Abstract
The anaerobic oxidation of methane (AOM) with sulfate controls the emission of the greenhouse gas methane from the ocean floor1,2. In marine sediments, AOM is performed by dual-species consortia of anaerobic methanotrophic archaea (ANME) and sulfate-reducing bacteria (SRB) inhabiting the methane–sulfate transition zone3,4,5. The biochemical pathways and biological adaptations enabling this globally relevant process are not fully understood. Here we study the syntrophic interaction in thermophilic AOM (TAOM) between ANME-1 archaea and their consortium partner SRB HotSeep-1 (ref. 6) at 60 °C to test the hypothesis of a direct interspecies exchange of electrons7,8. The activity of TAOM consortia was compared to the first ANME-free culture of an AOM partner bacterium that grows using hydrogen as the sole electron donor. The thermophilic ANME-1 do not produce sufficient hydrogen to sustain the observed growth of the HotSeep-1 partner. Enhancing the growth of the HotSeep-1 partner by hydrogen addition represses methane oxidation and the metabolic activity of ANME-1. Further supporting the hypothesis of direct electron transfer between the partners, we observe that under TAOM conditions, both ANME and the HotSeep-1 bacteria overexpress genes for extracellular cytochrome production and form cell-to-cell connections that resemble the nanowire structures responsible for interspecies electron transfer between syntrophic consortia of Geobacter9,10. HotSeep-1 highly expresses genes for pili production only during consortial growth using methane, and the nanowire-like structures are absent in HotSeep-1 cells isolated with hydrogen. These observations suggest that direct electron transfer is a principal mechanism in TAOM, which may also explain the enigmatic functioning and specificity of other methanotrophic ANME–SRB consortia.
Similar content being viewed by others
Main
The anaerobic oxidation of methane with sulfate (AOM) controls the emission of methane from the seabed1,3,4. At environmental conditions the net reaction CH4(aq) + SO42− → HS− + HCO3− + H2O allows an energy yield of only −20 to −40 kJ per mol of methane oxidized, shared between the two partner organisms. Generally, AOM consortia show exceptionally slow growth with generation times >2 months, which has so far impeded their cultivation6,11. Sulfate-coupled AOM in marine habitats is performed by members of three different ANME clades (ANME-1, -2 and -3), which associate physically with specific partner bacteria of the Desulfosarcina/Desulfococcus or the Desulfobulbus cluster11,12,13, indicating an obligate functional role of the SRB in AOM. Early studies had already suggested a syntrophic coupling of both partners via a transfer of reducing equivalents4,14, yet the underlying mechanisms remain unknown. Biochemically, the anaerobic oxidation of methane appears in the ANME and involves a reversal of the enzymatic machinery of the methanogenesis pathway15,16,17,18. However, reversing an energy-yielding process is per se endergonic, and hence AOM requires an efficient transfer of reducing equivalents from methane to sulfate, so that the ANME can gain energy by AOM14,19. Previous results indicate that the partner SRB4,20 act as electron sinks of AOM, but recently members of the ANME-2 clade were also suggested to perform incomplete sulfate reduction by an as yet unknown pathway21.
In this study we focus on the hypothesis of syntrophic growth in thermophilic AOM consortia by direct interspecies electron transfer, and test this and alternative hypotheses (mechanisms illustrated in Extended Data Fig. 1). The studied sediment-free TAOM enrichment was cultivated at 60 °C and supplied with 28 mM sulfate and 0.2 MPa methane, allowing an energy yield (ΔGR) of −34 kJ mol−1, and resulting in doubling times of approximately 68 days (Fig. 1a) and growth efficiencies of 2% (see Methods). The culture is dominated by consortia of ANME-1 and HotSeep-1 appearing in an approximate 1:1 stoichiometry. Owing to their larger size ANME account for around 75% of the consortial biomass (Fig. 1b and Extended Data Table 1). Using a dilution-to-extinction approach (1:10 to 1:109) with hydrogen (0.2 MPa) and sulfate (28 mM), we were able to separate a strain of HotSeep-1 that was identical to the partner bacterium of the TAOM consortium (>99% identity in 16S ribosomal RNA gene and internal transcribed spacer region, and (genomic) average nucleotide identity >99%, Extended Data Table 2). This strain grows without ANME-1 as single cells or in mono-species aggregates (Extended Data Fig. 2a) and with a single contaminant, Archaeoglobus sp. (1–5% of all cells), which does not form consortia with HotSeep-1. Substrate tests with the HotSeep-1 culture showed that it is an obligate chemolithoautotroph, with hydrogen and sulfate as the sole molecular redox couple and doubling times of 4 to 6 days (see Methods and Extended Data Fig. 2b, c). Although the supplied hydrogen (0.2 MPa) could provide a tenfold higher energy yield to HotSeep-1 than syntrophy in TAOM (ΔGR = −151 kJ mol−1 versus −17 kJ mol−1, the latter being half of the net energy yield of TAOM consortia), its carbon assimilation efficiency remained similarly low (approximately 1.5% of converted reducing equivalents).
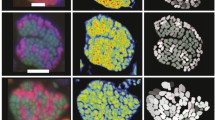
a, The exponential increase of sulfide production translates to a doubling time of 68 days (biological replicates n = 4). b, Representative fluorescence micrograph of TAOM consortia from a; red, ANME-1; green, HotSeep-1. Scale bar, 10 µm. Representative of 14 similar images recorded. c, Sulfide production under TAOM conditions (red circles, 0.07 mM sulfide per day) versus a control (white squares, 0.02 mM per day). Hydrogen (blue triangles), or hydrogen plus methane (purple stars) increased sulfide production (both 0.55 mM per day; biological replicates n = 3, symbols represent mean values, error bars are s.d.). d, Per cent of total RNA reads mapped to ANME-1 (red) and HotSeep-1 (green) after 3 days of incubation (c), biological replicates n = 3, data are mean values, for statistical analyses see Supplementary Table 1.
