Abstract
Implantation of the blastocyst is a developmental milestone in mammalian embryonic development. At this time, a coordinated program of lineage diversification, cell-fate specification, and morphogenetic movements establishes the generation of extra-embryonic tissues and the embryo proper, and determines the conditions for successful pregnancy and gastrulation. Despite its basic and clinical importance, this process remains mysterious in humans. Here we report the use of a novel in vitro system1,2 to study the post-implantation development of the human embryo. We unveil the self-organizing abilities and autonomy of in vitro attached human embryos. We find human-specific molecular signatures of early cell lineage, timing, and architecture. Embryos display key landmarks of normal development, including epiblast expansion, lineage segregation, bi-laminar disc formation, amniotic and yolk sac cavitation, and trophoblast diversification. Our findings highlight the species-specificity of these developmental events and provide a new understanding of early human embryonic development beyond the blastocyst stage. In addition, our study establishes a new model system relevant to early human pregnancy loss. Finally, our work will also assist in the rational design of differentiation protocols of human embryonic stem cells to specific cell types for disease modelling and cell replacement therapy.
Similar content being viewed by others
Main
Recently established in vitro implantation platforms in the mouse have expanded our knowledge of post-implantation development and recapitulated in vivo early developmental landmarks1,2. However, extrapolation from mouse to human is limited since mammalian embryos display species-specific differences in post-implantation morphology and many molecular markers have not been validated3,4,5. Therefore, to gain insights into the self-organizing abilities of human embryos, we sought to establish an in vitro attachment platform. To determine the dynamics of cell fate specification, we monitored the morphology and the expression of cell-type-specific markers in pre- and post-attachment embryos: the inner cell mass (ICM) and epiblast (Epi) markers OCT4 and NANOG, the ICM and primitive endoderm (PE) marker GATA6, and the trophectoderm (TE) markers CDX2 and GATA3 (ref. 6). Additionally, to distinguish TE subtypes, we used cytokeratin 7 (CK7) and β-human chorionic gonadotropin (HCGB)7. Embryos were fixed and stained every other day starting at day post-fertilization (d.p.f.) 6, and counterstained with 4′,6-diamidino-2-phenylindole (DAPI) to detect and count nuclei and phalloidin to delineate cell boundaries (Supplementary Video 1).
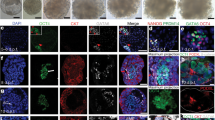
The pre-implantation human blastocyst (d.p.f. 6–7) is a hollow ball of cells composed of tightly packed, small nuclei in the ICM and larger cells in the TE (n = 8; Fig. 1a–c, Extended Data Figs 1a–c and 2a, Supplementary Table and Supplementary Videos 2 and 3). All ICM cells showed high intensity OCT4 staining (OCT4HI), with a subset having high GATA6 (GATA6HI) in a salt-and-pepper manner (Fig. 1d–f and Extended Data Fig. 2b–d). NANOG was observed exclusively in OCT4+ ICM cells, which did not stain for GATA6 (Fig. 1m–q). No GATA3 or CDX2 were detected in the ICM. All TE cell nuclei stained for GATA3, as well as low intensity OCT4 (OCT4LO), and GATA6 (GATA6LO; Fig. 1g–j) but not NANOG. Variable levels of CDX2 were also detected in TE (CDX2+; Fig. 1k–l and Extended Data Fig. 2e, f); however, in most cases, surprisingly, the signal was localized to the cytoplasm instead of the nucleus (Fig. 1k–l). No HCGB and weak, non-filamentous CK7 were detected (Extended Data Fig. 2g; n = 6). The ubiquitous OCT4 pattern we report is in agreement with previous staining of human blastocysts at d.p.f. 5–7 (refs 6, 8, 9), and stands in contrast to the mouse where it is ICM specific. GATA6 expression in the ICM is different from the mouse and previous reports in humans6,8,9. In the mouse, the salt-and-pepper ICM distribution becomes restricted to PE and physically sorted from the Epi population by late blastocyst10. Our data suggest that physical sorting between Epi and PE has not yet occurred in human d.p.f. 6, even if the transcriptional profile is consistent with specification of presumptive Epi and PE fate. While both CDX2 and GATA3 are TE specific, we find that GATA3 is a better marker of human TE because of its high signal and consistent nuclear localization. Our results also confirm the molecular profiles from single-cell RNA sequencing studies of laser-dissected blastocyst (d.p.f. 5–6)11, and expand them by providing new spatial resolution. Thus, at d.p.f. 6, we define two cell populations in the ICM: OCT4HI/NANOG+/GATA6− (presumptive Epi precursors), and OCT4HI/GATA6HI/NANOG− (presumptive PE precursors); we also define one population of TE cells: OCT4LO/GATA6LO/GATA3+/CDX2+.
