Abstract
Most human breast cancers have diversified genomically and biologically by the time they become clinically evident1,2,3. Early events involved in their genesis and the cellular context in which these events occur have thus been difficult to characterize. Here we present the first formal evidence of the shared and independent ability of basal cells and luminal progenitors, isolated from normal human mammary tissue and transduced with a single oncogene (KRASG12D), to produce serially transplantable, polyclonal, invasive ductal carcinomas within 8 weeks of being introduced either subrenally or subcutaneously into immunodeficient mice4. DNA barcoding5,6 of the initial cells revealed a dramatic change in the numbers and sizes of clones generated from them within 2 weeks, and the first appearance of many ‘new’ clones in tumours passaged into secondary recipients. Both primary and secondary tumours were phenotypically heterogeneous and primary tumours were categorized transcriptionally as ‘normal-like’. This system challenges previous concepts that carcinogenesis in normal human epithelia is necessarily a slow process requiring the acquisition of multiple driver mutations. It also presents the first description of initial events that accompany the genesis and evolution of malignant human mammary cell populations, thereby contributing new understanding of the rapidity with which heterogeneity in their properties can develop.
Similar content being viewed by others
Main
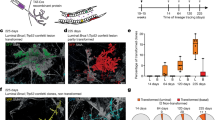
To investigate the susceptibility of different normal human mammary cell types to transformation under the influence of known oncogenes, we isolated CD49f+EpCAMlow basal cells (BCs), CD49f+EpCAM+ luminal progenitors (LPs), CD49f−EpCAM+ non-clonogenic luminal cells (LCs), and non-epithelial CD49f−EpCAM− stromal cells (SCs) at high purity (>97%) by fluorescence-activated cell sorting (FACS) from 37 normal human reduction mammoplasty samples depleted of endothelial and haematopoietic cells4,7,8 (Fig. 1a and Extended Data Tables 1 and 2). We then exposed one or more of these subsets to one or more oncogene-encoding lentiviral preparations (encoding complementary DNAs (cDNAs) for TP53R273C and green fluorescent protein (TP53R273C–GFP), PIK3CAH1047R and yellow fluorescent protein (PIK3CAH1047R–YFP)), and KRASG12D and mCherry (KRASG12D–mCherry) and, in some experiments, to a library of biologically neutral, barcoded lentiviral GFP vectors to allow subsequent clonal tracking of their progeny using a DNA sequencing approach5,6 (Extended Data Fig. 1a). The cells were then embedded in a collagen gel (0.3 × 105 to 16 × 105 cells per gel) and the gels transplanted into highly immunodeficient NOD-SCID Il2rg−/− (NSG) or NOD Rag1−/−Il2rg−/− (NRG) female mice.
a, Experimental design (n = number of experiments performed or tumours obtained). b, Examples of tumours compared with a control transplant which showed no evidence of a tumour when the mouse was killed. c, Bioluminescence measured over time in primary and secondary mice transplanted subcutaneously with oncogene- and luciferase-transduced BCs (blue) and LPs (red). SQ, subcutaneous; SR, subrenal.
In initial experiments, 2 × 105 irradiated C3H-10T1/2 fibroblasts were co-embedded in the gels which were then transplanted subrenally, followed by subcutaneous implantation of the recipients with a slow-release capsule containing 17-β-oestradiol and progestin (EP pellets)4, recognizing this would limit follow-up to 8 weeks because of incurred bladder toxicity. BCs and/or LPs isolated from 17 of 27 normal donors and exposed to all three oncogenic vectors produced tumours within 8 weeks (Fig. 1b) at similar overall frequencies (46% of BC isolates and 61% of LP isolates, respectively, Extended Data Fig. 1b). Identical treatment of LCs and SCs isolated from three of these samples did not produce any tumours in the same 8-week period. Both the BC- and LP-derived tumours resembled invasive ductal carcinomas (Fig. 2a and Extended Data Table 3) and were histologically very different from the organized bilayered structures generated in analogous xenografts of unmanipulated or simply barcoded normal human mammary cells4,6. Secondary female immunodeficient mice transplanted subcutaneously with a small piece (~25–33% of the initial tumour mass)5,9 from four of nine of these primary tumours (one BC- and eight LP-derived) developed palpable tumours within another 8 weeks (Extended Data Table 4).
a, Representative images of haematoxylin and eosin (H&E)- and antibody-stained sections from different primary BC- or LP- derived tumours arising from cells transplanted subrenally. Scale bar, 50 μm. b, Prevalence of cells in each tumour analysed expressing the indicated marker at different levels: white, negative; light grey, dim; dark grey, mid; black, bright. Each bar represents a single tumour. c, Unsupervised clustering (Spearman correlation and average linkage) of RNA sequencing data from de novo tumours compared with their matched cells of origin. d, Heat maps showing commonly differentially expressed genes between tumours and their matched cells of origin (top), and between BC- and LP-derived tumours in common with BCs and LPs (bottom). e, PAM50 and AIMS analyses of the RNA sequencing data from the de novo tumours.