We compared the activity of the TAOM consortia and the hydrogenotrophic HotSeep-1 by physiological experiments combined with metagenomic and metatranscriptomic analyses and electron microscopy of the involved organisms. A classical experiment for the study of syntrophy in dual-species consortia is the addition of potential intermediates that could be theoretically produced by the ANME as a by-product of methane oxidation, and consumed as the electron donor by the partner SRB (mechanisms illustrated in Extended Data Fig. 1). If these compounds were relevant in interspecies transfer of reducing equivalents, their addition to the medium should favour the electron-accepting partner, and should repress the electron transfer between the consortial partners14,21,22,23. By contrast, a model of direct electron transfer via nanowires as proposed in refs 7,8,10, would be insensitive to such external additions of potential intermediates if the additions do not represent an alternative, preferred substrate for one of the partner organisms.
With the exception of hydrogen, none of the potential intermediates added as sole electron donor caused significant microbial sulfide production in the TAOM enrichment (Extended Data Table 3). Carbon monoxide and methyl sulfide even inhibited sulfide production when added together with methane. Carbon monoxide is known to inhibit cytochrome c activity, which may play an important role in intra- and intercellular transfer of reducing equivalents in AOM17. Methylated substrates may interfere with the reverse, oxidative operation of the methanogenesis pathway24. The addition of colloidal zero-valent sulfur to the TAOM culture (supplied in concentrations from 1 to 25 mM, Extended Data Fig. 3a) did not result in the production of sulfide and sulfate as reported in a previous study with ANME2a/DSS consortia21. However, with hydrogen as an electron donor (0.16 MPa), sulfide production rates increased three- to eightfold compared to replicate incubations with methane as the sole electron donor at TAOM conditions (Fig. 1c and Extended Data Table 3). We investigated further the influence of hydrogen on the oxidation of methane using headspace-free incubations (Extended Data Fig. 4). In incubations with methane and hydrogen, hydrogen was first selectively consumed and methane oxidation was repressed. When hydrogen was consumed, methane oxidation rates recovered to the same level as in replicate incubations with only methane, suggesting an inhibition of methane oxidation in the presence of hydrogen. To investigate the influence of hydrogen on the consortial partners, we mapped total RNA reads to the genome drafts of ANME-1 and HotSeep-1 after exposure to different substrate conditions (Fig. 1d, for read numbers see Supplementary Table 1). Under TAOM conditions, relative RNA expression patterns reflected the biomass ratio between the ANME and their partner bacteria (3:1) (Fig. 1d). The addition of hydrogen caused a strong relative increase of HotSeep-1 over ANME gene expression, even in the presence of methane. This indicates that if the partner SRB does not act as an electron sink for reverse methanogenesis, ANME activity is repressed; an effect of syntrophic cooperation that was predicted previously14.
To test the hypothesis of hydrogen production by ANME as a direct intermediate in TAOM (Extended Data Fig. 1) that is consumed by HotSeep-1, we assessed the presence and production of hydrogen under TAOM conditions. Maximal hydrogen concentrations were only about 2 Pa in the TAOM enrichments, and re-established within 7 h after gas phase exchange (Fig. 2a). Thermodynamically, HotSeep-1 could thrive on these low hydrogen concentrations with an energy yield of approximately −24 kJ mol−1. However, the production of hydrogen in TAOM cultures corresponded to only ∼0.5% of the theoretical hydrogen production rates as reflected by sulfide production (according to the stoichiometry of reverse methanogenesis; Fig. 2b). This is insufficient to explain the consortial growth of HotSeep-1. Furthermore, we could not detect catalytic subunits of [FeFe] or [NiFe] hydrogenases in the ANME-1 draft genome. In conclusion, hydrogen appears to be an alternative growth substrate for HotSeep-1 when available externally, but is not provided by ANME-1 as an intermediate in TAOM.
a, Hydrogen gas pressure under TAOM conditions (methane, 0.2 MPa; sulfate, 28 mM; 5-day incubation (filled circle)). The dashed arrow depicts gas phase exchange with methane. Open circles show equilibration of hydrogen in the headspace (n = 1). b, Hydrogen production in 10 ml TAOM culture supplied with 0.2 MPa methane after headspace exchange and addition of 10 mM molybdate (final concentration) to inhibit hydrogen consumption. Open circles are replicate measurements with hydrogen production of 2 and 3 nmol l−1 min−1. Dotted line is predicted hydrogen production for reverse methanogenesis (CH4 + 2H2O → CO2 + 4H2) = 420 nmol H2 l−1 min−1 culture, for an observed sulfide production rate of 104 nmol l−1 min−1. Both experiments were repeated once with the same results.
An alternative explanation of the TAOM interaction could be direct interspecies electron transfer (DIET) between ANME-1 and HotSeep-1, also hypothesized as a principle mechanism for syntrophic growth of AOM consortia8,10,17,25. A switch from interspecies hydrogen transfer to DIET has been previously shown for the dual-species interaction between Geobacter sulfurreducens and Geobacter metallireducens, benefiting both consortial partners, as evidenced by their increased growth rates via DIET10,26. In the tightly packed Geobacter consortia, a dense network of cell-to-cell connections was detected by transmission electron microscopy and immunogold labelling, probably serving in electron transfer10. The functioning of electron transfer via conductive cell-to-cell connections (nanowires) is not fully understood, but apparently involves the expression of the pilin protein PilA of the type IV pili together with certain members of the cytochrome c family9,10,27,28. Recent findings of such cytochromes in the ANME genome, along with redox-dependent staining of the intercellular matrix of the ANME-2/SRB consortia8, suggest that DIET could also be relevant in AOM.
To find evidence for DIET in TAOM we analysed the genome and specific gene expression of ANME-1 and HotSeep-1, with the focus on similarities to the Geobacter consortia using DIET as the main electron transfer mechanism. The ANME-1 draft genome contains several potentially extracellular multi-haem cytochrome c proteins, some of which are highly expressed during TAOM, but no genes for pili formation (Extended Data Table 4, Supplementary Table 2). However, HotSeep-1 comprises the genes for the biosynthesis and assembly of type IV pili, as well as large multi-haem cytochrome c proteins, both with high amino acid similarity to respective proteins in Geobacter spp.29 (Extended Data Table 4 and Supplementary Tables 3 and 4). We further investigated expression patterns of these potentially DIET-related genes in comparison to genes for AOM (mcrA) and sulfate reduction (dsrA), representing key catabolic processes in ANME-1 and HotSeep-1 (Fig. 3a, for statistical analyses see Supplementary Table 5). In agreement with the results from total RNA expression (Fig. 1d), a switch from methane to hydrogen (or methane plus hydrogen) as an energy source caused an immediate drop in mcrA and cytochrome expression in ANME, as well as a reduction of the expression of the HotSeep-1 pili and cytochromes. Comparing relative gene expression of HotSeep-1 in consortial growth using methane, versus single growth using hydrogen, both pilA and cytochrome c are clearly overexpressed under TAOM conditions, this is also the case when compared to dsrA expression (Extended Data Fig. 5, for statistical analyses see Supplementary Table 5).
