Abstract
Nerve growth factor (NGF) is known to regulate both cancer cell survival and death signaling, depending on the cellular circumstances, in various cell types. In this study, we showed that NGF strongly upregulated the protein level of tropomyosin-related kinase A (TrkA) in TrkA-inducible SK-N-MC cancer cells, resulting in increases in various TrkA-dependent cellular processes, including the phosphorylation of c-Jun N-terminal kinase (JNK) and caspase-8 cleavage. In addition, NGF enhanced TrkA-induced morphological changes and cell death, and this effect was significantly suppressed by the JNK inhibitor SP600125, but not by the phosphatidylinositol 3-kinase (PI3K) inhibitor wortmannin. To investigate novel targets associated with the enhancement of TrkA-induced SK-N-MC cell death caused by NGF, we performed Coomassie Brilliant Blue staining and two-dimensional (2D) proteomic analysis in TrkA-inducible SK-N-MC cells. We identified 31 protein spots that were either greatly upregulated or downregulated by TrkA during NGF treatment using matrix-associated laser desorption/ionization time of flight/time of flight mass spectrometry, and we analyzed the effects of SP600125 and wortmannin on the spots. Interestingly, 11 protein spots, including heterogeneous nuclear ribonucleoprotein K (hnRNP K), lamin B1 and TAR DNA-binding protein (TDP43), were significantly influenced by SP600125, but not by wortmannin. Moreover, the NGF/TrkA-dependent inhibition of cell viability was significantly enhanced by knockdown of hnRNP K using small interfering RNA, demonstrating that hnRNP K is a novel target associated with the regulation of TrkA-dependent SK-N-MC cancer cell death enhanced by NGF.
Similar content being viewed by others
Introduction
Nerve growth factor (NGF) is known to regulate both cancer cell survival and death through its receptors tropomyosin-related kinase A (TrkA) and p75 neurotrophin receptor (p75NTR), depending on the tumor origin: NGF activates survival signaling through TrkA and p75NTR in breast cancer, whereas NGF activates cell death signaling through p75NTR in prostate cancer and through TrkA in glioblastoma and medulloblastoma.1 β-tubulin and a 43-kDa chromogranin B-derived fragment were identified as the major NGF-regulated proteins during neuronal differentiation of PC12 cells using a 2D proteomic analysis.2 Various NGF-regulated proteins, including 40S ribosomal protein S27a, 60S acidic ribosomal protein P1 and eukaryotic translation initiation factor 5A, were identified using a multiplex iTRAQ proteomic analysis of neurite outgrowth regulation of dorsal root ganglia (DRG) cells,3 suggesting that translation regulators may have an important role in NGF-induced morphological changes. Moreover, NGF induced c-Jun N-terminal kinase (JNK) activation to promote cell death and neurite outgrowth in PC12 cells, and NGF-induced JNK phosphorylation was significantly suppressed by specific inhibitors of the ERK1/2, p38 MAPK, JAK3, Src and Gi proteins, but not by the PI3K inhibitor wortmannin,4 suggesting that various signaling pathways, except for PI3K-Akt signaling, are involved in NGF-induced cell death and neuronal differentiation. Interestingly, JNK-associated neuronal death required nuclear but not cytosolic JNK activity, indicating that the nuclear localization of JNK is critical for neuronal death.5
Although it remains unclear why NGF functions as both an inducer and an inhibitor of cell death, NGF seems to promote cell death signaling rather than survival in situations of high ectopic TrkA expression. NGF is known to induce cell death in ectopic TrkA-expressing PC12 cells and medulloblastoma cells.6, 7 In addition, ectopic TrkA causes neuronal death in mouse embryonic stem (ES) cells and inhibits angiogenesis and tumor growth in SY5Y neuroblastoma cells by downregulation of angiogenic factors.8, 9 The ACTB protein, tropomyosin 3, dynein intermediate chain protein, vimentin, lamin A/C isoform 1 precursor, lamin B2, hnRNPKa and heat shock 70-kDa protein 5 have been identified as NGF-mediated TrkA-dependent effector proteins using a DIGE (fluorescence two-dimensional difference gel electrophoresis) proteomic analysis in SY5Y neuroblastoma cells stably expressing ectopic TrkA.10 However, little is known about the proteomic factors associated with the enhancement of TrkA-dependent cancer cell death caused by NGF.
