Abstract
A deeper understanding of the mechanisms that underlie aberrant signal transduction in B-cell cancers such as chronic lymphocytic leukemia (CLL) may reveal new treatment strategies. The lipid-activated nuclear receptor peroxisome proliferator-activated receptor delta (PPARδ) accounts for a number of properties of aggressive cancers and was found to enhance Janus kinase (JAK)-mediated phosphorylation of signal transducer and activator of transcription (STAT) proteins in B lymphoma cell lines and primary CLL cells. Autocrine production of cytokines such as IL10 and interferon-beta was not increased by PPARδ but signaling responses to these cytokines were amplified and associated with increased cholesterol biosynthesis and plasma membrane levels. Plasmalemmal cholesterol and STAT phosphorylation from type 1 interferons (IFNs) were increased by PPARδ agonists, transgenes and exogenous cholesterol, and decreased by cyclodextrin, PPARD deletion and chemical PPARδ inhibitors. Functional consequences of PPARδ-mediated perturbation of IFN signaling included impaired upregulation of co-stimulatory molecules. These observations suggest PPARδ modulates signaling processes in malignant B cells in part by altering cholesterol metabolism and changes the outcomes of signaling from cytokines such as IFNs. PPARδ antagonists may have therapeutic activity as anti-leukemic signal transduction modulators.
Similar content being viewed by others
Introduction
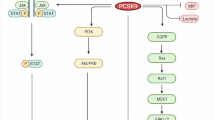
Peroxisome proliferator-activated receptor delta (PPARδ) is the least characterized member of the lipid-activated nuclear receptor family that includes PPARα, which regulates fatty acid oxidation, and PPARγ, which regulates adipocyte differentiation.1, 2 PPARδ changes conformation following activation by free fatty acids, bioactive lipids or synthetic agonists such as GW50516 and GW07422 and then mediates transcription of genes with peroxisome proliferator response elements in their promoters. PPARδ-regulated genes include PPARD itself and many that are involved in glucose, lipid and cholesterol metabolism.3, 4
There is increasing evidence that PPARδ confers aggressive properties on cancer cells.3, 5, 6, 7, 8, 9 We found that PPARδ allowed breast and B-cell cancer cell lines as well as primary chronic lymphocytic leukemia (CLL) cells to survive in harsh metabolic conditions containing cytotoxic drugs or low amounts of glucose and oxygen. PPARδ protected cancer cells by increasing their metabolic efficiency and ability to tolerate oxidative stress.3, 10
Another characteristic of aggressive cancers is aberrant signal transduction.11, 12 For example, transforming growth factor beta (TGFβ) is normally a tumor suppressor that causes growth arrest through SMAD proteins but it promotes tumor growth through p42/44 mitogen activated protein kinase (MAPK) in aggressive prostate cancer cells.11 The type 1 interferon (IFN)-pathway represents another example of corrupted signaling in aggressive cancer cells. IFNs signal through the interferon alpha receptor (IFNAR), composed of IFNAR1 and IFNAR2 subunits. Ligand binding causes these subunits to dimerize, resulting in phosphorylation and activation by Janus kinases (JAKs) of signal transducer and activator of transcription (STAT) proteins, especially STAT1. IFN-activated STAT1-mediated genes include co-stimulatory molecules such as CD80, CD86 and class I major histocompatibility complex genes that increase the immunogenicity of cancer cells and support anti-tumor immunity.12 However, IFN signaling is altered in aggressive CLL cells to cause prolonged expression of the immunosuppressive transcription factor tyrosine-phosphorylated STAT3 (pSTAT3).12
In the course of studying PPARδ in cancer cells,3, 10 we noted that increased pSTAT3 expression often accompanied increased PPARδ activity. The experiments in this manuscript were designed to try to understand this phenomenon by using model systems that included Daudi lymphoma cells and primary CLL cells transduced with PPARD expression vectors or treated with PPARδ agonists. The results indicate PPARδ modulates signaling processes in malignant B cells in part by altering cholesterol metabolism.
Materials and methods
Antibodies and reagents
Fluorescent CD80, CD83, CD86, TNFα and HLA-ABC antibodies were obtained from BD Biosciences (Bedford, MA, USA). Human IL10 and IL6 along with fluorescent IL10 receptor antibodies and IL10-blocking antibodies were from eBioscience (San Diego, CA, USA). Nile Red, resiquimod, Lovastatin, cyclodextrin, 2-mercaptoethanol, soluble cholesterol and β-actin antibodies were from Sigma-Aldrich (St Louis, MO, USA). Ruxolitinib was from SelleckChem (Houston, TX, USA). The IL6 receptor-blocking antibody Actemra (Roche Canada, Mississauga, ON, Canada), human IFNα2b (Schering Canada Inc., Pointe-Claire, QC, Canada), IL2 (Chiron, Corp., San Francisco, CA, USA), fludarabine sulfate (Berlex Canada Inc., Pointe-Claire, QC, Canada) and vincristine sulfate (Faulding Canada Inc., Kirkland, Quebec) were purchased from the hospital pharmacy. GW0742 and GW501516 (PPARδ agonists) and human anti-PPARδ antibodies (101720) were from Cayman Chemical (Ann Arbor, MI, USA). DG172 and NXT1551 (PPARδ antagonists) were described before3 and provided by Wibke Diederich (Marburg University, Germany) and Peppi Prasit (Inception, San Diego, CA, USA), respectively. Anti-IgM was from MP Biomedicals (Solon, OH, USA). Murine IFNβ and IFNAR antibodies were from PBL Assay Science (Piscataway Township, NJ, USA). 7-aminoactinomycin D (7AAD) and IL10 ELISA kits were from Biolegend (San Diego, CA, USA). LysoTracker Green DND-26 and antibodies to total and phospho-STAT3(Y705) (Cat. No. 9131), phospho-STAT1(Y701) (Cat. No. 9172), phospho-p44/42 MAPK(Erk1/2) (Thr 202/Tyr 204) (Cat. No. 9102), phospho-AKT (Thr 308) (Cat. No. 9275), phospho-AKT (Ser 473) (Cat. No. 9271), phospho-SMAD2(Ser465/467) (Cat. No. 3108) as well as anti-rabbit and anti-mouse IgG antibodies (Cat.7074 and 7076) were from Cell Signaling Technology (Beverly, MA, USA). Hexahistidine-tagged HIS6-GFP-D4 fusion proteins (HIS6X-GFP-D4) were produced as described previously.13 The C-terminal domain of perfringolysin O (PFO), a cytolysin from Clostridium perfringens is called domain 4 (D4) and binds exofacial cholesterol in the plasma membrane. RPMI-1640 cell culture media was from Wisent Bioproducts (St Bruno, QC, Canada). AIM-V serum-free media was from Thermofisher Scientific (Mississauga, ON, Canada).