a, Expression of archaeal (red) and bacterial (green) genes in incubations with hydrogen (darker shade), or hydrogen plus methane (lighter shade), relative to methane alone (expression under TAOM condition = 1; biological replicates n = 3; *P < 0.05; for statistics see Supplementary Table 5b). b–f, Micrographs of thin sections. Scale bars, 300 nm. b–d, TAOM consortia with HotSeep-1 cells (H; rod-shaped; approximately 1 × 0.5 µm) and ANME-1 cells (A; cylindrical shape with envelope30; 1.5 × 0.8 µm). Nanowires of 10 nm diameter and up to several thousand nanometres in length connect both species, representative of 70 recorded images. c, d, Arrows mark the apparent origin of the wires from the membrane of HotSeep-1 bacteria to the polar sites of ANME-1 cells. d, HotSeep-1 cell with nanowires crossing the cell membrane (marked by the arrow). e, f, Aggregated HotSeep-1 cells grown with hydrogen do not develop nanowires; representative of 48 images.
This observation is supported independently by transmission electron microscopy on thin sections of TAOM consortia. Using two different embedding techniques we found a dense network of pili-like structures connecting HotSeep-1 to ANME-1 cells (Fig. 3b, c and Extended Data Fig. 6a), resembling the nanowire structures found in Geobacter consortia (visualized with the same methods, see Extended Data Fig. 6b). In TAOM consortia the nanowires are larger, and appear more dense, with diameters of approximately 10 nm and apparent lengths of 100 to >1,000 nm. In agreement with the genomic patterns, these wires seem to be formed by the partner bacteria, connecting to the ANME-1 at their polar sides (Fig. 3b–d). In contrast, HotSeep-1 cells in mono-species aggregates isolated with hydrogen show smooth surfaces without such extracellular extensions (Fig. 3e, f), indicating that the observed intercellular structures are specific to consortial growth under TAOM conditions and not only related to cellular attachment.
In conclusion, our data show that consortial growth of thermophilic ANME-1 archaea and HotSeep-1 bacteria is probably based on similar principles as those in Geobacter consortia, where DIET is mediated by intercellular wiring made up of pili-like structures and outer-membrane multi-haem cytochromes. The underlying biophysics and biochemistry of intercellular wiring for direct electron transfer needs further investigation. If this mode of syntrophic cooperation between the electron-generating archaea and nanowire-producing bacteria is also the underlying mechanism for other types of AOM consortia as suggested recently8, it may explain the enigmatic specificity of dual-species partnerships in AOM in general.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Cultivation of TAOM consortia
Sediment-free, TAOM enrichment cultures were obtained after 1.5 years by semi-continuous incubation of hydrothermal vent sediments from Guaymas Basin with sulfate reducer medium31 and 0.225 MPa CH4 (+0.025 MPa CO2) as the sole energy source at 60 °C, as described in ref. 6. Culture medium was replaced and samples were diluted 1:2 when sulfide concentrations exceeded 12 mM. For the different experiments, subsamples of the main culture (biological replicates) were incubated in parallel.
DNA extraction, sequencing and phylogenetic classification of TAOM partners
Genomic DNA was extracted as described previously32 from an active TAOM culture. The protocol encompassed three cycles of freezing and thawing, chemical lysis in a high-salt extraction buffer (1.5 M NaCl) by heating of the suspension in the presence of sodium dodecyl sulfate and hexadecyltrimethylammonium bromide, and treatment with proteinase K. To amplify bacterial 16S ribosomal DNA genes the primer pair GM3/GM4 (ref. 33) was used. For archaeal 16S rDNA genes the primers 20F (ref. 34) and Arc1492R (ref. 35) were selected. PCR reactions were performed according to ref. 6. The phylogenetic affiliation was inferred with the ARB software package36 and release 115 of the ARB SILVA database37. Representative 16S rRNA gene sequences are deposited at NCBI with the accession numbers KT152859–KT152885.
Visualization of TAOM aggregates by fluorescence in situ hybridization
Cell aliquots were fixed in 2% formaldehyde for 2 h at room temperature, washed with 1 × PBS (pH 7.4). Fixed cell suspensions were treated with mild sonication (Sonoplus HD70; Bandelin) and aliquots of 50–250 µl were filtered onto GTTP filter (0.2 µm pore size, 20 mm diameter). CARD-FISH was performed as described previously38 with the following modifications: for cell wall permeabilization, filters were sequentially incubated in lysozyme solution (10 mg ml−1 lysozyme powder, 0.1 M Tris–HCl, 0.05 M EDTA, pH 8) for 15–30 min at 37 °C and proteinase K solution (0.45 mU ml−1, 0.1 M Tris–HCl, 0.05 M EDTA, pH 8, 0.5 M NaCl) for 2 min at room temperature. Endogenous peroxidases were inactivated by incubating the filters in 0.15% H2O2 in methanol (30 min, room temperature). The oligonucleotide probes ANME-1-350 and HotSeep-1-590 were applied with formamide concentrations according to ref. 6. For dual CARD-FISH, peroxidases of the first hybridization were inactivated by 0.3% H2O2 in methanol (30 min, room temperature). Catalysed reporter deposition was combined with the fluorochromes Alexa Fluor 488 and Alexa Fluor 594. Filters were stained with DAPI (4,6-diamidino-2-phenylindole). Micrographs were obtained by confocal laser scanning microscopy (LSM 780; Zeiss).
