Abstract
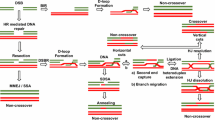
The alternative non-homologous end-joining (NHEJ) machinery facilitates several genomic rearrangements, some of which can lead to cellular transformation. This error-prone repair pathway is triggered upon telomere de-protection to promote the formation of deleterious chromosome end-to-end fusions1,2,3. Using next-generation sequencing technology, here we show that repair by alternative NHEJ yields non-TTAGGG nucleotide insertions at fusion breakpoints of dysfunctional telomeres. Investigating the enzymatic activity responsible for the random insertions enabled us to identify polymerase theta (Polθ; encoded by Polq in mice) as a crucial alternative NHEJ factor in mammalian cells. Polq inhibition suppresses alternative NHEJ at dysfunctional telomeres, and hinders chromosomal translocations at non-telomeric loci. In addition, we found that loss of Polq in mice results in increased rates of homology-directed repair, evident by recombination of dysfunctional telomeres and accumulation of RAD51 at double-stranded breaks. Lastly, we show that depletion of Polθ has a synergistic effect on cell survival in the absence of BRCA genes, suggesting that the inhibition of this mutagenic polymerase represents a valid therapeutic avenue for tumours carrying mutations in homology-directed repair genes.
Similar content being viewed by others
Main
Chromosome end-to-end fusions are inhibited by shelterin; a multi-subunit complex anchored to telomeric DNA by two Myb-containing proteins—TRF1 and TRF2 (ref. 4). Telomere fusions are executed by two independent end-joining pathways. Classical non-homologous end-joining (C-NHEJ), mediated by LIG4 and the Ku70/80 heterodimer, is primarily blocked by TRF2 (ref. 5). Conversely, alternative NHEJ (alt-NHEJ), which is dependent on LIG3 (ref. 6) and PARP1 (ref. 7), is repressed in a redundant manner2,3. Alt-NHEJ is fully unleashed after the simultaneous deletion of TRF1 and TRF2, and the creation of shelterin-free telomeres in cells deficient for Ku70 and Ku80 (also known as Xrcc6 and Xrcc5, respectively)2. This error-prone end-joining pathway mediates fusion of naturally eroded telomeres1, joining of switch regions during class-switch recombination8, and formation of chromosomal translocations in mouse cells9,10.
To characterize the differences between C-NHEJ and alt-NHEJ at dysfunctional telomeres, we determined whether the sequence of the junction between two fused telomeres differed depending on the type of repair pathway used. Telomere fusions by C-NHEJ were triggered by Cre-mediated depletion of TRF2 using previously described mouse embryonic fibroblasts (MEFs) (Trf2F/FCre-ERT2)11 (Extended Data Fig. 1a). To induce robust fusions by the alt-NHEJ pathway, we depleted the entire shelterin complex by deleting Trf1 and Trf2 from Trf1F/F Trf2F/F Ku80−/− Cre-ERT2 MEFs2 (Extended Data Fig. 1a). DNA was subjected to next-generation sequencing, and reads corresponding to telomeres were identified on the basis of the presence of at least three consecutive TTAGGG repeats. To detect rare reads containing fusion junctions, we exploited the novel sequence arrangement created by the ligation of the 3′ G-rich strand (TTAGGG-3′) to the 5′ C-rich strand (5′-CCCTAA) (Fig. 1a), and filtered reads that started with at least three G-rich repeats and ended with two or more C-rich repeats. We confirmed that this approach could successfully identify telomere fusions by comparing reads derived from Trf2-proficient and Trf2-deficient cells. Starting with a similar number of telomere-repeat containing reads, we identified >90 fusogenic events in Trf2-knockout MEFs, compared to only three events in wild-type cells (Fig. 1b). Sequence analysis of the junctions highlighted different permutations of TTAGGG/AATCCC sequences. Notably, the spectrum of the fusion junctions was different in shelterin-free settings, in which frequent non-telomeric nucleotide insertions (9 out of 46 events) were identified at fusion breakpoints (Fig. 1b–d and Supplementary Information).
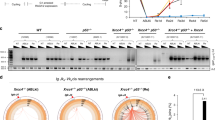
a, Schematic of the junction of a telomere fusion. The 3′ end of the telomeric G-rich strand of a chromosome (blue) is fused to the 5′ end of the C-rich strand of a different chromosome (red). b, Illumina sequencing to analyse telomere fusion junctions. Reads ≥3XTTAGGG consecutively were scored as derived from telomere fragments. Those with ≥3XTTAGGG on the 5′ end and ≥2XCCCTAA at the 3′ end were scored as telomere fusion junctions (see Supplementary Information). c, Examples of telomere fusions generated by C-NHEJ of TRF2-depleted telomeres. Light grey highlights fusion junctions, dark grey marks the flanking telomere repeats. d, Examples of insertions in shelterin-free Ku80-null MEFs. e, Telomere fusions in metaphase spreads from Trf1F/F Trf2F/F Ku80−/− Cre-ERT2 MEFs. Telomeres in red (peptide nucleic acid (PNA) probe) and chromosomes in blue (4′,6-diamidino-2-phenylindole; DAPI). f, Frequency of telomere fusions after the depletion of candidate polymerases. 4-OHT, 4-hydroxytamoxifen; shCtrl, control shRNA. Bars represent mean of n > 1,000 chromosome ends derived from one experiment.
To identify the enzyme that incorporated nucleotides at dysfunctional telomeres, we depleted known low-fidelity DNA polymerases in shelterin-free cells lacking Ku80, and analysed chromosome-end fusions on metaphase spreads. Notably, we observed a reduction in the frequency of telomere fusions in cells with reduced levels of polymerase theta (Polθ, encoded by Polq in mice) (Fig. 1e, f and Extended Data Fig. 1b). The activity of Polθ is specific to alt-NHEJ as its inhibition in Trf2-knockout cells did not affect the frequency of C-NHEJ (Fig. 2a, b and Extended Data Fig. 2a–c).
a, Metaphases from TRF2-depleted (Trf2F/F Cre-ERT2+ 4-OHT) and shelterin-free (Trf1F/F Trf2F/F Ku80−/− Cre-ERT2+ 4-OHT) MEFs infected with the indicated short hairpin RNA (shRNA). b, Quantification of telomere fusions in MEFs with the indicated treatment (mean values ± s.d. derived from six independent experiments. P = 0.003; two-tailed Student’s t-test). c, Design of the translocation assay in which DSBs are induced by Cas9-gRNA(Rosa26;H3f3b). Joining of DNA ends generates der(6) and der(11), detected by nested PCR9. d, Translocation frequency in Polq+/+ and Polq−/− cells 60 h after Cas9-gRNA(Rosa26;H3f3b) expression. Mean values ± s.d. derived from three independent experiments. P = 0.009; two-tailed Student’s t-test.