We recently reported that, in the absence of NGF treatment, SP600125 strongly suppressed TrkA-dependent cellular processes, including the phosphorylation of JNK and c-Jun, in TrkA-inducible SK-N-MC cancer cells, indicating that JNK is involved in TrkA-dependent signaling pathways.11 In the present study, we investigated the roles of NGF in the regulation of various TrkA-dependent cellular processes, morphological changes and cell death. In addition, we identified novel targets associated with the enhancement of TrkA-induced SK-N-MC cancer cell death caused by NGF through a 2D proteomic analysis, and we investigated the effects of small interfering RNA (siRNA) knockdown of these targets on the regulation of NGF/TrkA-dependent cell death.
Materials and methods
Reagents
Tetracycline (Tet) was obtained from Duchefa Biochemie (Haarlem, the Netherlands). DMEM, Tet-screened FBS and penicillin/streptomycin were from Life Technologies (Carlsbad, CA, USA). Blasticidin and zeocin were from Invitrogen (Carlsbad, CA, USA). Human recombinant NGF-β, 0.4% trypan blue solution, Coomassie Brilliant Blue (CBB) G, octyl β-D-thioglucopyranoside and α-cyano-4-hydroxycinnamic acid were from Sigma-Aldrich (St Louis, MO, USA). SP600125 was from Calbiochem (Billerica, MA, USA). Wortmannin was from Enzo Life Sciences (Farmingdale, NY, USA). Cell counting kit-8 (CCK-8) was from Dojindo (Kumamoto, Japan). Acrylamide/bis-acrylamide 37.5:1 solution (40%), urea, Tween-20, ammonium sulfate and DMSO were from Amresco (Solon, OH, USA). IPG strip (pH 4-7) 17 cm was from Bio-Rad (Quarry Bay, Hong Kong). IPG buffer (pH 4–7) was from Amersham Biosciences (Piscataway, NJ, USA). Recombinant DNase I (RNase-free) was from Takara (Shiga, Japan). Sequencing-grade modified trypsin was from Promega (Madison, WI, USA). SuperSignal West Pico stable peroxide solution was from Pierce (Rockford, IL, USA). G-fectin transfection reagent was from Genolution Inc. (Seoul, Korea). The following antibodies were used in this study: TrkA (763, Santa Cruz), phospho-TrkA (E-6, Santa Cruz, Dallas, TX, USA), phospho-tyrosine (PY20, Transduction Laboratories, San Jose, CA, USA), β-actin (Sigma-Aldrich), phospho-Y204-ERK1/2 (Bioworld Technology, Louis Park, MN, USA), ERK1 (Transduction Laboratories), phospho-T183/Y185-JNK1/2/3 (Bioworld Technology), JNK1 (D-6, Santa Cruz), phospho-S473-Akt (Cell Signaling, Danvers, MA, USA), α-tubulin (B-5-1-2, Sigma-Aldrich), PARP (Oncogene, Boston, MA, USA), active caspase-7 (Abcam, Cambridge, MA, USA), caspase-8 (Invitrogen), lamin B1 (H-90, Santa Cruz), heterogeneous nuclear ribonucleoprotein K (hnRNP K; D-6, Santa Cruz), TDP43 (E-10, Santa Cruz), GAPDH (FL-335, Santa Cruz), goat anti-rabbit horseradish peroxidase conjugate (Bio-Rad) and goat anti-mouse horseradish peroxidase conjugate (Thermo Scientific, Rockford, IL, USA).
Cell culture
SK-N-MC-TrkA cells, which can induce the expression of ectopic TrkA via the Tet-On system, were previously established from the human neuroblastoma SK-N-MC cell line (Korean Cell Line Bank No. 30010).12 The cells were maintained in medium A (10% Tet-screened FBS, 1% penicillin/streptomycin, DMEM) containing 1.25 μg ml−1 blasticidin and 25 μg ml−1 zeocin in a humidified 5% CO2 incubator at 37 °C. The ectopic expression of TrkA was induced by adding 2 μg ml−1 Tet to the medium A.
Western blot analysis
Whole cells were extracted with sodium dodecyl sulfate (SDS) sample buffer and were boiled for 5 min at 95 °C. The resultant proteins were separated using 10% SDS polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to an NC membrane. The blot was blocked for 1 h at room temperature in blocking buffer (3% skim milk, 0.1% Tween-20, phosphate-buffered saline (PBS)) and then incubated with primary antibody at 4 °C overnight. The blot was then washed with PBST (0.1% Tween-20, PBS) three times for 10 min and incubated with an horseradish peroxidase-conjugated secondary antibody in blocking buffer for 2 h. After being washed with PBST, the blot was analyzed with the SuperSignal ECL detection system.
