Abstract
Background:
Brown adipose tissue (BAT) thermogenesis is an adaptive process, essential for energy expenditure and involved in the control of obesity. Obesity is associated with abnormally increased autophagy in white adipose tissue. Autophagy has been proposed as relevant for brown-vs-white adipocyte differentiation; however, its role in the response of BAT to thermogenic activation is unknown.
Methods:
The effects of thermogenic activation on autophagy in BAT were analyzed in vivo by exposing mice to 24 h cold condition. The effects of norepinephrine (NE), cAMP and modulators of lysosomal activity were determined in differentiated brown adipocytes in the primary culture. Transcript expression was quantified by real-time PCR, and specific proteins were determined by immunoblot. Transmission electron microscopy, as well as confocal microscopy analysis after incubation with specific antibodies or reagents coupled to fluorescent emission, were performed in BAT and cultured brown adipocytes, respectively.
Results:
Autophagy is repressed in association with cold-induced thermogenic activation of BAT in mice. This effect was mimicked by NE action in brown adipocytes, acting mainly through a cAMP-dependent protein kinase A pathway. Inhibition of autophagy in brown adipocytes leads to an increase in UCP1 protein and uncoupled respiration, suggesting a repressing role for autophagy in relation to the activity of BAT thermogenic machinery. Under basal conditions, brown adipocytes show signs of active lipophagy, which is suppressed by a cAMP-mediated thermogenic stimulus.
Conclusions:
Our results show a noradrenergic-mediated inverse relationship between autophagy and thermogenic activity in BAT and point toward autophagy repression as a component of brown adipocyte adaptive mechanisms to activate thermogenesis.
Similar content being viewed by others
Introduction
Brown adipose tissue (BAT) is the main site of non-shivering thermogenesis in mammals. BAT activity in rodent models not only ensures appropriate adaptation to cold environment but also protects against obesity by promoting energy expenditure.1, 2 Recent studies have indicated that adult humans possess active BAT, contrary to the prevailing concept that the role of BAT in humans is restricted to neonates, and further showed that BAT activity is systematically reduced in obese patients.3, 4, 5
Brown adipocytes possess large amounts of mitochondria with a high oxidative capacity. Mitochondrial uncoupling protein-1 (UCP1), uniquely expressed in brown adipocytes, uncouples the respiratory chain from oxidative phosphorylation, thus leading to high oxidation rates and the use of metabolic energy to provide heat. Lipid stores in brown adipocytes, used for fueling mitochondrial oxidation, are stored as multilocular droplets that are in close proximity to mitochondria. To sustain the metabolic energy needs of heat production, brown adipocytes also possess strong enzymatic machinery for glucose and lipid uptake and oxidation.6
BAT is an extremely plastic tissue, in keeping with its need to adapt to changing thermal and dietary conditions. When thermogenesis is activated, existing brown adipocytes adapt by enhancing the amount of intracellular protein devoted to oxidative pathways, including UCP1 synthesis and mitochondrial biogenesis. Overall, sympathetic activation and noradrenergic stimuli appear to be the main mechanism involved in orchestrating BAT activation processes, although a growing number of non-adrenergic neuro-endocrine factors7 and regulatory molecules such as adenosine8 have also been identified.
Autophagy is a process that contributes to the maintenance of cellular homeostasis by removing damaged organelles and recycling intracellular substrates.9 Autophagy has an important role in cellular adaptive changes to physiological and pathological challenges, for example, providing substrates during nutrient deprivation, and degrading intracellular ectopic lipid accumulation; it also is involved in differentiation-associated remodeling events.9, 10, 11, 12 Macroautophagy (referred to hereafter as autophagy) involves the formation of double-membrane vesicles (autophagosomes) that engulf a portion of the cytosol and deliver the incorporated cargo to the lysosome for degradation.13
There is a close relationship between autophagic activity and energy metabolism.14, 15, 16 Complex intracellular pathways regulate autophagic activity by controlling different protein interactions, similar to those involving autophagy-related (Atg) protein conjugation cascades, as well as posttranslational protein modifications involved in autophagosome formation and substrate recognition. Recent data indicate that transcriptional regulation of genes encoding autophagy components also has a role in the regulation of autophagy by metabolic and endocrine signals.17 On the other hand, multiple studies are coincident in reporting that human obesity is associated with increased autophagy in white adipose tissue.14, 18, 19
Some reports have indicated that autophagy may have a role in regulating brown adipocyte differentiation. Impairment of autophagy through suppression of Atg7 in aP2+ adipocytes has been reported to cause acquisition of a BAT phenotype by white adipose tissue (browning) as well as increase BAT mass.20, 21 In addition, a more recent study has shown that suppression of Atg7 specifically in Myf5+ precursor cells disrupts brown adipocyte differentiation and function,22 suggesting that autophagy is necessary during brown adipocyte differentiation. However, the role of autophagy in the adaptation of BAT to thermogenic activation has not been determined. Here we report that thermogenic activation leads to strong repression of autophagic activity in BAT, via the noradrenergic, cAMP-mediated pathway of BAT regulation.