CLL cells
CLL cells were isolated as before from the blood of consenting patients attending the Sunnybrook CLL clinic.12 Cells were used within 4 h of isolation for all experiments. Patients were untreated for CLL for at least 6 months prior to blood collection. Patient characteristics are described in Supplementary table 1. Protocols were approved by the Sunnybrook Research Ethics Board (PIN 222-2014).
Cell lines
Vector control Daudi cells1 and PPARDhi Daudi cells were cultured in RPMI-1640 supplemented with 5% fetal bovine serum (FBS, Multicell, Toronto, ON, Canada) and 1 × penicillin–streptomycin (Multicell) in a humidified atmosphere containing 5% CO2 at 37 °C. MCF-7 and SKBR-3 cells were described before.3 Cells were cultured in DMEM media (Multicell) supplemented with 5% FBS and 1 × penicillin–streptomycin in a humidified atmosphere containing 5% CO2 at 37 °C.
Mouse spleen cell preparation
Spleens from PPARD−/−10, 14 and wild-type C57BL/6J mice were homogenized and filtered through a 40 μm cell strainer (BD Falcon, BD Biosciences). Spleen mononuclear cells were obtained by centrifugation over lympholyte-M (Cedarlane, Hornsby, ON, Canada) according to the manufacturer’s instructions. B cells were isolated by negative selection (RosetteSep mouse B Cell Enrichment Cocktail, StemCell Technologies, Vancouver, BC, Canada) using the manufacturer’s instructions.
Flow cytometric analysis of cell viability, intracellular lipid droplets, plasma membrane cholesterol and lysosomes
Viable cells were indicated by 7AAD exclusion as before.12 Nile Red reflects lipid droplets.15 HIS6X-GFP-D4 (also called ‘PFO’ in the manuscript) was used to measure plasma membrane cholesterol.13 One million cells were stained with 3 μl 7AAD for 10 min, 3 μM Nile Red for 20 min or 15 μg/ml PFO for 15 min in serum-free RPMI medium at room temperature, washed and resuspended with ice-cold PBS. Lysosome content was indicated by staining cells with 100 nM LysoTracker Green at 37 °C for 30 min. Cells were analyzed in a FACS Calibur (BD Bioscience) flow cytometer using CellQuest software (BD Bioscience). Data were analyzed with FlowJo software (Ashland, OR, USA). At least 10 000 events were collected for each experiment.
Retroviral and lentiviral infections
Human PPARD full-length complimentary DNA was obtained from Addgene (Cambridge, MA, USA) and sub-cloned into the XhoI and EcoRI sites of retroviral MSCV2.2 plasmids or into the XhoI and NotI sites of lentiviral pLemiR plasmids. Sequences of the constructs were confirmed before transfection. Replication-defective viruses were prepared by transfecting the MSCV-PPARD viral plasmid into the helper-free packaging cell line GP+A (B8), as described before.3 CD5+ Daudi cells at 2 × 106 cells/ml were infected with supernatants from the virus-producing cells. Stably transfected clones were obtained by limiting dilution and selection in G418 (Multicell). Transfection was performed with Lipofectamine 3000 according to the manufacturer's protocol (Invitrogen, Carlsbad, CA, USA). Cells infected with retroviruses containing the empty vectors but otherwise handled in the same way were used as controls and referred to in the text as ‘Vector control cells’.
To make lentiviruses (Lvs), 8 × 105 HEK293T cells were seeded into 6-well plates and transfected 24 h later with plemiR-PPARδ plasmids (1 μg) and package plasmids (0.8 μg 8.2VPR vector and 0.2 μg VSVG vector) using Lipofectamine 3000 according to the manufacturer's instructions. After 24 h, the media was replaced with 2 ml fresh media. After 48 h, the supernatants containing Lv particles were collected and used to infect CLL cells. Control cells were also made with the empty plasmids.
Lv titers were measured by infecting HEK293T cells with serial dilutions of the Lv-containing supernatants and counting infected, red fluorescent HEK293T cell colonies after 1 week of puromycin selection. Lv particles (~20–200 × 107) were mixed with 1 × 107 CLL cells in six-well plates with 2 ml AIM-V medium and 5 μg/ml polybrene, which was replaced with fresh AIM-V 24 h later. Whole cultures of infected CLL cells were used for further experiments after 72 h if more than 50% of the cells exhibited red fluorescence by fluorescence microscopy.
Immunoblotting
Protein extraction and immunoblotting were performed as before.12 Proteins were resolved in 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis and transferred to Immobilon-P transfer membranes (Millipore Corp., Billerica, MA, USA). Western blot analysis was performed according to the manufacturers’ protocol for each antibody. Chemiluminescent signals were created with SupersignalWest Pico Luminal Enhancer and Stable Peroxide Solution (Pierce, Rockford, IL, USA) and detected with a Syngene InGenius system (Syngene, Cambridge, UK). For additional signal, blots were stripped for 15 min at room temperature in Restore Western Blot stripping buffer (Pierce), washed twice in Tris-buffered saline plus 0.05% Tween-20 at room temperature, and reprobed as required. Densitometry was performed using Image J software (NIH). The densitometry value for each sample was normalized against the value for β-actin to obtain the intensities for pSTAT3, STAT1 and AKT reported in the figures.
Isolation of genomic DNA
Control and lentivirus-infected CLL cells were washed in PBS, suspended in 500 μl lysis buffer (10 mM Tris-HCL pH 7.5, 400 mM NaCl, 2 mM EDTA pH 8.0), and incubated overnight at 50 °C after adding 20% SDS (20 μl) and proteinase K (40 μl) (Roche Diagnostics GmbH, Mannheim, Germany). One volume of phenol:chloroform:isoamyl alcohol (25:24:1) was added to each sample, which was then centrifuged at room temperature for 10 min at 14 000 g. The upper aqueous phase was transferred to a fresh tube and the process repeated two times and then twice with chloroform alone. Sodium chloride (5 M) (1/10 volume) and ethanol (2 × volume) were then added and the mixture was shaken gently for 10 min. The precipitate was obtained by centrifugation at room temperature for 10 min at 14 000 g, rinsed with 70% ethanol, dissolved in ultra-pure water and stored at 4 °C until used for PCR analysis. DNA concentrations were determined in a NanoDrop 2000 Spectrophotometer (Thermo Scientific, Wilmington, DE, USA) at 260/280 nm.
