Abstract
Despite therapeutic advances, multiple myeloma (MM) remains an incurable disease, predominantly because of the development of drug resistance. The activator protein-1 (AP-1) transcription factor family has been implicated in a multitude of physiologic processes and tumorigenesis; however, its role in MM is largely unknown. Here we demonstrate specific and rapid induction of the AP-1 family member JunB in MM cells when co-cultured with bone marrow stromal cells. Supporting a functional key role of JunB in MM pathogenesis, knockdown of JUNB significantly inhibited in vitro MM cell proliferation and survival. Consistently, induced silencing of JUNB markedly decreased tumor growth in a murine MM model of the microenvironment. Subsequent gene expression profiling revealed a role for genes associated with apoptosis, DNA replication and metabolism in driving the JunB-mediated phenotype in MM cells. Importantly, knockdown of JUNB restored the response to dexamethasone in dexamethasone-resistant MM cells. Moreover, 4-hydroxytamoxifen-induced activation of a JunB-ER fusion protein protected dexamethasone-sensitive MM cells against dexamethasone- and bortezomib-induced cytotoxicity. In summary, our results demonstrate for the first time a specific role for AP-1/JunB in MM cell proliferation, survival and drug resistance, thereby strongly supporting that this transcription factor is a promising new therapeutic target in MM.
Similar content being viewed by others
Introduction
Multiple myeloma (MM) is characterized by clonal expansion of malignant plasma cells within the bone marrow (BM) compartment, monoclonal protein in blood and/or urine, bone lesions, renal compromise and immunodeficiency.1 Both genetic changes in the tumor cell and MM cell-induced disruption of the BM homeostasis support MM cell proliferation, survival and drug resistance through activation of various signaling pathways such as the phosphatidylinositol 3-kinase (PI3K)/Akt, Janus kinase/signal transducer and activator of transcription (JAK/Stat), Raf/MEK/mitogen-activated protein kinase (MAPK), nuclear factor (NF)-κB and Wnt pathways.2 Several therapeutic targets have been identified within these pathways and derived agents have been developed. Few of them have progressed to clinical trials; even fewer have been approved for MM therapy. However, those few compounds have significantly improved patient survival during the past 15 years to currently >8 years. Nevertheless, MM remains an incurable disease mainly because of the development of drug resistance. The identification of novel targets and the development of derived efficient anti-MM agents are therefore urgently needed.
Transcription factors (TFs) represent convergence points of oncogenic signaling pathways and are commonly deregulated in solid and hematologic malignancies. The TF family activator/activating protein-1 (AP-1) is characterized by highly conserved basic leucine zipper (bZIP) DNA-binding domains.3 Functionally, AP-1 dimers bind to the 12-O-tetradecanoylphorbol-13-acetate response element (5′-TGA(C/G)TCA-3′) and to related DNA elements including the cyclic AMP response element (5′-TGACGTCA-3′) and regulate not only physiologic processes, but also tumorigenesis, tumor cell promotion and progression in particular.4, 5, 6 Although AP-1 family members have been primarily considered to be oncogenic, it is now clear that AP-1 has double-edged activity.5 The decision of whether AP-1 is oncogenic or antioncogenic depends not only on the cell type, the genetic background of the tumor, its differentiation state and the tumor stage, but also on the tumor microenvironment.5 As a consequence of our increasing knowledge of the pathophysiologic role of AP-1 in tumorigenesis, members of the AP-1 family have emerged as actively pursued therapeutic targets among TFs over the past years.
In plasma cell biology and MM pathogenesis, little is known about the functional relevance of AP-1 TF family members. Recently, Grötsch et al.7 showed that Fra1 blocks plasma cell differentiation and immunoglobulin production. Conversely, the absence of Fra1 increases antiapoptotic activity in late germinal center B cells, and thereby facilitates plasma cell differentiation.8, 9 In MM, c-Maf enhances tumor cell growth via adhesion to the BM stroma, and increases angiogenesis through production of VEGF. Based on its frequent overexpression in MM cells, c-Maf has been proposed as a potential therapeutic target.10, 11, 12 Moreover, our own previous studies surprisingly showed that drug-induced inhibition of MM cell proliferation and apoptosis is, at least in part, mediated by robust upregulation of c-Jun and downstream caspase-dependent c-Abl cleavage.13
Here we identified a previously unknown role of JunB as the main AP-1 family member to induce MM cell proliferation, survival and drug resistance within the BM microenvironment. We therefore propose JunB as a promising new therapeutic target for the treatment of MM.
Materials and methods
Reagents
Recombinant human interleukin-6 (IL-6) protein was from R&D Systems (Minneapolis, MN, USA); Tocilizumab (RoActemra) was from Roche (Grenzach-Wyhlen, Germany); U0126, BAY 11-7085, LY294002, dexamethasone, doxycycline and 4-hydroxytamoxifen were from Sigma (St Louis, MO, USA); and bortezomib was from Selleck Chemicals (Houston, TX, USA). Antibodies against human JunB (C-11), c-Jun (H-79), c-Maf (M-153), extracellular signal-regulated kinase 2 (ERK2), p-ERK (E-4) and Mcl-1 (S-19) were from Santa Cruz Biotechnology (Dallas, TX, USA); antibodies against c-Fos (9F6), IκBα, phospho-IκBα (Ser32/36), Akt and phospho-Akt (Ser473) were from Cell Signaling Technology (Danvers, MA, USA); and anti-JunD was from Abcam (Cambridge, UK).
Cell culture and transfection
All human MM cell lines as well as KM-105 and HS-27A stromal cell lines were cultured in RPMI-1640 medium supplemented with 10% heat-inactivated fetal bovine serum, 1% penicillin/ streptomycin and 2 mM L-glutamine (all from Gibco, Thermo Fisher Scientific Inc., Waltham, MA, USA). MM.1S cells were transiently transfected with indicated plasmids using the Amaxa Cell Line Nucleofector Kit V (Lonza, Basel, Switzerland) according to the manufacturer’s instructions.
Primary human BMSCs and MM cells
Primary bone marrow stromal cells (BMSCs) and MM cells from MM patients were isolated from BM aspirates after informed consent was obtained in accordance with the Declaration of Helsinki. The collection and use of patient MM cells and BMSCs has been approved by the Ethics committee of the Medical Faculty, University of Heidelberg (approval number 229/2003 and S-152/2010). Isolation, purification and culture of BMSCs and primary MM cells were performed as described in Supplementary Methods.
Co-culture and conditioned medium assays
Co-culture experiments were performed as described previously.13 For details, see Supplementary Methods.
Cell lysis and western blot analysis
Whole-cell lysates were prepared in RIPA lysis buffer (150 mM NaCl, 10 mM Tris pH 7.2, 0.1% SDS, 1% Triton X-100, 1% deoxycholate and 5 mM EDTA) supplied with Halt Protease and Phosphatase Inhibitor Cocktail (Pierce, Darmstadt, Germany). Western blot analysis was performed as previously described.14
Quantitative reverse transcription-PCR
Quantitative reverse transcription-PCR was performed as previously described.14 The primers used are shown in Supplementary Table 1.
Oncomine analysis
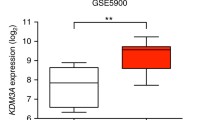
Oncomine analysis (www.oncomine.org) for JunB mRNA expression was performed using publicly available microarray data sets (GSE13591 and GSE5900)15, 16 in samples isolated from healthy individuals and samples isolated from monoclonal gammopathy of undetermined significance, smoldering MM and MM patients.