Materials and methods
Animals
Adult (5-month-old) male C57BL/6J Ola Hsd and Swiss ICR (CD-1) mice were obtained from Harlan Laboratories (Barcelona, Spain) and were maintained under a 12 h dark/light cycle with ad libidum access to food (2018 Teklad Global 18% Protein Rodent Diet; Harlan Laboratories) and water. All experiments were performed in accordance with the European Community Council directive 86/609/EEC, and experiments as well as the number of animals to be used were approved by the Institutional Animal Care and Use Committee of the University of Barcelona based on the expected effects size. C57BL/6J mice were randomly divided into two groups. One group (controls; n=8) was maintained at the standard animal facility temperature (22 °C) and the other group was exposed to cold (24 h at 4 °C; n=7).For fasting experiments, animals were randomly divided into fasted (24 h, n=6) and fed ad libitum (n=6) groups, and then killed by decapitation. Interscapular BAT pads were dissected, after which one piece was frozen in liquid nitrogen for mRNA and total protein extraction and another piece was fixed for electron microscopy (see below).
Cell culture and reagents
Primary cultures of brown adipocytes were obtained by first isolating preadipocytes from interscapular, cervical and axillary BAT depots from 3-week-old Swiss ICR (CD-1) or C57BL/6J mice. Isolated precursor cells were then plated and grown in Dulbecco’s modified Eagle’s medium/Ham's F12 Medium (1:1) supplemented with 10% fetal bovine serum, 1% Fungizone, 20 nM insulin, 2 nM T3 and 100 μM ascorbate. Experiments were performed on day 10 of culture, at which point 90% of cells were considered to be differentiated on the basis of lipid accumulation and brown adipocyte morphology. Three independent cell culture experiments were analyzed and assays were carried out in triplicate for every experimental condition at every cell culture set. Where indicated, cells were treated with 0.5 μM norepinephrine (NE), 1 mM dibutyryl cAMP, 100 μM leupeptin (Roche Diagnostics, Mannheim, Germany), 20 mM ammonium chloride, 10 mM 3-methyladenine, 10 μM H89 and/or 10 μM SB202190. Immortalized brown adipocytes from C57BL/6J mice, a gift from J Klein (Medical University of Lübeck, Lübeck, Germany), were cultured as previously described.23 For immunofluorescence assays, cells were cultured on poly-L-lysine-coated coverslip in 24-well plates at 3 × 104 cells per well. All reagents used for cell culture and cell treatments were from Life Technologies (Eugene, OR, USA) or Sigma-Aldrich (Saint Louis, MO, USA) unless otherwise stated.
Fluorescence microscopy
Adherent cells on coverslips were incubated when indicated with MitoTraker Orange CM-H2TMRos (Invitrogen, Eugene, OR, USA) or BODIPY 558/568 C12 (Life Technologies) rinsed with phosphate-buffered saline (PBS) and fixed with 4% paraformaldehyde in PBS. After fixation, cells were permeabilized with Triton X-100 in PBS and blocked by incubating with blocking buffer consisting of PBS containing 2% bovine serum albumin and 0.3% Triton X-100. Cells were incubated overnight at 4 °C with an anti-LC3B primary antibody (no. 2775S, Cell Signaling Technology, Danvers, MA, USA) and then rinsed and incubated for 2 h at room temperature with the Alexa-488-conjugated secondary antibody (A11070, Life Technologies). Cells were co-stained with 4’,6-diamidino-2-phenylindole (Life Technologies) and mounted with Mowiol (EMD Millipore Corp., Billerica, MA, USA). Images were acquired on a Leica TCS SP2 confocal microscope (Leica Microsystems, Wetzlar, Germany) using a × 63/1.32 oil-immersion objective. Confocal z-stacks (5-μm-thick sections) were acquired at equivalent exposure times, and images were analyzed using the ImageJ software (NIH, Bethesda, MD, USA). Cellular fluorescence, expressed as the corrected integrated density per cell, was quantified in maximum-intensity projections. Cells from three independent cell cultures were analyzed, and >25 cells from three replicates per condition were analyzed. Colocalized pixels were detected and depicted in white using the ‘colocalization finder’ plugin in ImageJ. Images were processed using Adobe Photoshop CS6 (Adobe Systems, San José, CA, USA) for brightness and contrast adjustments; the same changes were applied to all compared images.
Electron microscopy
BAT samples were fixed in 2.5% glutaraldehyde and 2% paraformaldehyde in 0.1 M phosphate buffer (pH 7.4) and postfixed in 1% osmium tetroxide and 0.8% FeCNK in phosphate buffer. After dehydration in a graded acetone series, tissue samples were embedded in Spurr resin. Ultrathin sections were stained with uranyl acetate and lead citrate and examined with a Jeol 1010 transmission electron microscope (Izasa Scientific, Barcelona, Spain).
RNA isolation and real-time quantitative PCR
RNA from tissues and cells was extracted using a NucleoSpin RNA Kit (Macherey-Nagel, Düren, Germany), and the levels of mRNA were determined by quantitative reverse transcription-PCR, using the corresponding TaqMan probes (Applied Biosystems, Foster City, CA, USA) (Supplementary Table S1). The mRNA level of each gene of interest was normalized to that of a housekeeping reference gene (18S rRNA) using the comparative (2−ΔCt) method.