Polθ is an A-family DNA polymerase that exhibits low fidelity on templated DNA12, and also displays a terminal transferase-like activity that catalyses nucleotide addition in a template-independent manner13. The relevance of these activities in vivo was highlighted in Drosophila melanogaster, in which Polθ was shown to stimulate nucleotide insertions during double-stranded break (DSB) repair by alt-NHEJ14. More recently, Polθ was shown to promote end-joining of replication-associated DSBs in Caenorhabditis elegans15, preventing large deletions around G-rich DNA16. The exact function of Polθ during DSB repair in mammalian cells remains elusive.
The crucial role for Polθ at dysfunctional telomeres prompted us to test whether it is required for DSB repair at non-telomeric loci. To this end, we tested whether the depletion of Polθ affects chromosomal translocations in the context of mouse pluripotent cells, reported to be mediated by alt-NHEJ10 in a LIG3-dependent manner9. To model chromosomal translocations, we induced DSBs in the Rosa26 and H3f3B mouse loci using the CRISPR/Cas9 system (Fig. 2c). When introduced into Polq+/+ and Polq−/− cells17, the Cas9-gRNA(Rosa26;H3f3b) expression plasmid induced simultaneous cleavage of both loci with comparable efficiencies (Extended Data Fig. 2d, e). Consistent with previous reports, 24% of translocation events in Polq+/+ cells were scarred by random insertions. Interestingly, the overall frequency of translocations in cells lacking Polq was significantly reduced (Fig. 2d). Sequence analysis of residual translocations in Polq−/− cells highlighted the absence of insertions, and a concomitant decrease in micro-homology at junctions (Fig. 2d and Extended Data Figs 2, 3, 4, 5). Notably, we observed similar results when assessing translocation frequency in cells expressing a catalytically inactive form of Polθ (Extended Data Fig. 2g–k). Altogether, our data indicate that the promiscuous activity of Polθ during DSB repair contributes to the increased mutagenicity of alt-NHEJ. Importantly, our results indicate that mammalian Polθ plays a critical part by stimulating the end-joining reaction.
We next investigated the upstream signalling event(s) required for the recruitment of Polθ to DNA damage sites, induced after micro-irradiation of HeLa cells expressing Myc-tagged Polθ. Accumulation of Polθ at laser-induced DNA breaks, discerned by its co-localization with the phosphorylated histone H2AX (γ-H2AX), occurred in ∼25% of cells that stained positive for Myc (Fig. 3a, b), and was independent of either ATM or ATR signalling (Extended Data Fig. 6). Instead, co-localization of Polθ with γ-H2AX was reduced after depletion of PARP1 with short interfering RNAs (siRNAs), or after the inhibition of PARP1 activity (using KU58948) (Fig. 3a, b). In a parallel experiment that used a recently developed U2OS-DSB reporter cell line18, we were able to ascertain the localization of Myc–Polθ to bona fide DSBs, induced after FOK1 cleavage of a LacO-tagged genomic locus (Extended Data Fig. 7). In conclusion, our data suggest that PARP1, previously known to be required for alt-NHEJ7,19, facilitates the recruitment of Polθ to DSBs.
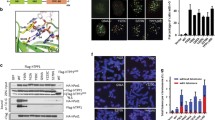
a, Myc–Polθ localization to DNA damage was monitored after laser micro-irradiation of HeLa cells. Cells were fixed and stained for γ-H2AX and Myc, 1 h after damage induction. b, Quantification of Polθ accumulation at sites of laser damage (mean values ± s.e.m. derived from two independent experiments). c, To test whether Polθ represses recombination at telomeres, we depleted the polymerase in shelterin-free and Lig4-deficient MEFs2, and both repair pathways were monitored using CO-FISH. White arrows indicate alt-NHEJ events, red arrows highlight HDR-mediated T-SCEs. d, Quantification of telomere fusion (alt-NHEJ) and T-SCE (HDR) in cells transduced with shRNAs against Polq, Lig3 or control shRNA. Error bars denote ± s.d. from three independent experiments. e, Immunofluorescence for RAD51 and γ-H2AX in the indicated MEFs 3 h after irradiation. f, Graph representing quantification of ionizing-radiation-induced RAD51 foci. Mean values ± s.d. derived from three independent experiments. P < 0.05, P < 0.01; two-tailed Student’s t-test.
Homology-directed repair (HDR) is prevalent during the S/G2 phase of the cell cycle, which coincides with the peak of alt-NHEJ activity, and these pathways also share the initial resection step mediated by MRE11 and CtIP20. To test whether inhibiting alt-NHEJ could potentially result in increased HDR, we depleted shelterin in Lig4-deficient MEFs, a genetic setting that is conducive to the activity of alt-NHEJ as well as HDR2. To investigate the relative contribution of the two repair pathways, we used a chromosome-orientation fluorescence in situ hybridization (CO-FISH) assay21, and monitored the exchange of telomeres between sister chromatids by HDR (telomere sister chromatid exchange, T-SCE), and, at the same time, measured the frequency of chromosome end–end fusion by end-joining (Fig. 3c). After depletion of shelterin from Trf1F/F Trf2F/F Lig4−/− Cre-ERT2 MEFs, ∼10% of the telomeres were processed by alt-NHEJ, whereas ∼5% of chromosome ends showed T-SCEs2 (Fig. 3c, d and Extended Data Fig. 8a–c). As expected, we observed a substantial reduction in the frequency of alt-NHEJ at shelterin-free telomeres in Lig4−/− cells that lack Polq or Lig3 (Fig. 3d and Extended Data Fig. 8a–e). Remarkably, Polq-depleted cells exhibited a concomitant increase in T-SCE, which was not evident in cells lacking Lig3 (Fig. 3d), thereby highlighting a unique role for Polθ in counteracting HDR. To gain insight into this novel Polθ function, we show that the promiscuous polymerase is not required for end-resection of DSBs (Extended Data Fig. 8f, g). Instead, its activity counteracts the accumulation of RAD51 foci (Fig. 3e, f and Extended Data Fig. 8h). To corroborate these findings, we used the traffic light reporter (TLR) system, designed to generate a flow-cytometric readout for HDR and end-joining at a site-specific DNA break induced by I-Sce1 (ref. 22). We observed that after knocking down Polq in Lig4−/− cells, resolution of the I-Sce1-induced DNA break by HDR is increased, in conjunction with a significant reduction in the frequency of alt-NHEJ (Extended Data Fig. 9).