Plasmids, retroviral/lentiviral constructs and transduction
The 3 × AP-1 reporter was a gift from Alexander Dent (Indiana University School of Medicine, Indianapolis, IN, USA; Addgene plasmid 40342).17 For lentivirus-mediated RNA interference, constructs carrying small hairpin RNA (shRNA) in the pLKO.1-puro vector were from RNAi Consortium shRNA Libraries (TRC, Broad Institute, Cambridge, MA, USA). The shRNA constructs used are shown in Supplementary Table 2. pLKO.1-puro-scramble (pLKO-SCR) was used as negative control. For lentivirus-mediated inducible shRNA knockdown, pRSIT17-U6Tet-sh-CMV-TetRep-2A-TagGFP2-2A-Puro vector was used and the JUNB-specific shRNA (shJunB #1) hairpin sequence was inserted (Tet-SCR and Tet-shJunB; Cellecta/Biocat, Heidelberg, Germany). For retrovirus-mediated gene overexpression, pMSCV-JunB-ER-IRES-GFP (JunB-ER) and control empty vector pMSCV-IRES-GFP (IRES-GFP) were used.18 Viral production and transduction are described in Supplementary Methods.
Cell fractionation and TransAM AP-1 transcription factor assay
Extracts of cytoplasmic and nuclear fractions were prepared using Nuclear Extract Kit (Active Motif, Carlsbad, CA, USA), and then TransAM assays (Active Motif) were performed according to the manufacturer’s instructions.
AP-1 reporter gene assay
AP-1 reporter gene assay was measured as previously described.14
Cell proliferation and viability assays
Cell proliferation was measured by [3H]-thymidine incorporation, as described previously;13 or by measuring green fluorescent protein (GFP)-positive cells using Tecan Infinite 200 PRO (Tecan, Männedorf, Schweiz). AlamarBlue assay was performed to assess the cell viability according to the manufacturer’s instructions (Invitrogen, Thermo Fisher Scientific Inc., Waltham, MA, USA). All experiments were performed in triplicate or quadruplicate.
Cell cycle assay
Propidium iodide (Sigma)-stained cells were analyzed on a FACSCanto II flow cytometer (BD Biosciences, San Diego, CA, USA), and the percentage of cells in sub-G1, G1, S and G2/M phases was determined using the BD FACSDiva (BD Biosciences) and FlowJo softwares (Ashland, OR, USA).
In vivo studies
The 6–8-week-old Fox Chase nonobese diabetic/severe combined immunodeficient (SCID/NOD) female mice (Harlan Laboratories, Inc., Indianapolis, IN, USA) were housed and monitored in the Animal Research Facility at Magna Græcia, Catanzaro, Italy. All experimental procedures and protocols have been approved by the institutional ethical committee at the Magna Græcia University, Catanzaro. In accordance with the institutional guidelines, mice were killed when their tumors reached 2 cm in diameter, or in the event of paralysis or major compromise in their quality of life. For details, see Supplementary Methods.
Microarray analysis
MM.1S cells were infected with pLKO-shJunB #1 or pLKO-SCR lentivirus. After puromycin selection, the cells were stimulated with 25 ng/ml IL-6 for 4 h or left unstimulated. Total RNAs for microarray analyses were extracted with the TRIzol Reagent (Invitrogen, Thermo Fisher Scientific Inc.) followed by DNase treatment. Then, the cleanup of RNA samples was performed using RNeasy Mini kit (Qiagen, Hilden, Germany). All samples were prepared in biological duplicate. Gene Expression Profiling was performed using Illumina HumanHT-12 v4 Expression BeadChip Kits (Illumina Inc., San Diego, CA, USA) at the DKFZ Genomics and Proteomics Core Facility (Heidelberg, Germany). Clustering was performed using R (http://www.R-project.org/) and GenePattern.19 Gene set enrichment analysis (GSEA) was performed as previously described20 (GSEA v2.0 at http://www.broad.mit.edu/gsea) using gene set as permutation type and 1000 permutations and signal to noise as a metric for ranking genes.
Statistics
Statistical analysis was performed using Student’s t-test. P-values are indicated by asterisks (*P<0.05 and **P<0.01).
Results
The bone marrow microenvironment specifically induces JunB
In order to increase our insight into the pathophysiologic function of AP-1 family members in MM, we first determined their expression profiles in a panel of MM cell lines. Our data show a heterogeneous pattern of expression among the most prominent AP-1 family members (JunB, c-Jun, JunD, c-Fos and c-Maf) in MM cell lines (Figure 1a). Next, we co-cultured MM.1S cells with BMSCs to examine whether the BM microenvironment plays a regulatory role in AP-1 expression in MM cells. Our data demonstrate that when co-cultured with BMSCs, JunB was the AP-1 member strongest and durable upregulated in a time-dependent manner at both the mRNA (Figure 1b) and protein levels (Figure 1c). In contrast, BMSC-mediated induction of other AP-1 family members (c-Jun, JunD, c-Fos, c-Maf) was moderate or undetectable in MM cells (Figures 1b and c). In BMSCs, AP-1 member expression levels did not change (data not shown). The specific induction of JunB in MM cells was observed in not only MM.1S cells (Figure 1; Supplementary Figure 1) but also other MM cell line cells (Figure 1d) when co-cultured with BMSCs isolated from MM patients. Importantly, BMSC-induced JunB upregulation was also observed in primary MM cells isolated from patients at initial diagnosis, but not in mononuclear cells isolated from a healthy donor (Figures 1e and f).
Co-cultures of MM cells with BMSCs rapidly and strongly induce sustained expression of JunB, but not of other AP-1 family members. (a) Protein levels of AP-1 members in MM cells. Cell lysates were prepared from the indicated MM cell lines and analyzed by western blotting with antibodies against JunB, c-Jun, JunD, c-Fos and c-Maf. ERK2 served as a loading control. (b, c) AP-1 profiling in MM.1S cells induced by MM/BMSC co-cultures. MM.1S cells were cultivated alone or with primary BMSCs from MM patients (Pat. BMSCs). At the indicated time points, MM cells were detached from BMSCs and lysed. The expression levels of JunB, c-Jun, JunD, c-Fos and c-Maf in MM cells were determined by quantitative PCR (qPCR) with B2M as an endogenous control (b), and by western blotting with ERK2 as a loading control (c). **Fold change >2.5 and P<0.01 as compared with control. (d) AP-1 profiling in additional MM cells induced by MM/BMSC co-cultures. Three human MM cell lines (NCI-H929, RPMI 8226 and DOX40) were cultivated alone or with patient BMSCs for 8 h. Then, expression of JunB, c-Jun, JunD, c-Fos and c-Maf in MM cells was analyzed by western blotting with ERK2 as a loading control. (e, f) JunB expression in primary MM cells. Primary CD138+ MM cells isolated from three patients at initial diagnosis (Pat. MM) or peripheral blood mononuclear cells (PBMCs) from a healthy donor (HD) were cultivated alone or with patient BMSCs for 8 h. Then, MM cells were harvested for analysis of JunB expression by qPCR with B2M as an endogenous control (e), and by western blotting with ERK2 as a loading control (f). Each value is shown as mean±s.d. of three independent experiments. **P<0.01 compared with control. (g) Oncomine analysis of publicly available microarray data sets for JunB mRNA expression in normal plasma cells (NPCs), cells from patients with monoclonal gammopathy of undetermined significance (MGUS), smoldering multiple myeloma (Smoldering MM) and MM. The numbers in the parentheses are the n numbers per group. P-values as indicated; NS, not significant.