Western blotting
Cell extracts were prepared by homogenization in a buffer consisting of 20 mM Tris HCl (pH 7.4), 1 mM EDTA, 1 mM EGTA, 1% Triton X-100, a protease inhibitor cocktail (Roche Diagnostics) and 2 mM sodium orthovanadate and 10 mM β-glycerophosphate as phosphatase inhibitors. Total protein from BAT was isolated by homogenization in a buffer consisting of 50 mM Tris HCl (pH 7.4), 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% sodium dodecyl sulfate, 0.1 mM EDTA, 0.1 mM EGTA, a protease inhibitor cocktail, 2 mM sodium orthovanadate and 10 mM β-glycerophosphate. Total protein (30 μg per lane) was resolved by sodium dodecyl sulfate-polyacrylamide gel electrophoresis on 12% or 15% gels and electrotransferred to Immobilon-P polyvinylidene difluoride membranes (GE Healthcare, Little Chalfont, UK). Membranes were incubated with primary antibodies specific for UCP1 (ab10983, Abcam, Cambridge, UK), LC3B (no. 2775S, Cell Signaling Technology, Danvers, MA, USA), p62/SQSTM1 (sc-25575, Santa Cruz Biotechnology Inc., Dallas, TX, USA), Total-OXPHOS Rodent WB antibody cocktail (MS601, MitoSciences, Eugene, OR, USA) and/or α-TUBULIN (T9026, Sigma-Aldrich) and then with horseradish peroxidase-conjugated anti-mouse IgG (170-6516, Bio-Rad, Hercules, CA, USA) or anti-rabbit IgG (711-035-152, Jackson Immunoresearch Laboratories Inc., West Grove, PA, USA), as appropriate. Signals were detected using a chemiluminescence horseradish peroxidase substrate (EMD Millipore). Densitometric analyses of digitalized images were performed using the Multi Gauge V3.0 (Fujifilm, Tokyo, Japan) software. Images were processed using Adobe Photoshop CS6 (Adobe Systems) for brightness and contrast adjustments, applied uniformly across the entire image.
Oxygen consumption assay
Oxygen consumption in brown adipocytes was recorded using the Oxygen Biosensor System (BD Biosciences, San Jose, CA, USA) following previously described procedures.24 Primary brown adipocytes differentiated in culture were treated either with or without lysosome inhibitors for 6 h. Cells were then trypsinized, harvested in Dulbecco’s modified Eagle’s medium and placed in a 96-well BD Oxygen Biosensor System plate. Three independent cell cultures were used, and for each condition, four wells containing 105 cells and 200 μl of media were analyzed. Uncoupled respiration was determined by adding oligomycin (10 μg ml−1) just before initiating the oxygen consumption recording. Fluorescent signal was read every 1 min for 60 min using a FLUOstar OPTIMA microplate reader (BMG Labtechnologies, Ortenberg, Germany). Fluorescent signal was normalized according to the manufacturer’s instructions (BD Biosciences). The relative oxygen consumption was calculated using the linear parts of the curves of fluorescence.
Statistical analysis
Two-tailed unpaired Student’s t-tests were used to test the statistical significance of differences between two experimental conditions. Welch’s correction was applied when unequal variances were detected by F-test, using the GraphPad statistical software (GraphPad Software Inc., La Jolla, CA, USA). Statistical significance was set with an α-value of P<0.05, and underlying assumptions for validity of all tests were assessed. Data are presented as mean±s.e.m.
Results
Thermogenic activation causes repression of autophagy in BAT
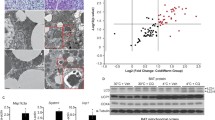
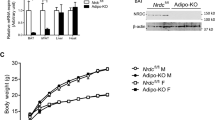
In order to determine how the autophagy cellular machinery responds to thermogenic activation of BAT, we first analyzed the effects of cold on the expression profile transcripts for autophagy components. Exposure of C57BL/6J mice to 4 °C for 24 h resulted in a significant reduction in Atg7, Atg5, Atg9a, Ulk1, Map1lc3b and Pik3c3 mRNA expression levels, accompanied by the expected increase in the expression of transcripts for the thermogenic marker genes Ppargc1a, Dio2 and Ucp1 (Figure 1a). Similar effects were found after cold exposure of mice from a distinct strain (Swiss ICR, CD-1) (Supplementary Table S2). To further determine whether this cold-induced reduction in transcript levels translated into a reduction in autophagic activity, we examined the levels of the phosphatidyletanolamine-conjugated microtubule-associated protein 1 light chain 3 beta (LC3B-II).25, 26 Immunoblotting analyses showed a significant decrease in total LC3B-II in BAT from cold-challenged mice compared with control mice, non-significant changes in LC3B-I protein along with an accumulation of the autophagic substrate of degradation p62 (Figure 1b), consistent with a reduction in autophagic flux. This was accompanied by the expected significant induction of UCP1 protein levels.
Autophagy and thermogenesis are oppositely regulated in BAT. (a) Relative mRNA levels of thermogenic and autophagy-related proteins in BAT from cold-exposed C57BL/6J mice (24 h at 4 °C; n=7) compared with control mice maintained at 22 °C (n=8). (b) Immunodetection of phosphatidylethanolamine-conjugated LC3B (LC3B-II), LC3B-I, p62 and UCP1. Left: Representative immunoblot for the indicated proteins. Right: Relative amounts of the indicated proteins, determined by densitometry and adjusted using α-TUBULIN (ATUB) as a loading control (n=6). (c) Relative mRNA levels of thermogenic and autophagy-related proteins in BAT from fed and fasted (24 h) C57BL/6J mice (n= 6). The bars represent means±s.e.m. (*P<0.05, **P<0.01, ***P<0.001).