a, DIC image of a d.p.f. 6 human blastocyst; scale bar, 100 μm. b, Cartoon of a d.p.f. 6 embryo with salt-and-pepper distribution of OCT4 (green) and OCT4/GATA6 (red) ICM cells, and GATA3 (blue); mural (arrow) and polar (arrowhead) TE cells. c–q, Immunostaining of d.p.f. 6 blastocysts. c, Three-dimensional rendering of the front half of a d.p.f. 6 blastocyst (left) and ICM-zoom (right). DAPI (white) and phalloidin (magenta, actin, virtual channel). d–j, Three-dimensional rendering of a d.p.f. 6 blastocyst (top) and ICM-zoom (bottom, box in j) stained for OCT4 (green), GATA6 (red, virtual channel), GATA3 (blue) (n = 3–8); arrows indicate high marker levels in ICM, arrowhead low levels in TE. k–l, Whole embryo (top) and ICM-zoom (bottom, box in l) of the d.p.f. 6 blastocyst from c; OCT4 (green), GATA6 (red, virtual channel), CDX2 (cyan) (n = 5). m–q, ICM-zoom of d.p.f. 6; OCT4 (green), NANOG (magenta), GATA6 and CD24 (red, n = 3). Scale bar, 100 μm for whole embryos, 20 μm for ICM-zooms.
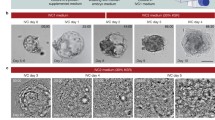
We next allowed blastocysts to attach using a recently established mouse in vitro implantation platform1,2. We found that most human blastocysts (70%, n = 91) attached between d.p.f. 7 and 8 (Fig. 2a, b). Interestingly, despite their random original orientation, by d.p.f. 7 the blastocysts always attached on the side of the polar TE, the portion of TE that is the closest to the ICM (Fig. 2c–e). At d.p.f. 8, all embryos adopted a flattened structure (Fig. 2b, e, f, Extended Data Fig. 3a, Supplementary Table and Supplementary Video 4). While the total cell number remained constant, the relative number of cells contributing to different territories changed (Extended Data Fig. 1a–c and Supplementary Table). Two discrete territories emerged from the ICM, and formed the Epi and PE lineages, respectively characterized by OCT4HI/ GATA6− and OCT4−/ GATA6HI (Fig. 2g–i). Epi cells formed a tight core at the centre of the attached embryos (Fig. 2g). Phalloidin staining was extremely faint in all embryos in two separate experiments, suggesting qualitatively different actin status in the immediate post-attachment period. In the TE, OCT4 and GATA6 staining completely disappeared. CDX2 remained at very low and variable levels and was restricted to weak nuclear staining of a subset of TE cells at the periphery (Fig. 2j–l), while GATA3 demarcated all OCT4− and GATA6− TE nuclei surrounding Epi and PE (Fig. 2m–q). At this stage, all GATA3+ cells also showed strong filamentous CK7 staining (Fig. 2r–s), a marker of general TE lineage and cytotrophoblast (CT), suggesting molecular progression of TE differentiation compared with d.p.f. 6; however, no HCGB, a marker of syncytiotrophoblast (ST), was detected (Extended Data Fig. 3b)7. Taken together, this evidence suggests that in humans, Epi/PE cell sorting occurs at a later stage than in the mouse, around the time of implantation.
a, b, DIC images of (a) attaching d.p.f. 7.5 embryos with the TE (top left) collapsing onto the embryo and (b) attached and compacted d.p.f. 8. Scale bar, 50 μm. c–e, Cartoons of blastocyst (c) floating, (d) orienting to attach on the side of the polar TE, and (e) attached; Epi (green), PE (red), TE (blue). f–p, Optical sections of d.p.f. 8 embryos stained with the indicated markers (n = 4). Scale bar, 20 μm. q, Three-dimensional rendering of the embryo in f–p with OCT4 (green), GATA6 (red), and GATA3 (blue), flanked by xz (top) and yz (side) views at coordinates indicated by yellow lines. Scale bar, 50 μm. r, s, Optical sections of d.p.f. 8 embryo stained with the indicated markers. Scale bar, 20 μm. t, u, Carnegie stage 5a section (t) and cartoon (u); Epi (green), PE (red).