Cell morphology analysis
SK-N-MC-TrkA cells were split in a six-well dish (3 × 105 cells) in the absence or presence of Tet for 12 h and were then left untreated or were treated with 50 ng ml−1 NGF, 1 μM wortmannin and 2 μg ml−1 SP600125 for 24 h. The cell morphology was analyzed using phase-contrast light microscopy (EVOS XL Core, Life Technologies) with × 150 amplification.
Detection of dead cells
SK-N-MC-TrkA cells were split in a six-well dish (3 × 105 cells) in the absence or presence of 2 μg ml−1 Tet for 12 h and then were left untreated or were treated with 50 ng ml−1 NGF, 1 μM wortmannin and 2 μg ml−1 SP600125 for 24 h, as indicated. Without removing the medium, the cells were stained with a 0.03% trypan blue staining solution for 20 min. Dead cells, which were stained a dark blue color, were visualized using bright-field phase-contrast light microscopy (EVOS XL Core, Life Technologies) with × 150 amplification. The color contrast of the images was adjusted with a Photoshop program.
Cell viability analysis
SK-N-MC-TrkA cells were split in a 24-well dish (5 × 104 cells) in the absence or presence of 2 μg ml−1 Tet for 12 h and then were left untreated or were treated with 50 ng ml−1 NGF, 1 μM wortmannin and 2 μg ml−1 SP600125 for 24 h. The cells were incubated with medium A containing 10% CCK-8 reagent for 2 h at 37 °C in a CO2 incubator. The reaction solution (100 μl each) was then transferred to a 98-well dish and was analyzed by measuring the absorbance at OD485 nm using a CHAMELEON microplate reader (Hidex).
Sample preparation, 2D electrophoresis and CBB staining
Whole cells (attached and floating cells) were washed with PBS and prepared for 2D electrophoresis as previously described.13 Protein samples (500 μg each) were subjected to one-dimensional isoelectric focusing, followed by 2D electrophoresis on a 8.5–14% sucrose gradient polyacrylamide gel. The gels were fixed in fixing solution (40% methanol, 5% ortho-phosphoric acid) at room temperature for 3 h and were stained with CBB staining solution (17% ammonium sulfate, 3% phosphoric acid, 34% methanol, 0.1% CBB G) for 24 h. After destaining with H2O, gel images were obtained using a UMAX scanner (PowerLook 2100XL).
Image analysis and statistical significance
A quantitative analysis of 16 CBB-stained images (8 groups: −Tet; +Tet; −Tet,+NGF; +Tet,+NGF; −Tet,+NGF,+wortmannin; +Tet,+NGF,+wortmannin; −Tet,+NGF,+SP600125; and +Tet,+NGF,+SP600125) was performed using PDQuest software (Bio-Rad) according to the manufacturer’s instructions. Among the total protein spots indicated by the spot detection wizard, 503 clear and significant protein spots were manually selected and matched by landmarking. The quantity of each spot was normalized to the total valid spot intensity and described as parts per million (p.p.m.) of the total integrated optical density. Thus, we found 31 protein spots that were more than two-fold upregulated or downregulated by TrkA with NGF. Among them, 11 protein spots were determined to by significantly affected (more than 20%) by SP600125, but not by wortmannin. The statistical significance of the image analysis was determined using Student's t-test (P<0.05 was considered statistically significant).
Mass spectrometric analysis and database search
The CBB-stained protein spots were excised and finely chopped, and the gel pieces were destained, reduced, alkylated and in-gel tryptic digested as previously described.13 After being sonicated for 2 min, the tryptic digested peptides were collected by two extraction with 66% acetonitrile/0.1% trifluoroacetic acid and were dried in a vacuum centrifuge. The peptides were resolved with 50% acetonitrile/0.1% trifluoroacetic acid solution, and 1.5 μl of the sample was mixed with 1.5 μl of matrix solution (10 mg of α-cyano-4-hydroxycinnamic acid, 50% acetonitrile, 0.1% trifluoroacetic acid). The mixed tryptic peptides (1 μl) were immediately spotted onto a 384-well target plate for mass spectrometric analysis. Peptide mass spectra were obtained using a ABI 4800 Plus time of flight/time of flight (TOF/TOF) mass spectrometer (Applied Biosystems, Framingham, MA, USA) set to 200 Hz neodymium-doped (ND): 355 nm yttrium aluminium garnet (YAG) laser operation.14 Ten intense ions with a signal/noise ratio>25 were selected for tandem mass spectrometry (MS/MS) analysis in 1 kV mode and 1000–1250 consecutive laser exposure. The resultant MS/MS spectra were analyzed using the NCBInr/UniProt database and Protein Pilot V.3.0 software with the following settings: carbamidomethylation of cysteine, oxidation of methionine, monoisotopic, peptide mass tolerance of ±100 p.p.m., fragment mass tolerance of±0.3 Da, and a maximum of one missed cleavages. Mascot protein scores >65% were considered statistically significant (P<0.05). The final decision regarding protein identification from among the statistically significant candidates was based on the number of matching peptide masses, the tryptic fragment coverage, and a comparison of the experimental and theoretical properties of the proteins.
