Abstract
The scientific literature is full of articles discussing poor reproducibility of findings from animal experiments as well as failures to translate results from preclinical animal studies to clinical trials in humans. Critics even go so far as to talk about a “reproducibility crisis” in the life sciences, a novel headword that increasingly finds its way into numerous high-impact journals. Viewed from a cynical perspective, Fett's law of the lab “Never replicate a successful experiment” has thus taken on a completely new meaning. So far, poor reproducibility and translational failures in animal experimentation have mostly been attributed to biased animal data, methodological pitfalls, current publication ethics and animal welfare constraints. More recently, the concept of standardization has also been identified as a potential source of these problems. By reducing within-experiment variation, rigorous standardization regimes limit the inference to the specific experimental conditions. In this way, however, individual phenotypic plasticity is largely neglected, resulting in statistically significant but possibly irrelevant findings that are not reproducible under slightly different conditions. By contrast, systematic heterogenization has been proposed as a concept to improve representativeness of study populations, contributing to improved external validity and hence improved reproducibility. While some first heterogenization studies are indeed very promising, it is still not clear how this approach can be transferred into practice in a logistically feasible and effective way. Thus, further research is needed to explore different heterogenization strategies as well as alternative routes toward better reproducibility in animal experimentation.
Similar content being viewed by others
Main
In October 2013, the biomedical research community was startled by the latest issue of The Economist running the headline “How science goes wrong”. In a short briefing, modern scientists were accused of doing “too much trusting and not enough verifying”, followed by a list of problems, pitfalls, and mistakes that currently limit the validity and reproducibility of research findings, mostly in the context of animal-based research1. However, this is not the first time that criticism has been expressed about common practices in the field of biomedical research. Already in 2005, John Ioannidis published a paper provocatively entitled “Why most published research findings are false”2, in which he pointed out that most studies are more likely to report a false finding than a true one. So, what is behind this criticism? Are these just alarming claims, or are there indeed problems with the translational value, the validity, and/or the reproducibility of research findings?
How self-correcting is science?
The scientific literature of animal-based research is indeed full of publications reporting or discussing poor reproducibility, as well as failures to translate results from preclinical animal experiments to clinical trials in humans3,4,5,6,7,8,9. In a 10-year review of drug development, for example, Kola and Landis pointed out that the success rate from first-in-man to registration for different therapeutic areas between 1991 and 2000 was on average 11%, indicating that only one in nine compounds made it through the complete development process and were approved by the regulatory authorities4. Notably, the success rate was even worse for trials in specific research areas, such as oncology or women's health4. Similarly, in a systematic evaluation of how well mice mimic human inflammatory responses, fundamental disparities in genomic responses between mice and men were detected. Among genes that changed significantly in humans, the murine orthologs were close to random in matching their human counterparts, questioning the translational value of current mouse models for severe inflammation10. Thus, despite the overall and widely recognized improvement in scientific and technological tools over the last years, novel compounds have been criticized to fail more often in clinical development today than in the 1970s11.
However, translational failures are not the only challenge the scientific community has to face. There is also an increasing concern about the rate at which published findings are reproducible. The current debate even goes so far as to generate a novel headword, i.e. “reproducibility crisis”, that increasingly finds its way in to numerous high-impact journals11,12,13,14,15. Against this background, it is not surprising that 90% of 1,576 interviewed life scientists believe that they are currently facing either a slight or a significant reproducibility crisis12.
By definition, “reproducibility” refers to the degree of accordance between results of the same experiment performed independently in the same or in a different laboratory16 (Table 1). Results that cannot be reproduced cast serious doubts on the quality of experiments and hinder scientific progress. In the context of animal experimentation, poor reproducibility is also an ethical issue, as the need for additional follow-up studies undermines the aim of reducing animal use. In that respect, it is an extremely serious matter that reproducibility problems seem to be most prevalent in those research areas that work with animal model systems17,18, although recent surveys indicate that they also occur in other fields, such as psychology, chemistry and physics12,19,20. Current estimates for irreproducibility in biomedical research are alarmingly high, ranging from 50 to 90%15,21. Begley and Ellis, for example, reported that only 6 out of 53 “landmark studies” in oncology could be replicated22, and Prinz and colleagues detected inconsistencies in 75 to 80% of 67 in-house projects in oncology, women's health and cardiovascular diseases18. From an economic perspective, these high irreproducibility rates have been associated with costs of approximately US$28 billion per year in the United States alone21. As indicated by these examples, basic science has lost a great deal of credibility over the last years, emphasizing the need for fundamental changes in the conduct and analysis of experiments. However, the causes of current limitations to translation and reproducibility need to be identified first, before changes can be adequately addressed.