Furthermore, we evaluated the expression of JunB mRNA in MM patient samples using the Oncomine software. Consistent with our results, we identified a significant induction of JunB mRNA levels progressing from normal plasma cells to cells from patients with monoclonal gammopathy of undetermined significance, smoldering MM and MM in two15, 16 independent gene expression profiling data sets (Figure 1g). In contrast, no significant changes were observed in the expression of other AP-1 family members (data not shown). Taken together, our data demonstrate a significant and specific upregulation of JunB in MM cells exposed to the BM microenvironment and indicate that JunB expression may correlate with MM pathogenesis.
Induction of JunB in MM cells is predominantly mediated by soluble factors secreted by the bone marrow microenvironment
We next explored whether direct MM/BMSC cell–cell contact is required for JunB upregulation. Conditioned medium from the BMSC line KM105 significantly induced JunB upregulation in MM cells (Figures 2a and b and Supplementary Figure 2a) at both the transcriptional and translational levels (Supplementary Figure 2b–d). Fixing BMSCs with paraformaldehyde before co-cultures completely abrogated JunB upregulation in MM cells (Figure 2c). Moreover, JunB upregulation in MM cells was also observed in a transwell system that does not allow direct BMSC/MM cell contact (Figure 2d). These data strongly support a role of secreted factors in BMSC-induced JunB upregulation in MM cells.
Induction of JunB expression in MM cells is predominantly mediated by soluble factors, IL-6 in particular, secreted by BMSCs rather than direct MM/BMSC contact. (a, b) Induction of JunB expression in MM cells mediated by soluble factors secreted by the KM-105 stromal cell line. MM.1S cells were cultivated alone, with KM-105 BMSCs or with KM-105 conditioned medium (CM). At the indicated time points, MM cells were harvested for analysis of JunB expression by western blotting with ERK2 as a loading control (a), and for analysis of JunB and c-Jun expression by quantitative PCR (qPCR) with B2M as an endogenous control (b). Each value is shown as mean±s.d. of three independent experiments. **P<0.01 compared with control. (c, d) Induction of JunB expression in MM cells is predominantly mediated by soluble factors secreted by BMSCs rather than direct MM/BMSC contact. MM.1S cells were cultivated alone, with paraformaldehyde-fixed or non-fixed patient BMSCs (c); or with patient BMSCs, using a non-contact co-culture system as described in the ‘Materials and methods’ (d). After 8 h, JunB expression in MM cells was analyzed by western blotting with ERK2 as a loading control. (e) IL-6 induces JunB upregulation in a dose- and time-dependent manner. MM.1S cells were treated with or without IL-6 at the indicated concentrations. At the indicated time points, MM cells were harvested for analysis of JunB expression by western blotting with ERK2 as a loading control. (f) Tocilizumab inhibits BMSC-induced upregulation of JunB in MM cells. MM.1S cells were pretreated with tocilizumab and then cultivated alone or with patient BMSCs. At the indicated time points, MM cells were harvested for analysis of JunB expression by western blotting with ERK2 as a loading control. (g) Tocilizumab inhibits BMSC-induced proliferation of MM cells. MM.1S cells or primary CD138+ MM cells isolated from three patients (Pat. MM) were pretreated with tocilizumab and then cultivated alone or with patient BMSCs. After 24 h, cell proliferation was evaluated by [3H]-thymidine incorporation assay. Each value is shown as mean±s.d. of three independent experiments. *P<0.05 and **P<0.01. (h) Inhibition of the MEK/ERK pathway by U0126 and the NF-κB pathway by BAY 11-7085, but not the PI3K pathway by LY294002, blocks JunB upregulation. MM.1S cells were pretreated with U0126, BAY 11-7085 or LY294002, and then cultivated alone or with Pat. BMSCs. After 2 h, MM cells were harvested for western blotting analysis using antibodies against JunB, p-ERK, p-IκBα and p-AKT. ERK2 served as a loading control.
Soluble factors significantly upregulated in the MM BM microenvironment include IL-6, insulin-like growth factor-1, vascular endothelial growth factor, macrophage inflammatory proteins, monocyte chemoattractant proteins and stromal cell-derived factor-1.21 IL-6 is the preeminent growth and survival factor in MM22, 23 predominantly produced and secreted by BMSCs and osteoblasts.2, 24 As shown in Figure 2e, IL-6 induced JunB upregulation in MM cells in a time- and dose-dependent manner. Conversely, tocilizumab, a humanized monoclonal IL-6 receptor antibody, time-dependently inhibited BMSC-induced JunB upregulation (Figure 2f). Remarkably, tocilizumab inhibited BMSC-induced cell proliferation in both the MM.1S cell line and primary MM cells (Figure 2g). Using U0126, BAY 11-7085 and LY294002, our data show that IL-6-triggered JunB upregulation is MEK/MAPK and NF-κB dependent, but PI3K/Akt independent (Figure 2h). Taken together, our data demonstrate that humoral factors secreted by the MM BM microenvironment, IL-6 in particular, trigger MEK/ERK- and NF-κB-dependent but PI3K-independent induction of JunB expression in MM cells.
JunB induces MM cell proliferation
The specific pathophysiologic relevance of JunB upregulation in MM was investigated next. After nuclear protein extraction, the binding of activated AP-1 family members JunB, c-Jun, JunD and c-Fos to the oligonucleotide of their consensus binding site was quantified using antibodies specific for the bound, active form of JunB, pSer73 of c-Jun, JunD, and c-Fos. MM/BMSC co-cultures induced significant binding of JunB to its oligo-containing TRE element in vitro, but only moderate or no binding of other AP-1 family members to their specific oligo elements (Figure 3a). Tocilizumab significantly decreased JunB binding in MM cells co-cultured with BMSCs (Figure 3b), further supporting a key role for IL-6 in regulating JunB expression. We next addressed AP-1 transactivation activity by a reporter assay. As shown in Figure 3c, IL-6 enhanced AP-1 transactivation activity, and doxycycline-induced silencing of JUNB significantly inhibited AP-1-reporter activity in MM cells exposed to exogenous IL-6.