FACS analysis showed most of the transduced cells in all primary tumours examined co-expressed all three fluorescent reporters, consistent with the high transduction efficiencies measured in separate cell aliquots maintained in vitro for 72 h after virus exposure, and a similar expression profile was maintained in the single secondary tumour similarly analysed (Extended Data Fig. 1c, d). Notably, when the three oncogene-encoding viruses were tested on their own, or in pairs, tumours were obtained with similar efficiency only when the KRASG12D vector was included, and even on its own (64/102 for all transductions that included KRASG12D compared with 1/12 when KRASG12D was not present; for more details, see Extended Data Fig. 1b and Extended Data Tables 1 and 2). PCR and Sanger sequencing confirmed the tumour cells contained the expected oncogene sequences including doubly and triply fluorescent cells isolated separately from tumours arising from cells initially exposed to three oncogenic vectors (Extended Data Fig. 2a–c).
Subsequent experiments demonstrated that invasive ductal carcinomas were also obtained at a similar frequency from both BCs and LPs (but not LCs or SCs from the same mammoplasty samples) when the transduced cells were transplanted subcutaneously without irradiated fibroblasts or EP pellets, even when the cells were exposed only to the KRASG12D vector (Extended Data Figs 1e and 2d and Extended Data Tables 1 and 2). These tumours could frequently also be serially passaged (Extended Data Fig. 2e and Extended Data Table 4) and their growth more accurately monitored by luciferase bioluminescence (Fig. 1c and Supplementary Table 1).
FACS analysis of 15 tumours showed that 48 ± 5% of the cells were human EpCAM+ and/or HLA+, with similar results for BC- and LP-derived tumours (Extended Data Fig. 1c, f). Immunohistochemical (IHC) analyses of tumour sections (Fig. 2a, b) showed 88% and 55% of primary BC- and LP-derived tumours contained >5% ERα+ cells (median = 58% and 8% ERα+ cells, respectively), but none contained >2% PR+ cells. HER2+ cells were present at similar frequencies (in 88% and 52% of BC- and LP-derived tumours, respectively). Frequencies of Ki67+ cells ranged from 2% to 30%, with only one secondary tumour containing as many as 70% Ki67+ cells. In contrast, cells expressing EGFR, MUC1 and K8/18 were prevalent in almost all tumours examined. High K5 expression, normally exclusive to BCs, was prevalent (median = 90% K5+ cells) in most LP-derived tumours, and less (median = 5% K5+ cells) in BC-derived tumours. Expression of CD44, a marker associated with undifferentiated epithelial cells, was also less prevalent in BC- compared with LP-derived tumour cells (median = 2% and 50%, respectively). K14, another marker of normal human BCs, was also variably detected in both BC- and LP-derived tumours.
Gene expression analyses (Extended Data Fig. 2f) showed that transcripts for vimentin (VIM) and N-cadherin (CDH2), normally found exclusively in BCs, were present at high levels in both LP- and BC-derived tumours, with similar results for E-cadherin (CDH1) and ELF5, genes normally expressed exclusively in LPs and LCs. However, transcript levels of SLUG (SNAI2), another BC marker, were strongly decreased in BC-derived tumours whereas transcript levels of both GATA3 and NOTCH3, two markers of LPs, showed little change. Cyclin-dependent kinase 1 (CDK1) was also highly expressed in all tumours, but other proliferation-associated genes, such as cyclin B1 (CCNB1) and PCNA, were highly expressed only in the LP-derived tumours. TERT transcripts remained at a similar level to that seen in the parental normal cell populations whereas those for VEGFA, HIF1A and MAPK3 were more variable.
RNA sequencing analysis was conducted on FACS-purified human cells isolated from six primary tumours generated from triply transduced cells (three from BCs and three from LPs) and the matched starting cells. Unsupervised clustering showed a closer relation of coding gene transcript levels in all six tumour populations to each other than to the normal cells from which the tumours had arisen (Fig. 2c). This prominent sharing of transcriptome changes in tumours derived both from BCs and from LPs suggests a key role of their mode of creation on their resultant molecular features. Specific differences in the gene expression changes that distinguished BC- and LP-derived tumours and their respective starting cell populations showed shared increased and decreased expression of 146 and 22 genes, respectively in both, indicative of a common gene signature in the transformants (Fig. 2d, top). Further analyses using either PAM50 (ref. 10) or AIMS classifier methodologies11 indicated the transcriptional profiles of the de novo tumours most closely resembled those of spontaneous human breast cancers classified as ‘normal-like’ (Fig. 2e).