We analyzed a second model of reciprocal change in BAT thermogenesis: 24-h starvation. Fasting is a known inhibitor of BAT thermogenic activity in mice.27 Consistent with this, we observed a strong repression of Ppargc1a and Ucp1 expression at the transcript level in starved mice (Figure 1c) and the corresponding reduction in UCP1 protein levels (Supplementary Figure S1). A parallel assessment of autophagy-related transcripts revealed a systematic induction of Atg7, Ulk1, Map1lc3b and Pik3c3 (Figure 1c). These results indicate an inverse relationship between thermogenic activation of BAT and induction of autophagic machinery components.
Noradrenergic repression of autophagy in brown adipocytes
In order to establish whether the effects of cold on the autophagic machinery observed in vivo occur in a cell autonomous manner and involve the noradrenergic pathway, the main mechanism of control of thermogenesis in BAT, we analyzed the effect of NE on brown adipocytes in culture. NE significantly reduced mRNA levels of the autophagy-related genes Atg7, Atg9a, Ulk1, Map1lc3b, Tfeb and Pik3c3 in primary cultures of brown adipocytes from Swiss ICR (CD-1) mice (Figure 2a). Similarly, primary cultures from C57Bl/6J mice also showed a reduction in the mRNA expression of Atg7, Atg9a, Map1lc3b, Tfeb, Ulk1 and Pik3c3 (Supplementary Table S3). An investigation of the involvement of peroxisome proliferator-activated receptors alpha (PPARα) and gamma (PPARγ) signaling, thermogenic activity regulatory pathways with a potential role in the transcriptional regulation of autophagy,28, 29 showed that GW7647 and rosiglitazone, activators of the corresponding receptors, had no such repressive effect (data not shown).
Noradrenergic repression of autophagy in brown adipocytes. (a) Relative mRNA levels of thermogenic and autophagy-related genes in primary brown adipocytes from Swiss ICR (CD-1) mice in culture treated with NE and harvested at the indicated time points (n=3). (b) Autophagic flux was assessed by measuring LC3B-II protein accumulation in differentiated brown adipocytes incubated with or without NE and treated with Leu/NH4Cl or 3-methyladenine. Left: Immunoblot for LC3B with Coomassie staining (CS) used as a loading control. Right: Relative amounts of LC3B-II determined by densitometry and adjusted using CS. The bars represent means±s.e.m. (*P<0.05, **P<0.01; ***P<0.001, treated with NE vs control; ###P<0.001, with vs without inhibitors).
To determine the effects of NE on autophagic flux, we incubated cells with or without NE and a combination of leupeptin and ammonium chloride (Leu/NH4Cl), added to inhibit lysosomal activity and degradation of autophagolysosome membrane-conjugated LC3B-II protein.26 3-Methyladenine, a general inhibitor of phosphoinositide 3-kinase (PI3K) activity, including that of PI3KC3, was also used to block earlier steps in autophagosome formation.26 NE significantly reduced the extent of LC3B-II protein accumulation in the presence of Leu/NH4Cl compared with that in control conditions, indicating that NE represses the net autophagic flux (Figure 2b) but acts in a PI3KC3-independent manner.
Effects of blockage of lysosomal activity on brown adipocytes
We also analyzed the effects of blocking lysosomal activity in brown adipocytes on the regulation of UCP1, the mitochondrial protein conferring to brown adipocytes their specific oxidative thermogenic function. We found that the inhibition of the lysosomal degradation with Leu/NH4Cl caused an increase in UCP1 protein levels both under basal and NE-sitmulated conditions (Figure 3a). However, the transcript levels of Ucp1 were not increased compared with control cells (Figure 3b). These data suggest a role of autophagy in UCP1 protein turnover.
Autophagic degradation of UCP1 in brown adipocytes. (a) UCP1 protein levels in primary brown adipocytes from Swiss ICR (CD-1) mice in culture treated or not with Leu/NH4Cl and/or NE. Immunoblot for UCP1 and quantification using Coomassie staining (CS) as loading control (n=2). (b) Relative mRNA levels of Ucp1 (n=3). (c) Total and uncoupled respiration of primary brown adipocytes in culture treated or not with Leu/NH4Cl (n=4). (d) LC3 and mitochondria colocalization in differentiated brown adipocytes treated with Leu/NH4Cl. Mitochondria were stained with MitoTraker (red), nuclei were stained with DAPI and endogenous LC3 was immunostained with anti-LC3 antibody (green). Colocalized pixels in a single z-stack are shown in white and highlighted with yellow arrowheads. The bars represent means±s.e.m. (*P<0.05, **P<0.01, treated with vs without NE; #P<0.05; ##P<0.01; ###P<0.001, treated with vs without inhibitors).