BMSC-induced AP-1 activity and cell proliferation is significantly inhibited by JUNB knockdown in MM cells. (a) BMSC-induced AP-1 family member binding activity in MM.1S cells. MM.1S cells were cultivated alone or with patient BMSCs. At the indicated time points, the nuclear extracts were prepared and DNA-binding activities of different AP-1 family members were measured using TransAM AP-1 family kit as described in the ‘Materials and methods’. Each value is shown as mean±s.d. of three independent experiments. *P<0.05 and ** P<0.01 as compared with control. (b) Tocilizumab inhibits BMSC-induced JunB-binding activity in MM.1S cells. MM.1S cells were pretreated with tocilizumab and then cultivated alone or with patient BMSCs. After 24 h, DNA-binding activity of JunB was determined using TransAM AP-1 family kit. The wild-type (wt) or mutated (mut) AP-1 consensus oligonucleotide was used as a competitor for JunB binding. Each value is shown as mean±s.d. of three independent experiments. **P<0.01. (c) Inhibition of IL-6-induced AP-1 activity by JUNB knockdown. Tet-shJunB/ MM.1S cells treated with or without doxycycline were transiently transfected with 3 × AP-1 reporter or pGL2 basic vector, together with pRL-CMV Renilla luciferase vector. Then, the cells were treated with IL-6 or left untreated. At the indicated time points, luciferase activity was measured by dual-luciferase reporter assay. The fold change of luciferase activity relative to control cells is shown as mean±s.d. from three independent experiments. *P<0.05 and **P<0.01 as compared with control. (d, e) Rapid and strong inhibition of MM.1S cell proliferation by JUNB knockdown. Silencing of AP-1 family members (JUNB, c-JUN, JUND, c-FOS) in MM cells was achieved by using a lentiviral delivery system. pLKO.1-infected MM.1S cells after 48 h of selection with 1 μg/ml puromycin were cultivated alone or with patient BMSCs. After 2 h, expression of AP-1 family members in MM cells was analyzed by western blotting with ERK2 as a loading control (d). Cell proliferation of MM.1S cells cultivated with patient BMSCs was evaluated by [3H]-thymidine incorporation assay. pLKO.1-SCR-infected MM.1S cells served as control. Each value is shown as mean±s.d. of two independent experiments. **P<0.01 compared with control (e). (f) Significant increase of the G2/M phase followed by a subsequent significant increase of the sub G1-phase after silencing of JUNB in MM.1S cells co-cultured with patient BMSCs. JUNB knockdown-induced changes in the cell cycle of MM cells was analyzed. The mean percentage of MM cells in each cell cycle phase is presented in the lower panel, with the s.d. values in parentheses. (g) JUNB knockdown-induced inhibition of cell proliferation in other MM cells. pLKO.1-shJunB #1-infected MM cells (RPMI 8226 and DOX40) after 48 h of selection with 1 μg/ml puromycin were then cultivated with patient BMSCs. After 24 h, cell proliferation was evaluated by [3H]-thymidine incorporation assay (left panel), and expression of JunB in MM cells was analyzed by western blotting with ERK2 as a loading control (right panel). Each value is shown as mean±s.d. of two independent experiments. **P<0.01 compared with control.
Knockdown of JUNB abrogated BMSC-induced JunB upregulation in MM cells. Importantly, shRNA-mediated knockdown of c-JUN, JUND and c-FOS did not influence BMSC-induced JunB upregulation (Figure 3d). Functionally, shRNA-mediated JUNB knockdown triggered robust and rapid inhibition of MM.1S cell proliferation in MM/BMSC co-cultures. Because of complete depletion of JunB by shJunB #1 but not shJunB #2, shJunB #1 was used for all subsequent experiments. Knockdown of JUND and c-FOS triggered delayed inhibition of MM.1S proliferation when co-cultured with BMSCs (Figure 3e). Cell cycle analysis revealed a significant increase of the G2/M phase followed by a subsequent significant increase of the sub-G1-phase after silencing of JUNB in MM.1S cells co-cultured with primary BMSCs (Figure 3f), KM-105 (Supplementary Figure 3a) or HS-27A BM stromal cell line cells (data not shown), respectively. Similar results were obtained in RPMI 8226 and DOX40 cells, when cultured alone (Supplementary Figure 3b) or in MM/BMSC co-cultures (Figure 3g). These results suggest an essential functional role of JunB in MM.
Induced silencing of JUNB inhibits MM cell proliferation in a murine MM xenograft model
Doxycycline-induced JUNB silencing (Supplementary Figure 4a) inhibited cell growth in Tet-shJunB/ MM.1S cells when cultured in the presence of BMSCs (Supplementary Figure 4b). In contrast, doxycycline did not induce inhibition of BMSC-triggered MM cell proliferation in MM.1S cells infected with scrambled shRNA (Supplementary Figure 4b). Inhibition of IL-6-induced MM cell proliferation by doxycycline-induced JUNB knockdown in Tet-shJunB/MM.1S cells was rescued by transfection with a JUNB expression vector (Supplementary Figure 4c). In vivo, Tet-SCR/MM.1S or Tet-shJunB/MM.1S were injected subcutaneously together with human-derived BMSCs and matrigel into the left and right flanks of mice, respectively (Figure 4a), and fed with doxycycline in their drinking water for 5 weeks. Treatment with doxycycline inhibited JunB protein levels in Tet-shJunB/MM.1S, but not in Tet-SCR/MM.1S (Supplementary Figure 4d). A significant reduction in growth was observed in tumors formed by Tet-shJunB/ MM.1S versus control cells (Figures 4a and b and Supplementary Figure 4e). Doxycycline did not induce toxicity, as evidenced by equal body weight of mice treated with doxycycline or those left untreated (Supplementary Figure 4f). Taken together, our data demonstrate for the first time that JunB plays a key role in MM growth.
Induced silencing of JUNB inhibits MM cell proliferation in a murine MM xenograft model. Tet-SCR/MM.1S or Tet-shJunB/MM.1S were injected subcutaneously together with human-derived BMSCs and matrigel into the left and right flanks of mice, respectively (injection scheme), and fed with doxycycline in their drinking water for 5 weeks. (a) Injection scheme and photographs of one representative mouse at days 18, 30 and 38 after cell inoculation. (b) Growth curves of MM.1S xenografts.
Effect of JUNB knockdown on gene expression in MM cells
To gain insight into molecular pathways driving JunB-mediated phenotypes, we next performed gene expression profiling of MM.1S cells infected with scrambled control and JUNB-targeting shRNA, with or without IL-6 stimulation. Unsupervised clustering segregated the samples in two sharply divided groups, suggesting a prominent role for JunB in MM biology, even more prominent than IL-6 stimulation (Figure 5a). We then sought to identify the subset of genes that were upregulated by IL-6 stimulation through JunB. To this end, we performed GSEA. Remarkably, the apoptotic pathway was among the most significantly deregulated pathways in MM cells after IL-6 treatment in JUNB knockdown cells (normalized enrichment score=1.64, false discovery rate=0.016) (Figure 5b). Among the genes deregulated following JUNB knockdown were CASPASE-3, CASPASE-8, FADD, BID, DFFB, APAF1 and BIRC3 (Figure 5c and Supplementary Figure 5). More specifically, JUNB knockdown had an impact on both the intrinsic and extrinsic apoptotic pathways (Figure 5d). Beyond apoptosis, genes of several additional pathways were deregulated following JUNB knockdown, including the PI3K, NF-κB, JAK/Stat and MAPK pathways (Figure 5d). Moreover, JUNB knockdown deregulated molecules associated with DNA replication, metabolism, cell cycle, cell surface proteins as well as the production of cytokines and growth factors (Figure 5e).
Effect of JUNB knockdown on gene expression in MM cells. (a) Unsupervised hierarchical clustering of MM1S cells infected with scrambled (SCR) or JUNB (shJunB) shRNAs in the presence or absence of IL-6 (IL6). (b) GSEA enrichment profile of the gene set Biocarta_death_pathway from MSigDB. The bar-code plot indicates the position of the genes belonging to the gene set on the expression data rank-sorted by its association with JUNB knockdown, with red and blue colors indicating over- and underexpression in the shRNA group. (c) Heat map of the ranked genes listed in the MSigDB gene set Biocarta_death_pathway. (d) Summary of genes associated with MM cell apoptosis that are modified by JUNB knockdown. Upregulated genes are in red; downregulated genes are shown in blue. (e) Overview of genes modified by JUNB knockdown. Upregulated genes are shown in underlined font, downregulated genes are shown in normal font.