Alt-NHEJ is often considered as a back-up choice for DSB repair, operating at the expense of genomic stability. Circumstantial evidence suggests that this pathway could be enhanced when HDR is impaired23,24. We therefore postulated that this error-prone mode of repair has an essential role in cells with compromised HDR activity. We tested this hypothesis by inhibiting Polq in cells lacking the breast cancer susceptibility genes—Brca1 and Brca2. Chromosome analysis revealed a fourfold increase in chromosomal aberrancies after Polq depletion in MEFs lacking either Brca1 or Brca2. Such aberrancies included chromatid and chromosome breaks, in addition to radial chromosome structures characteristic of Lig4-mediated processing of chromatid breaks via the C-NHEJ pathway (Fig. 4a, b and Extended Data Fig. 10a, b). Ultimately, the increased genomic instability in cells co-depleted for Polq and Brca genes compromised cellular survival. We observed that BRCA1-mutated human cells (Fig. 4c, d), and mouse cells lacking Brca1 (Extended Data Fig. 10c–f), displayed significantly reduced colony-forming capabilities after Polq impairment. Although we cannot exclude that Polθ performs additional activities required for the survival of Brca-deficient cells25, our data suggest that Polθ-mediated alt-NHEJ promotes the survival of cells with a compromised HDR pathway. In the absence of a safer means to repair breaks, alt-NHEJ may therefore prevent genomic havoc by resolving unrepaired lesions.
a, Analysis of genomic instability in metaphase spreads from Brca1F/FCre-ERT2 MEFs treated with shRNA against Polq or vector control. b, Quantification of breaks (chromatid and chromosome) and radials in Brca1F/FCre-ERT2 and Brca2F/FCre-ERT2 MEFs with the indicated treatment. Mean values are presented with error bars denoting ± s.d. from three independent experiments. c, Clonogenic survival after Polq depletion. Crystal violet staining of BJ-hTERT, MCF7 and HCC1937 cells treated with shRNA against Polq or vector control. d, Quantitative analyses of colony formation assay. Colonies in each control shRNA cell line were set to 100%. Colonies in shPolq-expressing cells are normalized to shCtrl. Mean values ± s.d. derived from three independent experiments. P < 0.05, P < 0.01; two-tailed Student’s t-test. e, Schematic depicting our model for the function of Polθ during DSB repair (see Supplementary Information).
Here we provide direct evidence linking Polθ to alt-NHEJ repair in mammalian cells (Fig. 4e). We also show that while Polθ hinders error-free repair by HDR, its activity is essential for the survival of HDR-deficient cells (Fig. 4e). The question remains as to how this promiscuous polymerase orchestrates DSB repair. After DSB formation in the S/G2 phase of the cell cycle, resection of DSBs by MRE11 and CtIP20 potentially exposes micro-homology that allows spontaneous annealing of broken DNA ends (Fig. 4e). The binding of RPA antagonizes this annealing step to promote HDR-mediated repair26,27. An opposing activity is likely to be exerted by Polθ. Placing our finding in the context of recent biochemical experiments and genetic studies in model organisms13,14,15,16, we ultimately propose a model in which Polθ exploits both its template-independent and template-dependent activities to stabilize the annealed intermediate structure and channel repair towards the alt-NHEJ pathway (Fig. 4e). Our findings that Polθ is critical for alt-NHEJ support this model, which provides a potential explanation as to how this polymerase counteracts HDR.
Finally, it is intriguing that although POLQ expression in normal human tissues is generally repressed28, it is upregulated in a wide range of human cancers and associates with poor clinical outcome in breast tumours29,30. Our findings that cells with compromised HDR activity depend on this mutagenic polymerase for survival establish a rationale for the development of Polθ-targeted approaches for cancer treatment.
Methods
Cell culture procedures
Trf2F/F Cre-ERT2, Trf1F/F Trf2F/F Ku80−/− Cre-ERT2 and Trf1F/F TrfF/F Lig4−/− Cre-ERT2 MEF lines were previously described2,11. Polq+/+ and Polq−/− MEFs were a gift from N. Shima17. Brca1F/FCre-ERT2 and Brca2F/FCre-ERT2 MEFs and U2OS-DSB reporter cells18 were a gift from R. Greenberg. Trf1F/F Trf2F/F Lig4−/− Cre-ERT2 MEFs were derived from mice that were deficient in p53. The remaining MEF lines were immortalized with pBabeSV40LargeT. MEFs were cultured in DMEM supplemented with 10–15% FBS (Gibco), 2 mM l-glutamine (Sigma), 100 U ml−1 penicillin (Sigma), 0.1 μg ml−1 streptomycin (Sigma), 0.1 mM non-essential amino acids (Invitrogen) and 1 mM sodium pyruvate (Sigma). Expression of Cre recombinase was induced by treating MEFs carrying the Cre-ERT2 allele with 0.5 μM 4-OHT (Sigma H7904) for 12 h. The t = 0 time point was set at the time of treatment with 4-OHT. BJ-hTERT and MCF7 cells were grown in DMEM supplemented with 10% FBS. HCC1937 cells were grown in RPMI medium (Gibco) containing 15% FBS. U2OS-DSB reporter cells were grown in DMEM supplemented with 10% BCS. Human HeLa cells were grown in DMEM supplemented with 10% FBS, 100 U ml−1 penicillin, 100 μg ml−1 streptomycin and 2 mM l-glutamine. Mouse embryonic stem cells were grown in DMEM supplemented with 15% FBS (ES-qualified FBS) (Gibco), 2 mM l-glutamine (Sigma), 100 U ml−1 penicillin (Sigma), 0.1 μg ml−1 streptomycin (Sigma), 0.1 mM non-essential amino acids (Invitrogen), leukaemia inhibitory factor (LIF) and 2-β-mercaptoethanol (Gibco 21985). For inhibitor experiments, PARPi (KU58948, Axon medchem), ATMi (KU-55933, Tocris) and ATRi (VE-821, Selleckchem) were all used at a final concentration of 10 μM, and were applied to culture medium 2–4 h before irradiation. For ionizing radiation treatment, cells were exposed to 1–10 Gy ionizing radiation by a Faxitron X-ray system (120 kV, 5 mA, dose rate 5 Gy min−1) and recovered for 4 h before immunofluorescence analysis.
Lentiviral delivery of shRNA
shRNA treatments were carried out before 4-OHT treatment. shRNAs (see below for a list of sequences) were introduced by two lentiviral infections at 12 h intervals using supernatant from transfected 293T cells. Parallel infection with pLK0.1 was used as a negative control. Cells were selected with puromycin for 3 days.
TLR assay
Lentiviral constructs coding for TLR (31482) and I-Sce1 with donor e-GFP (31476) were purchased from Addgene22. To avoid the confounding effect of classical-NHEJ on the repair of I-Sce1-induced DNA breaks, we stably integrated the TLR construct into Ku80−/− and Lig4−/− MEFs. The plasmid was transduced by two lentiviral infections at 12 h intervals using supernatant from transfected 293T cells. Cells with integrated TLR were selected with puromycin for 5 days. Cells were then transduced with concentrated Polq shRNA lentiviral particles followed by I-Sce1. Cells were collected 72 h later without further antibiotic selection and analysed on a BD LSRII. eGFP fluorescence, which reflects HDR repair, was measured using a 488-nm laser for excitation and a 530/30 filter for detection. mCherry fluorescence, indicative of alt-NHEJ was measured by using a 561-nm laser for excitation and a 610/20 filter for detection. Data were analysed using FloJo software.
Detection of telomeric fusions
To enrich for telomeric DNA, genomic DNA was digested with two frequent cutters (AluI and MboI) and fragments greater than 10 kilobases (kb) were isolated. The resulting DNA was used to generate a library using the NEBNext Ultra Library Prep Kit and the NEBNext Multiplex Oligos for Illumina (NEB) following the manufacturer instructions. The resulting library was run on an Illumina HiSeq platform generating 100-base-pair (bp) indexed pair-end reads.