Real-time PCR
RNA was prepared with the RNeasy mini kit (Qiagen, Valencia, CA, USA). complimentary DNA was synthesized from 2 μg of RNA using Superscript III reverse transcriptase (Life Technologies, Invitrogen, Burlington, ON, Canada) according to the manufacturer’s instructions. RNA transcripts were amplified with the following primers:
PCRs were carried out in a DNA engine Opticon System (MJ Research, Waltham, MA, USA) and cycled 34 times after initial denaturation (95 °C, 15 min) with the following parameters: denaturation at 94 °C for 20 s; annealing of primers at 58 °C for 20 s and extension at 72 °C for 20 s. Abundance of mRNA transcripts was evaluated by a standard amplification curve relating initial copy number to cycle number. Copy numbers were determined from two independent complimentary DNA preparations for each sample. The final result was expressed as the relative fold change of the target gene to HPRT.
Conditions for amplifying 5'-LTR regions in genomic DNA after initial denaturation (94 °C, 3 min) were: 30 cycles with denaturation at 94 °C for 45 s, primer annealing at 55 °C for 30 s and extension at 72 °C for 90 s in a final volume of 100 μl with 10 ng genomic DNA template, 2.5 U of Taq DNA polymerase, 0.5 mM of each 5'-LTR primer, 1.5 mM MgCl2, 0.2 mM dNTP and 1 × PCR buffer minus Mg2+. When cycling was complete, the sample was mixed with dye and loaded onto a 2% agarose gel, electrophoresed and then photographed on a UV light box. The PCR product was 200 bp.
Cytokine measurements
A kit for human interleukin10 (IL10) was used according to the manufacturer's instructions (eBioscience). Concentrations were determined from standard curves. The assay was linear between 30 and 1000 pg/ml of IL10.
Cholesterol manipulations
Methyl-β-cyclodextrin is a water soluble structure that forms soluble inclusion complexes with cholesterol.16 To load cells with cholesterol, water soluble cholesterol packaged inside a methyl-β-cyclodextrin ring structure was added to purified CLL cells at a concentration of 15 μM. To strip cholesterol from cell membranes, empty methyl-β-cyclodextrin was added at a concentration of 0.5 mM.
Statistical analysis
Experimental cultures were set up in triplicate and each experiment was repeated 2–3 times. Data are presented as mean±s.e. unless otherwise indicated. Unpaired two-tailed student t-tests were used to determine P-values for differences between sample means with P-values <0.05 considered significant.
Results
PPARδ increases phospho-STAT3 levels in cancer cells
In the course of studying the biology of PPARδ in cancer,3, 10 increased levels of tyrosine-pSTAT3 proteins were observed to accompany enhanced activity of PPARδ (Figure 1). This relationship was seen in Daudi lymphoma cells stably transfected with a PPARD expression vector (Figure 1a), in primary CLL cells transduced with Lvs expressing PPARD (Figure 1b), and also in PPARD-transgenic MCF-7 and SKBR-3 breast cancer cells17 (Figures 1d and e), suggesting it was not a unique feature of B-cell cancers.
Effect of PPARδ on JAK-mediated tyrosine phosphorylation of STAT3 in cancer cells. Vector control and PPARDhi Daudi cells were plated in RPMI-1640+5% FBS at 2 × 106 cells/ml for 12 h (a) Primary CLL cells from three different patients were infected with lentiviruses that expressed PPARD or a control vector as described in the Materials and methods section and then cultured in AIM-V serum-free medium for 3–4 days (b) Ruxolitinib (Ruxo) (400 nM) was added to exponentially growing vector control and PPARDhi Daudi cells for 30 min. (c) Vector control and PPARDhi MCF-7 (d) and SK-BR-3 (e) breast cancer cells were cultured in DMEM+5% FBS medium at 1 × 106 cells/ml for 24 h. Ruxolitinib (400 nM) was added to some cultures for 30 min. Cells were then harvested and phospho-STAT3(tyr705), STAT3 and PPARδ levels were determined by immunoblotting with β-actin and also used as a loading control. The blots were quantified by densitometry relative to β-actin and shown in the graphs on the right. The lines in the graphs in b represent individual patient samples.**P<0.001; NS, no significant difference.
STAT3 is often phosphorylated by JAKs that have been activated by cytokines.12, 18 Consistent with a link between PPARδ and cytokine signaling, pSTAT3 expression was abolished in Daudi (Figure 1c) and breast cancer cells (Figures 1d and e) by the JAK inhibitor Ruxolitinib.18
PPARδ does not increase production of STAT3-activating cytokines by Daudi cells
An initial hypothesis to explain these observations was that PPARδ increased autocrine production of cytokines that phosphorylate STAT3 through JAKs. We have previously characterized type 1 IFN, IL10 and IL6 as cytokines that could be made by CLL cells and phosphorylate STAT3.1, 12, 19 The IL6 promoter containing peroxisome proliferator response elements and IL6 could potentially increase following overexpression of PPARD.20 However pSTAT3 levels in PPARDhi Daudi cells were not changed significantly by IL6-blocking antibodies (Figure 2a). In contrast, blocking either IL10 (Figure 2a) or the IFN receptor (IFNAR) (Figure 2e) lowered pSTAT3 levels, suggesting autocrine IL10 and type 1 IFN contributed to STAT3 phosphorylation in PPARDhi Daudi cells. However, no evidence could be found for increased IL10 or type 1 IFN production by these cells. IL10 (Figure 2b) and IFNB1 (Figure 2f) messenger RNA expression in PPARDhi cells and control Daudi cells were similar and PPARDhi cells actually made less IL10 as measured by enzyme-linked immunosorbent assay (Figure 2c).
Effect of IL10 and type 1 interferon on STAT3 phosphorylation in Daudi cells. Vector control and PPARDhi Daudi cells were cultured in RPMI-1640+5% FBS with or without blocking antibodies to IL10, IL6 (a) or IFN receptors (e) at 1 μg/ml. Cells were collected after 12 h and pSTAT3 levels determined by immunoblotting and quantified by densitometry using β-actin as a loading control. Examples of immunoblots are shown on the left with densitometry values on the right. The experiments were repeated three times with similar results. (b, f). After 24 h, PPARD and IL10 (b) or IFNB (f) messenger RNA (mRNA) levels were measured by real-time PCR. (c) IL10 levels in culture supernatants were measured by enzyme-linked immunosorbent assay after 48 h. (d) Vector control cells were cultured with or without the PPARδ agonist GW0742 (1 μM) and PPARDhi Daudi cells with or without the PPARδ antagonist DG172 (1 μM). After 24 h, IL10 receptor (IL10R) expression was measured by flow cytometry and shown as percentages of IL10R+ cells. (g) Purified CLL cells were cultured at 2 × 106 cells/ml in AIM-V for 2–3 days with or without GW0742. Some cultures were stimulated with IL6 (60 ng/ml), IL10 (10 ng/ml) or IFNα2b (1000 U/ml). Total and phospho-STAT3(tyr705) were quantified by immunoblotting after 30 min. An example is shown and average densitometry values and s.e. for three different patient samples are plotted in the graph on the right. *P<0.01; **P<0.05; NS, no significant difference.