JunB protects MM cells against dexamethasone- and bortezomib-induced cell death
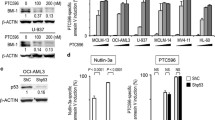
Despite recent advances in MM treatment, MM remains an incurable disease predominantly because of the development of drug resistance.1 In contrast to glucocorticoid (GC)-sensitive MM.1S cells, cells of the MM.1 subline MM.1R are GC resistant.25 Interestingly, we observed high levels of basal JunB expression in MM.1R versus MM.1S cells (Figure 1a), indicating a potential role of JunB in dexamethasone-resistant MM.1R cells. Our results show that shRNA-mediated knockdown of JUNB (Supplementary Figure 6) potently sensitize MM.1R cells to dexamethasone treatment in a dose-dependent manner leading to inhibition of cell proliferation (Figure 6a) and survival (Figure 6b).
JunB protects MM cells against dexamethasone- and bortezomib-induced MM cell death. (a, b) Depletion of JunB sensitizes dexamethasone-resistant MM.1R cells to dexamethasone. pLKO.1-shJunB #1 or SCR-infected MM.1R after 48 h of selection with 1 μg/ml puromycin were then treated with or without dexamethasone at the indicated concentrations. Cell proliferation was evaluated using the [3H]-thymidine incorporation assay after 48 h (a), and cell viability was evaluated by AlamarBlue assay after 72 h (b). Each value is shown as mean±s.d. of two independent experiments. *P<0.05 and **P<0.01. (c, d) 4-Hydroxytamoxifen (4-OHT)-induced JunB activity in JunB-ER/MM.1S but not IRES-GFP/ MM.1S cells. JunB-ER/ MM.1S and IRES-GFP/MM.1S cell lines were established as described in the ‘Materials and methods’. Cells were treated with or without 4-OHT at the indicated concentrations. After 8 h, expression of JunB-ER in MM cells was analyzed by western blotting with the antibody against JunB. ERK2 served as a loading control. *Indicates position of JunB-ER protein that is ~80 kDa (c). JunB-ER/MM.1S and IRES-GFP/ MM.1S cells were transiently transfected with 3 × AP-1 reporter together with pRL-CMV Renilla luciferase vector. Then, cells were treated with 4-OHT at the indicated concentrations or left untreated. After 6 h, luciferase activity was measured by dual-luciferase reporter assay. Fold change of luciferase activity relative to control cells is shown as mean±s.d. from three independent experiments. **P<0.01 compared with control (d). (e, f) 4-OHT-triggered JunB activity reduces dexamethasone-induced inhibition of proliferation in JunB-ER/ MM.1S but not IRES-GFP/MM.1S cells. JunB-ER/MM.1S (f) and IRES-GFP/MM.1S (e) cells cultivated in the presence or absence of 0.2 μM 4-OHT were treated with dexamethasone at the indicated concentrations or left untreated, and proliferation was determined by the [3H]-thymidine incorporation assay after 24 h. **P<0.01. (g, h) 4-OHT-triggered JunB activity reduces bortezomib-induced cell death in JunB-ER/MM.1S but not IRES-GFP/MM.1S cells. JunB-ER/MM.1S (h) and IRES-GFP/MM.1S (g) cells cultivated in the presence or absence of 0.2 μM 4-OHT were treated with bortezomib at the indicated concentrations or left untreated, and cell viability was determined by the AlamarBlue assay after 24 h. **P<0.01.
We next generated MM.1S cells stably expressing an inducible JunB fusion protein with the hormone-binding domain of the human estrogen receptor (JunB-ER) in order to further investigate the role of JunB in drug resistance of MM cells. The principle of this inducible system is that, although the chimeric protein is constitutively expressed, it is inactive in the absence of hormone and can be activated by the addition of 4-hydroxytamoxifen (4-OHT).18 As expected, whereas absolute protein levels of JunB did not change (Figure 6c), 4-OHT triggered a dose-dependent increase of transcriptional activity in JunB-ER/ MM.1S cells but not control MM.1S cells (Figure 6d). Importantly, activation of the JunB-ER fusion protein with 4-OHT significantly increased resistance to dexamethasone in JunB-ER/MM.1S cells, but not control MM.1S cells (Figures 6e and f and Supplementary Figure 7a). Similarly, activation of the JunB-ER fusion protein with 4-OHT also significantly increased resistance to bortezomib in JunB-ER/MM.1S cells, but not control MM.1S cells (Figures 6g and h and Supplementary Figure 7b).
In summary, these data demonstrate that JunB plays a key role in not only MM cell proliferation and survival, but also in drug resistance.
Discussion
Despite unprecedented therapeutic advances during the past two decades that led to major improvements of patient survival, MM remains an incurable disease with a continued need for new therapies. Although the TF family AP-1 has emerged as an actively pursued drug discovery target, little is known about its function in plasma cell biology and MM pathophysiology. Here we introduced JunB as the main AP-1 TF member involved in the pathogenesis of MM. This was first evidenced by rapid and strong upregulation of JunB but not other AP-1 family members in MM cells when co-cultured with BMSCs. Consistently, analysis of publicly available data sets revealed the progression of JUNB expression from normal plasma cells to cells isolated from monoclonal gammopathy of undetermined significance, smoldering MM and MM patients. Moreover, our data show that JunB upregulation was predominantly triggered in response to MM/BMSC-induced IL-6 secretion. In concordance with our data, previously obtained gene expression profiles in the IL-6-dependent MM cell lines INA-6 and XG-1 have shown significant upregulation of JunB in response to IL-6.26 Conversely, blockade of the IL-6 receptor by tocilizumab inhibited MM/BMSC-induced JunB upregulation in MM cells. Moreover, our data demonstrate that IL-6 triggers MEK/ERK- and NF-κB-dependent but PI3K-independent induction of JunB expression in MM cells. Supporting our own data, the involvement of the MEK/ERK and NF-κB pathways in the regulation of AP-1 family member expression and activity has been suggested before, for example, in Hodgkin’s lymphoma and anaplastic large-cell lymphoma.5, 27, 28, 29, 30 However, detailed mechanisms by which MEK/ERK and NF-κB pathways upregulate JunB in MM cells are currently unknown. Studies to particularly investigate the potential regulatory role of NF-κB itself as well as of Ets-1 and Elk-1 to induce JunB expression in MM cells are ongoing. Of note, IL-6-induced upregulation of JunB was independent of the Ras mutation status in MM cells, with RPMI 8226, DOX40, MM.1S and MM.1R cells carrying K-Ras mutations, and NCI-H929 cells carrying an N-Ras mutation.31, 32, 33 Ongoing studies investigate whether other cellular and noncellular components of the BM microenvironment also induce JunB upregulation in MM cells.