Test of potential AOM intermediates/alternative HotSeep-1 substrates
All experiments were performed with artificial seawater medium containing 30 mM of carbonate buffer at TAOM cultivation temperature (60 °C), except when specified otherwise. To ensure equilibration of gas phases, samples were agitated on shaking tables. Highly pure gases and chemicals were used as additions to the incubations. Standard TAOM conditions are defined here as 0.2 MPa methane and 28 mM sulfate. To test the TAOM enrichment for substrate-specific sulfide production, triplicate culture aliquots (10 ml in 20 ml Hungate tubes) were supplemented with different substrates (Extended Data Table 2) at concentrations of 20 mM, except methyl sulfide and carbon monoxide (both 0.05 MPa), and hydrogen (0.16 MPa) with and without methane (0.2 MPa). Zero-valent sulfur was prepared according to ref. 39 and was supplied as dissolved species. For this compound we additionally tested sulfide development via disproportionation in a concentration gradient from 1–12 mM final S0 concentration (Extended Data Fig. 6a). As positive reference, methane was provided at 0.2 MPa (at 60 °C roughly equivalent to 1.6 mM in solution). Sulfide production in the experiments was repeatedly measured every 3 to 4 days using the copper sulfide assay40 and absorption spectrometry at 480 nm. TAOM rates with methane as the sole energy source (0.2 MPa) reached approximately 0.100 ± 0.030 µM per day, compared to a negative control (nitrogen; <0.001 µM per day). Rates determined for other substrates were compared to those under TAOM conditions.
Influence of hydrogen addition on methane oxidation in TAOM cultures
To determine the effect of hydrogen addition on methane oxidation rates, TAOM culture aliquots were supplemented with methane and hydrogen (0.15 MPa and 0.05 MPa, respectively), or only methane as control (0.15 MPa). Cultures were incubated headspace-free at 50 °C for this experiment, because hydrogen was too rapidly consumed at 60 °C for time-course experiments. To determine concentrations of methane and hydrogen, 1 ml of medium was sampled with gas-tight glass syringes, and the sampled medium was concurrently replaced with substrate-free medium to avoid the formation of a headspace. The sampled medium was injected through the septum of 10 ml Exetainer filled with 1 ml NaOH and concentrations of CH4 and H2 were measured as described below.
Presence and production of hydrogen in active TAOM cultures
To determine hydrogen concentrations at TAOM conditions, 20 ml of culture was transferred into 156-ml bottles at 60 °C and gas phases were repeatedly sampled using glass syringes (1 ml) combined with direct measurements on the gas chromatograph. Cultures incubated for 3 or more days reached stable hydrogen concentrations. A comparison to molybdate addition is provided in Extended Data Fig. 6b, c. To quantify molecular hydrogen production in TAOM, 20 ml of culture was supplied with sodium molybdate (10 mM final concentration). This molybdate concentration assured complete inhibition of hydrogen-dependent sulfate reduction as shown in replicate incubations of TAOM culture (1 to 25 mM molybdate) with hydrogen (0.1 MPa) as the sole electron donor (Extended Data Fig. 6d). Samples were stored at 60 °C on a shaking table and repeatedly sampled by glass syringes. Concentrations of methane and hydrogen were measured via gas chromatography coupled to flame ionization detection (Focus GC, Thermo) and via reducing compound photometry (RCD; Peak Performer 1 RCP; Peak Laboratories).
Determination of carbon fixation by TAOM consortia
Replicate culture aliquots (n = 5) were incubated in 5-ml Hungate tubes supplemented with methane, sulfate and 14C-labelled inorganic carbon (380 kBq). AOM-independent carbon fixation was determined under N2 atmosphere (n = 5). To determine methane oxidation rates, replicate vials were incubated with 14C-methane (14 kBq). Incubations were performed at 60 °C for 24 h. Samples were blotted onto 0.2-µm mixed cellulose esters membrane filters (Millipore, Merck). Filters were dried and potential residual inorganic carbon was removed by exposing the filters to an HCl atmosphere for 24 h. Radioactivity in liquid aliquots (0.1 ml) and filters was determined by liquid scintillation counting (scintillation mixture; Filtercount; Perkin Elmer; scintillation counter 2900TR LSA; Packard).
Cultivation of HotSeep-1 on molecular hydrogen
To isolate the hydrogenotrophic sulfate reducers in the TAOM enrichment, aliquots were transferred to Hungate tubes (20 ml) and diluted 1:10 to 1:109 with marine sulfate reducer medium. All vials were amended with 0.2 MPa H2:CO2 (80:20) gas phase, and additionally stored in N2 atmosphere to prevent oxygen flux into the culture vials. Vials were stored at the TAOM temperature optimum (60 °C) and measured for sulfide production using the copper sulfate assay40. To identify cultivated microorganisms, the 16S rRNA gene of active hydrogenotrophic cultures was directly amplified from freeze-thawed pellets of culture aliquots (primer pair GM3/GM4) and sequenced as described above. The phylogenetic affiliation was inferred with the ARB software package36 and Release 115 of the ARB SILVA database37. Representative sequences are deposited at NCBI with the accession numbers KT152886 and KT152887.
Physiology experiments with HotSeep-1
Electron acceptor tests. Culture aliquots (1 ml tenfold-diluted in artificial anoxic seawater medium) were supplied with different potential electron acceptors (colloidal sulfur, sulfite or thiosulfate) with and without the addition of hydrogen. Potential growth on alternative carbon sources (that is, acetate, butyrate, peptone and methyl sulfide) was tested.
Growth rates. Growth rates were independently determined from the development of sulfide concentrations and cell counts (from DAPI-stained cells for total cell numbers and from fluorescence in situ hybridized cells for specific cell numbers) from replicate cultures (grown from 10% inoculum).
Growth efficiencies. Efficiencies were determined in a 14C-DIC radiotracer assay. Replicate cultures were spiked with 14C-DIC (∼ 5.4 MBq) and incubated with H2:CO2 or, as control, with N2:CO2 headspace. Sulfate-dependent hydrogen consumption was determined by the increase of sulfide (colourimetrically40) and by the decrease of sulfate (via ion chromatography) in the medium. Fixed carbon was measured from culture aliquots (5 ml volume) blotted on filters as described above. Concentrations of radioactivity on the filter and the medium were determined via scintillation counting. The total carbon fixation (mmol per ml culture) was calculated as 14C uptake into particulate organic carbon multiplied by total DIC [14C-POC (Bq per ml of culture)/14C-total (Bq per ml of culture) × DIC (mmol per ml culture)], and normalized to reducing equivalent transfer, values are compared with the consumption of sulfide.