However, the unsupervised clustering also indicated that the three BC- and three LP-derived tumours formed separate groups, suggesting some retained influence of their different origins. This was further supported by the finding that >20% (72) of the differentially expressed genes in the BC- and LP-derived tumours were similarly differentially expressed in the cells from which the tumours had arisen (Fig. 2d, bottom). Nevertheless, genes whose expression was upregulated in BC-derived tumours included several that are normally highly expressed in LPs and LCs but not BCs (for example, AR, ESR1, FOXA1, TOX3, EPCAM, EHF and ELF5). Conversely, the genes whose expression was upregulated in LP-derived tumours included several recognized BC-specific genes (for example, VIM, TP63, ACTA2, THY1 and CDH2, Supplementary Table 2).
Clonal analyses were performed on primary tumours obtained from 45 isolates of BCs and LPs both from DNA extracted directly from tumour tissue and from FACS-purified human cells (Extended Data Fig. 3). The results showed a high variability in the clone content of different tumours (up to 1,700 using a threshold of 70 cells per clone), regardless of the protocol used for their generation (Extended Data Fig. 4a, b). Calculated (minimal) frequencies of tumorigenic clone-forming cells (T-CFCs) using the total number of initial cells transplanted as the denominator, ranged from 1/23,000 to 1/150. Paired comparisons for tumours produced from BCs and LPs from the same donor also did not reveal any effect on T-CFC frequency (Fig. 3a). To estimate clone sizes, we first derived ‘relative’ clone size values by normalizing each tumour to the sum of its absolute clone sizes. We then pooled the data for all tumours in each group being assessed. The overall distribution of relative clone sizes, like the clone frequencies, was very broad and showed no evidence of any effect of the cell of origin, oncogene(s) used or the transplantation site (Fig. 3b).
a, Distribution of T-CFC frequencies (grey lines connect patient-matched BC- and LP-derived tumours, coloured lines show medians). b, Relative clone size distributions for all tumours in each group (left). Pie charts showing the clone size distributions in a representative tumour from each group (right). c, Distributions of the relative sizes of all clones pooled from all BC- and LP-derived secondary tumours (left). Pie charts showing the clone size distributions in a representative tumour from each group (right). d, Clones detected exclusively in primary (white) or secondary (grey) tumours, or in both (black). e, Clone size distributions of combined primary or secondary tumours subdivided into those detected exclusively in either primary (white) or secondary (grey) tumours or both (black).
Analysis of 15 secondary tumours showed their clonal content was often high but again very variable, regardless of their origin (Fig. 3c and Extended Data Fig. 5a, b). Calculated frequencies of secondary clones (with respect to the number of cells initially transplanted into primary mice) also yielded highly variable secondary T-CFC values but with no consistent difference from the calculated primary T-CFC frequencies. However, >75% of the clones detected in each secondary tumour were ‘new’; that is, not detected in the matching primary tumour (Fig. 3d). Moreover, most of the clones present in multiple sibling secondary tumours produced from a common primary tumour were also different from one another (two primary tumours analysed, Extended Data Fig. 5c). Overall the total measured T-CFC frequencies (calculated from the total number of different clones in the primary or secondary tumours combined) ranged from ~1/5,700 to ~1/120 (Extended Data Fig. 5d). The relative sizes of the clones present in secondary tumours were also highly variable (Fig. 3c). Interestingly, in secondary tumours, the median size of the ‘continuing’ clones (evident in both primary and secondary tumours) was significantly larger than the clones that first became detectable upon tumour passaging (P = 4 × 10−12, Mann–Whitney U-test, Fig. 3e right panels).
We then analysed the clonal composition of the cells produced from oncogene-transduced BCs and LPs after just 2 weeks in subrenal transplants, before tumours become grossly evident. The results showed the sizes as well as the numbers of clones detected at this time to be similar to those detected 6 weeks later in tumours derived from the same input cells (Fig. 4a and Extended Data Fig. 6a, b). The distributions of the relative clone sizes measured in the 2-week transplants both of BCs and of LPs were also similar (Fig. 4b). However, after 2 weeks, the absolute sizes of the clones derived from the KRASG12D-transduced LPs were already significantly larger than the sizes of the clones produced by matching transplants of control vector-transduced cells (median = 206 and 93, respectively, P = 3.3 × 10−8, Mann–Whitney U-test), with a slightly smaller effect apparent in the progeny of BCs from the same two donors (median = 112 and 94, respectively, P = 3.6 × 10−7, Mann–Whitney U-test, Fig. 4c).
a, Comparison of the frequencies of BCs (blue) and LPs (red) that made clones in 2-week grafts (CFCs) versus those that made clones found in tumours 6 weeks later (T-CFCs). Cells transduced with all three oncogenes are shown as triangles, and with KRASG12D only as circles (lines connect donor-matched samples). b, Distribution of relative clone sizes for grafts containing BCs and LPs transduced with three oncogenes or KRASG12D only (left). Pie charts showing the clone size distributions in a representative graft (right). c, Absolute clone size distributions of matched vector control-transduced BCs and LPs, and grafts containing BCs and LPs transduced with KRASG12D (n = 2 donors for each group).