In order to test whether this UCP1 protein accumulation is associated with an increase in cell respiration, we measured brown adipocyte oxygen consumption. Leu/NH4Cl treatment caused a 35% induction in total respiration. Moreover, brown adipocytes treated with Leu/NH4Cl displayed 52% increase in uncoupled respiration (measured as the oligomycin-resistant respiration) (Figure 3c). These results suggest that inhibiting lysosomal degradation not only cause a UCP1 protein accumulation but also an increase in mitochondrial oxidative activity and especially the uncoupled respiration in brown adipocytes. The levels of several respiratory chain/oxidative phosphorylation subunits tended to be induced as a consequence of Leu/NH4Cl treatment but to a much lower extent than UCP1 (Supplementary Figure S2).
We further analyzed the putative interactions between autophagosomes and mitochondria in brown adipocytes by immunodetection of endogenous LC3 after Leu/NH4Cl treatment. We found that autophagosomes, detected as LC3 puncta, colocalized with mitochondria (Figure 3d), consistent with accumulation of the UCP1 protein after inhibition of lysosomal activity, noted above. Mitochondrially associated LC3 puncta tended to accumulate in crown-like structures around lipid droplets (see below).
cAMP-mediated repression of autophagy in brown adipocytes
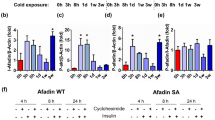
We determined whether the observed repressive effects of NE on autophagy were mediated by cAMP, the main intracellular signal transducer of noradrenergic action in brown adipocytes. Treatment with the cell-permeable cAMP analog dibutyryl-cAMP caused a significant reduction in transcript levels for Atg7, Atg5, Atg9a, Ulk1, Map1lc3b, Tfeb and Pik3c3 (Figure 4a), in association with the expected induction of Ucp1 mRNA, results similar to those caused by NE exposure. We next explored the potential involvement of protein kinase A (PKA) and the mitogen-activated protein kinase p38 (p38-MAPK), the major intracellular mediators of cAMP-dependent thermogenic regulation in brown adipocytes, in the autophagic pathway. Exposure of brown adipocytes to either SB202190, a p38-MAPK inhibitor, or H89, an inhibitor of PKA, significantly reduced the cAMP-dependent expression of Ucp1 mRNA as expected (Figure 4a). SB202190 and H89 also increased the basal mRNA levels of multiple autophagic genes. However, SB202190 did not suppress the cAMP-mediated reduction in the levels of autophagy-related transcripts, whereas H89 suppressed the inhibitory effects of cAMP on Atg7, Atg5, Atg9a, Ulk1 and Pik3c3 expression (Figure 4a).
cAMP-mediated repression of autophagy in brown adipocytes. Primary brown adipocytes from Swiss ICR (CD-1) mice in culture were treated with dibutyryl cyclic AMP (cAMP), H89 (PKA inhibitor) and/or SB202190 (p38-MAPK inhibitor) as indicated. (a) Relative mRNA levels of Ucp1 and autophagy-related genes (n=4). The bars represent means±s.e.m. (*P<0.05, treated with vs without cAMP; #P<0.05, treated with vs without inhibitors). (b) Autophagic flux was assessed by comparing relative amounts of LC3B-II and p62 using immunoblotting. α-TUBULIN (ATUB) was used as a loading control.
Moreover, dibutyryl-cAMP caused a reduction in LC3B-II protein levels accompanied by an increase in p62 accumulation (Figure 4b). These results confirmed that NE acts through cAMP to repress the autophagic machinery in brown adipocytes. Both kinase inhibitors increased LC3B-II protein levels under basal conditions and in the presence of cAMP (Figure 4b). Similar to other analyzed autophagic genes, Map1lc3b transcript levels were increased by treatment with SB202190 or H89 and reduced by treatment with dibutyryl-cAMP (Figure 4a), indicating that transcriptional regulation may contribute to the changes in LC3B protein levels. Inhibition of PKA or p38-MAPK did not prevent the cAMP-mediated reduction in Map1lc3b mRNA or LC3B-II protein.
Repression of autophagy by cAMP in brown adipocytes involves disruption of lipid droplet/LC3 multivesicular structures
We monitored LC3 in brown adipocytes by immunofluorescent detection under basal and cAMP-exposed conditions. Under basal conditions, scattered LC3 puncta were present in the cytosolic compartment of brown adipocytes (Figure 5a, left). cAMP caused a dramatic reduction in the intensity and number of LC3 puncta per cell (Figure 4a, right), indicating a reduction in autophagy in response to cAMP. A further examination of the subcellular localization of LC3 in control brown adipocytes revealed that LC3-positive vesicles were often found in association with lipid droplets, forming crown-like structures (Figure 5b, left). Exposure of cells to cAMP caused a dramatic loss of these structures (Figure 5b, right).