Metagenome sequencing and draft genome assembly of ANME-1 and HotSeep-1
Genomic DNA was extracted from TAOM and HotSeep-1 enrichment cultures (as described above) and prepared for Illumina sequencing using the Nextera mate pair sample preparation kit (Illumina), following the Gel-Plus protocol of the manufacturer’s user guide. DNA fragments with a length of approximately 5–8 kb were extracted from a preparative gel before circularization. Additionally a paired-end read library with insert size of 500 bp was constructed for the TAOM enrichment using the TruSeq library preparation kit. Libraries were sequenced on a MiSeq instrument (MiSeq, Illumina) in a 2 × 250 bases paired-end run. Quality-controlled mate pair reads were assembled using the SPAdes genome assembler v.3.5.0 (ref. 41) with default values of k and the -hqmp option. Assembled contigs from the TAOM metagenome were binned based on tetranucleotide frequency using the Metawatt software42. ANME-1- and HotSeep-1-specific bins were extracted for targeted reassembly using the SPAdes genome assembler v.3.5.041 with mapped mate pair and paired end read data and default values of k and subsequently were used as draft genomes. A HotSeep-1 draft genome was also obtained from the assembled contigs of the highly enriched HotSeep-1 culture metagenome (hydrogenotrophic HotSeep-1).
Draft genome analysis
Draft genomes were annotated with Prokka43, and the draft genome of HotSeep-1 (obtained from the hydrogenotrophic HotSeep-1 enrichment) was additionally annotated with an in-house pipeline and analysed using GenDB44 and JCoast45. The annotation of reported genes was manually curated. An expectation (E)-value cut-off of 1 × 10−5 was considered for all predictions of putative protein functions. Identity of the enriched hydrogenotrophic HotSeep-1 and the TAOM partner HotSeep-1 was evaluated by pairwise blast search of the nucleotide sequence of the 16S and 23S rRNA genes, functional and housekeeping genes and the intergenic spacer region (Extended Data Table 2) derived from the draft genome of the TAOM partner HotSeep-1 (query) versus the hydrogenotrophic HotSeep-1 (subject). To verify that the organisms belong to the same species the average nucleotide identity (ANI) and the tetranucleotide frequency correlation of the two draft genomes were determined using JSpecies46 (v.1.2.1). Analyses resulted in tetranucleotide frequency correlation of 0.999 and ANI of >99%. To check for absence of ANME-1 in the hydrogenotrophic HotSeep-1 culture metagenomic reads were mapped to the SILVA SSU 119 reference database (bbmap v.35 and pyhloflash v.1.5) for phylogenetic classification at minimum identities of 90%, 95% and 97% resulting in approximately 3,500, 2,100 and 1,500 classified 16S rRNA gene fragments, respectively, which were screened for hits to ANME related sequences.
To identify potential cytochrome c and type IV pili (T4P) genes in the draft genomes of ANME-1 and HotSeep-1, protein domains were predicted using hmmscan (HMMER 3.047) with the PfamA48 and TIGRFAM49 reference databases. Potential cytochromes were identified by the CXXCH motive and cytochrome c-specific protein domain models. Potential T4P genes were identified using protein models related to T4P. ANME-1 cytochrome and HotSeep-1 cytochrome and pili genes were compared for their best matching hits in the G. sulfurreducens (strain PCA) and G. metallireducens genome and the NCBI non-redundant protein database using blastp. Cytochrome annotation based on detected protein domains in PfamA, pili annotation based on detected protein domains and amino acid sequence. Subcellular localization was predicted with PSORTb50 (v.3.0.2). For cytochromes the number of potential haem-binding sites was derived from the abundance of the CXXCH motif. For sequence comparison to the NCBI non-redundant protein database and Geobacter spp. and for details on protein domains and subcellular localization prediction see Supplementary Table 2a, b, (ANME-1 cytochromes), Supplementary Table 3a, b, (HotSeep-1 cytochromes) and Supplementary Table 4a, b (HotSeep-1 Type IV pili biogenesis). Representative sequences are deposited in GenBank under the accession numbers KT759143–KT759147, KT795302–KT795321, KT795322 and KT795323.
The ANME-1 draft genome was searched for genes encoding catalytic subunits of hydrogenases using blastp search against known genes of catalytic subunits of [NiFe] and [FeFe] hydrogenases (mvhA, echA, frhA, vhuA, vhtA, ehaO, hymC). Annotation of genes with hits was evaluated by blastp search against the NCBI non-redundant protein database for best matching reference sequences related to hydrogenases, but none were found.
Transcriptome analysis of TAOM and HotSeep-1
To collect cells for transcriptome analyses a 3.5-day experiment with replicates of 20 ml culture in 60-ml vials was carried out (Fig. 1). From triplicate TAOM cultures incubated with methane as control, with hydrogen, with methane/hydrogen mixture, or nitrogen as negative control, ∼80% of the enrichment medium was removed and RNA was preserved using pre-heated RNA later (Life Technologies, ThermoFisher Scientific). Total RNA was extracted using the Quick-RNA MiniPrep kit (Zymo Research), treated with DNase I (Roche) and purified using the RNeasy MinElute Cleanup kit (Qiagen) following the manufacturer’s recommendations. Removal of rRNA was omitted and total RNA was prepared for sequencing using the TruSeq stranded mRNA library prep kit (Illumina) following the manufacturer’s guidelines. The cDNA library was sequenced on a MiSeq instrument (MiSeq, Illumina) generating between 2 to 3 million 150-bp single-end reads per library. Quality-controlled reads were mapped to the draft genomes of HotSeep-1 and ANME-1 using bbmap (v.35) with a minimum mapping identity of 98%. To quantify gene expression unambiguously mapped reads per gene were counted using bedtools multicov (v.2.24.0). To compare relative expression patterns within each organism across treatments, read counts per feature were converted to transcripts per million (TPM), which is the abundance of a specific gene (i) relative to the abundance and length of all other transcribed genes (j) observed in one million sequenced reads calculated according to ref. 51:

where X = counts and l = length (bp) per gene. Relative changes in expression of selected genes were calculated by comparing TPM normalized expression data of the H2 and H2 + CH4 treatment to those under TAOM (control) conditions. Differential expression (P value, fold change and effect size) between control (TAOM condition) and treatment (H2 or H2 + CH4) was computed with the aldex2 R package52 for ANOVA-like differential expression analysis. Raw read numbers, read mapping data and statistical analysis are provided in Supplementary Table 1 (total expression) and Supplementary Table 5 (specific gene expression).