These studies provide new insights into the earliest phases of malignant transformation in vivo of cells isolated directly from normal human mammary tissue. Four findings are particularly noteworthy. The first is the rapidity and efficiency, albeit with high variability, with which this process can be induced in prospectively purified, biologically distinct types of normal human mammary epithelial cells using a single transducing oncogene (KRASG12D). This finding challenges previous assumptions of a requirement for a slow, multi-step selective process to accrue the genetic and/or epigenetic changes needed to obtain a continuously growing tumour. Interestingly, we did not obtain tumours from LCs or SCs subjected to the same protocols, in contrast to a recent report of highly ERα+ tumours generated by transduction of EpCAM+CD49f− LCs with SV40/Ras12.
The second important finding was the considerable heterogeneity displayed in the numbers, phenotypes and growth behaviour of clonally tracked human cells with tumorigenic activity in vivo within 2–8 weeks. This result suggests that a similar range and speed of perturbations may accompany the spontaneous development of some breast cancers in patients.
A third and unexpected finding was the lack of a strong influence of the human mammary cell type initially transduced with the frequency of clones generated, the histopathology of the tumours produced or their loss of lineage-specific expression profiles. Taken together, this suggests a greater effect of the potent transforming role of the KRASG12D oncogene in these cells.
The fourth finding was the frequent delayed activation of clonal growth observed in secondary tumours. This latency could either be biologically determined, reflecting an origin of these late appearing clones from their normal counterparts with similar features5, or simply reflective of a stochastic process, as previously indicated for established human breast cancer cell lines passaged in vivo4.
These results set the stage for examining the molecular basis of the biological heterogeneity now revealed that can occur during the earliest stages of breast cancer formation, the role of additional modifiers and how these may influence the acquisition of treatment response and resistance13,14.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Cells
Reduction mammoplasty tissue was collected with informed consent, as approved by the University of British Columbia Research Ethics Board, and dissociated to obtain organoid-rich pellets which were then viably cryopreserved4. Thawed organoids were rinsed with Hank’s Balanced Salt Solution supplemented with 2% FBS (HF), and the cells then dissociated in 2.5 mg ml−1 trypsin with 1 mM EDTA and 5 mg ml−1 dispase (STEMCELL Technologies) with 100 μg ml−1 DNaseI (Sigma), washing with HF between each step. The resulting cell suspension was filtered through a 40 μm mesh and BCs isolated by FACS according to their CD45−CD31−EpCAMloCD49f+ (or CD45−CD31−CD10+CD90+CD49f+) phenotype, LPs according to their CD45−CD31−EpCAMhiCD49f+ (or CD45−CD31−CD10−CD90−CD49f+) phenotype, LCs according to their CD45−CD31−EpCAMhiCD49f− phenotype and SCs according to their CD45−CD31−EpCAM−CD49f− phenotype. In each case, a small aliquot of cells was immediately re-analysed to measure the purity of each sorted population (routinely >97%). Supplementary Table 3 lists the fluorochrome-labelled antibodies used.
Lentiviral constructs and transduction
Variations of the MNDU3-PGK–GFP lentiviral construct15 encoding YFP or mCherry in place of the GFP reporter were generated and KRASG12D, PIK3CAH1047R and TP53R273C mutant cDNAs then cloned into these, using flanking AscI and PacI restriction sites downstream of the MNDU3 promoter. Human KRAS cDNA was cloned from a human cell line, and altered by site-directed mutagenesis to obtain the G12D mutant. The TP53R273C mutant was cloned directly from a human cell line harbouring this mutation, and the human PIK3CAH1047R cDNA was obtained from A. Weng (Terry Fox Laboratory, BC Cancer Agency, Vancouver, BC, Canada). All cDNAs were sequence-verified before ligation into the lentiviral constructs. Clones confirmed to contain the mutant genes in the correct orientation were then selected for plasmid purification. Lentiviral supernatants containing ~109 infectious units per millilitre were produced16 and added at a final dilution of 1:100 (for each) to cell suspensions containing 1 × 106 to 2 × 106 cells per 100 μl. The library of barcoded lentiviruses (titre of ~109 infectious units per millilitre)6 was added to cells at a 1:200 dilution (~5 × 105 infectious units per 100 μl), to achieve an ~30% transduction efficiency.