Another potential explanation for increased pSTAT3 was that cytokine receptor expression was increased by PPARδ. However, IL10R levels measured by flow cytometry were similar in PPARDhi and vector control Daudi cells, and not increased when control cells were treated with the PPARδ agonist GW0742 or decreased when PPARDhi cells were treated with the PPARδ antagonist DG172 (Figure 2d). These observations suggested the possibility that cytokine signaling responses had been amplified by overexpression of PPARD. Consistent with this idea, STAT3 phosphorylation was increased after 30 min by IL10 and IFNα2b in CLL cells pre-treated with GW0742 (Figure 2g). Interestingly, IL6-induced pSTAT3 levels did not appear to be affected by increased PPARδ activity (Figure 2g).
PPARδ enhances IFN signaling in B-cell cancers
IFN signaling was chosen for more detailed study using a number of different models. First, control and PPARDhi Daudi cells were serum-starved to lower background signaling and then treated with IFNα2b. IFN signaling, indicated by the magnitude of pSTAT1 and pSTAT3 expression after 30 min and 4 h, was stronger in PPARDhi cells (Figure 3a).
Enhanced IFNAR signaling in PPARDhi Daudi and CLL cells. (a) Vector control and PPARDhi Daudi cells were serum-starved (SS) for 4 h, stimulated with IFNα2b (1000 U/ml) and protein extracts made at 0, 0.5 and 4 h. (b) Vector control Daudi cells were cultured for 2–3 days in RPMI-1640+5% FBS with or without GW0742 (1 μM), SS and stimulated with IFNα2b. Protein extracts were made 30 min later. (c) Primary CLL cells were infected with lentiviruses that expressed PPARD or a control vector and cultured in AIM-V medium for 3–4 days. Some cultures at 2 × 106 cells/ml were treated with IFNα2b (1000 U/ml) and protein extracts made after 30 min. (d) Purified CLL cells were cultured at 2 × 106 cells/ml in AIM-V for 2–3 days with or without GW0742 (1 μM). Some cultures were stimulated with IFNα2b and protein extracts made at 0, 0.5 and 4 h. Total and phospho-STAT1(tyr701) and phospho-STAT3(tyr705) along with PPARδ were then measured by immunoblotting and quantified by densitometry using β-actin as a loading control. Examples of blots are shown on the left and densitometry values are shown in graphs on the right. For a and b, the averages and s.e. of three and five separate experiments, respectively, are shown. For c and d, the lines represent results with four individual samples. *P<0.05; **P<0.025; ***P<0.01; NS, no significant difference.
Vector control Daudi cells were then treated for 2–3 days with the PPARδ agonist GW0742 to determine if non-genetic enhancement of PPARδ activity affected IFN signaling. Levels of pSTAT3 particularly increased following treatment with IFNα2B in cells with higher PPARδ activity (Figure 3b).
To determine if similar results occurred in primary B-cell tumors, CLL cells were transduced with PPARD-expressing Lvs3, 10 and maintained in serum-free AIM-V media for several days before stimulation with IFNα2B. Alternatively, CLL cells were treated with GW0742 for 2–3 days to increase PPARδ activity prior to stimulation. Consistent with the observations in Daudi cells, pSTAT3 and pSTAT1 levels increased in CLL cells with higher PPARδ activity (Figures 3c and d).
Since upregulation of PPARδ expression is associated with situations that cause cell death,3, 10 it is possible these results were not caused directly by PPARδ but were due to toxic effects of GW0742, spontaneous death of CLL cells in culture21 or killing by Lvs.22 Indeed, cytotoxic drugs like vincristine and fludarabine killed CLL cells and increased PPARD messenger RNA expression (Supplementary Figure 3B). However, spontaneous death of CLL cells is minimal in serum-free AIM-V media (Supplementary Figure 3C, top panel) in contrast to serum-containing media23, 24 and was not increased by GW0742 (Supplementary Figure 3C, bottom panel). More cells died following infection with Lvs but there were no significant differences with PPARD expression vectors or control vectors (Supplementary Figure 4A). Taken together, these observations suggest the signaling effects (Figures 2g and 3) were caused directly by PPARδ and not indirectly from toxicity.
PPARδ increases signaling responses to serum and B-cell receptor cross-linking
Responses to other signals were assessed to determine if the effects of PPARδ were restricted to cytokines. Control and PPARDhi Daudi cells were serum-starved for 4 h and exposed to 5% FBS, which contains many biological factors including TGFβ.25 After 4 h, levels of phospho-STAT3 and phospho-SMAD2, indicative of TGFβ-signaling,26 were higher in PPARDhi cells. However, constitutive activation of ERK was not affected significantly (Supplementary Figure 5A).
Antigenic signaling through the B-cell receptor is a central pathogenic process in CLL and involves phosphorylation of STAT3 and AKT.27, 28 Phosphorylation of AKT on threonine 308 was increased after 30 min following cross-linking with IgM antibodies in CLL cells treated with GW0742 for 2–3 days to increase PPARδ activity (Supplementary Figure 5B). Tyrosine phosphorylation of STAT3 was also higher at 4 h but not different at later times (Supplementary Figure 5C). Interestingly, phosphorylation of AKT on serine 473 was not increased in CLL cells with higher PPARδ activity (Supplementary Figure 5C). Taken together, the results suggested that many, but not all, signaling responses were affected by PPARδ.
PPARδ increases membrane cholesterol in leukemic B cells
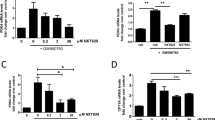
Cholesterol in plasma membranes can affect signaling complexes and alter signal transduction.29 Since PPARδ has been linked to cholesterol metabolism,30 Nile red and HIS6X-GFP-D4 (PFO)13 dyes were used to measure neutral lipid and plasma membrane cholesterol content, respectively, by flow cytometry as a function of PPARδ activity. Control cells treated for 2–3 days with GW0742 to increase PPARδ activity as well as PPARDhi Daudi cells had higher Nile red staining than control cells (Figure 4a). Primary CLL cells treated with GW0742 also exhibited higher Nile red and PFO staining along with greater PPARD expression (Figure 4b).