At d.p.f. 10, the attached embryos increased in size while maintaining a flattened morphology (Fig. 3a–c, Extended Data Fig. 1a–c and 3b, Supplementary Table and Supplementary Video 5). The OCT4HI Epi at the centre of the embryo was more compacted, and formed a cavity delineated by phalloidin: the putative amniotic cavity (Fig. 3d–h, Extended Data Figs 4a–r and 5 and Supplementary Videos 5, 6, 7). To ask if the Epi population had progressed in its differentiation, we stained for CD24, which has been reported to increase in primed (Epi-derived) versus ‘naive’ mouse and human embryonic stem cells (ESCs)12. CD24 was selectively observed on the surface of human Epi cells starting only at d.p.f. 10 (n = 4 of 4; Fig. 3v–w and Extended Data Fig. 4s–x), and was not detected at d.p.f. 6 (Fig. 1o; n = 3 of 3) or d.p.f. 8 (Extended Data Fig. 3c; n = 4 of 4), suggesting a developmental transition. All Epi nuclei were also NANOG+ (Extended Data Fig. 4s–x). It is therefore the d.p.f. 10 Epi population (OCT4+/NANOG+/CD24+), and not the d.p.f. 6 or d.p.f. 8 Epi, that most closely matches the profile of human ESCs12, which supports a recent hypothesis on the developmental origin of human ESCs9. GATA6HI PE cells were distributed in a layer juxtaposed to one side of the Epi, delineating the bi-laminar germ disc (Fig. 3c–p and Extended Data Fig. 4a–r). On the other side of the PE cells, a second, larger cavity self-organized: the putative yolk sac cavity (Fig. 3i–p, Extended Data Figs 4l–r and 5 and Supplementary Videos 5 and 6). A novel population of CDX2+/GATA6LO/OCT4LO cells, never described before in other mammals, lined the yolk sac cavity in all embryos examined (n = 4). While it is possible that these cells represent parietal endoderm, we believe that they are a novel population because of their unique molecular signature. We therefore named these cells ‘yolk sac trophectoderm’ or ysTE (Fig. 3p). GATA3+ cells continued to demarcate the entire TE (Extended Data Fig. 4a–k), which also co-stained for CK7 (Fig. 3q–u). However, a subset of GATA3+/CK7+ cells now stained for HCGB (Fig. 3t), suggesting the emergence of the HCGB+ ST lineage7. Collectively, the architecture of the embryo at this stage closely resembles Carnegie stage 5b–c13, with amniotic cavity, yolk sac cavity, bi-laminar disc in between, and TE differentiation (Fig. 3x–y). This demonstrates that the embryo alone can direct both lineage specification and diversification, as well as tissue morphogenesis and architectural organization, without maternal input.
a, b, DIC image of a d.p.f. 10 embryo (scale bar, 50 μm) (a) and cartoon of a section through the embryo (b); Epi (green), amniotic cavity (arrow), PE (red), yolk sac cavity (arrowhead), ysTE (orange), TE (blue). c, Three-dimensional rendering of a d.p.f. 10 embryo stained with OCT4 (green), GATA6 (red), and GATA3 (blue) (n = 4), flanked by xz (top) and yz (side) views at coordinates indicated by the yellow lines, with amniotic (arrow) and yolk sac cavities (arrowhead). Scale bar, 50 μm. d, Optical section of the central region from c; OCT4 (green), GATA6–phalloidin (red), and CDX2 (cyan) (n = 4). Scale bar, 20 μm. e–h, Optical section of the dashed box in d, but on a z-plane 6 μm higher, with the amniotic cavity at the centre (arrow) lined by PE (arrowhead). Scale bar, 20 μm. i–p, Optical section of the box in d, but on a z-plane 4 μm below with the yolk sac cavity at the centre lined by PE (white arrowhead) and ysTE (yellow arrowhead). Scale bar, 20 μm; q–u, Optical section of a d.p.f. 10 embryo stained with the indicated markers (n = 4). Scale bar, 100 μm. v, w, Optical section of the Epi/PE area of a d.p.f. 10 embryo stained with OCT4 (green) and GATA6–CD24 (red) (n = 4). Scale bar, 20 μm. x, y, Carnegie stage 5b section (x) and cartoon (y) with Epi (green), amniotic cavity, PE (red), yolk sac cavity, and ysTE (orange). ph, phalloidin.
At d.p.f. 12, the amniotic and yolk sac cavities, albeit present, appeared collapsed (Fig. 4a–l, Extended Data Figs 6 and 7a–j, Supplementary Table and Supplementary Video 8). Epi and PE expression profiles were unchanged from d.p.f. 10, while TE showed further differentiation and organization. HCGB staining increased in intensity and clusters expanded to generate multinucleated cells with lacunae, the in vivo site of maternal blood vessel invasion, therefore adding functional cytological characteristics of ST lineage progression (Fig. 4n–q and Extended Data Fig. 7k–o). TE subtypes self-organized in concentric rings, with a ring of CT surrounding the Epi, multiple foci of ST towards the edge of the embryo, and a layer of CT adjacent to the substrate (Fig. 4r–v). These features closely mimic and provide the first molecular signature of histological samples13 (Carnegie stage 5b–c; Fig. 4w–x). We concluded our experiments at d.p.f. 14, in accordance with internationally recognized bioethical guidelines14,15. Outgrowths at this late stage lost interpretable relation to in vivo correlates13 (n = 8), suggesting the limitations of our two-dimensional culture environment.
a, b, DIC image of a d.p.f. 12 embryo (a) (scale bar, 50 μm) and cartoon of a side view (b) with Epi (green), amniotic and yolk sac cavities, PE (red), ysTE (orange), and TE (blue). c, Three-dimensional rendering of a d.p.f. 12 embryo stained for OCT4 (green), GATA6 (red), and GATA3 (blue) (n = 2); flanked xz (top) and yz (side) views at indicated coordinates. d–m, Epi zoom of c stained with the markers indicated; the arrow in e points at the amniotic cavity. Scale bar, 20 μm. n–q, Optical section of the embryo in c re-stained with the indicated markers (n = 2); scale bar, 100 μm. r–v, Appearance of ST phenotypes; multinucleated ST cells, DAPI (white), GATA3 (blue), phalloidin (red), HCGB (yellow), and nascent lacuna (arrow). Scale bar, 20 μm. w–x, Carnegie stage 5c section (w), and cartoon (x) of Epi (green), amniotic and yolk sac cavities, PE (red), ysTE (orange), CT, ST, and lacunae in TE (blue).