Functionally, the Jun protein family has been proposed to be the limiting component that determines whether AP-1 promotes or inhibits cell proliferation.34 Early studies suggested antagonistic roles for Jun proteins in proliferation and transformation, with c-Jun enhancing and JunB inhibiting transformation. However, accumulating evidence shows that both c-Jun and JunB can exert distinct functions depending on the cellular origin, the cell-cycle stage and environmental conditions. For example, JunB is a gatekeeper in acute and chronic myeloid leukemia;35, 36 in contrast, it is a positive regulator in renal cell carcinoma,37 Hodgkin’s lymphoma, anaplastic large-cell lymphoma, lymphomatoid papulosis and cutaneous lymphomas.27, 28, 38, 39, 40 Moreover, amplification and/or overexpression of JUNB has been associated with cervical, endometrial and colorectal cancers.41, 42, 43 In this report, the pathophysiologic relevance of BMSC/IL-6-induced JunB upregulation in MM was evidenced by increased JunB/DNA binding and JunB transcriptional activity. Moreover, the predominant effect of JunB among other AP-1 family members on MM cell growth and survival was confirmed by shRNA-induced knockdown. Importantly, our in vitro data could also be translated into a murine xenograft model using doxycycline-inducible Tet-shJunB/MM.1S cells. Molecules deregulated upon JUNB knockdown in MM cells are associated with not only cell apoptosis but also DNA replication, metabolism, cell cycle, cell surface proteins as well as the production of cytokines and growth factors. Mechanisms by which AP-1 family members achieve this functional diversity dependent on the cell type are not completely understood. The availability of external and internal survival factors, the complex interactions between different dimerization partners as well as the interaction of the AP-1 complex with other transcription factors are presumably critical.5, 18, 34, 44, 45, 46, 47, 48 Our ongoing studies in MM aim to identify functional relevant binding partners of JunB. Although our data demonstrate a key role for JunB, a functional role of JunD, c-Fos and other more rare AP-1 family members in MM pathogenesis cannot be excluded. Further studies to identify and delineate their respective roles are needed.
Finally, MM remains an incurable disease predominantly because of the development of drug resistance. Of note, our data demonstrate the involvement of JunB in mediating resistance to not only dexamethasone but also bortezomib in MM cells. In accordance with a role of JunB in drug resistance, JunB was among a limited subset of 170 out of 15 906 human cDNAs tested for their ability to confer resistance to kinase inhibitors in a recent global screening study.49 In order to delineate mechanisms that mediate JunB-induced resistance of MM cells to dexamethasone, we performed additional experiments investigating the effect of JUNB knockdown on the GC receptor. Specifically, GC receptor expression levels were compared in MM.1R cells transfected with SCR control or shJunB using quantitative PCR. Equally low levels of GC receptor expression were observed in both SCR/MM.1R and shJunB/MM.1R cells. We therefore excluded a direct transcriptional effect of JunB on GC receptor expression as a cause for transferring dexamethasone resistance to MM.1R cells. JunB-mediated modulation of molecules that inhibit GC receptor activity may represent a likely alternative mechanism. For example, Hsp27 has been proposed to overcome IL-6-mediated protection of dexamethasone-induced apoptosis.50 Whether JunB regulates Hsp27 expression in MM cells is currently unknown. Furthermore, previous studies have shown that inhibition of deubiquitinating enzymes resensitizes MM cells to bortezomib.51, 52 Whether JunB mediates resistance of MM cells to bortezomib via induction of deubiquitinating enzymes is under investigation. Moreover, using two- and three-dimensional co-culture systems, we currently evaluate whether JunB also confers resistance to a panel of other investigational, novel and conventional anti-MM agents including lenalidomide, pomalidomide, carfilzomib and doxorubicin.
Besides targeting molecules within oncogenic signaling pathways (for example, cytokine/growth factor receptors, that is, IL-6 receptor), which converge at a specific TF or downstream effectors, approaches to directly target TFs are among the most promising novel anticancer strategies with a potentially high therapeutic index. We and others hypothesize that inhibition of TFs as the terminal effectors of oncogenic signal transduction leads to selective tumor cell death, whereas normal cells tolerate the loss of TFs with little consequences because of redundancies in normal signaling pathways. Indeed, drugs that target nuclear hormone receptors, including dexamethasone, tamoxifen, fulvestrant, bicalutamide and enzalutamide, belong to the most successful targeted therapies in oncology.53 Among non-nuclear hormone receptors TFs, TP53 and MYC, which encode the TFs p53 and c-Myc, are commonly altered genes across all cancers, including MM.54 Large surface areas for protein–DNA and protein–protein interactions, which are difficult to target, and the predominant nuclear localization, which makes the TF less accessible to therapeutic agents, remain a challenge.55 Preclinical studies that investigate the anti-MM activity of agents that target TFs such as c-Myc,56 Hif-1α57, 58 and SP1(ref. 59) showed already promising results. In addition, AP-1 family members have emerged as actively pursued therapeutic targets over the past years. Worldwide academic and industrial efforts are dedicated to the development of selective AP-1 inhibitors.60 Strategies to target JunB and other TFs include the inhibition of their expression (RNA interference or miRNAs) and DNA binding (oligodeoxynucleotide decoys or pyrrole-imidazole polyamides), as well as more recently the disruption of critical protein–protein interactions, and the restricted binding at the epigenetic level by modulation of the chromatin accessibility. However, no effective AP-1 member inhibitor has yet been approved for clinical use.
We also explored the possible role of JunB on patient survival. High JUNB expression levels were associated with a better prognosis when compared with patients with low JUNB levels (Supplementary Figure 8a). We surmise that MM cells, in which JunB is upregulated, rely extensively on the support of the microenvironment (including IL-6). Once cells become independent from their microenvironment surroundings, they render more aggressive and do not any longer rely on JunB so extensively. To further corroborate this hypothesis, we used an additional MM patient data set that includes gene expression profiles at the baseline (780 patient samples) and at relapse (255 patient samples) after various therapies. Remarkably, the expression levels of JUNB were significantly higher at the baseline than at relapse of the disease (Supplementary Figure 8b).
In summary, our data demonstrate for the first time a critical role for JunB/AP-1 in MM cell growth, survival and drug resistance and strongly propose JunB as a novel therapeutic target in MM.
References
Fairfield H, Falank C, Avery L, Reagan MR . Multiple myeloma in the marrow: pathogenesis and treatments. Ann NY Acad Sci 2016; 1364: 32–51.
Podar K, Chauhan D, Anderson KC . Bone marrow microenvironment and the identification of new targets for myeloma therapy. Leukemia 2009; 23: 10–24.
Eychène A, Rocques N, Pouponnot C . A new MAFia in cancer. Nat Rev Cancer 2008; 8: 683–693.
Bernstein LR, Colburn NH . AP1/jun function is differentially induced in promotion-sensitive and resistant JB6 cells. Science 1989; 244: 566–569.
Eferl R, Wagner EF . AP-1: a double-edged sword in tumorigenesis. Nat Rev Cancer 2003; 3: 859–868.
Matthews CP, Colburn NH, Young MR . AP-1 a target for cancer prevention. Curr Cancer Drug Targets 2007; 7: 317–324.
Grötsch B, Brachs S, Lang C, Luther J, Derer A, Schlötzer-Schrehardt U et al. The AP-1 transcription factor Fra1 inhibits follicular B cell differentiation into plasma cells. J Exp Med 2014; 211: 2199–2212.
Krum SA, Chang J, Miranda-Carboni G, Wang C-Y . Novel functions for NFκB: inhibition of bone formation. Nat Rev Rheumatol 2010; 6: 607–611.
De Silva NS, Simonetti G, Heise N, Klein U . The diverse roles of IRF4 in late germinal center B-cell differentiation. Immunol Rev 2012; 247: 73–92.
Chesi M, Nardini E, Lim RS, Smith KD, Kuehl WM, Bergsagel PL . The t(4;14) translocation in myeloma dysregulates both FGFR3 and a novel gene, MMSET, resulting in IgH/MMSET hybrid transcripts. Blood 1998; 92: 3025–3034.