siRNA transfection
To induce the knockdown of NGF/TrkA-dependent targets by siRNA transfection, negative control and hnRNP K siRNAs were synthesized and validated using read-time PCR 24 h after the transfection of HeLa cells by Genolution, Inc.: negative control siRNA, (sense) 5′-CCUCGUGCCGUUCCAUCAGGUAGUU-3′ and (antisense) 5′-CUACCUGAUGGAACGGCACGAGGUU-3′; hnRNP K siRNA #1 (96.6% reduction), (sense) 5′-GAGUAUCAGUGCUGAUAUUUU-3′ and (antisense) AAUAUCAGCACUGAUACUCUU-3′; hnRNP K siRNA #2 (91.6% reduction), (sense) 5′-CGAUGAAACCUAUGAUUAUUU-3′ and (antisense) 5′-AUAAUCAUAGGUUUCAUCGUU-3′. To transfect these siRNAs into SK-N-MC-TrkA cells, the annealed siRNAs (10 nM each) were incubated with G-fectin in PBS at room temperature for 15 min and were spread onto 6-well and 24-well dishes containing trypsin-treated SK-N-MC-TrkA cells. After 24 h in a 37 °C CO2 incubator, the cells were treated with 0.5 μg ml−1 Tet and 10 ng ml−1 NGF for 24 or 48 h and analyzed using western blot or a cell viability analysis.
Results
NGF enhances TrkA-dependent cellular processes in a JNK-dependent manner
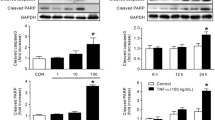
To better understand NGF-mediated TrkA-dependent signaling pathways during cancer cell death, we investigated the effects of NGF on the regulation of TrkA and TrkA-dependent cellular proteins in TrkA-inducible SK-N-MC cancer cells by western blot analysis. As shown in Figure 1a, ectopic TrkA was increased depending on the amount of NGF, and this TrkA upregulation resulted in increases in TrkA tyrosine-490 phosphorylation and the TrkA-dependent tyrosine phosphorylation of cellular proteins. In addition, the TrkA-dependent phosphorylation of ERK, JNK and Akt was upregulated 4, 8 and 12 h after NGF treatment, except for Akt phosphorylation at 12 h (Figure 1b). Notably, TrkA-dependent JNK phosphorylation was upregulated 4 h after NGF treatment and was significantly suppressed by SP600125, but not by wortmannin, indicating a specific inhibition of JNK signaling by SP600125 (Figure 1c). In comparison, TrkA-dependent Akt phosphorylation was upregulated 4 h after NGF treatment, and it was completely blocked by wortmannin and suppressed by SP600125 via an unknown mechanism, indicating a strong inhibitory effect of wortmannin on PI3K-Akt signaling (Figure 1c). Moreover, the upregulation of TrkA tyrosine-490 phosphorylation by NGF was not altered by either wortmannin or SP600125 4 h after NGF treatment, indicating a drug specificity of SP600125 and wortmannin. However, the upregulation of TrkA tyrosine-490 phosphorylatin by NGF was significantly downregulated during the long-term inactivation of JNK caused by SP600125 treatment for 12 h, but not by wortmannin, suggesting a role of JNK in the maintenance of TrkA tyrosine-490 phosphorylation enhanced by NGF (Figure 1c). In addition, TrkA-dependent cellular processes such as ERK phosphorylation, JNK phosphorylation, PARP cleavage, caspase-7 activation and caspase-8 cleavage were strongly enhanced by NGF and were significantly suppressed by SP600125, but not by wortmannin, indicating that NGF enhances TrkA-dependent apoptotic signaling in a JNK-dependent manner (Figure 1d).