Effect of PPARδ on lipid content of malignant B cells. (a, b) Vector control Daudi cells with or without GW0742 (1 μM) and PPARDhi Daudi cells were cultured in RPMI-1640+5% FBS (a) and purified CLL cells (2 × 106 cells/ml) were cultured with or without GW0742 in AIM-V for 2–3 days. (b) PPARD messenger RNA (mRNA) expression was measured by real-time PCR and shown for CLL cells (b, left panel). Nile red and PFO staining were then measured by flow cytometry. Examples for Daudi cells are shown in a and mean fluorescence intensities (MFIs) are shown for CLL cells in b, with each line representing a different patient sample. (c) Oncomine analysis of the Haferlach data set indicates PPARD expression is more characteristic of CLL than acute lymphocytic leukemia (ALL), acute myeloid leukemia (AML), chronic myelogenous leukemia (CML) and myelodysplastic syndrome (MDS), and significantly correlated with genes in the cholesterol biosynthesis pathway, including isopentenyl-diphosphate delta isomerase 1 (IDI1), cytochrome b5 reductase 3 (CYB5R3), mevalonate decarboxylase (MVD) and 3-hydroxy-3-methylglutaryl-CoA synthase 1 (HMGCS1). (d) PPARD and HMGCR were measured by RT-PCR in vector control and PPARDhi Daudi cells after 24 h. Averages and s.e. of three separate measurements are shown. (e) Vector control Daudi cells and CLL cells from five different patients were cultured with or without (con) GW0742 (GW) for 2–3 days. Lovastatin (200 nM) was added to some cultures for 12 h and PFO staining then measured by flow cytometry. Dot plots for Daudi cells are shown on the left and average MFIs normalized to values for control cultures and s.e. for five CLL samples are shown in the right graph. (f) Vector control Daudi cells were cultured with or without cholesterol (25 μM) and PPARDhi Daudi cells were cultured with or without cyclodextrin (1 mM) for 12 h in RPMI-1640+5% FBS. IFNα2b (1000 U/ml) was added to some cultures for 30 min. Phospho-STAT3(tyr705) and -STAT1(tyr701) levels were then measured by immunoblotting and quantified by densitometry with β-actin as a loading control. Examples of immunoblots are shown and similar results were seen in three separate experiments. Aliquots of cells were analyzed by PFO staining to confirm membrane cholesterol was changed appropriately. Average MFIs and s.e. from three separate experiments are shown in the graph on the right. *P<0.05; **P<0.001; NS, no significant difference.
Peroxisome proliferator response elements are present in some genes of the cholesterol biosynthesis pathway and could explain higher cholesterol levels in cells with higher PPARδ activity.20 For insight into a possible relationship between PPARD expression and cholesterol metabolism in CLL cells, data sets in the public Oncomine database31 (http://www.oncomine.com/) were queried with the search terms: concept: cholesterol metabolism—GO biological process; Gene: PPARD; cancer type: chronic lymphocytic leukemia. A number of cholesterol biosynthesis genes were co-expressed with PPARD (Figure 4c), as shown by the results of querying the Haferlach data set.32 Interestingly, the relationship between PPARD and cholesterol biosynthesis appeared to be unique to CLL compared to other hematologic cancers such as acute lymphoblastic and myeloid leukemia, chronic myeloid leukemia and myelodysplastic syndrome (Figure 4c).
Consistent with these findings, expression of HMGCR, which encodes the rate-limiting enzyme of cholesterol biosynthesis 3-hydroxy-3-methyl-glutaryl-coenzyme A reductase (HMGCoAR), was nearly four-fold higher in PPARDhi Daudi cells than control cells (Figure 4d). PFO staining did not increase in Daudi (Figure 4e, left panel) or CLL cells (Figure 4e, right panel) treated with GW0742 in the presence of the HMGCoAR inhibitor Lovastatin (Figure 4e), suggesting the changes in plasma membrane cholesterol were related to increased cholesterol synthesis.
Consistent with the possibility that altered signaling in neoplastic B cells with high PPARδ activity (Figures 1, 2, 3) might be related to changes in plasma membrane cholesterol, addition of free cholesterol, solubilized in methyl-β-cyclodextrin, increased STAT3 phosphorylation in control Daudi cells (Figure 4f, left). Conversely, IFN-mediated STAT1 and STAT3 phosphorylation in PPARDhi Daudi cells were reduced by removing cholesterol with empty methyl-β-cyclodextrin (Figure 4f, middle). Uptake and stripping of cholesterol were confirmed by PFO staining (Figure 4f, right).
PPARδ inhibitors decrease cellular cholesterol and IFN-signaling
Nile red and PFO staining were lower in spleen cells from PPARD−/− mice compared to wild-type mice (Supplementary Figure 1A), consistent with a coupling of lipid content and membrane cholesterol to PPARδ activity. The PPARδ agonist GW501516 also increased Nile red staining in PPARD+/+ mice but not PPARD−/− mice (Supplementary Figure 1A, left upper panel). Induction of pSTAT1 and pSTAT3 by murine IFN-beta in PPARD−/− spleen cells were also lower compared to wild-type spleen cells (Supplementary Figure 1B).
These observations suggested chemical PPARδ inhibitors might affect IFN signaling by altering cellular lipid and plasma membrane cholesterol content. Clinically relevant PPARδ inhibitors are not available3, 10 but DG172 exhibits high-binding affinity and potent inverse agonistic properties (that is, binds PPARδ as an agonist but decreases basal target gene expression)33 and NXT1511 inhibits PPARδ at low micromolar concentrations.3 Treatment with DG172 decreased both Nile red and PFO staining in PPARDhi Daudi cells (Supplementary Figure 1C) along with IFN-induced pSTAT1 and pSTAT3 levels (Supplementary Figure 1D). Similar results were obtained with NXT1511 (Supplementary Figure 1D).
PPARδ alters IFN signaling outcomes in CLL cells
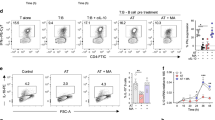
Studies were then carried out to determine if the effects of PPARδ on IFN signaling (Figures 3 and 4; Supplementary Figure 1) had functional consequences. Outcomes of type I IFN signaling in CLL cells include enhanced antigen presentation12 and toll-like receptor (TLR) responses.34 Expression of the co-stimulatory molecules C80 and CD86 were generally increased on CLL cells treated with IFNα2B for 2–3 days (Supplementary Figure 2A). Activation of PPARδ with GW0742 did not change baseline expression but upregulation of these molecules by IFNα2b was impaired by PPARδ activation (Supplementary Figure 2A). Upregulation of class I major histocompatibility antigens (HLA-A, B, C) by IFNα2B tended also to be impaired by activated PPARδ (Supplementary Figure 2A).