Threats to translation and reproducibility
Besides possible shortcomings in the clinical trials that may contribute to high attrition rates, translational failures have been attributed to biased research approaches, overoptimistic conclusions, or the lack of external validity in preclinical studies5,23 (Table 1). Similarly, to explain poor reproducibility in animal experimentation, most explanatory approaches have concentrated on methodological issues, such as the inadequate choice of experimental designs and control groups or different types of biases5,24,25,26 (Table 1). Knowledge of treatment assignment, for example, may consciously or unconsciously affect the outcome assessment, a phenomenon recognized for the first time at the beginning of the 20th century. Here, a horse named Hans drew worldwide attention as the first animal with “numeracy skills”. By tapping its hoof, the horse seemed to solve arithmetic operations, read the clock, or recognize playing cards. A few years later, however, it turned out that the horse was only able to respond correctly to these tasks in the presence of the questioning person. If this person was absent or did not know the answer, the horse suddenly seemed to lose these skills. Thus, instead of being able to solve math problems, the horse was simply receptive to subtle cues present in the human questioners. Today, known as “Clever Hans Effect” or “Experimenter Bias”, this simple example illustrates how non-conscious cues from experimenters can introduce bias into testing. Similarly, so-called “Selection Biases” (i.e., biased allocation to treatment groups) may lead to selective exclusion or inclusion of animals to treatment groups, resulting in systematic differences in the baseline characteristics between groups3,5.
Steps can be taken to reduce the risk of bias. But, where risks of bias have been systematically assessed in reviews of in vivo studies, an alarmingly low reporting rate of measures against risks of bias has been found. For example, a systematic review of studies reporting on functional outcome in animal models of acute ischemia found that random treatment allocation was reported in only 42% of the studies, blinded administration of the treatment in 22%, and blinded assessment of outcome in 40% (ref. 27). Similarly, a meta-analysis published in 2015 revealed that out of 2,671 publications reporting drug efficacy in eight different disease models, randomization was reported in only 662 publications (24.8%), blinded assessment of outcome in 788 (29.5%), and a sample size calculation in 20 cases (0.7%) (ref. 25). Notably, reporting rates of such quality criteria are not only low at the publication level, but also at the level of applications for animal experiments, (i.e., before the studies have been conducted). A recent meta-analysis published in PLoS Biology indicated that out of 1,277 applications for animal experiments in Switzerland, only 3.2% included a statement about blinding, 12.6% about randomization, and 7.9% about a sample size calculation28. Reporting guidelines have therefore become a major tool in overcoming risks of bias29.
As one important step toward improved reporting standards in animal experimentation, the ARRIVE guidelines (Animal Research: Reporting of In Vivo Experiments) have been introduced in 2010 (refs. 30,31). Based on a 20-item checklist of information to be reported in publications, the guidelines aim at maximizing the availability and utility of the information gained from every animal and every experiment. However, although the ARRIVE guidelines have been endorsed by over 1,000 journals since their introduction, little improvement in reporting standards has been observed28,32. Nevertheless, overall awareness seems to have risen, as Macleod and colleagues showed that reporting rates in at least specific research areas in the biomedical sciences have increased over time25.
Poor reproducibility has also been linked to manifold failures in the statistical analyses and the choice of the experimental unit (i.e., the smallest physical unit that can be randomly assigned to a treatment condition)33,34,35,36. If, for example, a pregnant female animal is subjected to an experimental treatment, but the scientific interest is in the individual offspring, analyses are often based on individual pups37. Because pups within a litter represent highly dependent entities, treating each pup as an independent experimental unit results in artificially large sample sizes associated with a substantial inflation of the nominal 0.05 alpha level38. In fact, simulation studies have shown that an increase of the sample size by treating two pups per litter as independent measurements can almost triple the nominal 0.05 alpha level39. Referred to as “litter or cage effects”, such misconceptions contribute to an overrepresentation of false positives in the scientific literature and, hence, hamper reproducibility33,40. Similar problems arise in multiple testing situations. If more than one statistical test is performed on a given data set, the chance of drawing at least one false conclusion increases rapidly with the number of tests applied. Thus, failures to control the familywise error rate and adjust the p-value adequately also contribute to poor reproducibility by increasing the chance of producing and publishing false positive findings41.