In this study we show that by simply providing an attachment substrate, human blastocysts self-organize to recapitulate many key features of in vivo development, surprisingly independently of maternal input at least up to d.p.f. 12. Additionally, the differences between human and mouse embryos, including unique architecture, cell types, and tissue organization, emphasize the necessity of working with human embryos to understand human development. By defining the molecular composition of cell lineages in the early embryo, our data provide a new ex vivo reference to reinterpret controversial discrepancies between mouse and human stem cells and their derivatives. This platform also constitutes a new model for a variety of poorly understood placental and embryonic disorders and early pregnancy loss. These and future studies on the origin and regulation of early embryonic cell types will lead to more developmentally based and rational approaches to reprogramming, disease modelling, and cell replacement therapies.
Methods
Ethics statement
This work was approved by The Rockefeller University Institutional Review Board and was in accord with University policy for research involving human embryos. The informed consent process for embryo donation complied with the NAS Guidelines for Human Embryonic Stem Cell Research and the Tri-Institutional Stem Cell Research Operating Procedures for ESCRO Reviewed Research. The Tri-Institutional Stem Cell Initiative Embryonic Stem Cell Research Oversight (Tri-SCI ESCRO) Committee determined that the use of human embryos for basic developmental studies did not require review and approval by the TriSCI ESCRO Committee because it did not involve derivation of, or research with, human pluripotent stem cells.
Embryo culturing protocol
Cryopreserved d.p.f. 5 embryos were thawed using a THAW-KIT1 (Vitrolife) according to the manufacturer’s instructions. Briefly, embryos were thawed by placing the cryovials in a 37 °C water bath for about 30–60 s, collected, transferred, and incubated in ETS1 for 5 min at room temperature, 22–23 °C transferred and incubated in ETS2 for 5 min at room temperature, transferred and incubated in ETS3 for 7.5 min at room temperature, transferred and incubated in cryo-PBS for 5 min at room temperature and 5 min at 37 °C (not in the incubator). Embryos were morphologically scored upon thaw and 24 h after culture. To remove the zona, embryos were briefly exposed to acidic Tyrode’s solution (Sigma). Embryos were seeded on ibiTreat microscopy μ plates (Ibidi), filled with pre-equilibrated IVC1 medium (Cell Guidance Systems). Half of the medium was replaced after 24 and 48 h in culture. Zona-free embryos typically attached 48–60 h after seeding. At this point, the medium was exchanged to IVC2 (Cell Guidance Systems), and embryos were cultured for the indicated times, with half of the medium being replaced with fresh medium every 24 h. The embryo culturing and imaging protocol has been deposited on Nature Protocol Exchange, doi 10.1038/protex.2016.022.
Immunofluorescence
Embryos were fixed in 4% PFA in PBS for 30 min at 4 °C, washed three times in PBS, quenched in wash buffer (PBS + 0.1% Triton X-100) plus 100 mM glycine, blocked (wash buffer + 0.1% sodium azide + 10% normal donkey serum), stained for 3–12 h with primary antibodies in blocking buffer, washed, stained with labelled secondary antibodies (Alexa Fluor 488, 555, and 647), DAPI, and phalloidin 647, and imaged (see below). To maximize the molecular information from each embryo, we followed several sequential staining strategies. First, in preliminary stains we found that GATA6 was exclusively nuclear and generated almost no background staining; therefore, to maximize information content from each embryo, we stained all embryos with phalloidin-647 (the same fluorescence channel with GATA6) since these could be segregated during image analysis using a three-dimensional segmentation mask on the DAPI to generate a phalloidin-only or a GATA6-only virtual channel. Floating blastocysts were stained with either DAPI–CDX2–OCT4–GATA6 and phalloidin