Xenografts
Transduced human mammary epithelial cells were suspended in a neutralized rat tail collagen preparation4,17 with 2 × 105 irradiated (15 Gy) C3H-10T1/2 mouse fibroblasts as indicated per 20 μl gel and the gels allowed to solidify at 37 °C for 30 min (refs 4, 17) before being implanted either subrenally or subcutaneously in 5- to 8-week-old virgin female NSG or NRG mice, that were then also implanted under the dorsal skin with a slow-release EP pellet, as indicated. Mice were bred, maintained and followed under specific-pathogen-free conditions in the Animal Resource Centre in the British Columbia Cancer Research Centre in accordance with protocols approved by the University of British Columbia Animal Care Committee. This included the monitoring of tumour growth for the periods indicated, or their removal earlier whenever a tumour reached a size of 1 cm3. The length along the tumour’s longest axis was measured at the time of removal, and was immediately categorized as small (<5 mm), medium (5–10 mm) or large (10–15 mm). No other measurements were recorded. For some of the primary tumours, a small mechanically dissociated fragment or an enzymatically dissociated cell suspension was prepared and transplanted subcutaneously with 50% (v/v) matrigel, with or without an EP pellet, as indicated, into secondary recipients. To measure tumour bioluminescence from luciferase expression, mice were injected intraperitoneally with 150 mg kg−1 body weight D-luciferin (Promega) and 10 min later imaged using a Xenogen IVIS Lumina system with Living Image version 3.0 software (Caliper Life Sciences). To prepare cell suspensions from tumours, the tissue was minced with a scalpel, incubated at 37 °C in DMEM/Ham’s F12 media, supplemented with 5% FBS and 300 U ml−1 collagenase and 100 U ml−1 hyaluronidase for 2–4 h with periodic vortexing, washed with HF, and treated with 2.5 mg ml−1 trypsin with 1 mM EDTA and 5 mg ml−1 dispase with 100 μg ml−1 DNaseI. Human cells were sorted after staining with anti-human-specific antibodies directed against EpCAM and HLA with simultaneous depletion of mouse cells stained with anti-mouse-specific antibodies directed against BP-1, CD140a, CD45 and CD31 (Supplementary Table 3).
Histopathology and immunohistochemistry
Collagen gels or pieces of tumours obtained from mice were fixed in 10% buffered formalin (Fisher), washed in 70% ethanol and embedded in paraffin. A tissue microarray using 1.5 mm dual cores per sample was constructed for all primary tumours analysed. From this tissue microarray, 4 μm sections were obtained. All secondary tumours were embedded in paraffin and sections prepared directly from these blocks. The sections of both primary and secondary tumours were either stained directly with H&E, or were first treated with Target Retrieval solution (DAKO) and then a cytomation serum-free protein block (DAKO) followed by staining with either an anti-K14 antibody, an anti-MUC1 antibody, an anti-K5 antibody, an anti-K8/18 antibody, an anti-ER antibody, an anti-CD44 antibody, an anti-Ki67 antibody, an anti-HER2 antibody, an anti-PR antibody, or an anti-EGFR antibody. Use of a secondary rabbit antibody conjugated to horseradish peroxidase and treatment with 3,3′-diaminobenzidine (DAB, DAKO) was used to obtain a positive brown staining. Supplementary Table 3 provides details of the antibodies used and their sources. A negative control using one of the tumour samples, and a positive control using normal reduction mammoplasty tissue, was included for each marker analysed. Tumour sections stained with H&E were reviewed for histomorphological analysis. Each of the tumours was classified into one of the histological subtypes according to the World Health Organization human breast tumour classification18. Nuclear grade was scored on a three-point scale according to the National Surgical Adjuvant Study of Breast Cancer (NSAS-BC) grading system19, and tubular formation, nuclear atypia, mitotic counts and histological grade were scored on a three-point scale according to the Nottingham grading system20. For each of the markers examined by IHC, intensity of staining (weak, intermediate or strong) and percentage of stained invasive tumour cells (0–100%) were scored. All of the histological and IHC parameters were scored blinded to the sample identity by a trained pathologist (T.O.).
Barcode analysis
Extracted genomic DNA was transferred to a 96-well plate in which researchers were blinded to their identity, although the order of the samples was not randomized. The samples were then treated identically, as previously described6. Defined numbers of control cells (102–106) containing a known barcode sequence at a single copy number per cell were analysed alone, and following their addition to each experimental sample, where they served as an internal normalization standard from which the relationship between fractional read value and cell numbers could be derived to calculate clone sizes (in absolute cell numbers). A threshold corresponding to a fractional read value equivalent of 70 cells was applied.