Effects of NGF on TrkA protein level and TrkA-dependent cellular processes. SK-N-MC-TrkA cells were cultured in a 10-cm dish in the absence (−) or presence (+) of Tet for 12 h. (a) Cells were left untreated or were treated with the indicated amounts of NGF for 12 h. (b) Cells were left untreated or were treated with 50 ng ml−1 NGF for the indicated times. (c) Cells were left untreated or were treated with 50 ng ml−1 NGF, 1 μM wortmannin (WM) and 2 μg ml−1 SP600125 (SP) for the indicated times. (d) Cells were left untreated or were treated with 50 ng ml−1 NGF, 1 μM WM, and 2 μg ml−1 SP600125 for 12 h. After the medium was removed, the cells were immediately extracted with SDS sample buffer, followed by a western blot analysis using the indicated antibodies (a–d).
NGF enhances TrkA-dependent cancer cell death in a JNK-dependent manner
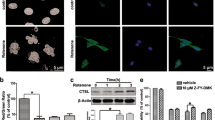
We next investigated the effects of NGF on TrkA-dependent morphological changes and cell death using phase-contrast light microscopy. As shown in Figure 2a, ectopic TrkA led to morphological changes in SK-N-MC cells: the attached cells were larger and had a modified cell body and extracellular membrane; the floating cells were somewhat fragmented and aggregated (compare panels a and b); and the morphological changes caused by TrkA were strongly enhanced by NGF (compare panels b and d). Notably, the enhancement of the TrkA-induced morphological changes caused by NGF was significantly suppressed by SP600125, but not by wortmannin (Figure 2a; compare panels d, f and h). Consistent with these results, the number of TrkA-dependent dead cells stained by trypan blue was greatly increased by NGF, and this effect was significantly suppressed by SP600125, but not by wortmannin, indicating that NGF enhances TrkA-dependent cancer cell death in a JNK-dependent manner (Figure 2b). In addition, the ability of TrkA to reduce cell viability was strongly upregulated by NGF and was significantly suppressed by SP600125, but not by wortmannin (Figure 2c).
Effects of wortmannin (WM) and SP600125 on the enhancement of TrkA-induced cell death caused by NGF. SK-N-MC-TrkA cells were cultured in the absence or presence of Tet for 12 h and then were left untreated or were treated with 50 ng ml−1 NGF, 1 μM WM and 2 μg ml−1 SP600125 for 24 h. (a) The cell morphology was analyzed by phase-contrast light microscopy. (b) After trypan blue staining, dead cells were detected by bright-field phase-contrast light microscopy. (c) Cell viability was determined with a CCK-8 assay. Each bar represents the mean±s.d. of three experiments, and the data significance was evaluated with Student’s t-test, *P<0.05.
Identification of TrkA-dependent protein spots regulated by NGF
To better understand the NGF-mediated TrkA-dependent cellular processes and cell death mechanism, we investigated TrkA-dependent targets regulated by NGF in TrkA-inducible SK-N-MC cells through CBB staining and 2D proteomic analysis. A PDQuest image analysis was performed with two sets of well-separated 2D images; representative 2D images are shown in Supplementary Figure 1. Protein spots that were more than two-fold upregulated or downregulated by TrkA with NGF were identified via a matrix-associated laser desorption/ionization (MALDI)-TOF/TOF MS/MS analysis, and the statistically significant results (P<0.05) are shown in Table 1 and Figure 3.
The 2D electrophoresis profile of TrkA-dependent protein spots regulated by NGF. SK-N-MC-TrkA cells were cultured on a 15-cm dish in the absence or presence of Tet for 12 h and then were left untreated or were treated with 50 ng ml−1 NGF, 1 μM wortmannin and 2 μg ml−1 SP600125 for 24 h. The cells were analyzed by 2D electrophoresis and CBB staining. The 18 upregulated (a) and 13 downregulated (b) TrkA-dependent protein spots regulated by NGF were determined using a PDQuest image analysis of two sets of well-separated 2D electrophoresis images and were identified by a MALDI-TOF/TOF MS/MS analysis. Each protein spot is indicated by an arrow and SSP number on a representative 2D image.
Identification of novel targets associated with the enhancement of TrkA-induced cancer cell death caused by NGF
Figure 2b demonstrates that the enhancement of TrkA-induced cell death caused by NGF was significantly suppressed by SP600125 but not by wortmannin. Thus, among the spots regulated by TrkA with NGF, we investigated protein spots that were significantly affected (more than 20%) by SP600125, but not by wortmannin. As shown in Figure 4a, the upregulation of the hnRNP C1/C2 and PP2Acβ spots by TrkA was enhanced by NGF and was significantly suppressed by SP600125 compared with the effects of wortmannin. In addition, the downregulation of the SRSF3, PSα1 and septin-2 spots by TrkA was enhanced by NGF and was significantly suppressed by SP600125, but not by wortmannin (Figure 4b). Moreover, the lamin B1, SRSF1, hnRNP K, BAF53A and TDP43 spots were greatly downregulated by TrkA with NGF, but not by TrkA alone; this effect was significantly suppressed by SP600125, but not by wortmannin (Figure 4c). The remaining protein spots were not significantly affected by either wortmannin or SP600125. The quantitative analysis results are shown in Supplementary Figure 2.