in the same channel, or DAPI–GATA3–OCT4–GATA6 and phalloidin in the same channel since CDX2 and GATA3 antibodies were of the same species. After analysis and quantification of all attached embryos (d.p.f. 8–14), CDX2 cell numbers were found to be very small and weakly stained (except for the ysTE cells at d.p.f. 10–14); therefore all attached embryos were then re-stained with GATA3 (same species, then same colour (Alexa Fluor 555) as CDX2) and reimaged. The staining for GATA3 was substantially brighter than the previous mouse CDX2-555 signal, except for the ysTE cells at d.p.f. 10–14. Therefore an extra virtual channel could be superimposed on the same embryos to maximize spatial information, indicating bona fide GATA3 expression in previously CDX2− or CDX2low TE cells. Similarly, since GATA6 nuclear staining was absent at d.p.f. 14, we re-stained two embryos each with rabbit anti-OTX2 and rabbit anti-SOX2. To further enhance molecular resolution, we then probed TE lineage diversification by staining previously stained attached embryos or fresh day 7 blastocysts. For previously stained attached embryos, we observed that phalloidin completely detached by 2 weeks after staining and was no longer detectable, therefore leaving the 647 channel open except for GATA6+ nuclei. We therefore blocked embryos with proteolytic Fc fragments (mouse and rabbit), then with unlabelled donkey anti-mouse and -rabbit IgG Fab1, then stained with 647- and 594-labelled donkey anti-mouse and -rabbit IgG Fab1 (all Jackson), and imaged the embryos to measure background and residual crosstalk. We next stained with primary antibodies to trophoblast markers CK7 and HCGB followed by 647- and 594-labelled donkey anti-mouse and -rabbit IgG Fab1 and reimaged. Primary antibodies used were the following: goat-OCT4, Santa Cruz 8628, 1:500; rabbit-GATA6, Cell Signaling 23345, 1:1,000; mouse-NANOG, BD 560482, 1:500; mouse-GATA3, Pierce MA1-028, 1:100; mouse-CDX2, BioGenex CDX2-88, 1:10; rabbit-CK7 Abcam 181598, 1:400; mouse-HCGB, Abcam ab9582, 1:100; mouseCD24-PE conjugate, Abcam 7729, 1:5; rabbit-SOX2, Cell Signaling 35795, 1:200; rabbit-OTX2, Abcam 114138, 1:200. Labelled secondary antibodies, DAPI, and phalloidin-Alexa Fluor 647 (Life) were incubated for 3–12 h and washed before imaging the samples.
Image acquisition, analysis, and rendering
Z-stack images were acquired on a Leica SP8 inverted confocal microscope at 12 bits in 1024 pixels × 1024 pixels using an HCX PL APO CS × 20/0.75 numerical aperture air-immersion or an HC PL APO CS2 × 40/1.10 numerical aperture water-immersion objective, at 1 Airy unit pinhole diameter and 0.985–1.9 μm spacings with manual laser-power Z intensity-compensation, and avoiding detector saturation with laser excitation/HyD detector emission settings, in nanometres, as follows: 405 diode laser 405/415–486, WLL 492/500–550, WLL 552/560–621, and WLL 649/657–710 for DAPI, Alexa Fluor 488, 555, 594, and 647 respectively. When 594 was imaged, the detector setting for Alexa Fluor 555 was adjusted to 560–580, and a fifth track was added, 594 WLL 594/602–631. Images were then deconvolved with a three-dimensional blind algorithm (ten iterations) using AutoDeblur X software (Autoquant). Images were rendered and visualized in Imaris, where total and marker-positive nuclei were quantified by a spot finding algorithm with manual adjustment for fluorescence intensity threshold and quality. Where antibodies or tissue generated non-specific background staining, a nuclear mask was first generated from the DAPI signal, and spots were counted on masked channels as above. Final images were processed and assembled using ImageJ. For the three-dimensional renderings and movies, individual channels were gamma-adjusted to enhance visualization.
Statistical analysis
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Errors and error bars represent the s.e.m. from a minimum of four independent embryos unless otherwise indicated. The figures display representative results. Unless otherwise specified, the results were the same across all the embryos analysed.