Apart from these methodological issues, animal welfare constraints, poor training of researchers in experimental design and conduct17,42, as well as current publication ethics5,43,44, have all been linked to poor reproducibility and translational failures in animal experimentation. In particular the first point has been aptly summarized at a very early stage by Trevor Poole with his article “Happy animals make good science”45. The idea here is that it is not only better for the animal to be in a good welfare state, but also for the quality of the scientific results derived from experiments with animals of “normal” behavior and physiology46,47. Furthermore, current teaching standards have been criticized, because only few scientists get formal training in experimental techniques and statistical analyses. Many experiments are therefore planned and conducted on the basis of “lab traditions” rather than “good laboratory practices”, resulting in wrong and irreproducible findings42. Regarding publication ethics, key words like “publication bias”, “selective reporting”, or “p-hacking” have dominated the debate48,49. Particularly, the overweighting of positive results leads to a subsequent overestimation of effect sizes in meta-analyses and systematic reviews. Sena and colleagues, for example, came to the conclusion that publication bias accounts for one third of the effect observed in animal stroke studies43, and Simmons and colleagues argued that even before publication, researchers are remarkably adept at reaching those conclusions that mesh with their desires50. Current publication standards that emphasize positive results are thus likely to further exacerbate the reproducibility problems described in the life sciences.
Standardization in animal experimentation: necessity or fallacy?
More recently, the concept of standardization has gained attention as an additional source of irreproducible findings, especially in the context of behavioral phenotyping studies13. Standardization within experiments aims at reducing variation in the data, thereby increasing test sensitivity and reducing animal use51. Furthermore, standardization between experiments aims at reducing between-experiment variation, thereby improving the comparability and reproducibility of results between studies52. In light of the reproducibility crisis, however, the question arises, whether standardization really is a prerequisite for good reproducibility.
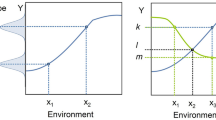
In practice, the concept of standardization has led to rigorous homogenization of the animals' genotype (for example, by inbreeding), the laboratory environment (for example, by using uniform cage enrichment), the daily routines (for example, by standardized handling procedures), and the test situation (for example, by defining the time of testing). The idea here is to isolate the variables of interest, minimize the background noise, and maximize the detection of even subtle treatment effects. However, while this approach may indeed allow for exploring condition-restricted hits effectively, it can reduce information gain at the same time. Fully effective homogenization would thus decrease inter-individual variation within a study population to zero, leading to statistically significant, but possibly irrelevant results that lack generalizability to slightly different conditions (referred to as the “standardization fallacy”53,54, (Fig. 1).
Usually, an animal experiment is conducted in a specific laboratory that narrowly standardizes the environmental conditions to an arbitrary local constellation of environmental factors (indicated by the dashed lines and the colored background). Because experimental treatments (indicated by the solid lines) can interact with some known or unknown environmental background variables, standardization to a specific environmental window leads to significant, but possibly irrelevant findings that cannot be reproduced in a second laboratory that standardizes the conditions to slightly different environmental conditions. By reducing within-experiment variation, standardization thus limits the inference to the specific environmental conditions, thereby counteracting the concept of phenotypic plasticity.
Ironically, this standardization fallacy can be best demonstrated by poor reproducibility in the scientific literature. In a groundbreaking study involving three different laboratories, Crabbe and colleagues conducted a series of common behavioral tests in eight different mouse strains that were delivered, housed, reared, handled and tested under highly homogenized conditions. Notwithstanding this extreme level of standardization between facilities, the authors observed interactions between genotype and laboratory (i.e., genetically identical mice behaved differently depending on site). Based on these results, the authors hypothesized that “experiments characterizing mutants may yield results that are idiosyncratic to a particular laboratory”55. Subsequently to these initial findings, several other multi-laboratory studies confirmed difficulties in reproducing behavioral strain differences across labs56,57,58, clearly showing that reproducibility problems arise despite rigorous standardization regimes. Because different laboratories inevitably standardize to different local constellations of experimental conditions (i.e., many factors, such as the experimenter, room architecture, or daily routines, cannot be standardized between laboratories), within-laboratory standardization will always exceed between-laboratory standardization. It is therefore not surprising that increasingly rigorous standardization within labs produces results that are increasingly distinct between laboratories (Fig. 1).
Previous approaches to solve the issue of poor reproducibility in behavioral phenotyping studies have focused on the search for tests yielding robust results across experiments and laboratories59,60. Suggestions for improving the situation range from establishing one “golden standard” test for each domain, to using a battery of tests all believed to measure the same construct to assess the robustness of measures61. Other approaches have focused on the experimenter as a major source of experimental noise, leading to the development of human-free testing environments, such as the SmartCube, the IntelliCage62 or modern touchscreen-based procedures63,64. Automated testing minimizes the need for human intervention and its accompanying stress and is therefore believed to reduce inter-individual variation65. However, because home cage testing is still under way, currently existing test systems are often too complex for high-throughput approaches.