Induction of thermogenesis reduces LC3 puncta and lipophagy in brown adipocytes. (a) Left: Immunofluorescence detection of endogenous LC3 (green) in immortalized brown adipocytes from C57BL/6J mice incubated or not with dibutyryl cyclic AMP (cAMP); nuclei were stained with DAPI (blue). Right: LC3 fluorescence and LC3 puncta per cell, measured using the ImageJ software. Images were obtained using confocal microscopy (n=30). (b) Left: Confocal images of a single z-stack showing LC3 crowns surrounding lipid droplets (LD) in differentiated brown adipocytes. LD were stained with BODIPY 558/568 C12 (red), and after fixation, cells were immunostained for LC3 (green) and stained with DAPI (blue). Right: Percentage of cells containing LD-LC3 vesicle crowns in confocal images (n=5). More than 25 cells for each condition were analyzed. (c) Representative transmission electron microscopic images of BAT from control and cold-exposed C57BL/6J mice (24 h at 4 °C). Yellow arrowheads indicate autophagolysosomes containing lipids. Blue arrowheads indicate autophagolysosomes containing cytoplasmic substrates without lipids. The bars represent means±s.e.m. (*P<0.05, **P<0.01; ***P<0.001).
Finally, we examined brown adipocyte subcellular structure by transmission electron microscopy. In non-thermogenically stimulated BAT, we observed the presence of multiple degradation vesicles consistent with an autophagolysosome structure (Figure 5c). Many of these vesicles contained small fat droplets, indicative of active lipophagy. However, this type of lipid degradation vesicle was largely absent in transmission electron microscopic images of BAT from cold-exposed mice (Figure 5c).
Discussion
In the present study, we report for the first time that repression of autophagy is a component of the cellular response of brown adipocytes to thermogenic activation. This repressive effect of noradrenergic stimuli is mediated by cAMP and activation of downstream intracellular kinases, mainly PKA. This indicates that autophagy repression is tightly and reciprocally coordinated with thermogenic activation of brown adipocytes, as both processes share similar intracellular pathways of regulation.
To date, the role of autophagy in BAT biology has been considered primarily in relation to in vivo differentiation/transdifferentiation processes in adipose depots. Previous reports have indicated that targeted ablation of autophagy machinery components in adipose tissues promotes enrichment of brown-vs-white adipocytes in adipose depots.20, 21 Our current finding that autophagy is repressed in response to thermogenic activation indicates a distinct role for autophagy in differentiated brown adipocytes that are tasked with acutely activating thermogenesis in response to a noradrenergic stimulus. In fact, this is consistent with early reports indicating repression of general proteolysis in brown adipocytes in response to NE, a finding suggestive of the involvement of autophagy.30 Other biological processes induced during differentiation of brown adipocytes but repressed by noradrenergic stimulation of already differentiated cells (for example, PPARγ expression) have been reported previously.31
The coordinated repression of the expression of autophagy-related genes, in addition to the repression of autophagic flux itself, suggests that the negative regulation of autophagy in response to thermogenic activation involves transcription-mediated events. The operation of transcriptional processes in controlling autophagy in response to metabolic adaptations has been recently recognized in the liver, involving transcription factors such as the cAMP response element-binding protein (CREB).17, 32, 33 In contrast to the stimulatory role in hepatocytes,33 we found a repressive effect of cAMP on autophagy in brown adipocytes, similarly to other non-hepatic cells.34, 35 This marked cell-specific regulation is consistent with the distinct physiological roles of cAMP in these two different cell types: in hepatocytes, cAMP mediates starvation responses, whereas in brown adipocytes cAMP mediates thermogenic responses. On the other hand, the fact that experimentally induced repression of autophagy increases the levels of UCP1 and uncoupled mitochondrial respiration suggests that naturally occurring repression of autophagy by thermogenic stimuli contributes to the adaptive enrichment of cellular components required for heat production.
Noradrenergic, cAMP-mediated, repression of autophagy causes a marked reorganization of the intracellular autophagy machinery. Under basal conditions, the autophagy vesicles were tightly associated with lipid droplets and cells contained multiple intracellular structures with lipophagic vesicle-like features. Thermogenic activation caused a dramatic loss of these structures, consistent with a repression of lipophagy. Noradrenergic, cAMP-mediated induction of classical, hormone-sensitive lipolysis is a key process in brown adipocytes during activation of thermogenesis. When thermogenesis is activated, free fatty acids derived from lipolysis of lipid droplets are rapidly utilized by mitochondria to sustain uncoupled respiration6 The intracellular organization of brown adipocytes, with tight physical interactions between lipid droplets and mitochondria and long stretches of mitochondria embracing lipid droplet, likely favor this process.36 Our findings are consistent with a scenario in which lipophagy, a mechanism of fat processing that is not directly linked to mitochondrial oxidation, is disfavored when fatty acid metabolism in lipid droplets is induced to provide fatty acids for direct use by mitochondria for oxidation. Further research will be needed to confirm this scenario of reciprocal activation of classical hormone-sensitive lipolysis and lipophagy in response to thermogenesis in BAT.
Lipophagy has been previously studied and found to be highly relevant for management of lipids in hepatic fat droplets.10, 37 However, it should be noted that the biological role of lipid droplets is markedly different in hepatocytes and brown adipocytes. Accumulation of lipid droplets is a pathogenic event in hepatocyte steatosis38 that is often associated with lipotoxicity,39 whereas it is a phenomenon associated with intrinsic biological function in brown adipocytes. Considering the much higher activity of hormone-sensitive lipolysis in BAT than in the liver,40 it is likely that lipophagy has a less prominent role in fat catabolism in BAT than in the liver.