Real-time PCR
Total RNA was extracted from cryopreserved tumour samples using a mirVana miRNA isolation kit (Life Technologies) and cDNA then synthesized using SuperScript II Reverse Transcriptase (Life Technologies). Real-time PCR was performed using a SYBR Green master mix (Applied Biosystems) and samples were run in triplicate with the custom-designed primers listed in Supplementary Table 4. Human-specific primers were validated before use by testing for lack of reactivity with RNA from a panel of mouse tissues. Test gene cycle threshold (Ct) values were normalized by subtracting the geometric mean of ΔCt values obtained for GAPDH and EIF4A1 (control) genes.
RNA sequencing
RNA was extracted from snap-frozen cells using the mirVana miRNA Isolation Kit (Life Technologies, AM1560) followed by ethanol precipitation. RNA was quantified using an Agilent Bioanalyzer (Life Technologies) and 100 ng of total RNA was ribosomal RNA (rRNA) depleted using a NEBNext rRNA Depletion Kit (New England BioLabs, E6310L). First strand cDNA was generated using a Maxima H minus First Strand cDNA Synthesis Kit (Thermo Scientific, K1652) with added Actinomycin D (1 μg, Sigma, A9415). The product was purified using in-house-prepared 20% PEG in 1 M NaCl Sera-Mag bead solution at a 1.8× ratio and then eluted in 35 μl of Qiagen EB buffer. Second-strand cDNA was synthesized in a 50 μl volume using SuperScript Choice System for cDNA Synthesis (Life Technologies, 18090-019) with 12.5 mM GeneAmp dNTP Blend with dUTP. Double-stranded cDNA was purified with 20% PEG in 1 M NaCl Sera-Mag bead solution at a 1.8× ratio and eluted in 40 μl of Qiagen EB buffer, and fragmented using Covaris E220 (55 s, 20% duty factor, 200 cycles per burst). Sheared cDNA was end repaired/phosphorylated, single A-tailed, and adaptor ligated using custom reagent formulations (New England BioLabs, E6000B-10) and in-house-prepared Illumina forked adaptor. PEG (20%) in 1 M NaCl Sera-Mag bead solution was used to purify the template between each of the enzymatic steps. To complete the process of generating strand directionality, adaptor-ligated template was digested with 5 U of AmpErase Uracil N-Glycosylase (Life Technologies, N8080096). Libraries were then indexed and PCR amplified using Phusion Hot Start II High Fidelity Polymerase (Thermo Scientific, F 549-L). An equal molar pool was sequenced on an Illumina MiSeq platform, which produced between 3 × 106 and 4 × 106 aligned sequence reads.
Adaptor sequences were stripped from the resulting 125-nucleotide sequence reads and the sequences uniformly trimmed to 75 nucleotides. Trimmed reads were aligned using BWA (version 0.5.7)21 to a transcriptome reference22 consisting of genomic sequence (GRCh37-lite July 2010) supplemented by read-length-specific exon–exon junction sequences. SAMtools (version 0.1.13)23 was used to sort the alignment bam files. The sorted bam files were repositioned to GRCh37-lite using JAGuaR (version 2.0.3)22 to assign sequences that aligned across exon–exon junctions to their correct ‘split’ genomic coordinates. An in-house RNA quality control and analysis pipeline24 was used to generate a report (Supplementary Table 5) and calculate a normalization constant for computing RPKM values (reads per kilobase per million mapped reads). The normalization constant was inferred from the total number of exonic reads (excluding mitochondrial reads, reads from ribosomal genes and reads from highest 0.5% expressed exons).
Pairwise comparisons between different sample types within the same donor were performed to identify differentially expressed genes using a custom DEfine matlab tool24 (false discovery rate cutoff = 0.015, differentially expressed in at least two of three samples). PAM50 classification was performed following the methods described in ref. 10 using the R-Bioconductor script available at genome.unc.edu/pubsup/breastGEO/. AIMS classification11 was performed using the Web tool available at www.bci.mcgill.ca/AIMS/.
Accession codes
Data deposits
Final transcriptome data has been deposited in the European Genome-phenome Archive (www.ebi.ac.uk/ega) under accession number EGAS00001001310.