Systematic heterogenization rather than rigorous standardization?
A central fact in biology is that living organisms do vary. Such phenotypic plasticity relies on complex gene-by-environment interactions that shape the individual phenotype. With the aim of reducing such variation, standardization neglects individual phenotypic plasticity, thereby counteracting the widely adopted idea of “reaction norms”13 (i.e., pattern of phenotypes produced by a given genotype under different environmental conditions66,67). Instead of spiriting this biological variation away, however, inter-individual differences may be key to making study populations more representative13. Thus, it may be advantageous, rather than detrimental, to use samples varied across genetic and/or environmental conditions to increase the external validity of the results, and improve the reproducibility of research findings. So, how can this logic be transferred into practice?
Common practice to identify idiosyncratic results is to run independent replicate experiments16,68. Ideally, a replicate experiment is not a mere repetition of the original experiment, but should extend the scope by varying a particular set of factors16. If a replication study then fails to confirm the results, either the replicate study, the original study (or both) may have produced false or spurious results of limited external validity69. Although this method indeed provides information on the robustness of a finding, it raises practical and ethical questions, because the need for replicate studies may easily inflate the number of animals needed to confirm a “true” effect. It thus seems to be preferable to incorporate such a “robustness check” directly in the experimental design. In this respect, the concept of “systematic heterogenization” has been proposed to be a powerful tool to extract robust and hence reproducible findings in animal experiments70,71,72.
The underlying idea of systematic heterogenization is to introduce variation systematically into a single experiment to make study populations more representative and findings more robust across the variation that inevitably exists between experiments. In line with this idea, a recent simulation study revealed greater variation of treatment effects between different single-laboratory studies in comparison to different multi-laboratory studies. Furthermore, reproducibility was improved from less than 50% to over 80% in studies involving as few as three labs7. These findings clearly indicate that the inevitable increase in environmental variation in a multi-laboratory situation benefits the external validity and hence the reproducibility of treatment effects. Since it is unlikely, however, that all single experiments will be replaced by multi-laboratory approaches in the near future, systematic heterogenization aims at transferring this logic to a single-lab situation by increasing the variation within each single experiment.
Including variation in a non-systematic and uncontrolled way may bear the risk of inflating the number of animals needed for each experiment. It is thus important to combine the approach with adequate analytical techniques and experimental designs, such as split-plot, factorial, or randomized block designs that control for the introduced variation without reducing test sensitivity and statistical power70,73,74. The potential value of split-plot designs, for example, has recently been demonstrated by a study investigating mixed-strain housing. Co-housing individuals of different strains increased the external validity of the experiment, without exerting negative effects on the data variability and the statistical power74. Originally, these designs derived from agricultural research, where the experimental area was divided into heterogeneous blocks of land73. However, such techniques are likely to have a much wider applicability in laboratory animal science, since they allow combining animals of, for example, different ages, batches, strains, litters, cages, or environmental conditions within a single experiment75.
Overall, such systematic and controlled forms of heterogenization may replace or at least complement many of the conventionally used standardized approaches in animal experimentation. While in some cases a highly standardized experiment can be useful to identify single condition-restricted treatment effects, a heterogenized approach may help to detect more universally applicable conclusions. The best approach, however, clearly depends on the specific research goal. From an animal-ethical point of view, systematic heterogenization contributes to the refinement and reduction of animal experimentation by either reducing the number of experiments needed to detect a meaningful result or by increasing information gained based on the same number of animals.
Toward better reproducibility: applying a heterogenization strategy
Some first heterogenization studies are indeed promising. In a series of three experiments, standardization was found to increase the incidence of spurious results in behavioral tests, accounting for poor reproducibility, while systematic heterogenization attenuated spurious results, thereby improving reproducibility71,72,76. Here, systematic heterogenization was achieved by varying two defined environmental factors that are known to interact with mouse genotype: (1) cage enrichment and (2) test age. According to a 2 × 2 factorial design, each factor was varied across two factor levels A and B, resulting in four different factor combinations (1A, 1B, 2A, 2B). Each heterogenized experiment was thus composed of mice that were kept and tested in four different ways.