Transient transfection of cells and laser micro-irradiation
Full-length human POLQ was cloned into pLPC-Myc vectors. HeLa cells were plated on glass-bottomed dishes (Willco Wells). Myc–Polθ constructs were transfected into HeLa and U2OS-DSB reporter cells cell with HilyMax (Dojindo) according to the manufacturer’s instruction. Then, 24 h after transfection, cells were pre-sensitized with 10 μM 5-bromo-2′-deoxyuridine (BrdU) in normal DMEM medium for 20 h. After indicated treatments, cells were damaged by laser micro-irradiation as previously described31. After laser micro-irradiation, cells were incubated for 1 h, then fixed and analysed by immunofluorescence and microscopic imaging as described below. For PARP1 siRNA experiments, cells were transfected with siCtrl (non-targeting pool, Thermo Scientific) or siPARP1 (GGGCAAGCACAGUGUCAAAUU, Sigma), for 24 h before POLQ transfections and subsequent treatments as described above.
Immunofluorescence and confocal microscopy
After the indicated treatments, cells were processed and analysed for immunofluorescence as previously described32. In brief, cells were fixed with 2% (v/v) paraformaldehyde for 15 min at room temperature. Cells were washed with PBS, permeabilized with 0.5% (v/v) Triton X-100 for 10 min, and blocked with PBS containing 3% BSA. Cells were incubated with the same buffer containing primary antibodies for 1 h at room temperature followed by secondary antibodies incubations for 1 h at room temp. Cells were imaged and analysed with Z-stacked setting using the FV10-ASW3.1 software on a Fluoview 1000 confocal microscope (Olympus). For laser line quantification, >50 cells were counted for all conditions from two independent experiments. The primary antibodies used for immunofluorescence were γ-H2AX (p Ser139) (rabbit polyclonal, Novus, NB100-384) and c-Myc (mouse monoclonal, Santa Cruz, sc-40). The secondary antibodies used for immunofluorescence were Alexa Fluor 594 (rabbit) (Invitrogen, A11037) and Alexa Fluor 488 (mouse) (Invitrogen, A11029). To analyse the recruitment of Polθ to double-stranded breaks, U2OS-DSB reporter cells expressing Myc–Polθ were analysed 4 h after treatment with shield and tamoxifen. Lastly, to analyse RAD51 foci formation and its co-localization with γ-H2AX (p Ser139) after ionizing radiation treatment, cells were treated with 0.2% Triton X-100 (in PBS) for 5 min on ice before fixation with paraformaldehyde. The primary antibodies used for RAD51 immunofluorescence were γ-H2AX (p Ser139) (mouse monoclonal, Novus, NB100-384) and RAD51 (rabbit polyclonal, Santa Cruz, sc-8349).
FISH
Cells were collected at 96 h after treatment with 4-OHT to analyse the frequency of telomere fusions. In brief, ∼80% confluent MEFs were incubated for 2 h with 0.2 μg ml−1 colcemid (Sigma). The cells were collected by trypsinization, resuspended in 0.075 M KCl at 37 °C for 30 min, and fixed overnight in methanol/acetic acid (3:1) at 4 °C. The cells were dropped onto glass slides and the slides were dried overnight. The next day, the slides were rehydrated with PBS for 15 min then fixed with 4% formaldehyde for 2 min at room temperature. Slides were digested with 1 mg ml−1 pepsin, pH 2.2, at 37 °C for 10 min, washed three times with PBS and fixed again in 4% formaldehyde for 2 min at room temperature. After three PBS washes, the slides were incubated consecutively with 75%, 95% and 100% ethanol and allowed to air dry for 30 min before applying hybridization solutions (70% formamide, 1 mg ml−1 blocking reagent (Roche), 10 mM Tris-HCl, pH 7.2) containing TAMRA-OO-(TTAGGG)3 PNA probes (Applied Biosystems). Slides were denatured by heating for 3 min at 80 °C and hybridized for 2 h at room temperature. The next day, the slides were washed twice for 15 min each in 70% formamide, 10 mM Tris-HCl, followed by three 5-min washes in 0.1 M Tris-HCl, pH 7.0, 0.15 M NaCl and 0.08% Tween-20. Chromosomal DNA was counterstained with DAPI during the second PBS wash. Slides were mounted in antifade reagent (ProLong Gold, Invitrogen) and images were captured with a Nikon Eclipse TI microscope (see http://delangelab.rockefeller.edu/protocols).
CO-FISH
Cells were labelled with BrdU:BrdC (3:1, final concentration 10 μM) for 14–16 h. Two hours before collection by trypsinization, 0.2 μg ml−1 colcemid was added to the media. To fix the cells and drop metaphases on a glass slide, the same procedure that was applied for FISH was followed. Slides were treated with 0.5 mg ml−1 RNase A (in PBS, DNase-free) for 10 min at 37 °C, incubated with 0.5 μg ml−1 Hoechst 33258 (Sigma) in 2×SSC for 15 min at room temperature, and exposed to 365-nm ultraviolet light (Stratalinker 1800 UV irradiator) for 30 min. The slides were then digested twice with 800 U exonuclease III (Promega) at room temperature for 10 min each, washed with PBS and dehydrated through an ethanol series of 70%, 95% and 100%. After air-drying, slides were hybridized with Tamra-OO-(TTAGGG)3 PNA probe in hybridization solution (70% formamide, 1 mg ml−1 blocking reagent (Roche) and 10 mM Tris-HCl, pH 7.2) for 2 h at room temperature. The slides were then washed for a few seconds with 70% formamide and 10 mM Tris-HCl, pH 7.2, and incubated with FITC-OO-(CCCTAA)3 PNA probe in hybridization solution for 2 h. Slides were washed and mounted as described for FISH (see http://delangelab.rockefeller.edu/protocols).
Western blot analysis
Cells were collected by trypsinization, lysed in 2× Laemmli buffer (100 mM Tris-HCl, pH 6.8, 200 μM dithiothreitol, 3% SDS, 20% glycerol and 0.05% bromophenol blue) at 1 × 104 cells per microlitre. The lysate was denatured for 10 min at 95 °C, and sheared by forcing it through a 28-gauge insulin needle ten times. Lysate from 1 × 105 cells was loaded on an SDS–PAGE and transferred to a nitrocellulose membrane. The membrane was blocked in 5% milk in TBS with 0.1% Tween-20 and incubated with primary antibody in TBS, 5% milk and 0.1% Tween-20 for 2 h at room temperature. The following primary antibodies were used: Polθ (ab80906, Abcam); TRF1 (1449, rabbit polyclonal); RAP1 (1252, rabbit polyclonal); phospho-CHK2 (Thr68) (rabbit polyclonal, Cell Signaling); CHK2 (rabbit polyclonal, Cell Signaling); phospho-CHK1 (Ser 345) (mouse monoclonal, Cell Signaling); CHK1 (mouse monoclonal, Santa Cruz); LIG3 (mouse monoclonal, Santa Cruz); Myc (9E10; Calbiochem); and γ-tubulin (clone GTU-88, Sigma); PARP1 (polyclonal, Cell signaling). (See http://delangelab.rockefeller.edu/protocols.)