For HotSeep-1 transcriptomes total RNA was extracted from triplicate cultures (50 ml) grown on hydrogen/CO2 following the same procedure as described for TAOM enrichments (see above). Removal of rRNA was omitted and total RNA was prepared for sequencing using the TruSeq stranded mRNA library prep kit (Illumina) following the manufacturer’s guidelines. The cDNA library was sequenced on a MiSeq instrument (MiSeq, Illumina) generating between 6.4 to 6.9 million 75-bp paired-end reads per library. Quality-controlled reads were mapped to the draft genome of HotSeep-1 using bbmap (v.35) with a minimum mapping identity of 98%. To quantify gene expression unambiguously mapped reads per gene were counted using featureCount53 (part of Subread, v.1.4.6.) with the -p option to count fragments instead of reads. Fragment counts per gene were converted to transcripts per million (TPM) as described above for TAOM transcriptome analyses.
Cultivation of Geobacter consortia
Active cultures of G. sulfurreducens (strain PCA; DSM 12127) and G. metallireducens (strain GS-15; DSM 7210) were mixed in fresh medium (DSM Medium 826) supplied with Na2-fumarate (50 mM) and ethanol (20 mM) according to ref. 10 and cultivated anaerobically at 33 °C. After subsequent transfers (1% inoculum) a well-growing culture consisting of reddish microbial aggregates developed, which was used for thin sectioning and electron microscopy.
Transmission electron microscopy
The cell material was harvested at 2,000 r.p.m. using a Stat Spin Microprep 2 table-top centrifuge. After centrifugation the pellet was fixed by immersion using 2% glutaraldehyde in 0.1 M cacodylate buffer at pH 7.4. Fixation was performed for 60 min at room temperature. The fixed pellet was immobilized with 2% agarose in cacodylate buffer at pH 7.4. The pellet was cubed and the pieces carefully washed with buffer and further fixed in 1% osmium tetroxide. After pre-embedding staining with 1% uranyl acetate, samples were dehydrated and embedded in Agar 100 (Epon 812 equivalent). As an independent complementary method (shown in Extended Data Fig. 5a), samples were placed in aluminium platelets of 150 µm depth containing 1-hexadecene (ref. 54). The platelets were frozen using a Leica Em HPM100 high pressure freezer (Leica Mikrosysteme Vertrieb GmbH). The frozen samples were transferred to an Automatic Freeze Substitution Unit Leica EM AFS2 and substituted at −90 °C in a solution containing anhydrous acetone, 0.1% tannic acid for 24 h and in anhydrous acetone, 2% OsO4, 0.5% anhydrous glutaraldehyde (EMS Electron Microscopical Science) for an additional 8 h. After a further incubation over 20 h at −20 °C samples were warmed up to + 4 °C and washed with anhydrous acetone subsequently. The samples were embedded at room temperature in Agar 100 at 60 °C over 24 h. Thin sections (30–60 nm) were counterstained with uranyl acetate and lead citrate and examined using a Philips CM 120 transmission electron microscope (Philips Inc.). In total, we recorded more than 200 views on TAOM consortia, 64 views on HotSeep-1 and 90 views of Geobacter consortia.
Thermodynamic calculations
Free energy yields (ΔGrxn) were calculated according to the equation:

including the gas constant R, the temperature T (K) and the measured activities/partial pressures of the respective products Pi and reactants Ri in their respective stoichiometric appearance (n) in the reaction. Values consider activities and fugacity of respective compounds. The temperature-corrected standard free energy  were determined according to ref. 55.
were determined according to ref. 55.
Accession codes
Primary accessions
BioProject
GenBank/EMBL/DDBJ
Data deposits
Representative full-length 16S rRNA gene sequences of TAOM and HotSeep-1 enrichment have been submitted to NCBI GenBank under accession numbers KT152859–KT152885 and KT759143–KT759147, functional genes under the accession numbers KT152886, KT152887 and KT795302–KT795321, and genomic 16S rRNA genes under the accession numbers KT795322 and KT795323. Sequencing projects are registered at NCBI under the BioProject accession numbers PRJNA286178 (TAOM enrichment) and PRJNA276404 (HotSeep-1 enrichment).
References
Reeburgh, W. S. Oceanic methane biogeochemistry. Chem. Rev. 107, 486–513 (2007)
Boetius, A. & Wenzhöfer, F. Seafloor oxygen consumption fuelled by methane from cold seeps. Nat. Geosci. 6, 725–734 (2013)
Hinrichs, K.-U., Hayes, J. M., Sylva, S. P., Brewer, P. G. & DeLong, E. F. Methane-consuming archaebacteria in marine sediments. Nature 398, 802–805 (1999)
Boetius, A. et al. A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 407, 623–626 (2000)
Orphan, V. J. et al. Comparative analysis of methane-oxidizing archaea and sulfate-reducing bacteria in anoxic marine sediments. Appl. Environ. Microbiol. 67, 1922–1934 (2001)
Holler, T. et al. Thermophilic anaerobic oxidation of methane by marine microbial consortia. ISME J. 5, 1946–1956 (2011)
Thauer, R. K. & Shima, S. Methane as fuel for anaerobic microorganisms. Ann. NY Acad. Sci. 1125, 158–170 (2008)
McGlynn, S. E., Chadwick, G. L., Kempes, C. P. & Orphan, V. J. Single cell activity reveals direct electron transfer in methanotrophic consortia. Nature http://dx.doi.org/10.1038/nature15512 (this issue)
Reguera, G. et al. Extracellular electron transfer via microbial nanowires. Nature 435, 1098–1101 (2005)
Summers, Z. M. et al. Direct exchange of electrons within aggregates of an evolved syntrophic coculture of anaerobic bacteria. Science 330, 1413–1415 (2010)
Knittel, K. & Boetius, A. Anaerobic oxidation of methane: progress with an unknown process. Annu. Rev. Microbiol. 63, 311–334 (2009)
Niemann, H. et al. Novel microbial communities of the Haakon Mosby mud volcano and their role as a methane sink. Nature 443, 854–858 (2006)
Schreiber, L., Holler, T., Knittel, K., Meyerdierks, A. & Amann, R. Identification of the dominant sulfate-reducing bacterial partner of anaerobic methanotrophs of the ANME-2 clade. Environ. Microbiol. 12, 2327–2340 (2010)
Hoehler, T. M., Alperin, M. J., Albert, D. B. & Martens, C. S. Field and laboratory studies of methane oxidation in an anoxic marine sediment: evidence for a methanogen-sulfate reducer consortium. Glob. Biogeochem. Cycles 8, 451–463 (1994)
Krüger, M. et al. A conspicuous nickel protein in microbial mats that oxidize methane anaerobically. Nature 426, 878–881 (2003)
Hallam, S. J. et al. Reverse methanogenesis: testing the hypothesis with environmental genomics. Science 305, 1457–1462 (2004)