References
Stephens, P. J. et al. Oslo Breast Cancer Consortium (OSBREAC). The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012)
Sørlie, T. et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl Acad. Sci. USA 98, 10869–10874 (2001)
Curtis, C. et al. METABRIC Group. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012)
Eirew, P. et al. A method for quantifying normal human mammary epithelial stem cells with in vivo regenerative ability. Nature Med. 14, 1384–1389 (2008)
Nguyen, L. V. et al. DNA barcoding reveals diverse growth kinetics of human breast tumour subclones in serially passaged xenografts. Nature Commun. 5, 5871 (2014)
Nguyen, L. V. et al. Clonal analysis via barcoding reveals diverse growth and differentiation of transplanted mouse and human mammary stem cells. Cell Stem Cell 14, 253–263 (2014)
Kannan, N. et al. Glutathione-dependent and -independent oxidative stress-control mechanisms distinguish normal human mammary epithelial cell subsets. Proc. Natl Acad. Sci. USA 111, 7789–7794 (2014)
Kannan, N. et al. The luminal progenitor compartment of the normal human mammary gland constitutes a unique site of telomere dysfunction. Stem Cell Rep. 1, 28–37 (2013)
Eirew, P. et al. Dynamics of genomic clones in breast cancer patient xenografts at single-cell resolution. Nature 518, 422–426 (2015)
Parker, J. S. et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J. Clin. Oncol. 27, 1160–1167 (2009)
Paquet, E. R. & Hallett, M. T. Absolute assignment of breast cancer intrinsic molecular subtype. J. Natl Cancer Inst. 107, 357 (2015)
Keller, P. J. et al. Defining the cellular precursors to human breast cancer. Proc. Natl Acad. Sci. USA 109, 2772–2777 (2012)
Kreso, A. et al. Variable clonal repopulation dynamics influence chemotherapy response in colorectal cancer. Science 339, 543–548 (2013)
Bhang, H. E. et al. Studying clonal dynamics in response to cancer therapy using high-complexity barcoding. Nature Med. 21, 440–448 (2015)
Logan, A. C. et al. Factors influencing the titer and infectivity of lentiviral vectors. Hum. Gene Ther. 15, 976–988 (2004)
Imren, S. et al. High-level β-globin expression and preferred intragenic integration after lentiviral transduction of human cord blood stem cells. J. Clin. Invest. 114, 953–962 (2004)
Eirew, P., Stingl, J. & Eaves, C. J. Quantitation of human mammary epithelial stem cells with in vivo regenerative properties using a subrenal capsule xenotransplantation assay. Nature Protocols 5, 1945–1956 (2010)
Lakhani, S. R., Ellis, I. O., Schnitt, S. J., Tan, P. H. & van de Vijver, M. J. WHO Classification of Tumours of the Breast 4th edn, Ch. 2 (World Health Organization, 2012)
Tsuda, H., Akiyama, F., Kurosumi, M., Sakamoto, G. & Watanabe, T. ; Japan National Surgical Adjuvant Study of Breast Cancer (NSAS-BC) Pathology Section. Establishment of histological criteria for high-risk node-negative breast carcinoma for a multi-institutional randomized clinical trial of adjuvant therapy. Jpn. J. Clin. Oncol. 28, 486–491 (1998)
Elston, C. W. & Ellis, I. O. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 19, 403–410 (1991)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Butterfield, Y. S. et al. JAGuaR: junction alignments to genome for RNA-seq reads. PLoS One 9, e102398 (2014)
Li, H. et al. 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Gascard, P. et al. Epigenetic and transcriptional determinants of the human breast. Nature Commun. 6, 6351 (2015)
Acknowledgements
We thank D. Wilkinson, G. Edin and M. Hale for technical support, E. Bovill, J. Boyle, S. Bristol, P. Gdalevitch, A. Seal, J. Sproul and N. van Laeken for access to discarded reduction mammoplasty tissue, T. Nielsen and N. Poulin for discussions, the Centre for Translational and Applied Genomics (BC Cancer Agency) for assistance with IHC, and T. MacDonald for assistance with rodent husbandry. This work was supported by grants from the Canadian Cancer Society Research Institute, the Canadian Breast Cancer Foundation and the Canadian Breast Cancer Research Alliance. L.V.N. received a Vanier Canada Graduate Scholarship from the Canadian Institutes of Health Research (CIHR), and N.K. was supported by a MITACS Elevate Fellowship. T.O. was supported by a Molecular Oncologic Pathology Fellowship from CIHR and the Terry Fox Foundation, and by grants from the Sumitomo Life Welfare and Culture Foundation, the Mochida Memorial Foundation for Medical and Pharmaceutical Research, and the Takashi Tsuruo Memorial Fund. S.A. is supported by a Canada Research Chair.
Author information
Authors and Affiliations
Contributions
L.V.N., D.P. and C.J.E. designed the project, drafted the manuscript and were assisted by S.L., C.L.C., W.K. and S. Balani in performing the experiments. M.M. and M.H. oversaw the generation of sequence data, and L.V.N., D.P., A.C. and M.B. analysed it. All authors contributed to the interpretation of the results, and read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Quantification of human cells containing different vector reporters in tumours derived from triply transduced starting populations.
a, Lentiviral constructs used. CBR-Luc, click beetle red luciferase. b, Frequencies of donor samples producing at least one tumour subrenally from BCs or LPs exposed to different combinations of oncogene-encoding vectors. c, Representative FACS profiles of a cell suspension prepared from a tumour produced from cells transduced with all three genes and sorted for human EpCAM and HLA using human-specific antibodies. d, Percentage of cells expressing different lentiviral reporters in cells maintained in vitro for 72 h after transduction, and in primary and secondary tumours (G, GFP; Y, YFP; mCh, mCherry). e, Frequencies of donor samples producing at least one tumour under various transplantation conditions using BCs or LPs transduced with KRASG12D. f, Percentages of human cells in BC- and LP-derived tumours detected by FACS on the basis of their expression of human EpCAM and/or HLA.