Chromosomal aberrancies
Cells were collected and dropped on microscope slides as described for the FISH protocol. After the slides had dried overnight, they were re-hydrated in PBS, stained with 0.25 μg ml−1 DAPI, dehydrated in a 70%, 95% and 100% ethanol series, mounted and imaged using Nikon Eclipse TI microscope. Aberrancies were scored as a percentage of chromatid breaks, chromosome breaks, and chromosome radial structures compared to total number of chromosomes.
Chromosomal translocation assay
Induced pluripotent stem cells were derived from primary Polq+/+ and Polq−/− MEFs according to standard Yamanaka protocol33. To perform the translocation assay, Polq+/+ and Polq−/− induced pluripotent stem cells were transfected with 2 μg of Cas9-gRNA(Rosa26;H3f3b) plasmid per million cell. We constructed Cas9-gRNA(Rosa26;H3f3b) by introducing two guide RNAs (5′-GTTGGCTCGCCGGATACGGG-3′ for H3f3b; 5′-ACTCCAGTCTTTCTAGAAGA-3′ for Rosa26) into pX330 (Addgene, 42230). After transfection, 1 × 104 cells were seeded per well in a 96-well plate, and lysed 3 days later in 40 μl lysis buffer (10 mM Tris, pH 8.0, 0.45% Nonidet P-40 and 0.45% Tween 20). The lysate was incubated with 200 μg ml−1 proteinase K for 2 h at 55 °C. Translocation detection was performed according to previously established protocol9, using nested PCR. The primers used for the first PCR reaction were Tr6-11-Fwd: 5′-GCGGGAGAAATGGATATGAA-3′ and Tr6-11-Rev: 5′- TTGACGCCTTCCTTCTTCTG-3′ for der(6), and Tr11-6-Fwd: 5′-AACCTTTGAAAAAGCCCACA-3′ and Tr11-6-Rev: 5′-GCACGTTTCCGACTTGAGTT-3′, for der(11). For the second round of PCR amplification we used the primers Tr6-11NFwd: 5′-GGCGGATCACAAGCAATAAT-3′ and Tr6-11NRev: 5′-CTGCCATTCCAGAGATTGGT-3′, and Tr11-6NFwd: 5′-AGCCACAGTGCTCACATCAC-3′ and Tr11-6NRev: 5′TCCCAAAGTCGCTCTGAGTT-3′. The number of PCR-positive wells was used to calculate the translocation frequency as previously described9. Amplified products from positive wells were sequenced to verify translocations and determine the junction sequences.
Surveyor assay
Forty-eight hours after transfection, genomic DNA was extracted with GE Healthcare Illustra Genomic Prep Mini Spin Kit (28-9042-76). The genomic region encompassing the guide RNA target sites was amplified using Q5 High-Fidelity DNA polymerase (New England BioLabs) with the primers Rosa26-Fwd: 5′-TAAAACTCGGGTGAGCATGT-3′ and Rosa26-Rev: 5′-GGAGTTCTCTGCTGCCTCCTG-3′, and H3f3b-Fwd: 5′-GCGGCGGCTTGATTGCTCCAG-3′ and H3f3b-Rev: 5′-AGCAACTTGTCACTCCTGAGCCAC-3′. PCR fragments were gel purified and the surveyor assay was performed using a detection kit (Transgenomic), according to manufacturer’s instructions. Agarose gels (2%) were used to visualize the bands after surveyor digestion.
Colony formation assay
After lentiviral transduction with shCtrl or (sequences listed below), cells were selected with puromycin (BJ: 0.5 μg ml−1; MCF7 and HCC1937: 1 μg ml−1; MEFs: 2 μg ml−1) for 72 h and plated in 6-cm dishes (1,000 and 10,000 cells per plate). After 10–14 days, colonies were fixed with 3% paraformaldehyde (5 min), rinsed with PBS, and stained with crystal violet (Sigma-Aldrich).
CRISPR targeting to mutate Polq gene in mouse embryonic stem cells
To generate cells carrying a catalytic dead Polθ, two mutations at residues Asp2494Gly and Glu2495Ser (ref. 34) were introduced in the endogenous Polq locus in mouse embryonic stem cells using CRISPR/Cas9 gene targeting. Two guides RNAs were co-transfected with a Cas9-nickase (pX335-U6-Chimeric_BB-CBh-hSpCas9n(D10A)), and a donor cassette that introduces a SacII restriction site while replacing the two amino acid residues. Clonal cells lines were derived and genotyped to determine successful targeting. Two independent clonally derived lines were used for the analysis of translocation.
shRNA target sequence (pLK0.1 vector)
shPolm: 5′-CTCACCTCTCACACACCATAA-3′; shPolk: 5′-GCCCTTAGAAATGTCTCATAA-3′; shPolh: 5′-GCTCGATTCTCCAGCTTACAA-3′; shPoli: 5′-AGTGAAGAAGATACGTTTAAA-3′; shPolb: 5′-CCAAAGTTGTTACATCGTGTT-3′; shPoln: 5′-CCTACTCACATGAAGGACATT-3′; shLig3 (mouse): 5′-CCAGACTTCAAACGTCTCAAA-3′; shLIG3 (human): 5′-CCGGATCATGTTCTCAGAAAT-3′; shPolq-1 (mouse): 5′-CGGCGGAGTATGAGAACTATT-3′; shPolq-2(mouse): 5′-CCAGGAATCAAAGACGACAAT-3′; shPolq-3(mouse): 5′-CCTGGCTGAATGCTGAACTTT-3′; shPOLQ (human): 5′-CGGGCCTCTTTAGATATAAAT-3′; shBrca1(mouse): 5′-GCTCAGTGTATGACTCAGTTT-3′.