During the preparation of this article, it was reported that 1 h exposure of mice to cold elicits autophagy activation in BAT due to central and peripheral mechanisms.41 The apparent contradiction of this observation with our current findings will deserve further research. However, the complex intracellular machinery associated with thermogenic activation in brown adipocytes (including UCP1 protein increase) is known to be minimally developed in such a very short-time cold exposure setting, and in fact, shivering is the major response to very short-time acute exposure to cold.6, 42
Moreover, it may happen that autophagy repression does not necessarily have a significant role in situations of induction of BAT thermogenesis less intense or not as long as the 1 day cold exposure used in the current study. In fact, autophagy regulation is expectable to have a relevant role when overt remodeling of BAT in response to sustained changes in thermogenic activation is taking place.
In summary, we report that thermogenic activation of BAT is associated with repressed autophagy. This results in increased levels of thermogenic machinery components and repression of the autophagic pathway of lipid management (lipophagy) in the brown adipocyte. The inverse relationship between autophagy and BAT-mediated energy expenditure is consistent with numerous reports of abnormally high autophagy in adipose tissue from obesity,14, 18, 19 a condition associated with reduced BAT activity. Thus, in light of the growing recognition on the relevance of BAT thermogenesis in the control of obesity, exploring autophagy regulation in BAT may provide novel tools for management of energy balance and their abnormalities to favor metabolic health.
References
Lowell BB, S-Susulic V, Hamann A, Lawitts JA, Himms-Hagen J, Boyer BB et al. Development of obesity in transgenic mice after genetic ablation of brown adipose tissue. Nature 1993; 366: 740–742.
Feldmann HM, Golozoubova V, Cannon B, Nedergaard J . UCP1 ablation induces obesity and abolishes diet-induced thermogenesis in mice exempt from thermal stress by living at thermoneutrality. Cell Metab 2009; 9: 203–209.
Cypess AM, Lehman S, Williams G, Tal I, Rodman D, Goldfine AB et al. Identification and importance of brown adipose tissue in adult humans. N Engl J Med 2009; 360: 1509–1517.
van Marken Lichtenbelt WD, Vanhommerig JW, Smulders NM, Drossaerts JM, Kemerink GJ, Bouvy ND et al. Cold-activated brown adipose tissue in healthy men. N Engl J Med 2009; 360: 1500–1508.
Virtanen KA, Lidell ME, Orava J, Heglind M, Westergren R, Niemi T et al. Functional brown adipose tissue in healthy adults. N Engl J Med 2009; 360: 1518–1525.
Cannon B, Nedergaard J . Brown adipose tissue: function and physiological significance. Physiol Rev 2004; 84: 277–359.
Villarroya F, Vidal-Puig A . Beyond the sympathetic tone: the new brown fat activators. Cell Metab 2013; 17: 638–643.
Gnad T, Scheibler S, von Kügelgen I, Scheele C, Kilić A, Glöde A et al. Adenosine activates brown adipose tissue and recruits beige adipocytes via A2A receptors. Nature 2014; 516: 395–399.
Levine B, Klionsky DJ . Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell 2004; 6: 463–477.
Dong H, Czaja MJ . Regulation of lipid droplets by autophagy. Trends Endocrinol Metab 2011; 22: 234–240.
Klionsky DJ . The molecular machinery of autophagy and its role in physiology and disease. Semin Cell Dev Biol 2010; 21: 663.
Mizushima N, Levine B . Autophagy in mammalian development and differentiation. Nat Cell Biol 2010; 12: 823–830.
Feng Y, He D, Yao Z, Klionsky DJ . The machinery of macroautophagy. Cell Res 2014; 24: 24–41.
Kovsan J, Bluher M, Tarnovscki T, Kloting N, Kirshtein B, Madar L et al. Altered autophagy in human adipose tissues in obesity. J Clin Endocrinol Metab 2011; 96: E268–E277.
Maixner N, Kovsan J, Harman-Boehm I, Bluher M, Bashan N, Rudich A . Autophagy in adipose tissue. Obes Facts 2012; 5: 710–721.
Singh R, Cuervo AM . Autophagy in the cellular energetic balance. Cell Metab 2011; 13: 495–504.
Settembre C, Ballabio A . Cell metabolism: autophagy transcribed. Nature 2014; 516: 40–41.
Jansen HJ, van Essen P, Koenen T, Joosten LA, Netea MG, Tack CJ et al. Autophagy activity is up-regulated in adipose tissue of obese individuals and modulates proinflammatory cytokine expression. Endocrinology 2012; 153: 5866–5874.
Ost A, Svensson K, Ruishalme I, Brannmark C, Franck N, Krook H et al. Attenuated mTOR signaling and enhanced autophagy in adipocytes from obese patients with type 2 diabetes. Mol Med 2010; 16: 235–246.
Singh R, Xiang Y, Wang Y, Baikati K, Cuervo AM, Luu YK et al. Autophagy regulates adipose mass and differentiation in mice. J Clin Invest 2009; 119: 3329–3339.
Zhang Y, Goldman S, Baerga R, Zhao Y, Komatsu M, Jin S . Adipose-specific deletion of autophagy-related gene 7 (atg7) in mice reveals a role in adipogenesis. Proc Natl Acad Sci USA 2009; 106: 19860–19865.