Quantitative analysis of TrkA-dependent protein spots regulated by NGF and significantly affected by SP600125. (a) These protein spots were upregulated by TrkA, which was enhanced by NGF. This upregulation by TrkA with NGF was significantly suppressed by SP600125. Compare the protein spots located at the center of each panel: −T (−Tet); +T (+Tet); WM (wortmannin); SP (SP600125). The relative intensities of the protein spots determined using PDQuest software are shown in the right panels. Each bar represents the mean±s.d. of two independent experiments, and the data significance was evaluated with Student’s t-test, *P<0.05. (b) These protein spots were downregulated by TrkA, which was strongly enhanced by NGF. This downregulation by TrkA with NGF was significantly suppressed by SP600125. (c) These protein spots were downregulated by TrkA with NGF, but not by TrkA alone, which was significantly suppressed by SP600125.
Confirmation of SP600125-controlled NGF/TrkA-regulated targets
Figure 4 shows the potential JNK-associated NGF/TrkA-regulated targets that were significantly affected by SP600125, as suggested by the 2D proteomic analysis. To confirm these results, samples prepared for 2D electrophoresis were analyzed by western blot analysis. As shown in Figure 5, TrkA was upregulated 36 h after Tet treatment, and this upregulation was strongly enhanced by NGF treatment for 24 h. Among the identified SP600125-controlled NGF/TrkA-regulated targets, lamin B1, hnRNP K and TDP43 were significantly downregulated by NGF/TrkA, but not by TrkA alone, and this effect was strongly suppressed by SP600125, but not by wortmannin, consistent with the results in Figures 4c and 5. Moreover, the cleaved forms of lamin B1 and hnRNP K were induced by NGF/TrkA via an unknown mechanism, although this was not significantly affected by SP600125 (Figure 5, indicated by asterisks).
Confirmation of the effects of wortmannin (WM) and SP600125 on the identified NGF/TrkA-regulated targets. SK-N-MC-TrkA cells were cultured in the absence or presence of Tet for 12 h and then were left untreated or were treated with 50 ng ml−1 NGF, 1 μM WM and 2 μg ml−1 SP600125 for 24 h. Protein samples (10 μg each) prepared for 2D electrophoresis were separated by 10% SDS-PAGE and analyzed using a western blot analysis with the indicated antibodies. The asterisks represent potential cleaved products of lamin B1 and hnRNP K. For a loading control, the 2D samples (10 μg each) were separated using 10% SDS-PAGE and CBB staining, as described in the Materials and Methods section.
Knockdown of hnRNP K enhances NGF/TrkA-dependent inhibition of cell viability
To determine whether the NGF/TrkA-regulated targets are involved in the regulation of TrkA-dependent cell death during NGF treatment, we performed siRNA transfection in SK-N-MC-TrkA cells, as described in the Materials and Methods section, followed by a western blot analysis. Because transfection efficiency is very low in SK-N-MC cells compared with other cells, the knockdown of hnRNP K protein by hnRNP K siRNAs #1 and #2 was not clearly detectable in untreated SK-N-MC-TrkA cells (Figure 6a, lanes 1–3 and 10–12). However, hnRNP K was somewhat downregulated by both hnRNP siRNAs #1 and #2 in the presence of 0.5 μg ml−1 Tet treatment, leading to the upregulation of TrkA expression 48 h after Tet treatment but not 24 h after Tet treatment (lanes 4–6 and 13–15). Moreover, the knockdown of hnRNP K by siRNA was clearly detectable after treatment with both 0.5 μg ml−1 Tet and 10 ng ml−1 NGF, which induced significant cleavages of TrkA and hnRNP K via an unknown mechanism (lanes 7–9 and 16–18). Interestingly, the TrkA-dependent downregulation of cell viability was enhanced by both hnRNP K siRNA #1 and #2 at 48 h after Tet treatment, but was not significantly enhanced at 24 h (Figure 6b). As expected, NGF enhanced the TrkA-dependent downregulation of cell viability, which was further strengthened by hnRNP K siRNAs. These results indicate that the inhibition of NGF/TrkA-dependent cell death by hnRNP K is mediated, at least in part, by the influence of hnRNP K on TrkA expression and modification.