Pretreatment with IFNα2b for 2 days generally increases responses of CLL cells to the TLR7 agonist resiquimod, as measured by TNFα production after 4 h (Supplementary Figure 2B).34 Pretreatment with the PPARδ agonist GW0742, which has anti-inflammatory activity,35 had limited effects on subsequent induction of TNFα protein expression by resiquimod (Supplementary Figure 2B). However, GW0742 caused IFNα2b to suppress subsequent resiquimod-induced TNFα production. Impaired production of TNFα in response to TLR signaling is associated with a state of ‘TLR-tolerance’.34 Taken together, these results suggested immunogenic responses to type 1 IFN were ‘switched off’ in CLL cells by PPARδ.
Discussion
The observations in this manuscript suggest PPARδ increases the strength of signaling by cytokines such as type 1 IFNs (Figures 1, 2, 3) in malignant B cells. The effects are associated with, and potentially caused by, increased neutral lipids and plasma membrane cholesterol (Figure 4) and can alter cellular decisions. Specifically, PPARδ can change immunostimulatory IFN signaling responses to immunosuppressive ones (Supplementary Figure 2). A schematic diagram is shown in Supplementary Figure 2C.
These studies were motivated by a consistent finding of increased pSTAT3 levels with overexpression of PPARD in Daudi lymphoma cells, primary CLL cells and even breast cancer cells (Figure 1). These levels were blocked almost completely by Ruxolitinib (Figures 1c–e), suggesting they were caused by JAKs that are generally activated by cytokines. Both IL10 and IFNAR-blocking antibodies lowered pSTAT3 levels in Daudi cells transduced with PPARD expression vectors (Figures 2a and e) without evidence for increased autocrine cytokine production (Figures 2b, c and f) or cell surface receptor expression (Figure 2e). These negative results suggested responses to autocrine IFN or IL10 were amplified by PPARδ and this hypothesis was confirmed by studying specific responses to IFNα2b in Daudi cells and primary CLL cells transduced with PPARD expression vectors or treated with PPARδ agonists (Figure 3). Since cellular lipids can alter the sensitivity of receptors36, 37 and PPARδ regulates lipid metabolism,38, 39 we considered PPARδ might change intracellular cholesterol and signal transduction amplitudes. PPARδ activity was associated with upregulation of cholesterol biosynthesis genes (Figure 4c) and increased neutral lipids and membrane cholesterol (Figures 4a and e), which were lowered by statins (Figure 4d) and genetic ablation (Supplementary Figure 1A). Cytokine signaling was increased by adding cholesterol and decreased by stripping cholesterol (Figure 4f), PPARD gene knockout (Supplementary Figure 1B) and chemical PPARδ inhibitors (Supplementary Figure 1D). These findings suggested membrane cholesterol was critical for the effect of PPARδ on IFNα2b-mediated pSTAT3 levels. Functional consequences of the signaling changes associated with PPARδ included decreased co-stimulatory molecule expression and TLR responses (Supplementary Figures 2A and B).
Similar observations have been reported for TGFβ in prostate cancer,11 where PPARδ switches TGFβ1 signaling from tumor-suppressing SMAD activation to tumor-promoting ERK activation. These signaling effects are also associated with changes in membrane cholesterol attributed to PPARδ-mediated upregulation of ABCA1-cholesterol transporter activity and caveolin-1 (Cav1) expression, causing increased endocytosis of TGFβ receptors. In this paper, a direct relationship of membrane cholesterol with PPARδ activity was observed in malignant B cells (Figure 4; Supplementary Figure 1). However, the mechanism is suggested to be through increased expression of cholesterol biosynthesis genes (Figures 4c and d).20 Consistent with this idea, PPARδ-mediated increases in membrane cholesterol were prevented by statins (Figure 4e).
In addition to biosynthesis, membrane cholesterol is affected by the processes of efflux and storage as cholesterol esters in lipid droplets. Cholesterol efflux through ABCA1 and ABCG1 transporters or scavenger receptor class B member 1 (SCARB1) requires acceptors such as high-density lipoproteins or lipid-poor apoA-I proteins.40 Many of the experiments in this manuscript were performed in serum-free media where absence of acceptor molecules could produce artifactual increases in membrane cholesterol. However, similar changes were noted in vivo in PPARD−/− mice (Supplementary Figure 1A), where acceptor molecules would be present.
Another possible explanation for the effects on intracellular cholesterol is provided by our finding that PPARδ increases glucose oxidation and metabolic efficiency in Daudi and CLL cells.10 If ATP generation from glucose oxidation is optimized by PPARδ, more citrate would be available for anaplerotic reactions to generate cytoplasmic acetyl-coA41 and fuel cholesterol biosynthesis. Consistent with this possibility, pSTAT3 levels in PPARDhi Daudi cells decreased within a few hours in low-glucose culture conditions (Supplementary Figure 6).
The results in this paper suggest PPARδ affects membrane cholesterol changes that impact signal transduction in malignant B cells, especially from cytokines such as type 1 IFNs (Figure 3). It is not clear exactly how IFN signaling is altered by membrane cholesterol. Lipid rafts or membrane nanodomains may enlarge due to increased membrane cholesterol and facilitate dimerization of IFNAR1 and IFNAR2, which may then increase STAT phosphorylation.36, 42 Consistent with this idea, West Nile virus decreases plasma membrane cholesterol and IFN-induced STAT1 activation by decreasing lipid rafts.29 Alternatively, increased membrane cholesterol increases endocytosis,36 which can enhance STAT phosphorylation activity of the IFNAR.42, 43 Consistent with such a mechanism, endocytosis gene expression correlates with PPARD in CLL cells and PPARDhi Daudi cells, and GW0742-treated CLL cells express higher staining with LysoTracker Green, a marker of lysosomal activity (Supplementary Figure 7).