References
Bedzhov, I., Leung, C. Y., Bialecka, M. & Zernicka-Goetz, M. In vitro culture of mouse blastocysts beyond the implantation stages. Nature Protocols 9, 2732–2739 (2014)
Bedzhov, I. & Zernicka-Goetz, M. Self-organizing properties of mouse pluripotent cells initiate morphogenesis upon implantation. Cell 156, 1032–1044 (2014)
Rossant, J. Mouse and human blastocyst-derived stem cells: vive les differences. Development 142, 9–12 (2015)
Rossant, J., Chazaud, C. & Yamanaka, Y. Lineage allocation and asymmetries in the early mouse embryo. Phil. Trans. R. Soc. Lond. B 358, 1341–1349 (2003)
Berg, D. K. et al. Trophectoderm lineage determination in cattle. Dev. Cell 20, 244–255 (2011)
Niakan, K. K. & Eggan, K. Analysis of human embryos from zygote to blastocyst reveals distinct gene expression patterns relative to the mouse. Dev. Biol. 375, 54–64 (2013)
Li, Y. et al. BMP4-directed trophoblast differentiation of human embryonic stem cells is mediated through a DeltaNp63+ cytotrophoblast stem cell state. Development 140, 3965–3976 (2013)
Roode, M. et al. Human hypoblast formation is not dependent on FGF signalling. Dev. Biol. 361, 358–363 (2012)
O’Leary, T. et al. Tracking the progression of the human inner cell mass during embryonic stem cell derivation. Nature Biotechnol. 30, 278–282 (2012)
Schrode, N., Saiz, N., Di Talia, S. & Hadjantonakis, A. K. GATA6 levels modulate primitive endoderm cell fate choice and timing in the mouse blastocyst. Dev. Cell 29, 454–467 (2014)
Blakeley, P. et al. Defining the three cell lineages of the human blastocyst by single-cell RNA-seq. Development 142, 3151–3165 (2015)
Shakiba, N. et al. CD24 tracks divergent pluripotent states in mouse and human cells. Nature Commun. 6, 7329 (2015)
O’Rahilly, R. & Müller, F. Developmental stages in human embryos: revised and new measurements. Cells Tissues Organs 192, 73–84 (2010)
National Research Council Human Embryonic Stem Cell Research Advisory Committee, Board on Life Sciences, Board on Health Sciences Policy, National Research Council, and Institute of Medicine. Final Report of the National Academies’ Human Embryonic Stem Cell Research Advisory Committee and 2010 Amendments to the National Academies’ Guidelines for Human Embryonic Stem Cell Research (National Academies Press, 2010)
International Society for Stem Cell Research. Guidelines for the Conduct of Human Embryonic Stem Cell Research, Version 1 (ISSCR, Northbrook, Illinois, 2006)
Acknowledgements
We thank the members of the Brivanlou laboratory for their advice and criticisms, in particular C. Nchako, S. Tse for technical assistance, and members of the Zernicka-Goetz laboratory for their advice on how to culture embryos through attachment. We also thank A.K. Hadjantonakis for discussions, A. Wilkerson for support, and A. Brivanlou and P. Carleton-Evans for their comments on the manuscript. This work was supported by a STARR Foundation grant (number 2013-026) and Rockefeller Private funds. Images were obtained using instrumentation in The Rockefeller University Bio-Imaging Resource Center purchased with grant funds from the Sohn Conference Foundation. The Carnegie stage images are used with permission from the Virtual Human Embryo Project (http://virtualhumanembryo.lsuhsc.edu). We give special thanks for technical advice on imaging to A. North, K. Thomas, and P. Ariel, and on image analysis and rendering to T. Tong. This work would not have been possible without the generosity of the people who consented to donate their embryos to research, to whom we are indebted.
Author information
Authors and Affiliations
Contributions
A.D., G.C., and L.P. performed experiments; A.D. and G.C. analysed experiments; M.Z.-G. was instrumental in teaching and transferring knowledge on the mouse technology to A.D.; E.S. provided criticism of the work and manuscript; A.H.B. conceived and designed the project, established contact with the source of the biological material, provided guidance and advice throughout the work, and interfaced with the Institutional Review Board at The Rockefeller University; all authors contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Quantification of cells positive for transcriptional markers over d.p.f. 6–12 of in vitro culture.
a, Time course of total cell numbers via DAPI quantification. Confocal data sets for each embryo were scored for total cell numbers in Imaris. The mean ± s.e.m. is shown for d.p.f. 6, 8, 10, 12 (n = 8, 4, 4, 2, embryos per time point respectively over n = 2, 2, 2, 1 independent experiments). A total of 91 embryos were thawed and 64 (70%) showed developmental progression and attached; 26 of 64 (40%) embryos showed normal size and morphology and were analysed for markers of cell fate. The remaining embryos all showed abnormal development, represented by few (0–5) and scattered OCT4 cells, and were excluded from analysis as obvious failures of development. Subsequent to staining and analysis of this set of embryos with these core markers, six d.p.f. 7 and five d.p.f. 10 embryos showing normal OCT4 development were also stained and analysed for additional markers. b, Total cells expressing cell type markers, mean ± s.e.m. for OCT4, GATA6, CDX2, and GATA3: d.p.f. 6 (n = 8, 7, 5, 3 respectively); d.p.f. 8 and 10 (n = 4 embryos per marker); d.p.f. 12 (n = 2 embryos per marker). The line subdividing the OCT4 population identifies the two subpopulation of OCT4+ cells: OCT4HI (bottom of the bar), passing a threshold for high-intensity staining; and OCT4LO (top of the bar), passing a low threshold (but not the higher one for OCT4HI) for staining. At d.p.f. 6, OCT4HI cells were confined to the ICM while the OCT4LO cells represented the TE; between d.p.f. 8 and 12, OCT4LO cells were also co-expressing CDX2 (ysTE cells), while OCT4HI cells were confined to the Epi. GATA6 was scored for high or low (HI/LO) expression at d.p.f. 6 only, where GATA6HI cells (bottom of the bar) localized to the ICM and GATA6LO cells localized to the TE. See also Supplementary Table. c, Number of cells expressing lineage-specific markers expressed as a percentage of total nuclei; each bar shows mean ± s.e.m. for OCT4, GATA3, GATA6, and CDX2. The line subdividing the OCT4 bars at d.p.f. 6–12 and the GATA6 bar at d.p.f. 6 represents the high or low (HI/LO) expressing cells as explained above.