Interestingly, this simple form of systematic environmental variation was sufficient to guarantee almost perfect reproducibility of behavioral strain differences between replicate experiments within a single laboratory72 (but see also refs. 77,78,79). Between laboratories, however, the observed improvement was not as strong as in the single-lab situation76. Although heterogenization improved reproducibility compared to standardization, differences in the size and direction of strain effects occurred in both experimental approaches. Thus, despite the increasing awareness of reproducibility problems, the experimental design of animal experiments is still in need of refinement. While the strict homogenization of experimental conditions obviously does not cure poor reproducibility and translational failures, it is still not clear which types of systematic heterogenization may improve the situation and how this approach can be transferred into practice
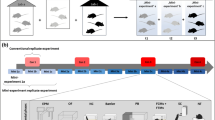
Richard Paylor suggested splitting experiments into small batches of animals that are tested some time apart instead of testing them all at once in just one large batch80. The underlying idea here is very close to the proposed concept of systematic heterogenization. Because each single small batch is supposed to rely on a unique time-dependent constellation of environmental and testing conditions, combining several “mini-experiments” in one big experiment is assumed to mimic a multi-laboratory situation within a single experiment and therefore to result in findings that are more robust. The approach reflects a kind of “systematic heterogenization over time” or “batch heterogenization” (Fig. 2). This is also in line with findings from computational approaches that have identified and ranked sources of variability in nociceptive responses in mice, showing that both season and time of day greatly influences the outcome measures81,82. Similarly, Karp and colleagues conducted an analysis of data from phenotyping studies, showing that batch (i.e., the time point of testing) explains about a quarter of the observed variation in mouse phenotypes83.
Batch heterogenization aims to split experiments into small batches of animals that are tested some time apart (heterogenized design) instead of testing them at once in just one large batch (standardized design). Combining these “mini-experiments” in one big experiment is then assumed to increase representativeness of the whole study population, resulting in findings that are more reproducible between experiments and laboratories.
Furthermore, the experimenter has been shown to be one of the most important factors influencing the outcome of an experiment57,81,82. Precisely what differentiates the experimenters between studies remains unknown, but recent work has shown that even the gender of the experimenter can affect baseline responses in behavioral testing to a significant extent84. Involving multiple experimenters for testing, instead of using only one, may suffice to make the study populations more representative and therefore less prone to variation between studies. Alternatively, genetic rather than environmental variation may represent a promising strategy to increase the external validity and the reproducibility of research findings. Because genetic background has been found to strongly modulate mutant phenotypes85, the systematic variation of different strains or genotypes within a single study may also contribute to increased generalizability. Testing these different strategies will reveal whether such minor variations are indeed sufficient to significantly improve the reproducibility of research findings, especially in the context of behavioral phenotyping studies3.
Conclusions
Poor reproducibility and translational failures in animal experimentation can be attributed to deficiencies on many different levels. While most researchers have linked these problems to poor experimental design and conduct, poor reporting standards, and animal welfare constraints, they may also result from strict homogenization regimes that are widely practiced in biomedical research. Instead, a systematically heterogenized experimental approach that takes biological variation into account might help to improve representativeness of study populations and contribute to improved external validity and reproducibility of research findings. However, despite some first efforts toward heterogenized experimental strategies, there is still no “golden solution” for the conduct of single laboratory experiments, highlighting the need for further improvement strategies and innovative research approaches.
References
Unreliable research. Trouble at the lab. The Economist (2013).
Ioannidis, J.P. Why most published research findings are false. PLoS Med. 2, e124 (2005).
Bailoo, J.D., Reichlin, T.S. & Würbel, H. Refinement of experimental design and conduct in laboratory animal research. ILAR J. 55, 383–391 (2014).
Kola, I. & Landis, J. Can the pharmaceutical industry reduce attrition rates? Nat. Rev. Drug Discov. 3, 711–716 (2004).
Van der Worp, H.B. et al. Can animal models of disease reliably inform human studies? PLoS Med. 7, e1000245 (2010).
Mogil, J.S. Laboratory environmental factors and pain behavior: the relevance of unknown unknowns to reproducibility and translation. Lab Anim. (NY) 46, 136–141 (2017).
Würbel, H. More than 3Rs: the importance of scientific validity for harm-benefit analysis of animal research. Lab Anim. (NY) 46, 164–166 (2017).
Garner, J.P., Gaskill, B.N., Weber, E.M., Ahloy-Dallaire, J. & Pritchett-Corning, K.R. Introducing Therioepistemology: the study of how knowledge is gained from animal research. Lab Anim. (NY) 46, 103–113 (2017).
Jarvis, M.F. & Williams, M. Irreproducibility in preclinical biomedical research: perceptions, uncertainties, and knowledge gaps. Trends Pharmacol. Sci. 37, 290–302 (2016).