Meyerdierks, A. et al. Metagenome and mRNA expression analyses of anaerobic methanotrophic archaea of the ANME-1 group. Environ. Microbiol. 12, 422–439 (2010)
Kojima, H., Moll, J., Kahnt, J., Fukui, M. & Shima, S. A reversed genetic approach reveals the coenzyme specificity and other catalytic properties of three enzymes putatively involved in anaerobic oxidation of methane with sulfate. Environ. Microbiol. 16, 3431–3442 (2014)
Thauer, R. K. Anaerobic oxidation of methane with sulfate: on the reversibility of the reactions that are catalyzed by enzymes also involved in methanogenesis from CO2 . Curr. Opin. Microbiol. 14, 292–299 (2011)
Milucka, J., Widdel, F. & Shima, S. Immunological detection of enzymes for sulfate reduction in anaerobic methane-oxidizing consortia. Environ. Microbiol. 15, 1561–1571 (2013)
Milucka, J. et al. Zero-valent sulphur is a key intermediate in marine methane oxidation. Nature 491, 541–546 (2012)
Schink, B. Energetics of syntrophic cooperation in methanogenic degradation. Microbiol. Mol. Biol. Rev. 61, 262–280 (1997)
Stams, A. J. & Plugge, C. M. Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nature Rev. Microbiol. 7, 568–577 (2009)
Moran, J. J. et al. Methyl sulfides as intermediates in the anaerobic oxidation of methane. Environ. Microbiol. 10, 162–173 (2008)
Stokke, R., Roalkvam, I., Lanzen, A., Haflidason, H. & Steen, I. H. Integrated metagenomic and metaproteomic analyses of an ANME‐1‐dominated community in marine cold seep sediments. Environ. Microbiol. 14, 1333–1346 (2012)
Nagarajan, H. et al. Characterization and modelling of interspecies electron transfer mechanisms and microbial community dynamics of a syntrophic association. Nature Commun. 4, 2809 (2013)
Rotaru, A.-E. et al. A new model for electron flow during anaerobic digestion: direct interspecies electron transfer to Methanosaeta for the reduction of carbon dioxide to methane. Energ. Environ. Sci. 7, 408–415 (2014)
Malvankar, N. S. et al. Structural basis for metallic-like conductivity in microbial nanowires. MBio 6, e00084–e00015 (2015)
Methé, B. A. et al. Genome of Geobacter sulfurreducens: metal reduction in subsurface environments. Science 302, 1967–1969 (2003)
Reitner, J. et al. Concretionary methane-seep carbonates and associated microbial communities in Black Sea sediments. Palaeogeogr. Palaeoclimatol. Palaeoecol. 227, 18–30 (2005)
Widdel, F. & Bak, F. in The Prokaryotes Vol. 4 (eds Trüper, H. G., Balows, A., Dworkin, M., Harder, W. & Schleifer, K. H. ) 3352–3378 (Springer, 1992)
Zhou, J., Bruns, M. A. & Tiedje, J. M. DNA recovery from soils of diverse composition. Appl. Environ. Microbiol. 62, 316–322 (1996)
Muyzer, G., Teske, A., Wirsen, C. O. & Jannasch, H. W. Phylogenetic relationships of Thiomicrospira species and their identification in deep-sea hydrothermal vent samples by denaturing gradient gel electrophoresis of 16S rDNA fragments. Arch. Microbiol. 164, 165–172 (1995)
Massana, R., Murray, A., Preston, C. & DeLong, E. Vertical distribution and phylogenetic characterization of marine planktonic Archaea in the Santa Barbara Channel. Appl. Environ. Microbiol. 63, 50–56 (1997)
Teske, A. et al. Microbial diversity of hydrothermal sediments in the Guaymas Basin: evidence for anaerobic methanotrophic communities. Appl. Environ. Microbiol. 68, 1994–2007 (2002)
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004)
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35, 7188–7196 (2007)
Pernthaler, A., Pernthaler, J. & Amann, R. Fluorescence in situ hybridization and catalyzed reporter deposition for the identification of marine bacteria. Appl. Environ. Microbiol. 68, 3094–3101 (2002)
Steudel, R., Göbel, T. & Holdt, G. The molecular composition of hydrophilic sulfur sols prepared by decomposition of thiosulfate. Z. Naturforsch. B Chem. Sci. 43, 203–218 (1988)
Cord-Ruwisch, R. A quick method for the determination of dissolved and precipitated sulfides in cultures of sulfate-reducing bacteria. Microbiol. Meth. 4, 33–36 (1985)
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012)
Strous, M., Kraft, B., Bisdorf, R. & Tegetmeyer, H. E. The binning of metagenomic contigs for microbial physiology of mixed cultures. Front. Microbiol. 3, 410 (2012)
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014)
Meyer, F. et al. GenDB—an open source genome annotation system for prokaryote genomes. Nucleic Acids Res. 31, 2187–2195 (2003)
Richter, M. et al. JCoast — a biologist-centric software tool for data mining and comparison of prokaryotic (meta) genomes. BMC Bioinformatics 9, 177 (2008)
Richter, M. & Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl Acad. Sci. USA 106, 19126–19131 (2009)
Eddy, S. HMMER User’s Guide. Biological sequence analysis using profile hidden Markov models (Howard Hughes Medical Institute, 2003)
Finn, R. D., Miller, B. L., Clements, J. & Bateman, A. iPfam: a database of protein family and domain interactions found in the Protein Data Bank. Nucleic Acids Res. 42, D364–D373 (2014)
Haft, D. H., Selengut, J. D. & White, O. The TIGRFAMs database of protein families. Nucleic Acids Res. 31, 371–373 (2003)
Yu, N. Y. et al. PSORTb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26, 1608–1615 (2010)
Li, B., Ruotti, V., Stewart, R. M., Thomson, J. A. & Dewey, C. N. RNA-Seq gene expression estimation with read mapping uncertainty. Bioinformatics 26, 493–500 (2010)
Fernandes, A. D. et al. Unifying the analysis of high-throughput sequencing datasets: characterizing RNA-seq, 16S rRNA gene sequencing and selective growth experiments by compositional data analysis. Microbiome 2, 1–13 (2014)
Liao, Y., Smyth, G. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014)
Studer, D., Michel, M. & Müller, M. High pressure freezing comes of age. Scanning Microsc., Suppl. 3, 253–268 (1989)
Conrad, R. & Wetter, B. Influence of temperature on energetics of hydrogen metabolism in homoacetogenic, methanogenic, and other anaerobic bacteria. Arch. Microbiol. 155, 94–98 (1990)
Acknowledgements
We thank K. Harding, I. Kattelmann, A. Ellrott and M. Meiners for technical support, and M. Richter, H. Gruber-Vodicka and P. Luigi Buttigieg for bioinformatic support, and K. Knittel, N. Dubilier, M. M. M. Kuypers and F. Widdel for discussions. Furthermore we thank A. Teske and the RV ATLANTIS and ALVIN team of cruise AT15-56 in 2009 for providing the initial sediment material. The project was funded by the DFG Leibniz program to A.B., and by the DFG excellence cluster MARUM, Center of Marine Environmental Sciences, Bremen. Further support was provided by the Max Planck Society to D.R. and A.B.
