20 μm (a), 10 μm (b), 5 μm (c, d), 15 μm (f–i). Data represent mean ± s.d.
Extended Data Figure 10 Analysis of the metabolic phenotype of FltpZV/ZV mice.
a, Quantification of β-cell number in islets of Langerhans from Fltp+/+ and FltpZV/ZV mice at P11 or adult (n (mice) = 3 (P11 Fltp+/+, 6,556 Nkx6.1+ cells and FltpZV/ZV, 6,791 Nkx6.1+ cells), n (mice) = 6 (adult Fltp+/+, 3,202 Nkx6.1+ cells) and n (mice) = 4 (adult FltpZV/ZV, 6,263 Nkx6.1+ cells)). b, Intraperitoneal GTT in FltpZV/ZV and Fltp+/+ mice (Fltp+/+ n (mice) = 7 and FltpZV/ZV n (mice) = 7; **P = 0.0022, two-sided unpaired t-test). c, d, Concentration of glucagon (c) and insulin (d) in pancreatic tissue from FltpZV/ZV and Fltp+/+ mice (Fltp+/+ n (mice) = 5 and FltpZV/ZV n (mice) = 5). e, ITT in FltpZV/ZV and Fltp+/+ mice (Fltp+/+ n (mice) = 5 and FltpZV/ZV n (mice) = 3). f, Plasma insulin level in fasted Fltp+/+ and FltpZV/ZV mice (Fltp+/+ n (mice) = 11 and FltpZV/ZV n (mice) = 10; *P = 0.0405, two-sided unpaired t-test). g, GSIS in isolated islets from Fltp+/+ and FltpZV/ZV mice (Fltp+/+ n (mice) = 5 and FltpZV/ZV n (mice) = 5; *P = 0.0263, one-way ANOVA, Bonferroni’s multiple comparison test). Data represent mean ± s.e.m.
Supplementary information
Supplementary Information
This file contains the Raw Data files for Figures 2k, 4h, 4j and Extended Data Figures 7m and 8f. (PDF 218 kb)
Supplementary Tables
This file contains Supplementary Tables 1-5. (PDF 1661 kb)
Source data
Rights and permissions
About this article
Cite this article
Bader, E., Migliorini, A., Gegg, M. et al. Identification of proliferative and mature β-cells in the islets of Langerhans. Nature 535, 430–434 (2016). https://doi.org/10.1038/nature18624
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18624
- Springer Nature Limited
This article is cited by
-
Capillary contact points determine beta cell polarity, control secretion and are disrupted in the db/db mouse model of diabetes
Diabetologia (2024)
-
Differential CpG methylation at Nnat in the early establishment of beta cell heterogeneity
Diabetologia (2024)
-
The role of wnt signaling in diabetes-induced osteoporosis
Diabetology & Metabolic Syndrome (2023)
-
Single-nucleus RNA sequencing of human pancreatic islets identifies novel gene sets and distinguishes β-cell subpopulations with dynamic transcriptome profiles
Genome Medicine (2023)
-
Integrated transcriptomics contrasts fatty acid metabolism with hypoxia response in β-cell subpopulations associated with glycemic control
BMC Genomics (2023)