IL10 signaling appeared to be affected by PPARδ in a similar manner as type 1 IFN (Figures 2a–d and g) in contrast to IL6 (Figures 2a and g). IL10 and type 1 IFN are both class 2 cytokines44 but it is unclear why IL6 receptor signaling should be relatively independent of PPARδ. Perhaps the IL6 receptor is not as dependent on lipid rafts or endocytosis in CLL cells as class 2 cytokine receptors. Similarly, it is not clear why PPARδ should result in greater numbers of AKT molecules that are phosphorylated on Thr 308 but not Ser 473 following Ig-cross-linking (Supplementary Figure 5C). An association of increased PPARδ expression with greater phosphorylation of AKT at Thr 308 relative to Ser 473 has been noted previously45 and may be due to the fact that phosphoinositide-dependent kinase 1 (PDPK1), the kinase responsible for Thr 308 phosphorylation, is transcriptionally upregulated by PPARδ.3 AKT molecules phosphorylated only at Thr 308 exhibit attenuated ability to phosphorylate FOXO isoforms while retaining the ability to phosphorylate down-stream targets such as mTORC1 and glycogen synthase kinase 3 (GSK3).46 Given the role of mTORC1 in glucose uptake and lipid synthesis,47 it is possible that PPARδ may mediate enhanced proliferative responses of CLL cells following activation of the B-cell receptor through increased phosphorylation of AKT on Thr 308, but this possibility requires further investigation.
Production of type 1 IFN and antiviral activity are known to be increased by inhibiting cholesterol biosynthesis.48 Our results suggest cholesterol biosynthesis may also tune the level and affect the outcome of IFN signaling (Supplementary Figure 2C). PPARδ agonists may be able to increase STAT phosphorylation activity, particularly for STAT3, and decrease pro-inflammatory effects of IFN. A role for PPARδ antagonists in cancer therapy has been previously proposed, based on their ability to decrease metabolic efficiency, anti-oxidant defenses and survival of cancer cells in harsh microenvironmental conditions.3, 10 The results reported here suggest PPARδ antagonists may also have therapeutic benefits as signal transduction modulators that can promote the anti-tumor immune activity of IFNs.
References
Spaner D, Lee E, Shi Y, Wen F, Li Y, Tung S et al. PPAR-alpha is a therapeutic target for chronic lymphocytic leukemia. Leukemia 2013; 27: 1090–1099.
Harmon GS, Lam MT, Glass CK . PPARs and lipid ligands in inflammation and metabolism. Chem Rev 2011; 111: 6321–6340.
Wang X, Wang G, Shi Y, Sun L, Gorczynski R, Li YJ et al. PPAR-delta promotes survival of breast cancer cells in harsh metabolic conditions. Oncogenesis 2016; 5: e232.
Khozoie C, Borland MG, Zhu B, Baek S, John S, Hager GL et al. Analysis of the peroxisome proliferator-activated receptor-beta/delta (PPARbeta/delta) cistrome reveals novel co-regulatory role of ATF4. BMC Genomics 2012; 13: 665.
Abdollahi A, Schwager C, Kleeff J, Esposito I, Domhan S, Peschke P et al. Transcriptional network governing the angiogenic switch in human pancreatic cancer. Proc Natl Acad Sci USA 2007; 104: 12890–12895.
Beyaz S, Mana MD, Roper J, Kedrin D, Saadatpour A, Hong SJ et al. High-fat diet enhances stemness and tumorigenicity of intestinal progenitors. Nature 2016; 53: 53–58.
Mao F, Xu M, Zuo X, Yu J, Xu W, Moussalli MJ et al. 15-Lipoxygenase-1 suppression of colitis-associated colon cancer through inhibition of the IL-6/STAT3 signaling pathway. FASEB J 2015; 29: 2359–2370.
Wang D, Fu L, Ning W, Guo L, Sun X, Dey SK et al. Peroxisome proliferator-activated receptor delta promotes colonic inflammation and tumor growth. Proc Natl Acad Sci USA 2014; 111: 7084–7089.
Zuo X, Xu M, Yu J, Wu Y, Moussalli MJ, Manyam GC et al. Potentiation of colon cancer susceptibility in mice by colonic epithelial PPAR-delta/beta overexpression. J Natl Cancer Inst 2014; 106, dju052.
Li YJ, Sun L, Shi Y, Wang G, Wang X, Dunn SE et al. PPAR-delta promotes survival of CLL cells in energetically unfavorable conditions. Leukemia 2017; e-pub ahead of print 31 January 2017; doi:10.1038/leu.2016.395.
Her NG, Jeong SI, Cho K, Ha TK, Han J, Ko KP et al. PPARdelta promotes oncogenic redirection of TGF-beta1 signaling through the activation of the ABCA1-Cav1 pathway. Cell Cycle 2013; 12: 1521–1535.
Tomic J, Lichty B, Spaner DE . Aberrant interferon-signaling is associated with aggressive chronic lymphocytic leukemia. Blood 2011; 117: 2668–2680.
Maekawa M, Fairn GD . Complementary probes reveal that phosphatidylserine is required for the proper transbilayer distribution of cholesterol. J Cell Sci 2015; 128: 1422–1433.
Dunn SE, Bhat R, Straus DS, Sobel RA, Axtell R, Johnson A et al. Peroxisome proliferator-activated receptor delta limits the expansion of pathogenic Th cells during central nervous system autoimmunity. J Exp Med 2010; 207: 1599–1608.
Listenberger LL, Brown DA . Fluorescent detection of lipid droplets and associated proteins. Curr Protoc Cell Biol 2016; 71: 4.31.1–4.31.14.
Zidovetzki R, Levitan I . Use of cyclodextrins to manipulate plasma membrane cholesterol content: evidence, misconceptions and control strategies. Biochim Biophys Acta 2007; 1768: 1311–1324.
Neve RM, Chin K, Fridlyand J, Yeh J, Baehner FL, Fevr T et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 2006; 10: 515–527.
Spaner DE, Wang G, McCaw L, Li Y, Disperati P, Cussen M et al. Activity of the janus kinase inhibitor Ruxolitinib in chronic lymphocytic leukemia: results of a phase II trial. Haematologica 2016; 101: e192–e195.
Li Y, Shi Y, McCaw L, Li YJ, Zhu F, Gorczynski R et al. Microenvironmental interleukin-6 suppresses toll-like receptor signaling in human leukemia cells through miR-17/19A. Blood 2015; 126: 766–778.
Fang L, Zhang M, Li Y, Liu Y, Cui Q, Wang N . PPARgene: a database of experimentally verified and computationally predicted PPAR target genes. PPAR Res 2016; 2016: 6042162.
Collins RJ, Verschuer LA, Harmon BV, Prentice RL, Pope JH, Kerr JF . Spontaneous programmed death (apoptosis) of B-CLL cells following their culture in vitro. Br J Haematol 1989; 71: 343–350.
Lévy C, Frecha C, Costa C, Rachinel N, Salles G, Cosset FL et al. Lentiviral vectors and transduction of human cancer B cells. Blood 2010; 116: 498–500.
Hammond C, Shi Y, Mena J, Tomic J, Cervi D, He L et al. Effect of serum and antioxidants on the immunogenicity of protein kinase C-activated CLL cells. J Immunother 2005; 28: 28–39.