Seok, J. et al. Genomic responses in mouse models poorly mimic human inflammatory diseases. Proc. Natl. Acad. Sci. USA 110, 3507–3512 (2013).
Scannell, J.W. & Bosley, J. When quality beats quantity: decision theory, drug discovery, and the reproducibility crisis. PLoS ONE 11, e0147215 (2016).
Baker, M. 1,500 scientists lift the lid on reproducibility. Nature 533, 452–454 (2016).
Voelkl, B. & Würbel, H. Reproducibility crisis: are we ignoring reaction norms? Trends Pharmacol. Sci. 37, 509–510 (2016).
Peng, R. The reproducibility crisis in science: A statistical counterattack. Significance 12, 30–32 (2015).
Begley, C.G. & Ioannidis, J.P. Reproducibility in science. Circ. Res. 116, 116–126 (2015).
van der Staay, F.J., Arndt, S.S. & Nordquist, R.E. Evaluation of animal models of neurobehavioral disorders. Behav. Brain Funct. 5, 11 (2009).
Collins, F.S. & Tabak, L.A. Policy: NIH plans to enhance reproducibility. Nature 505, 612–613 (2014).
Prinz, F., Schlange, T. & Asadullah, K. Believe it or not: how much can we rely on published data on potential drug targets? Nat. Rev. Drug Discov. 10, 712 (2011).
Open Science Collaboration. Estimating the reproducibility of psychological science. Science 349, aac4716 (2015).
Ioannidis, J.P. et al. Repeatability of published microarray gene expression analyses. Nat. Genet. 41, 149–155 (2009).
Freedman, L.P., Cockburn, I.M. & Simcoe, T.S. The economics of reproducibility in preclinical research. PLoS Biol. 13, e1002165 (2015).
Begley, C.G. & Ellis, L.M. Raise standards for preclinical cancer research. Nature 483, 531–533 (2012).
Giles, J. Animal experiments under fire for poor design. Nature 444, 981 (2006).
Ioannidis, J.P. et al. Increasing value and reducing waste in research design, conduct, and analysis. Lancet 383, 166–175 (2014).
Macleod, M.R. et al. Risk of bias in reports of in vivo research: a focus for improvement. PLoS Biol. 13, e1002273 (2015).
Reichlin, T.S., Vogt, L. & Würbel, H. The researchers' view of scientific rigor—survey on the conduct and reporting of in vivo research. PLoS ONE 11, e0165999 (2016).
van der Worp, H.B., de Haan, P., Morrema, E. & Kalkman, C.J. Methodological quality of animal studies on neuroprotection in focal cerebral ischaemia. J. Neurol. 252, 1108–1114 (2005).
Vogt, L., Reichlin, T.S., Nathues, C. & Würbel, H. Authorization of animal experiments is based on confidence rather than evidence of scientific rigor. PLoS Biol. 14, e2000598 (2016).
McNutt, M. Journals unite for reproducibility. Science 346, 679 (2014).
Kilkenny, C., Browne, W., Cuthill, I.C., Emerson, M. & Altman, D.G. Animal research: reporting in vivo experiments: the ARRIVE guidelines. Br. J. Pharmacol. 160, 1577–1579 (2010).
Kilkenny, C., Browne, W.J., Cuthill, I.C., Emerson, M. & Altman, D.G. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. PLoS Biol. 8, e1000412 (2010).
Baker, D., Lidster, K., Sottomayor, A. & Amor, S. Two years later: journals are not yet enforcing the ARRIVE guidelines on reporting standards for pre-clinical animal studies. PLoS Biol. 12, e1001756 (2014).
Lazic, S.E. & Essioux, L. Improving basic and translational science by accounting for litter-to-litter variation in animal models. BMC Neurosci. 14, 37 (2013).
Festing, M.F. Design and statistical methods in studies using animal models of development. ILAR J. 47, 5–14 (2006).
Halsey, L.G., Curran-Everett, D., Vowler, S.L. & Drummond, G.B. The fickle P value generates irreproducible results. Nat. Methods 12, 179–185 (2015).
Goodman, S.N. Aligning statistical and scientific reasoning. Science 352, 1180–1181 (2016).
Wainwright, P.E. Issues of design and analysis relating to the use of multiparous species in developmental nutritional studies. J. Nutr. 128, 661–663 (1998).
Zorrilla, E.P. Multiparous species present problems (and possibilities) to developmentalists. Dev. Psychobiol. 30, 141–150 (1997).