Effects of hnRNP K siRNA on TrkA expression and cell viability. (a) Negative control and hnRNP siRNA #1 and #2 (10 nM each) were transfected using G-fectin into 3.5 × 105 SK-N-MC-TrkA cells in a six-well dish for 24 h and then were treated with 0.5 μg ml−1 Tet and 10 ng ml−1 NGF for 24 or 48 h. Whole cells were extracted with SDS sample buffer and were quantified using the Bradford Reagent. Total proteins (30 μg each) were separated on 10% SDS-PAGE and were analyzed by western blot using the indicated antibodies. (b) Negative control and hnRNP siRNA #1 and #2 (10 nM each) were transfected using G-fectin into 5 × 104 SK-N-MC-TrkA cells in a 24-well dish for 24 h and then were treated with 0.5 μg ml−1 Tet and 10 ng ml−1 NGF for 24 or 48 h. Cell viability was determined with a CCK-8 assay. The asterisks represent potential cleaved products of hnRNP K and TrkA.
Discussion
NGF-mediated TrkA activation is known to regulate cell survival and differentiation through the ERK, PI3K-Akt and phospholipase C-γ (PLC-γ) signaling pathways, whereas NGF-mediated p75NTR activation induces cell death through JNK signaling.15 However, NGF-mediated TrkA activation is also known to induce cell death in glioblastoma and medulloblastoma.1, 7 To better understand the NGF-mediated TrkA-dependent signaling pathways, we investigated the roles of NGF in TrkA-dependent cellular processes, morphological changes and cell death in TrkA-inducible SK-N-MC cells. Interestingly, the enhancement of TrkA-dependent apoptotic cellular processes and cell death caused by NGF was significantly suppressed by the JNK inhibitor SP600125, but not by the PI3K inhibitor wortmannin, indicating a JNK-dependent role of NGF in TrkA-induced cell-death signaling (Figures 1 and 2).
To better understand the mechanism of NGF-mediated TrkA-dependent cell death, we identified TrkA-dependent targets regulated by NGF through CBB staining and a 2D proteomic analysis. Thus, 18 upregulated and 13 downregulated protein spots by TrkA with NGF were identified using a MALDI-TOF/TOF MS/MS analysis (Figure 3). Because the enhancement of TrkA-dependent cell death caused by NGF was significantly controlled by SP600125, we performed a quantitative analysis to select protein spots that were significantly affected by SP600125 from among the 31 protein spots. Thus, the hnRNP C1/C2, PP2Acβ, SRSF3, PSα1, septin-2, lamin B1, SRSF1, hnRNP K, BAF53A and TDP43 protein spots were identified as potential targets associated with the enhancement of TrkA-induced neuroblastoma cell death caused by NGF (Figure 4). Moreover, the quantitative analysis results for lamin B1, hnRNP K and TDP43 were confirmed by a western blot analysis (Figure 5). Very interestingly, the knockdown of hnRNP K by siRNA transfection enhanced the NGF/TrkA-dependent downregulation of cell viability, indicating that hnRNP K inhibits NGF/TrkA-dependent cell death (Figure 6). Through phase-contrast light microscopy after trypan blue staining, we confirmed that the cell death caused by NGF/TrkA was significantly increased by knockdown of hnRNP K (data not shown).
The transcription cofactor hnRNP K is known to regulate transcriptional silencing by the lysine methyltransferase SETDB1 in embryonic stem cells,16 hnRNP C1/C2 regulates 1,25-dihydroxyvitamin D-directed transcriptional events in osteoblasts,17 the splicing factor SRSF1 regulates HIV transcription,18 SRSF3 is involved in enhanced splicing of glucose-6-phosphate dehydrogenase RNA in response to nutrients and hormones in the liver,19 the BAF53a subunit of ATP-dependent SWI/SNF-like BAF chromatin remodeling complexes is essential for hemopoietic stem cell function,20 and TDP-43 is involved in mRNA metabolism and regulates β-adducin (Add2) transcript stability.21 In addition, it is known that lamin B1 has an important role in the maintenance of nuclear structure and function,22 the proteasome subunit alpha type 3 (PSMA3) regulates p21/Cip1 protein stability through proteasomal degradation,23 septin 2 binds and hydrolyzes GTP similar to the Ras superfamily of GTPases,24 and PP2A regulates cell proliferation, signal transduction and apoptosis by functioning as a serine/threonine phosphatase to dephosphorylate many critical cellular proteins.25 These facts suggest that NGF-mediated TrkA activation could have important roles in the regulation of transcription via hnRNP K, hnRNP C1/C2, SRSF1, SRSF3, BAF53A and TDP43; in nuclear structure via lamin B1; in protein stability via PSα1; and in enzymatic modification via septin-2 and PP2Acβ in a JNK-dependent manner in SK-N-MC cancer cell death signaling.