Extended Data Figure 2 Human-specific transcriptional profiles of ICM and TE in d.p.f. 6 human blastocysts.
a, Additional three-dimensional rendering of the front half of a d.p.f. 6 blastocyst (left), and ICM-zoom (right). DAPI (white) identifies all nuclei and phalloidin (magenta, actin, virtual channel) shows intercellular boundaries between TE cells. A three-dimensional segmentation mask on the DAPI channel was subtracted from the raw GATA6–phalloidin channel to yield a phalloidin-only virtual channel for these panels. This embryo is the same as displayed in Fig. 1d–j. Scale bar, 100 μm for whole embryos, 20 μm for ICM-zooms. b–f, Top, whole embryo, and bottom, ICM-zooms, stained for OCT4 (green), GATA6 (red, virtual channel), CDX2 (cyan), and serial merges. b, c, Arrows show that ICM cells have high-intensity OCT4 with or without GATA6 staining; arrowheads show low-level expression of OCT4 and GATA6 in TE cells; e, f, CDX2 staining is weak and predominantly cytoplasmic. The GATA6-only channel is the result of an inverse mask using three-dimensional segmentation on DAPI to remove extra-nuclear phalloidin staining, resulting in nuclear the GATA6-only virtual channel. This embryo is the same as displayed in Fig. 1k–l. Scale bar, 100 μm for whole embryos, 20 μm for ICM-zooms. g, No HCGB and very weak CK7 signal was observed in d.p.f. 7 embryos (n = 6). Shown is a section through a d.p.f. 7 embryo stained with DAPI (white), OCT4 (green), HCGB (yellow), and CK7 (magenta). Scale bar, 100 μm.
Extended Data Figure 3 Molecular signature of d.p.f. 8–10 embryos.
a, Embryo reconstruction of a d.p.f. 8 embryo oriented at a 30° pitch; individual channels are shown in greyscale and then merged as OCT4 (green), GATA6–phalloidin (red), GATA3 (blue). Phalloidin staining of all d.p.f. 8 embryos was exceptionally weak compared with all other d.p.f. (all embryos analysed in two separate experiments); scale bar, 50 μm. b, No HCGB staining was detected at d.p.f. 8 (n = 4). Section through a d.p.f. 8 embryo showing DAPI (white), OCT4 (green), HCGB (yellow), and CK7 (magenta). Scale bar, 50 μm. c, No CD24 staining was detected at d.p.f. 8 (n = 4). Section through a d.p.f. 8 embryo showing DAPI (white), OCT4 (green), CD24 (red). Scale bar, 50 μm. d, Embryo reconstruction of a d.p.f. 10 embryo oriented at a 30° pitch; individual channels are shown in greyscale and then merged as OCT4 (green), GATA6–phalloidin (red), GATA3 (blue). Phalloidin staining of this embryo was so intense that the original underlying GATA6 nuclear stain is not visible in this reconstruction; scale bar, 50 μm.
Extended Data Figure 4 Human-specific transcriptional profile of d.p.f. 10 embryos.
D.p.f. 10 in vitro attached embryos recapitulate the principal landmarks of Carnegie stage 5b (in vivo d.p.f. 9). a–k, Confocal z-sections of a d.p.f. 10 embryo stained for DAPI (white), OCT4 (green), GATA6 and phalloidin (GATA6-Ph, red), CDX2 (cyan), GATA3 (blue), and combinatorial merges. The box in f indicates the area including amniotic and yolk sac cavities, which is shown in Fig. 3d; scale bar, 50 μm. l–r, Additional confocal z-section of the boxed region in f, stained for DAPI (white), OCT4 (green), GATA6–phalloidin (red), CDX2 (cyan), and combinatorial merges, showing amniotic cavity formed by OCT4+-only Epi cells (arrow in m and n), flanking GATA6+-only PE cells (white arrowhead in n), and CDX2+/GATA6+/OCT4LO ysTE cells lining the yolk sac cavity (yellow arrowhead in n, o); the CDX2+/GATA6+/OCT4LO merge of this series is shown in Fig. 3d. Scale bar, 20 μm. s–x, Confocal z-section of the Epi/PE area of a d.p.f. 10 embryo stained for DAPI (white), OCT4 (green), GATA6 and CD24 (red), NANOG (magenta). This is the same embryo as in Fig. 3v, w. Scale bar, 20 μm.
Extended Data Figure 6 Tilted three-dimensional reconstruction of a d.p.f. 12 embryo.
The embryo reconstruction is oriented at a 30° pitch, with individual channels shown in greyscale and then merged as OCT4 (green), GATA6–phalloidin (red), GATA3 (blue). Phalloidin staining of this embryo was so intense upon re-staining that the original underlying GATA6 nuclear stain is not visible in this data set. Scale bar, 50 μm.