Zent CS, Chen JB, Kurten RC, Kaushal GP, Lacy HM, Schichman SA . Alemtuzumab (CAMPATH 1H) does not kill CLL cells in serum free medium. Leuk Res 2004; 28: 495–507.
Childs CB, Proper JA, Tucker RF, Moses HL . Serum contains a platelet-derived transforming growth factor. Proc Natl Acad Sci USA 1982; 79: 5312–5316.
Nakao A, Imamura T, Souchelnytskyi S, Kawabata M, Ishisaki A, Oeda E et al. TGF-beta receptor-mediated signalling through Smad2, Smad3 and Smad4. EMBO J 1997; 16: 5353–5362.
Longo PG, Laurenti L, Gobessi S, Sica S, Leone G, Efremov DG . The Akt/Mcl-1 pathway plays a prominent role in mediating antiapoptotic signals downstream of the B-cell receptor in chronic lymphocytic leukemia B cells. Blood 2008; 111: 846–855.
Rozovski U, Wu JY, Harris DM, Liu Z, Li P, Hazan-Halevy I et al. Stimulation of the B-cell receptor activates the JAK2/STAT3 signaling pathway in CLL cells. Blood 2014; 123: 3797–3802.
Mackenzie JM, Khromykh AA, Parton RG . Cholesterol manipulation by West Nile virus perturbs the cellular immune response. Cell Host Microbe 2007; 2: 229–239.
Skogsberg J, Kannisto K, Cassel TN, Hamsten A, Eriksson P, Ehrenborg E . Evidence that peroxisome proliferator-activated receptor delta influences cholesterol metabolism in men. Arterioscler Thromb Vasc Biol 2003; 23: 637–643.
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 2004; 6: 1–6.
Haferlach T, Kohlmann A, Wieczorek L, Basso G, Kronnie GT, Bene MC et al. Clinical utility of microarray-based gene expression profiling in the diagnosis and subclassification of leukemia: report from the International Microarray Innovations in Leukemia Study Group. J Clin Oncol 2010; 28: 2529–2537.
Lieber S, Scheer F, Meissner W, Naruhn S, Adhikary T, Muller-Brusselbach S et al. DG172: an orally bioavailable PPARbeta/delta-selective ligand with inverse agonistic properties. J Med Chem 2012; 55: 2858–2868.
Shi Y, White D, He L, Miller RL, Spaner DE . Toll-like receptor-7 tolerizes malignant B cells and enhances killing by cytotoxic agents. Cancer Res 2007; 67: 1823–1831.
Barish GD, Atkins AR, Downes M, Olson P, Chong LW, Nelson M et al. PPARdelta regulates multiple proinflammatory pathways to suppress atherosclerosis. Proc Natl Acad Sci USA 2008; 105: 4271–4276.
Lange Y, Steck TL . Active membrane cholesterol as a physiological effector. Chem Phys Lipids 2016; 199: 74–93.
Fielding CJ, Fielding PE . Membrane cholesterol and the regulation of signal transduction. Biochem Soc Trans 2004; 32 (Pt 1): 65–69.
Beyaz S, Yilmaz O . Molecular pathways: dietary regulation of stemness and tumor initiation by the PPARδ pathway. Clin Cancer Res 2016; 22: 5636–5641.
Holst D, Luquet S, Nogueira V, Kristiansen K, Leverve X, Grimaldi PA . Nutritional regulation and role of peroxisome proliferator-activated receptor delta in fatty acid catabolism in skeletal muscle. Biochim Biophys Acta 2003; 1633: 43–50.
Rader DJ . Molecular regulation of HDL metabolism and function: implications for novel therapies. J Clin Invest 2006; 116: 3090–3100.
Zaidi N, Swinnen JV, Smans K . ATP-citrate lyase: a key player in cancer metabolism. Cancer Res 2012; 72: 3709–3714.
Schreiber G, Piehler J . The molecular basis for functional plasticity in type I interferon signaling. Trends Immunol 2015; 36: 139–149.
Marchetti M, Monier MN, Fradagrada A, Mitchell K, Baychelier F, Eid P et al. Stat-mediated signaling induced by type I and type II interferons (IFNs) is differentially controlled through lipid microdomain association and clathrin-dependent endocytosis of IFN receptors. Mol Biol Cell 2006; 17: 2896–2909.
Kotenko SV, Pestka S . Jak-Stat signal transduction pathway through the eyes of cytokine class II receptor complexes. Oncogene 2000; 19: 2557–2565.
Pollock CB, Yin Y, Yuan H, Zeng X, King S, Li X et al. PPARδ activation acts cooperatively with 3-phosphoinositide-dependent protein kinase-1 to enhance mammary tumorigenesis. PLoS ONE 2011; 6: e16215.
Jacinto E, Facchinetti V, Liu D, Soto N, Wei S, Jung SY et al. SIN1/MIP1 maintains rictor-mTOR complex integrity and regulates Akt phosphorylation and substrate specificity. Cell 2006; 127: 125–137.
Manning BD, Toker A . AKT/PKB signaling: navigating the network. Cell 2017; 169: 381–405.
York AG, Williams KJ, Argus JP, Zhou QD, Brar G, Vergnes L et al. Limiting cholesterol biosynthetic flux spontaneously engages type I IFN signaling. Cell 2015; 163: 1716–1729.
Acknowledgements
This work was supported by CIHR grant MOP1304, CIHR grant MOP 110952, the Leukemia and Lymphoma Society of Canada (DS), CIHR grant MOP133656 (GDF), NSFC (China) grant 81372456 and the Fund for the 8th group of Fostering Talents in Jilin Province of China JRZX8 (Y-JL). LS was supported by the China Scholarship Council (CSC 201506170134). We thank Peppi Prasit (Inception Sciences, San Diego, CA) for NXT1511 and Prof Dr Wibke E Diederich (Center for Tumor Biology and Immunology, Core Facility Medicinal Chemistry, Philipps-Universität Marburg) for DG172.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Sun, L., Shi, Y., Wang, G. et al. PPAR-delta modulates membrane cholesterol and cytokine signaling in malignant B cells. Leukemia 32, 184–193 (2018). https://doi.org/10.1038/leu.2017.162
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2017.162
- Springer Nature Limited
This article is cited by
-
Derivation and validation of a lipid-covered prognostic model for mature T-cell lymphomas
Cancer Cell International (2021)
-
Mesenchymal stem cells alleviate experimental immune-mediated liver injury via chitinase 3-like protein 1-mediated T cell suppression
Cell Death & Disease (2021)