Hurt EM, Wiestner A, Rosenwald A, Shaffer AL, Campo E, Grogan T et al. Overexpression of c-maf is a frequent oncogenic event in multiple myeloma that promotes proliferation and pathological interactions with bone marrow stroma. Cancer Cell 2004; 5: 191–199.
Annunziata CM, Hernandez L, Davis RE, Zingone A, Lamy L, Lam LT et al. A mechanistic rationale for MEK inhibitor therapy in myeloma based on blockade of MAF oncogene expression. Blood 2011; 117: 2396–2404.
Podar K, Raab MS, Tonon G, Sattler M, Barilà D, Zhang J et al. Up-regulation of c-Jun inhibits proliferation and induces apoptosis via caspase-triggered c-Abl cleavage in human multiple myeloma. Cancer Res 2007; 67: 1680–1688.
Fan F, Tonon G, Bashari MH, Vallet S, Antonini E, Goldschmidt H et al. Targeting Mcl-1 for multiple myeloma (MM) therapy: Drug-induced generation of Mcl-1 fragment Mcl-1(128-350) triggers MM cell death via c-Jun upregulation. Cancer Lett 2014; 343: 286–294.
Agnelli L, Mosca L, Fabris S, Lionetti M, Andronache A, Kwee I et al. A SNP microarray and FISH-based procedure to detect allelic imbalances in multiple myeloma: an integrated genomics approach reveals a wide gene dosage effect. Genes Chromosomes Cancer 2009; 48: 603–614.
Zhan F, Barlogie B, Arzoumanian V, Huang Y, Williams DR, Hollmig K et al. Gene-expression signature of benign monoclonal gammopathy evident in multiple myeloma is linked to good prognosis. Blood 2007; 109: 1692–1700.
Vasanwala FH, Kusam S, Toney LM, Dent AL . Repression of AP-1 function: a mechanism for the regulation of Blimp-1 expression and B lymphocyte differentiation by the B cell lymphoma-6 protooncogene. J Immunol 2002; 169: 1922–1929.
Bakiri L, Lallemand D, Bossy-Wetzel E, Yaniv M . Cell cycle-dependent variations in c-Jun and JunB phosphorylation: a role in the control of cyclin D1 expression. EMBO J 2000; 19: 2056–2068.
Reich M, Liefeld T, Gould J, Lerner J, Tamayo P, Mesirov JP . GenePattern 2.0. Nat Genet 2006; 38: 500–501.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005; 102: 15545–15550.
McMillin DW, Negri JM, Mitsiades CS . The role of tumour-stromal interactions in modifying drug response: challenges and opportunities. Nat Rev Drug Discov 2013; 12: 217–228.
Kawano M, Hirano T, Matsuda T, Taga T, Horii Y, Iwato K et al. Autocrine generation and requirement of BSF-2/IL-6 for human multiple myelomas. Nature 1988; 332: 83–85.
Klein B, Zhang XG, Lu ZY, Bataille R . Interleukin-6 in human multiple myeloma. Blood 1995; 85: 863–872.
Giuliani N, Rizzoli V, Roodman GD . Multiple myeloma bone disease: pathophysiology of osteoblast inhibition. Blood 2006; 108: 3992–3996.
Greenstein S, Krett NL, Kurosawa Y, Ma C, Chauhan D, Hideshima T et al. Characterization of the MM.1 human multiple myeloma (MM) cell lines: a model system to elucidate the characteristics, behavior, and signaling of steroid-sensitive and -resistant MM cells. Exp Hematol 2003; 31: 271–282.
Brocke-Heidrich K, Kretzschmar AK, Pfeifer G, Henze C, Löffler D, Koczan D et al. Interleukin-6-dependent gene expression profiles in multiple myeloma INA-6 cells reveal a Bcl-2 family-independent survival pathway closely associated with Stat3 activation. Blood 2004; 103: 242–251.
Staber PB, Vesely P, Haq N, Ott RG, Funato K, Bambach I et al. The oncoprotein NPM-ALK of anaplastic large-cell lymphoma induces JUNB transcription via ERK1/2 and JunB translation via mTOR signaling. Blood 2007; 110: 3374–3383.
Mathas S, Hinz M, Anagnostopoulos I, Krappmann D, Lietz A, Jundt F et al. Aberrantly expressed c-Jun and JunB are a hallmark of Hodgkin lymphoma cells, stimulate proliferation and synergize with NF-kappa B. EMBO J 2002; 21: 4104–4113.
Watanabe M, Sasaki M, Itoh K, Higashihara M, Umezawa K, Kadin ME et al. JunB induced by constitutive CD30-extracellular signal-regulated kinase 1/2 mitogen-activated protein kinase signaling activates the CD30 promoter in anaplastic large cell lymphoma and reed-sternberg cells of Hodgkin lymphoma. Cancer Res 2005; 65: 7628–7634.
Watanabe M, Itoh K, Togano T, Kadin ME, Watanabe T, Higashihara M et al. Ets-1 activates overexpression of JunB and CD30 in Hodgkin’s lymphoma and anaplastic large-cell lymphoma. Am J Pathol 2012; 180: 831–838.
Lodé L, Moreau P, Ménard A, Godon C, Touzeau C, Amiot M et al. Lack of BRAF V600E mutation in human myeloma cell lines established from myeloma patients with extramedullary disease. Blood Cancer J 2013; 3: e163.
Bolick SCE, Landowski TH, Boulware D, Oshiro MM, Ohkanda J, Hamilton AD et al. The farnesyl transferase inhibitor, FTI-277, inhibits growth and induces apoptosis in drug-resistant myeloma tumor cells. Leukemia 2003; 17: 451–457.
Suzuki R, Kikuchi S, Harada T, Mimura N, Minami J, Ohguchi H et al. Combination of a Selective HSP90α/β Inhibitor and a RAS-RAF-MEK-ERK Signaling Pathway Inhibitor Triggers Synergistic Cytotoxicity in Multiple Myeloma Cells. PLoS One 2015; 10: e0143847.
Shaulian E, Karin M . AP-1 as a regulator of cell life and death. Nat Cell Biol 2002; 4: E131–E136.
Passegué E, Jochum W, Schorpp-Kistner M, Möhle-Steinlein U, Wagner EF . Chronic myeloid leukemia with increased granulocyte progenitors in mice lacking junB expression in the myeloid lineage. Cell 2001; 104: 21–32.
Ott RG, Simma O, Kollmann K, Weisz E, Zebedin EM, Schorpp-Kistner M et al. JunB is a gatekeeper for B-lymphoid leukemia. Oncogene 2007; 26: 4863–4871.
Kanno T, Kamba T, Yamasaki T, Shibasaki N, Saito R, Terada N et al. JunB promotes cell invasion and angiogenesis in VHL-defective renal cell carcinoma. Oncogene 2012; 31: 3098–3110.
Rassidakis GZ, Thomaides A, Atwell C, Ford R, Jones D, Claret F-X et al. JunB expression is a common feature of CD30+ lymphomas and lymphomatoid papulosis. Mod Pathol 2005; 18: 1365–1370.
Mao X, Orchard G, Lillington DM, Russell-Jones R, Young BD, Whittaker SJ . Amplification and overexpression of JUNB is associated with primary cutaneous T-cell lymphomas. Blood 2003; 101: 1513–1519.
Mao X, Orchard G, Lillington DM, Child FJ, Vonderheid EC, Nowell PC et al. BCL2 and JUNB abnormalities in primary cutaneous lymphomas. Br J Dermatol 2004; 151: 546–556.