Extended Data Figure 7 Additional images from a d.p.f. 12 embryo.
a–e, Confocal z-section of the d.p.f. 12 embryo in Fig. 4c stained for DAPI (white), OCT4 (green), GATA6–phalloidin (GATA6-Ph, red), GATA3 (blue), and CDX2 (cyan). Scale bar, 100 μm. f–o, Confocal sections of the same d.p.f. 12 embryo stained and imaged first (f–j) for DAPI (white), OCT4 (green), GATA6–phalloidin (red), and GATA3 (blue) and then (k–o) re-stained and re-imaged for DAPI (white), OCT4 (green), CK7 (magenta), and HCGB (yellow). DAPI and OCT4 were used as landmarks to identify the same z-plane and the same cells between the two staining and imaging rounds. The arrow in j and o indicates an example of nascent lacuna, typical of ST cells at Carnegie stage 5c in vivo; the box indicates an example of multinucleated cells characteristic of ST lineage progression; zoom-ins of the area in the box are presented in Fig. 4r–v. Scale bar, 20 μm.
Supplementary information
Supplementary Table
This file contains Supplementary Table 1, which shows Cell scoring by marker and cell type. (PDF 109 kb)
Timecourse of in vitro human embryo attachment from DPF6 to DPF14
OCT4 (green), GATA3 (blue), GATA6-Phalloidin (red). For the 3D rendering of DPF14 only, OCT4 and GATA3 channels were thresholded using the spot-finding algorithm in Imaris and then masked to remove non-nuclear background. (MOV 29120 kb)
Example of DPF6 embryo
DAPI (blue), OCT4 (green), CDX2 (cyan), GATA6 (red), Phalloidin (white). For this 3D rendering animation, a DAPI mask was generated by Imaris spot finding and then used to remove nuclear GATA6 from the original GATA6-phalloidin channel (creating Phalloidin-only and GATA6-only virtual channels). The Phalloidin virtual channel was then normalized across z using default settings in Imaris. A DAPI nuclear mask was also used to isolate nuclear OCT4 signal from non-nuclear background staining. A few pieces of debris outside of the embryo were manually cropped in 3D. This embryo was imaged at 40x. (MOV 28454 kb)
Example of DPF6 embryo
OCT4 (green), GATA3 (blue), GATA6-Phalloidin (red) (MOV 24352 kb)
Example of DPF8 embryo.
OCT4 (green), GATA3 (blue), GATA6-Phalloidin (red) (MOV 17985 kb)
Example of DPF10 embryo and cavitation
DAPI (blue), OCT4 (green), CDX2 (cyan), GATA6-Phalloidin (red). A 3D DAPI mask was used to exclude non-nuclear CDX2 and OCT4 stainings from the raw channels. Video breakdown by seconds: 0-12, 3D render of DAPI, nuclear CDX2, nuclear OCT4, and GATA6-Phalloidin; 12-16, z-sections (2µm step) from the bottom to the top of stack; 16-35, OCT4 (unmasked channel) and GATA6-Phalloidin as z-sections from the top to the bottom of the stack; 35-45, CDX2 (unmasked channel) is added at the bottom z-plane and the slice traverses from the bottom to the top of the stack showing bright CDX2 co-staining with GATA6 and OCT4 in ysTE cells; 45-50, CDX2 (unmasked channel) was replaced with nuclear (masked) CDX2, the z-slice returns to the bottom of the stack leaving progressive overlaying sections on display; 50-55 zoom out of 3D render; 55-58 DAPI staining faded back in. (MOV 25859 kb)
Example of amniotic and yolk sac cavities in a DPF10 embryo
OCT4 (green), CDX2 (cyan), GATA6-Phalloidin (red). 3D reconstruction of a cropped volume containing EPI, amniotic and yolk sac cavities, and ysTE cells. An oblique, 1.5µm virtual section is imposed and the 3D rendering is removed to visualize the signal from the plane passing through the structure and back. This image was obtained at 40x. (MOV 26976 kb)
Example of amniotic cavity in a DPF10 embryo
OCT4 (green), CDX2 (cyan), GATA6-Phalloidin (red); Z-series fly-through reconstruction. (AVI 23591 kb)
Example of cytotrophoblast and syncytiotrophoblast phenotypes in a DPF12 embryo
OCT4 (green), GATA3 (blue), GATA6-Phalloidin (red). (MOV 24149 kb)
Source data
Rights and permissions
About this article
Cite this article
Deglincerti, A., Croft, G., Pietila, L. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016). https://doi.org/10.1038/nature17948
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17948
- Springer Nature Limited
This article is cited by
-
Distinct pathways drive anterior hypoblast specification in the implanting human embryo
Nature Cell Biology (2024)
-
Hypoblast from human pluripotent stem cells regulates epiblast development
Nature (2024)
-
Self-renewing human naïve pluripotent stem cells dedifferentiate in 3D culture and form blastoids spontaneously
Nature Communications (2024)
-
Self-patterning of human stem cells into post-implantation lineages
Nature (2023)
-
How to build a virtual embryo
Nature Methods (2023)