Author information
Authors and Affiliations
Contributions
G.W., V.K. and A.B designed the experiments; G.W., V.K., H.E.T. and D.R. performed experiments and data analyses; G.W., V.K. and A.B. wrote the manuscript with contributions from D.R. and H.E.T.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Models of possible species interaction mechanisms in TAOM tested in this study.
a, Transfer of molecular intermediates such as hydrogen. b, Incomplete reduction of sulfate in ANME and zero-valent sulfur transfer to the partner bacteria. c, Direct interspecies electron transfer via conductive nanowires.
Extended Data Figure 2 Visualization of and growth experiments with HotSeep-1.
a, Representative fluorescence micrograph of HotSeep-1 culture (probe HotSeep-1-590; 22 similar images obtained). Cells are solitary or form small aggregates. Scale bar, 10 µm. b, c, Semi-logarithmic illustration of the development of sulfide (b) or numbers of cells and resulting doubling times (c) (doubling time = ln(2)/exponential factor of the regression curve) during incubation of the HotSeep-1 culture with hydrogen as the sole energy source and sulfate. Biological replicates n = 3; data is presented as mean ± s.d., lines of best fit defined by least squares method.
Extended Data Figure 3 Effect of zero-valent elemental sulfur and molybdate additions on TAOM.
a, Sulfide production in response to zero-valent (colloidal) sulfur addition versus TAOM conditions; zero-valent sulfur did not cause sulfide formation. b, c, Monitoring of hydrogen partial pressures at TAOM conditions (open circles) versus extra addition of 10 mM molybdate (filled circles) for either the full times series (b) or the first 10 h (c). Molybdate addition caused tenfold higher hydrogen concentrations than the TAOM condition. d, Inhibition of methane-dependent sulfide production at different molybdate concentrations. Biological replicates n = 3; symbols represent mean values; error bars are s.d.; b and c show a single time series with the same culture.
Extended Data Figure 4 Effect of hydrogen on microbial methane oxidation.
a, Methane (0.15 MPa) supplied as the sole electron source was steadily consumed over time by TAOM. b, When both methane (0.15 MPa) and hydrogen (0.05 MPa) were added, hydrogen was rapidly consumed (grey bars), whereas methane consumption was reversely inhibited (green line) until hydrogen was fully consumed. Afterwards methane consumption occurred at the same rate as in the control with only methane (a). Methane, technical replicates n = 3; symbols represent mean values; error bars are s.d.; hydrogen, single measurements. Experiment was replicated once in the laboratory.
Extended Data Figure 5 Relative expression of marker genes of HotSeep-1 in consortial growth on methane (TAOM) versus enrichment on hydrogen.
Genes encoding proteins apparently involved in direct interspecies electron transfer (CytC and PilA) are strongly overexpressed during TAOM (red) compared to hydrogenotrophic growth (green). Gene expression given in terms of TPM (transcripts per million). Biological replicates n = 3; error bars are s.d.
Extended Data Figure 6 Thin-sections of TAOM and dual species Geobacter spp. aggregates.
a, TAOM culture. High-pressure frozen ANME-1 cells (A) have a cylindrical shape and a size of 1.5 × 0.8 µm, appearing circular in cross-section, and rectangular when cut along the axis. Their cell content shows a high contrast. A* indicates cell envelopes. HotSeep-1 cells (H) are smaller (approximately 1 × 0.5 µm), of rod-like shape, and have lower contrast. The matrix between the cells is largely filled with filaments. Representative of 24 images. Scale bar, 3 µm. b, Thin section of Geobacter consortium with intercellular nanowires using the same embedding techniques as for TAOM consortia, representative of 20 images. Scale bar, 300 nm.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-5. (PDF 162 kb)
Rights and permissions
About this article
Cite this article
Wegener, G., Krukenberg, V., Riedel, D. et al. Intercellular wiring enables electron transfer between methanotrophic archaea and bacteria. Nature 526, 587–590 (2015). https://doi.org/10.1038/nature15733
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15733
- Springer Nature Limited
This article is cited by
-
Mechanisms of extracellular electron transfer in anaerobic methanotrophic archaea
Nature Communications (2024)
-
Enhancement of exogenous riboflavin on microbiologically influenced corrosion of nickel by electroactive Desulfovibrio vulgaris biofilm
npj Materials Degradation (2023)
-
Candidatus Alkanophaga archaea from Guaymas Basin hydrothermal vent sediment oxidize petroleum alkanes
Nature Microbiology (2023)
-
A critical review on nanotechnological advancement in biogas production from organic waste
Biomass Conversion and Biorefinery (2023)
-
Removing 10 cm of degraded peat mitigates unwanted effects of peatland rewetting: a mesocosm study
Biogeochemistry (2023)