Holson, R.R. & Pearce, B. Principles and pitfalls in the analysis of prenatal treatment effects in multiparous species. Neurotoxicol. Teratol. 14, 221–228 (1992).
Lazic, S.E. The problem of pseudoreplication in neuroscientific studies: is it affecting your analysis? BMC Neurosci. 11, 5 (2010).
Noble, W.S. How does multiple testing correction work? Nat. Biotechnol. 27, 1135–1137 (2009).
Festing, M.F. We are not born knowing how to design and analyse scientific experiments. Altern. Lab. Anim. 41, 19–21 (2013).
Sena, E.S., Van Der Worp, H.B., Bath, P.M., Howells, D.W. & Macleod, M.R. Publication bias in reports of animal stroke studies leads to major overstatement of efficacy. PLoS Biol. 8, e1000344 (2010).
Cumming, G. The new statistics why and how. Psychol. Sci. 25, 7–29 (2014).
Poole, T. Happy animals make good science. Lab. Anim. 31, 116–124 (1997).
Garner, J.P. Stereotypies and other abnormal repetitive behaviors: potential impact on validity, reliability, and replicability of scientific outcomes. ILAR J. 46, 106–117 (2005).
Prescott, M.J. & Lidster, K. Improving quality of science through better animal welfare: the NC3Rs strategy. Lab Anim. (NY) 46, 152–156 (2017).
Nuzzo, R. Statistical errors. Nature 506, 150 (2014).
Head, M.L., Holman, L., Lanfear, R., Kahn, A.T. & Jennions, M.D. The extent and consequences of p-hacking in science. PLoS Biol. 13, e1002106 (2015).
Simmons, J.P., Nelson, L.D. & Simonsohn, U. False-positive psychology: undisclosed flexibility in data collection and analysis allows presenting anything as significant. Psychol. Sci. 22, 1359–1366 (2011).
Festing, M.F. Reduction of animal use: experimental design and quality of experiments. Lab. Anim. 28, 212–221 (1994).
Beynen, A.C., Baumans, V. & Van Zutphen, L.F.M. in Principles of Laboratory Animal Science (eds. L.F.M. Van Zutphen, V. Baumans & A.C. Beynen) 103–110 (Elsevier, Amsterdam, 2001).
Würbel, H. Behaviour and the standardization fallacy. Nat. Genet. 26, 263 (2000).
Würbel, H. Behavioral phenotyping enhanced–beyond (environmental) standardization. Genes Brain Behav. 1, 3–8 (2002).
Crabbe, J.C., Wahlsten, D. & Dudek, B.C. Genetics of mouse behavior: interactions with laboratory environment. Science 284, 1670–1672 (1999).
Mandillo, S. et al. Reliability, robustness, and reproducibility in mouse behavioral phenotyping: a cross-laboratory study. Physiol. Genomics 34, 243–255 (2008).
Lewejohann, L. et al. Environmental bias? Effects of housing conditions, laboratory environment and experimenter on behavioral tests. Genes Brain Behav. 5, 64–72 (2006).
Wolfer, D.P. et al. Laboratory animal welfare: cage enrichment and mouse behaviour. Nature 432, 821–822 (2004).
Wahlsten, D. Standardizing tests of mouse behavior: reasons, recommendations, and reality. Physiol. Behav. 73, 695–704 (2001).
Wahlsten, D. et al. Different data from different labs: lessons from studies of gene–environment interaction. J. Neurobiol. 54, 283–311 (2003).
Crabbe, J.C. & Morris, R.G. Festina lente: late-night thoughts on high-throughput screening of mouse behavior. Nat. Neurosci. 7, 1175–1179 (2004).
Galsworthy, M.J. et al. A comparison of wild-caught wood mice and bank voles in the Intellicage: assessing exploration, daily activity patterns and place learning paradigms. Behav. Brain Res. 157, 211–217 (2005).
Talpos, J. & Steckler, T. Touching on translation. Cell Tissue Res. 354, 297–308 (2013).
Richter, S.H. et al. Touchscreen-paradigm for mice reveals cross-species evidence for an antagonistic relationship of cognitive flexibility and stability. Front. Behav. Neurosci. 8, 154 (2014).
Richardson, C.A. Automated homecage behavioural analysis and the implementation of the three Rs in research involving mice. Altern. Lab. Anim. 40, 7–9 (2012).
Dingemanse, N.J., Kazem, A.J., Réale, D. & Wright, J. Behavioural reaction norms: animal personality meets individual plasticity. Trends Ecol. Evol. 25, 81–89 (2010).