Extended Data Figure 2 Molecular characterization of the tumours.
a, Examples of PCR evidence of all three vectors in DNA extracts obtained from a subset of tumours analysed with vector-specific primers. b, A representative Sanger sequencing chromatograph showing the expected point mutations in the tumour cells analysed. c, PCR evidence of the three vectors in FACS-purified doubly and triply transduced cells. d, e, Representative images of H&E- and IHC-stained sections of primary tumours (d, arising from cells transplanted subcutaneously) and secondary tumours (e, all arising subcutaneously) derived from either BCs or LPs. Scale bar, 50 μm. f, Relative expression (negative ΔCt values, mean ± s.e.m.) of gene transcripts typically associated with mesenchymal/basal or epithelial/luminal phenotypes, or associated with proliferation and cell growth.
Extended Data Figure 3 Threshold set for detection of barcoded clones for the two sequencing runs from which barcode data were acquired.
a, The relationship between the fractional read value (FRV) and the number of cells per clone. Spiked-in controls only and spiked-in controls added to experimental samples are shown as red and grey points, respectively. The shaded grey box represents distribution of false positive barcodes. b, Sensitivity and specificity data for controls compared with experimental samples for different sized clones.
Extended Data Figure 4 Clonal analyses of primary barcoded tumours.
a, Numbers of clones and frequencies of T-CFCs in primary tumours. b, Relative clone size distributions for individual primary tumours grouped by the cell type initially manipulated and the oncogene(s) used. Each column represents a single tumour. Each rectangle represents one clone. Its relative clone size is indicated by the shade of green, and its proportional contribution within each tumour is indicated by its length on the y axis.
Extended Data Figure 5 Clonal analyses of secondary barcoded tumours.
a, Numbers of clones and frequencies of T-CFCs in secondary tumours. b, Relative clone size distributions for individual secondary tumours grouped by the cell type initially manipulated. Each column represents a single tumour. Each rectangle represents one clone. Its relative clone size is indicated by the shade of green, and its proportional contribution within each tumour is indicated by its length on the y axis. c, Clonal landscape of replicate secondary tumours generated from single primary tumours in two separate experiments. Clones present in sibling tumours are shown above one another and unique clones are shown in the same horizontal bar. Increasing clone sizes are indicated by a grey intensity scale. d, Numbers of clones and T-CFC frequencies of combined primary and secondary tumours.
Extended Data Figure 6 Clonal analyses of transduced cells transplanted subrenally after 2 weeks in vivo.
a, Number of clones and frequency of CFCs in xenografts of transduced cells assessed after 2 weeks in vivo. b, Relative clone size distributions of individual 2-week transplants grouped by the cell type initially manipulated and the oncogene(s) used. Each column represents a single transplant. Each rectangle represents one clone. Its relative clone size is indicated by the shade of green, and its proportional contribution within each tumour is indicated by its length on the y axis.
Supplementary information
Supplementary Table 1
This file contains Luciferase activity values. (XLSX 11 kb)
Supplementary Table 2
This file contains lists of differentially expressed genes. (XLSX 113 kb)
Supplementary Table 3
This file shows Antibodies used for FACS and IHC analyses. (XLSX 9 kb)
Supplementary Table 4
This file contains Primers used for RT-qPCR. (XLSX 8 kb)
Supplementary Table 5
This file contains quality control report of RNA-seq data. (XLSX 15 kb)
Rights and permissions
About this article
Cite this article
Nguyen, L., Pellacani, D., Lefort, S. et al. Barcoding reveals complex clonal dynamics of de novo transformed human mammary cells. Nature 528, 267–271 (2015). https://doi.org/10.1038/nature15742
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15742
- Springer Nature Limited
This article is cited by
-
Experimental and spontaneous metastasis assays can result in divergence in clonal architecture
Communications Biology (2023)
-
Know thy cells: commonly used triple-negative human breast cancer cell lines carry mutations in RAS and effectors
Breast Cancer Research (2022)
-
De novo and cell line models of human mammary cell transformation reveal an essential role for Yb-1 in multiple stages of human breast cancer
Cell Death & Differentiation (2022)
-
Mastering the use of cellular barcoding to explore cancer heterogeneity
Nature Reviews Cancer (2022)
-
Multi-color clonal tracking reveals intra-stage proliferative heterogeneity during mammary tumor progression
Oncogene (2021)