Primers for quantitative PCR
BRCA1Fwd: 5′-CTGCCGTCCAAATTCAAGAAGT-3′ and BRCA1Rev: 5′-CTTGTGCTTCCCTGTAGGCT-3′; BRCA2Fwd: 5′-TGTGGTAGATGTTGCTAGTCCGCC-3′ and BRCA2Rev: 5′-GCTTTTCTCGTTGTAGTACTGCC-3′; POLbFwd: 5′-TGAACCATCATCAACGAATTGGG-3′ and POLbRev: 5′-CCATGTCTCCACTCGACTCTG-3′; POLmFwd: 5′-AGGCTTCCGCGTCCTAGAT-3′ and POLmRev: 5′-GTGGGGAGAGCATCCATGTT-3′; POLkFwd: 5′-AGCTCAAATTACCAGCCAGCA-3′ and POLkRev: 5′-GGTTGTCCCTCATTTCCACAG-3′; POLhFwd: 5′-ATCGAGTGGTTGCTCTTGTAGA-3′ and POLhRev: 5′-CCAAATGCTCGGGCTTCATAG-3′; POLiFwd: 5′-GCAGTCAAGGGCCACCTAC-3′ and POLiRev: 5′-AGGTCTGTCCTTTAATTCTGGGT-3′; POLnFwd: 5′-AGCTGATGGATGCTCTCAAGCAGG-3′ and POLnRev: 5′-GAGTCAGAGTGCTGTTGCCTACATGG-3′; LIG3(mouse)Fwd: 5′-GAAGAAAGCTGCTGTCCAGG-3′ and LIG3(mouse)Rev: 5′-CAGAGTTGTTGGGTTTTGCTG-3′; LIG3(human)Fwd: 5′-GAAGAGCTGGAAGATAATGAGAAGG-3′ and LIG3(human)Rev: 5′-AGTGGTTGTCAACTTAGCCTGG-3′; POLQ(mouse)Fwd: 5′-CAAGGTTTCATTCGGGTCTTGG-3′ and POLQ (mouse)Rev: 5′-CGAGCAGGAAGATTCACTCCAG-3′; POLQ(human) Fwd: 5′-CAGCCCTTATAGTGGAAGAAGC-3′ and POLQ(human) Rev: 5′-GCACATGGATTCCATTGCACTC-3′.
Statistical analysis
Results are presented as mean ± s.d. of two or three independent experiments unless otherwise stated. P < 0.05 was considered statistically significant, and P values were calculated using the two-tailed Student’s t-test. No statistical methods were used to predetermine sample size.
Accession codes
Primary accessions
BioProject
Data deposits
Sequence has been deposited with the BioProject database under accession PRJNA269507.
References
Capper, R. et al. The nature of telomere fusion and a definition of the critical telomere length in human cells. Genes Dev. 21, 2495–2508 (2007)
Sfeir, A. & de Lange, T. Removal of shelterin reveals the telomere end-protection problem. Science 336, 593–597 (2012)
Rai, R. et al. The function of classical and alternative non-homologous end-joining pathways in the fusion of dysfunctional telomeres. EMBO J. 29, 2598–2610 (2010)
Sfeir, A. Telomeres at a glance. J. Cell Sci. 125, 4173–4178 (2012)
van Steensel, B., Smogorzewska, A. & de Lange, T. TRF2 protects human telomeres from end-to-end fusions. Cell 92, 401–413 (1998)
Wang, H. et al. DNA ligase III as a candidate component of backup pathways of nonhomologous end joining. Cancer Res. 65, 4020–4030 (2005)
Audebert, M., Salles, B., Weinfeld, M. & Calsou, P. Involvement of polynucleotide kinase in a poly(ADP-ribose) polymerase-1-dependent DNA double-strand breaks rejoining pathway. J. Mol. Biol. 356, 257–265 (2006)
Yan, C. T. et al. IgH class switching and translocations use a robust non-classical end-joining pathway. Nature 449, 478–482 (2007)
Simsek, D. et al. DNA ligase III promotes alternative nonhomologous end-joining during chromosomal translocation formation. PLoS Genet. 7, e1002080 (2011)
Simsek, D. & Jasin, M. Alternative end-joining is suppressed by the canonical NHEJ component Xrcc4-ligase IV during chromosomal translocation formation. Nature Struct. Mol. Biol. 17, 410–416 (2010)
Okamoto, K. et al. A two-step mechanism for TRF2-mediated chromosome-end protection. Nature 494, 502–505 (2013)
Arana, M. E., Seki, M., Wood, R. D., Rogozin, I. B. & Kunkel, T. A. Low-fidelity DNA synthesis by human DNA polymerase theta. Nucleic Acids Res. 36, 3847–3856 (2008)
Hogg, M., Sauer-Eriksson, A. E. & Johansson, E. Promiscuous DNA synthesis by human DNA polymerase θ. Nucleic Acids Res. 40, 2611–2622 (2012)
Chan, S. H., Yu, A. M. & McVey, M. Dual roles for DNA polymerase theta in alternative end-joining repair of double-strand breaks in Drosophila. PLoS Genet. 6, e1001005 (2010)
Roerink, S. F., van Schendel, R. & Tijsterman, M. Polymerase theta-mediated end joining of replication-associated DNA breaks in C. elegans. Genome Res. 24, 954–962 (2014)
Koole, W. et al. A Polymerase Theta-dependent repair pathway suppresses extensive genomic instability at endogenous G4 DNA sites. Nature Commun. 5, 3216 (2014)
Shima, N. et al. Phenotype-based identification of mouse chromosome instability mutants. Genetics 163, 1031–1040 (2003)
Tang, J. et al. Acetylation limits 53BP1 association with damaged chromatin to promote homologous recombination. Nature Struct. Mol. Biol. 20, 317–325 (2013)
Wang, M. et al. PARP-1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nucleic Acids Res. 34, 6170–6182 (2006)
Truong, L. N. et al. Microhomology-mediated End Joining and Homologous Recombination share the initial end resection step to repair DNA double-strand breaks in mammalian cells. Proc. Natl Acad. Sci. USA 110, 7720–7725 (2013)
Bailey, S. M., Cornforth, M. N., Kurimasa, A., Chen, D. J. & Goodwin, E. H. Strand-specific postreplicative processing of mammalian telomeres. Science 293, 2462–2465 (2001)
Certo, M. T. et al. Tracking genome engineering outcome at individual DNA breakpoints. Nature Methods 8, 671–676 (2011)
Bindra, R. S., Goglia, A. G., Jasin, M. & Powell, S. N. Development of an assay to measure mutagenic non-homologous end-joining repair activity in mammalian cells. Nucleic Acids Res. 41, e115 (2013)
Sakai, W. et al. Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers. Nature 451, 1116–1120 (2008)
Fernandez-Vidal, A. et al. A role for DNA polymerase θ in the timing of DNA replication. Nature Commun. 5, 4285 (2014)
Deng, S. K., Gibb, B., de Almeida, M. J., Greene, E. C. & Symington, L. S. RPA antagonizes microhomology-mediated repair of DNA double-strand breaks. Nature Struct. Mol. Biol. 21, 405–412 (2014)
Chen, H., Lisby, M. & Symington, L. S. RPA coordinates DNA end resection and prevents formation of DNA hairpins. Mol. Cell 50, 589–600 (2013)
Kawamura, K. et al. DNA polymerase theta is preferentially expressed in lymphoid tissues and upregulated in human cancers. Int. J. Cancer 109, 9–16 (2004)
Higgins, G. S. et al. Overexpression of POLQ confers a poor prognosis in early breast cancer patients. Oncotarget 1, 175–184 (2010)
Lemée, F. et al. DNA polymerase theta up-regulation is associated with poor survival in breast cancer, perturbs DNA replication, and promotes genetic instability. Proc. Natl Acad. Sci. USA 107, 13390–13395 (2010)
Shee, C. et al. Engineered proteins detect spontaneous DNA breakage in human and bacterial cells. Elife 2, e01222 (2013)
Leung, J. W. et al. Nucleosome acidic patch promotes RNF168- and RING1B/BMI1-dependent H2AX and H2A ubiquitination and DNA damage signaling. PLoS Genet. 10, e1004178 (2014)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Prasad, R. et al. Human DNA polymerase θ possesses 5′-dRP lyase activity and functions in single-nucleotide base excision repair in vitro. Nucleic Acids Res. 37, 1868–1877 (2009)
Celli, G. B. & de Lange, T. DNA processing is not required for ATM-mediated telomere damage response after TRF2 deletion. Nature Cell Biol. 7, 712–718 (2005)
Sfeir, A., Kabir, S., van Overbeek, M., Celli, G. B. & de Lange, T. Loss of Rap1 induces telomere recombination in the absence of NHEJ or a DNA damage signal. Science 327, 1657–1661 (2010)
Acknowledgements
We thank T. de Lange, R. Greenberg, J. Shay, N. Shima, C. Cazaux and R. Wood for providing key reagents for this study. We are grateful to M. Ji, L. Walton Masters, A. Phillips, A. Pinzaru, F. Yeung, P. Tonzi and J. Wong for technical assistance. We thank S. Kabir and F. Lottersberger for critical reading of the manuscript. This work was supported by a grant from the Breast Cancer Alliance (A.S.), V-foundation (A.S.), Department of Defense Breast Cancer Research Program BC134020 (P.A.M.-G.), Pew-Stewart Scholars Award (A.S.), Pew Scholars Award (E.L.-D.), Novartis Advanced Discovery Institute (E.L.-D.), and a grant from the National Institutes of Health (NIH) AG038677 (E.L.-D.). The A.S. laboratory was supported by start-up funds from the Helen L. and Martin S. Kimmel Center for Stem Cell Biology. The K.M.M. laboratory was supported in part by start-up funds from the University of Texas at Austin and from the Cancer Prevention Research Institute of Texas (CPRIT, R116). K.M.M. is a CPRIT scholar.