Martinez-Lopez N, Athonvarangkul D, Sahu S, Coletto L, Zong H, Bastie CC et al. Autophagy in Myf5+ progenitors regulates energy and glucose homeostasis through control of brown fat and skeletal muscle development. EMBO Rep 2013; 14: 795–803.
Whittle AJ, Carobbio S, Martins L, Slawik M, Hondares E, Vazquez MJ et al. BMP8B increases brown adipose tissue thermogenesis through both central and peripheral actions. Cell 2012; 149: 871–885.
Hondares E, Rosell M, Gonzalez FJ, Giralt M, Iglesias R, Villarroya F . Hepatic FGF21 expression is induced at birth via PPARalpha in response to milk intake and contributes to thermogenic activation of neonatal brown fat. Cell Metab 2010; 11: 206–212.
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 2000; 19: 5720–5728.
Klionsky DJ, Abdelmohsen K, Abe A, Abedin MJ, Abeliovich H, Acevedo Arozena A et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 2016; 12: 1–222.
Trayhurn P, Jennings G . Nonshivering thermogenesis and the thermogenic capacity of brown fat in fasted and/or refed mice. Am J Physiol 1988; 254: R11–R16.
Barbera MJ, Schluter A, Pedraza N, Iglesias R, Villarroya F, Giralt M . Peroxisome proliferator-activated receptor alpha activates transcription of the brown fat uncoupling protein-1 gene. A link between regulation of the thermogenic and lipid oxidation pathways in the brown fat cell. J Biol Chem 2001; 276: 1486–1493.
Sears IB, MacGinnitie MA, Kovacs LG, Graves RA . Differentiation-dependent expression of the brown adipocyte uncoupling protein gene: regulation by peroxisome proliferator-activated receptor gamma. Mol Cell Biol 1996; 16: 3410–3419.
Moazed B, Desautels M . Differentiation-dependent expression of cathepsin D and importance of lysosomal proteolysis in the degradation of UCP1 in brown adipocytes. Can J Physiol Pharmacol 2002; 80: 515–525.
Lindgren EM, Nielsen R, Petrovic N, Jacobsson A, Mandrup S, Cannon B et al. Noradrenaline represses PPAR (peroxisome-proliferator-activated receptor) gamma2 gene expression in brown adipocytes: intracellular signalling and effects on PPARgamma2 and PPARgamma1 protein levels. Biochem J 2004; 382: 597–606.
Lee JM, Wagner M, Xiao R, Kim KH, Feng D, Lazar MA et al. Nutrient-sensing nuclear receptors coordinate autophagy. Nature 2014; 516: 112–115.
Seok S, Fu T, Choi SE, Li Y, Zhu R, Kumar S et al. Transcriptional regulation of autophagy by an FXR-CREB axis. Nature 2014; 516: 108–111.
Cherra SJ 3rd, Kulich SM, Uechi G, Balasubramani M, Mountzouris J, Day BW et al. Regulation of the autophagy protein LC3 by phosphorylation. J Cell Biol 2010; 190: 533–539.
Shahnazari S, Namolovan A, Mogridge J, Kim PK, Brumell JH . Bacterial toxins can inhibit host cell autophagy through cAMP generation. Autophagy 2011; 7: 957–965.
Cinti S . The Adipose Organ. Editrice Kurtis: Milan, Italy, 1999.
Singh R, Kaushik S, Wang Y, Xiang Y, Novak I, Komatsu M et al. Autophagy regulates lipid metabolism. Nature 2009; 458: 1131–1135.
Czaja MJ . Autophagy in health and disease. 2. Regulation of lipid metabolism and storage by autophagy: pathophysiological implications. Am J Physiol Cell Physiol 2010; 298: C973–C978.
Wang DQ, Portincasa P, Neuschwander-Tetri BA . Steatosis in the liver. Compr Physiol 2013; 3: 1493–1532.
Zechner R, Zimmermann R, Eichmann TO, Kohlwein SD, Haemmerle G, Lass A et al. Fat signals—lipases and lipolysis in lipid metabolism and signaling. Cell Metab 2012; 15: 279–291.
Martinez-Lopez N, Garcia-Macia M, Sahu S, Athonvarangkul D, Liebling E, Merlo P et al. Autophagy in the CNS and periphery coordinate lipophagy and lipolysis in the brown adipose tissue and liver. Cell Metab 2016; 23: 113–127.
Yu XX, Lewin DA, Forrest W, Adams SH . Cold elicits the simultaneous induction of fatty acid synthesis and beta-oxidation in murine brown adipose tissue: prediction from differential gene expression and confirmation in vivo. FASEB J 2002; 16: 155–168.
Acknowledgements
This work was supported by grants from MINECO (SAF2014-55725), Instituto de Salud Carlos III-FEDER (PI11/00376 and PI14/00063), EU (FP7 project BETABAT, grant HEALTH-F2-2011-277713) and Generalitat de Catalunya (2014SGR-141). MC is a FPI fellow from MINECO, Spain.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on International Journal of Obesity website
Rights and permissions
About this article
Cite this article
Cairó, M., Villarroya, J., Cereijo, R. et al. Thermogenic activation represses autophagy in brown adipose tissue. Int J Obes 40, 1591–1599 (2016). https://doi.org/10.1038/ijo.2016.115
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ijo.2016.115
- Springer Nature Limited