Choo KB, Huang CJ, Chen CM, Han CP, Au LC . Jun-B oncogene aberrations in cervical cancer cell lines. Cancer Lett 1995; 93: 249–253.
Bamberger AM, Milde-Langosch K, Rössing E, Goemann C, Löning T . Expression pattern of the AP-1 family in endometrial cancer: correlations with cell cycle regulators. J Cancer Res Clin Oncol 2001; 127: 545–550.
Wang H, Birkenbach M, Hart J . Expression of Jun family members in human colorectal adenocarcinoma. Carcinogenesis 2000; 21: 1313–1317.
Halazonetis TD, Georgopoulos K, Greenberg ME, Leder P . c-Jun dimerizes with itself and with c-Fos, forming complexes of different DNA binding affinities. Cell 1988; 55: 917–924.
Angel P, Imagawa M, Chiu R, Stein B, Imbra RJ, Rahmsdorf HJ et al. Phorbol ester-inducible genes contain a common cis element recognized by a TPA-modulated trans-acting factor. Cell 1987; 49: 729–739.
Chinenov Y, Kerppola TK . Close encounters of many kinds: Fos-Jun interactions that mediate transcription regulatory specificity. Oncogene 2001; 20: 2438–2452.
Macián F, López-Rodríguez C, Rao A . Partners in transcription: NFAT and AP-1. Oncogene 2001; 20: 2476–2489.
van Dam H, Castellazzi M . Distinct roles of Jun: Fos and Jun: ATF dimers in oncogenesis. Oncogene 2001; 20: 2453–2464.
Johannessen CM, Johnson LA, Piccioni F, Townes A, Frederick DT, Donahue MK et al. A melanocyte lineage program confers resistance to MAP kinase pathway inhibition. Nature 2013; 504: 138–142.
Chauhan D, Li G, Hideshima T, Podar K, Mitsiades C, Mitsiades N et al. Hsp27 inhibits release of mitochondrial protein Smac in multiple myeloma cells and confers dexamethasone resistance. Blood 2003; 102: 3379–3386.
Tian Z, D’Arcy P, Wang X, Ray A, Tai Y-T, Hu Y et al. A novel small molecule inhibitor of deubiquitylating enzyme USP14 and UCHL5 induces apoptosis in multiple myeloma and overcomes bortezomib resistance. Blood 2014; 123: 706–716.
Chauhan D, Tian Z, Nicholson B, Kumar KGS, Zhou B, Carrasco R et al. A small molecule inhibitor of ubiquitin-specific protease-7 induces apoptosis in multiple myeloma cells and overcomes bortezomib resistance. Cancer Cell 2012; 22: 345–358.
Bhagwat AS, Vakoc CR . Targeting transcription factors in cancer. Trends Cancer 2015; 1: 53–65.
Chesi M, Bergsagel PL . Advances in the pathogenesis and diagnosis of multiple myeloma. Int J Lab Hematol 2015; 37 (Suppl 1): 108–114.
Yeh JE, Toniolo PA, Frank DA . Targeting transcription factors: promising new strategies for cancer therapy. Curr Opin Oncol 2013; 25: 652–658.
Delmore JE, Issa GC, Lemieux ME, Rahl PB, Shi J, Jacobs HM et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 2011; 146: 904–917.
Ria R, Catacchio I, Berardi S, De Luisi A, Caivano A, Piccoli C et al. HIF-1α of bone marrow endothelial cells implies relapse and drug resistance in patients with multiple myeloma and may act as a therapeutic target. Clin Cancer Res 2014; 20: 847–858.
Borsi E, Terragna C, Brioli A, Tacchetti P, Martello M, Cavo M . Therapeutic targeting of hypoxia and hypoxia-inducible factor 1 alpha in multiple myeloma. Transl Res 2015; 165: 641–650.
Fulciniti M, Amin S, Nanjappa P, Rodig S, Prabhala R, Li C et al. Significant biological role of sp1 transactivation in multiple myeloma. Clin Cancer Res 2011; 17: 6500–6509.
Ye N, Ding Y, Wild C, Shen Q, Zhou J . Small molecule inhibitors targeting activator protein 1 (AP-1). J Med Chem 2014; 57: 6930–6948.
Acknowledgements
We thank the microarray unit of the German Cancer Research Center (DKFZ) Genomics and Proteomics Core Facility for providing the Illumina Whole-Genome Expression Beadchips and related services. We thank Dr Peter Angel, Dr Marina Schorpp-Kistner from DKFZ as well as Dr Jochen Hess from University Heidelberg and DKFZ for providing the luminometer; and Haifan Huang, Jian Xu and Yingying Liang from the Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan, Hubei, China, for technical help. This work was supported by B Braun Stiftungs-Grant (to KP). MHB is the recipient of the DAAD-Indonesian German Scholarship Programme for doctoral fellowship. GT is supported by the Associazione Italiana per la Ricerca sul Cancro (AIRC; Investigator Grants and Special Program Molecular Clinical Oncology, 5 per mille no. 9965). AS and DH are supported by grants from the Deutsche Forschungsgemeinschaft, SFB/TRR79 (Bonn, Germany), and the EU 7th framework program (OverMyR). CRB and HG are supported by the German Research Council (DFG; KFO 227), the Baden-Württemberg Stiftung and the DFG Collaborative Research Center 873. HG is supported by the Dietmar Hopp Stiftung, the German Ministry of Education and Science and the DFG. EFW is funded by the Banco Bilbao Vizcaya Argentaria (BBVA) Foundation and a European Research Council Advanced Grant (ERC FCK/2008/37). PT is the recipient of the Italian Association for Cancer Research (AIRC) ‘Special Program Molecular Clinical Oncology - 5 per mille’ no. 9980, 2010/15. DJ is supported by the Heidelberg University Association Grant, DKFZ—HIPO H034, Dietmar Hopp Stiftung and the DFG.
Author contributions
FF conceived and designed the experiments, performed experiments, analyzed data and wrote the manuscript. MHB performed experiments and analyzed data. EM performed in vivo experiments. GT participated in data analysis and interpretation. SM, SV and MJ performed experiments. AS, DH, LB, CS and YH contributed essential experimental tools, participated in designing experiments and interpreted data. CRB, HG, MS, HG, EFW, PT and DJ participated in conceiving the study and in the interpretation of data. KP conceived the study, designed experiments, analyzed and interpreted data and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Fan, F., Bashari, M., Morelli, E. et al. The AP-1 transcription factor JunB is essential for multiple myeloma cell proliferation and drug resistance in the bone marrow microenvironment. Leukemia 31, 1570–1581 (2017). https://doi.org/10.1038/leu.2016.358
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2016.358
- Springer Nature Limited
This article is cited by
-
Dual therapeutic targeting of MYC and JUNB transcriptional programs for enhanced anti-myeloma activity
Blood Cancer Journal (2024)
-
SH3RF2 contributes to cisplatin resistance in ovarian cancer cells by promoting RBPMS degradation
Communications Biology (2024)
-
Bone marrow stromal cells induce chromatin remodeling in multiple myeloma cells leading to transcriptional changes
Nature Communications (2024)
-
Distinct transcriptomes and autocrine cytokines underpin maturation and survival of antibody-secreting cells in systemic lupus erythematosus
Nature Communications (2024)
-
JUNB affects hair follicle development and regeneration by promoting the proliferation of dermal papilla cells in goat
Chemical and Biological Technologies in Agriculture (2023)