Author information
Authors and Affiliations
Contributions
A.S., E.L.-D. and P.A.M.-G. conceived the experimental design. P.A.M.-G. and A.S. performed the experiments and analysed the data. E.L.-D. and N.N. performed telomere-sequencing experiments. F.G. and K.M.M. performed experiments related to Polθ localization at DNA breaks. A.S. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Polθ promotes alt-NHEJ repair at dysfunctional telomeres.
(Related to Fig. 1.) a, Immunoblots for TRF1 and RAP1 after 4-OHT-induced depletion of TRF2 from Trf2F/FCre-ERT2 MEFs and co-depletion of TRF1 and TRF2 from Trf1F/F Trf2F/F Ku80−/− Cre-ERT2 cells. Loss of TRF2 is confirmed by the disappearance of RAP1; a TRF2-interacting protein the stability of which depends on TRF2 (refs 35, 36). b, To validate the effect of Polq depletion on alt-NHEJ we monitored the frequency of telomere fusions in shelterin-free Ku80-null cells treated with three independent shPolq vectors. shPolq-1 was used in Fig. 2. Mean values are presented with error bars denoting ± s.e.m. from two independent experiments.
Extended Data Figure 2 Polθ drives chromosomal translocations in mouse cells.
(Related to Fig. 2.) a, Immunobloting for Polθ in MEFs with the indicated genotype and treatment. b, Immunoblot for TRF1 in MEFs with the indicated genotype. Cells were analysed 96 h after Cre induction. c, RAP1 immunoblot (similar to b). d, Western blot analysis for Polθ and Flag–Cas9 in lysates prepared from Polq−/− and Polq+/+ cells after Cas9 expression. Tubulin serves as a loading control. e, Surveyor nuclease assay for Polq−/− and Polq+/+ cells expressing Cas9-gRNA(Rosa26;H3f3b) plasmid. Genomic DNA isolated from cells with the indicated genotype was used as a template to amplify across the cleavage site at either the Rosa26 or the H3f3b locus to assess intra-chromosomal NHEJ. Amplification products were denatured and then re-annealed to form heteroduplexes between unmodified and modified sequences from imprecise NHEJ. The mismatched duplex was selectively cleaved by the Surveyor nuclease at the loops that form at mismatches. f, Signature of translocations in Polq−/− and Polq+/+ cells (see Extended Data Figs 3–4, 5 for complete list of sequences). Table records the total number of translocation events identified following CRISPR-Cas9 induced-cleavage. On average, the same number of nucleotides was deleted at the fusion junction in Polq−/− and Polq+/+ cells. No nucleotide insertions were found in the absence of Polq. Lastly, the percentage of junctions exhibiting microhomology was significantly reduced in cells lacking Polq. g, Scheme depicting Polθ domains. CRISPR/Cas9 gene targeting was used to create two base substitutions at Asp2494Gly and Glu2495Ser, and generate a catalytic-dead polymerase34. h, Sequence analysis of targeted cells. i, Genotyping PCRs of Polq+/+ and PolqCD (catalytically dead allele of Polq) after SacII digestion. j, Immunoblotting to analyse Cas9 expression in Polq+/+ and two independently derived PolqCD clonal cell lines. k, Frequency of chromosomal translocations (der(6)) in Polq+/+ and PolqCD cells. Bars represent mean of four independent experiments ± s.d. (two experiments per clonal cell line). P = 0.006; two-tailed Student’s t-test. PCR products were sequenced to confirm translocation and identify possible insertions.
Extended Data Figure 3 Sequence analysis of translocation junctions in Polq+/+ cells.
(Related to Fig. 2.) Sequences of der(11) breakpoint junction from Polq+/+ cells. Predicted fusion breakpoint based on CRISPR cutting indicated by an arrow. Reference sequence highlighted at the top. The remaining lines represent individual translocations recovered by PCR and subject to Sanger sequencing. Nucleotide insertions are marked in red. In cases where insertions extended beyond the sequence included in the lane, the length of the insertion was noted in parenthesis (red). Gaps in the sequence represent nucleotide deletions. The average length of the deletions was noted in Extended Data Fig. 2f. Micro-homology is denoted by blue boxes. Micro-homology embedded in DNA extending beyond the enclosed sequence was noted in parentheses (blue).
Extended Data Figure 4 Sequence analysis of translocation junctions in Polq+/+ cells.
(Related to Fig. 2.) Sequences of der(6) breakpoint junction from Polq+/+ cells. Predicted fusion breakpoint based on CRISPR cutting indicated by an arrow. Reference sequence highlighted at the top. The remaining lines represent individual translocations recovered by PCR and subject to Sanger sequencing.
Extended Data Figure 5 Sequence analysis of translocation junctions in Polq−/− cells.
(Related to Fig. 2.) Sequences of der(11) and der(6) breakpoint junction from Polq−/− cells. Predicted fusion breakpoint based on CRISPR cutting indicated by an arrow. Reference sequence is highlighted at the top. The remaining lines represent individual translocations recovered by PCR and subject to Sanger sequencing. It is important to note that insertions were completely lacking at the fusions junctions in Polq−/− cells.