Sarkar, S. From the Reaktionsnorm to the adaptive norm: the norm of reaction, 1909–1960. Biol. Philos. 14, 235–252 (1999).
van der Staay, F.J. Animal models of behavioral dysfunctions: basic concepts and classifications, and an evaluation strategy. Brain Res. Rev. 52, 131–159 (2006).
Muma, J.R. The need for replication. J. Speech Lang. Hear. Res. 36, 927–930 (1993).
Würbel, H. & Garner, J.P. Refinement of rodent research through environmental enrichment and systematic randomization. NC3Rs 9, 1–9 (2007).
Richter, S.H., Garner, J.P. & Wurbel, H. Environmental standardization: cure or cause of poor reproducibility in animal experiments? Nat. Methods 6, 257–261 (2009).
Richter, S.H., Garner, J.P., Auer, C., Kunert, J. & Würbel, H. Systematic variation improves reproducibility of animal experiments. Nat. Methods 7, 167–168 (2010).
Grafen, A. & Hails, R. Modern statistics for the life sciences (Oxford University Press, Oxford, 2002).
Walker, M. et al. Mixed-strain housing for female C57BL/6, DBA/2, and BALB/c mice: validating a split-plot design that promotes refinement and reduction. BMC Med. Res. Methodol. 16, 11 (2016).
Festing, M.F. & Altman, D.G. Guidelines for the design and statistical analysis of experiments using laboratory animals. ILAR J. 43, 244–258 (2002).
Richter, S.H. et al. Effect of population heterogenization on the reproducibility of mouse behavior: a multi-laboratory study. PLoS ONE 6, e16461 (2011).
Würbel, H., Richter, S.H. & Garner, J.P. Reply to: “Reanalysis of Richter et al. (2010) on reproducibility”. Nat. Methods 10, 374 (2013).
Jonker, R.M., Guenther, A., Engqvist, L. & Schmoll, T. Does systematic variation improve the reproducibility of animal experiments? Nat. Methods 10, 373 (2013).
Wolfinger, R.D. Reanalysis of Richter et al. (2010) on reproducibility. Nat. Methods 10, 373–374 (2013).
Paylor, R. Questioning standardization in science. Nat. Methods 6, 253–254 (2009).
Chesler, E.J., Wilson, S.G., Lariviere, W.R., Rodriguez-Zas, S.L. & Mogil, J.S. Identification and ranking of genetic and laboratory environment factors influencing a behavioral trait, thermal nociception, via computational analysis of a large data archive. Neurosci. Biobehav. Rev. 26, 907–923 (2002).
Chesler, E.J., Wilson, S.G., Lariviere, W.R., Rodriguez-Zas, S.L. & Mogil, J.S. Influences of laboratory environment on behavior. Nat. Neurosci. 5, 1101–1102 (2002).
Karp, N.A., Melvin, D., Mott, R.F. & Project, S.M.G. Robust and sensitive analysis of mouse knockout phenotypes. PLoS ONE 7, e52410 (2012).
Sorge, R.E. et al. Olfactory exposure to males, including men, causes stress and related analgesia in rodents. Nat. Methods 11, 629–632 (2014).
Sittig, L.J. et al. Genetic background limits generalizability of genotype-phenotype relationships. Neuron 91, 1253–1259 (2016).
Acknowledgements
Current research on heterogenization and reproducibility is funded by the German Research Foundation (DFG, RI 2488/3-1). Furthermore, I would like to thank Norbert Sachser, Hanno Würbel, Sara Hintze, Niklas Kästner, and Vanessa von Kortzfleisch for their helpful comments on earlier drafts of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Rights and permissions
About this article
Cite this article
Richter, S. Systematic heterogenization for better reproducibility in animal experimentation. Lab Anim 46, 343–349 (2017). https://doi.org/10.1038/laban.1330
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/laban.1330
- Springer Nature America, Inc.
This article is cited by
-
Challenging current scientific practice: how a shift in research methodology could reduce animal use
Lab Animal (2024)
-
A paradigm shift in translational psychiatry through rodent neuroethology
Molecular Psychiatry (2023)
-
Predictive validity in drug discovery: what it is, why it matters and how to improve it
Nature Reviews Drug Discovery (2022)
-
Comparative Analysis of the Pharmacological Activity of Bis(3,5-di-Tert-Butyl-4-Hydroxyphenylthiolate)Dimethylol in Different Modes of Administration on a Mouse Model of Melanoma B16 Tumor Growth
Bulletin of Experimental Biology and Medicine (2022)
-
Mapping the past, present and future research landscape of paternal effects
BMC Biology (2020)