Extended Data Figure 6 Polθ recruitment to DNA breaks.
(Related to Fig. 3.) a, Laser micro-irradiation experiment using HeLa cells expressing Myc–Polθ and treated with ATM inhibitor (KU55933), ATR inhibitor (VE-821) or PARP inhibitor (KU58948). b, Western blot analysis for CHK1 and CHK2 phosphorylation. Cells with the indicated treatment were analysed 2 h after irradiation. c, Immunoblot for PARP1. HeLa cells were treated with PARP1 siRNA and analysed 72 h after siRNA transfection for efficiency of knockdown.
Extended Data Figure 7 PARP1-dependent Polθ recruitment to DNA double-stranded breaks (DSBs).
(Related to Fig. 3.) a, Results from immunofluorescence performed 4 h after induction (1 µM Shield1 ligand, Clontech 631037; 0.5 μM 4-OH tamoxifen) of DSBs by mCherry-LacI-FokI in the U2OS-DSB reporter cells18 transfected with the Myc–Polθ and treated with PARP inhibitor (KU58948). The mCherry signal is used to identify the area of damage and to assess the recruitment of Myc–Polθ to cleaved LacO repeats. b, Table displaying quantification related to a.
Extended Data Figure 8 Polθ suppresses homology-directed repair at dysfunctional telomeres.
(Related to Fig. 3.) a, Western blot analysis for Polθ and LIG3 in shelterin-free Lig4-null MEFs. b, Western blot for TRF1 and RAP1 after 4-OHT treatment of shelterin-free Lig4-deficient cells. c, Metaphase spreads from Trf1F/F Trf2F/F Lig4−/− Cre-ERT2 MEFs, with the indicated shRNA treatment, 96 h after Cre expression. CO-FISH assay was performed using a FITC-OO-(CCCTAA)3 PNA probe (green) and a Tamra-OO-(TTAGGG)3 PNA probe (red). DAPI in blue. Examples of alt-NHEJ-mediated fusion and T-SCE events (HDR) are indicated by white and red arrows, respectively. Examples of T-SCE events reflective of increased HDR in cells treated with shPolq are on the right. d, e, Quantification of telomere fusions by alt-NHEJ in MEFs with the indicated genotype and shRNA treatment. Bars represent mean of two independent experiments ± s.e.m. f, Representative in-gel hybridization to assess 3′ overhang of Trf1F/F Trf2F/F Lig4−/− Cre-ERT2 MEFs with the indicated shRNA treatment after Cre deletion. g, Quantification of the gel in f. The single-stranded DNA/total signal ratios of the ‘+Cre’ samples are expressed relative to the ‘−Cre’ samples for each shRNA treatment. Mean of two independent experiments. h, Graph representing RAD51 accumulation after ionizing radiation treatment of PolqCD, Polq+/+ and Polq−/− embryonic stem cells. Bars represent mean of two independent experiments. P >0.05; two-tailed Student’s t-test.
Extended Data Figure 9 Polθ promotes alt-NHEJ and inhibits homology-directed repair at I-SceI-induced DNA breaks.
(Related to Fig. 3.) a, Polθ represses recombination at DSBs induced by I-Sce1. The TLR system was used to measure the relative ratio of end-joining (mCherry) and HDR (enhanced green fluorescent protein (eGFP)) repair of a DSB. A diagram of the TLR is represented. b, The TLR construct was stably integrated into Lig4−/ and Ku80−/− MEFs to avoid the confounding effect of C-NHEJ, and limit end-joining reactions to the alt-NHEJ pathway. Expression of mCherry and eGFP was assessed by flow cytometry 72 h after I-Sce1 and 5′ eGFP donor transduction in cells with the indicated shRNA construct. Percentages of cells are indicated in the plot. c, Quantification of alt-NHEJ and HDR of TLR containing Ku80−/−MEFs after expression of I-Sce1 and 5′ eGFP together with the indicated shRNA construct. Bar graphs represent the mean of three independent experiments ± s.d. P = 0.03; two-tailed Student’s t-test. d, Real-time PCR to monitor the knockdown efficiency of Polq in Ku80−/− and Lig4−/− MEFs. The FACS analysis reported in e and f was carried out without selecting for cells expressing the shRNA-containing plasmid.
Extended Data Figure 10 Polθ is required for survival of recombination-deficient cells.
(Related to Fig. 4.) a, Accumulation of chromosomal aberrancies after Brca1 and Brca2 knockdown in Polq−/− and Polq+/+ MEFs. Quantification of chromosomal aberrancies (chromatid breaks, chromosome breaks and radials) in MEFs stably transduced with lentiviral vectors expressing the indicated shRNA. b, Real-time PCR to confirm the knockdown of Brca1 and Brca2 as in a. c, Quantitative analysis of colony formation in Brca1F/F Cre-ERT2 and Lig4−/− cells after Polq depletion. The number of colonies in control shRNA-treated cells was set to 100%. Mean values are presented with error bars denoting ± s.d. from three independent experiments. d, Real-time PCR to measure the knockdown efficiency of human POLQ in BJ-hTERT, MCF7 and HCC1937 cells and mouse Polq in Brca1F/F Cre-ERT2 cells. e, Quantitative analyses of colony formation in BJ-hTERT, MCF7 and HCC1937 cells after LIG3 inhibition. The number of colonies in control-shRNA-treated cells was set to 100%. The knockdown efficiency for Lig3 was ∼85%. Bars represent mean of two independent experiments ± s.e.m. f. Quantitative analyses of colony formation in PolqCD and Polq+/+ embryonic stem cells after BRCA1 inhibition. The number of colonies in control-shRNA-treated cells was set to 100%. The knockdown efficiency for BRCA1 was >80%. Bars represent mean of two independent experiments ± s.e.m. P = 0.05; two-tailed Student’s t-test.
Supplementary information
Supplementary Information
This file contains Supplementary Data including sequence analysis of telomere fusions using illumina technology and C‐NHEJ junction sequences. (PDF 233 kb)
Source data
Rights and permissions
About this article
Cite this article
Mateos-Gomez, P., Gong, F., Nair, N. et al. Mammalian polymerase θ promotes alternative NHEJ and suppresses recombination. Nature 518, 254–257 (2015). https://doi.org/10.1038/nature14157
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14157
- Springer Nature Limited
This article is cited by
-
POLQ inhibition attenuates the stemness and ferroptosis resistance in gastric cancer cells via downregulation of dihydroorotate dehydrogenase
Cell Death & Disease (2024)
-
The ALT pathway generates telomere fusions that can be detected in the blood of cancer patients
Nature Communications (2024)
-
CATI: an efficient gene integration method for rodent and primate embryos by MMEJ suppression
Genome Biology (2023)
-
ZSCAN4 interacts with PARP1 to promote DNA repair in mouse embryonic stem cells
Cell & Bioscience (2023)
-
Clinical and translational advances in ovarian cancer therapy
Nature Cancer (2023)