Taken together, the results of this work reveal novel targets associated with the enhancement of TrkA-induced cancer cell death caused by NGF in TrkA-inducible SK-N-MC cells through a 2D proteomic analysis. These findings suggest that these targets could have an important role in NGF-mediated TrkA-induced cancer cell death signaling via a novel JNK-dependent mechanism.
References
Molloy NH, Read DE, Gorman AM . Nerve growth factor in cancer cell death and survival. Cancers 2011; 3: 510–530.
Huang CM, Shui HA, Wu YT, Chu PW, Lin KG, Kao LS et al. Proteomic analysis of proteins in PC12 cells before and after treatment with nerve growth factor: increased levels of a 43-kDa chromogranin B-derived fragment during neuronal differentiation. Brain Res Mol Brain Res 2001; 92: 181–192.
Liu T, D'mello V, Deng L, Hu J, Ricardo M, Pan S et al. A multiplexed proteomics approach to differentiate neurite outgrowth patterns. J Neurosci Methods 2006; 158: 22–29.
Tso PH, Morris CJ, Yung LY, Ip NY, Wong YH . Multiple Gi proteins participate in nerve growth factor-induced activation of c-Jun N-terminal kinases in PC12 cells. Neurochem Res 2009; 34: 1101–1112.
Björkblom B, Vainio JC, Hongisto V, Herdegen T, Courtney MJ, Coffey ET . All JNKs can kill, but nuclear localization is critical for neuronal death. J Biol Chem 2008; 283: 19704–19713.
Yan C, Liang Y, Nylander KD, Schor NF . TrkA as a life and death receptor: receptor dose as a mediator of function. Cancer Res 2002; 62: 4867–4875.
Li C, MacDonald JI, Hryciw T, Meakin SO . Nerve growth factor activation of the TrkA receptor induces cell death, by macropinocytosis, in medulloblastoma Daoy cells. J Neurochem 2010; 112: 882–899.
Nikoletopoulou V, Lickert H, Frade JM, Rencurel C, Giallonardo P, Zhang L et al. Neurotrophin receptors TrkA and TrkC cause neuronal death whereas TrkB does not. Nature 2010; 467: 59–63.
Eggert A, Grotzer MA, Ikegaki N, Liu XG, Evans AE, Brodeur GM . Expression of the neurotrophin receptor TrkA down-regulates expression and function of angiogenic stimulators in SH-SY5Y neuroblastoma cells. Cancer Res 2002; 62: 1802–1808.
Sitek B, Apostolov O, Stühler K, Pfeiffer K, Meyer HE, Eggert A et al. Identification of dynamic proteome changes upon ligand activation of Trk-receptors using two-dimensional fluorescence difference gel electrophoresis and mass spectrometry. Mol Cell Proteomics 2005; 4: 291–299.
Jung EJ, Park HC, Chung KH, Kim CW . Proteomic analysis of SP600125-controlled TrkA-dependent targets in SK-N-MC neuroblastoma cells: inhibition of TrkA activity by SP600125. Proteomics 2014; 14: 202–215.
Jung EJ, Kim DR . Apoptotic cell death in TrkA-overexpressing cells: kinetic regulation of ERK phosphorylation and caspase-7 activation. Mol Cells 2008; 26: 12–17.
Jung EJ, Lee SY, Kim CW . Proteomic analysis of novel targets associated with TrkA-mediated tyrosine phosphorylation signaling pathways in SK-N-MC neuroblastoma cells. Proteomics 2013; 13: 355–367.
Kwon YS, Ryu CM, Lee S, Park HB, Han KS, Lee JH et al. Proteome analysis of Arabidopsis seedlings exposed to bacterial volatiles. Planta 2010; 232: 1355–1370.
Reichardt LF . Neurotrophin-regulated signalling pathways. Philos Trans R Soc Lond B Biol Sci 2006; 361: 1545–1564.
Thompson PJ, Dulberg V, Moon KM, Foster LJ, Chen C, Karimi MM et al. hnRNP K coordinates transcriptional silencing by SETDB1 in embryonic stem cells. PLoS Genet 2015; 11: e1004933.
Lisse TS, Vadivel K, Bajaj SP, Chun RF, Hewison M, Adams JS . The heterodimeric structure of heterogeneous nuclear ribonucleoprotein C1/C2 dictates 1,25-dihydroxyvitamin D-directed transcriptional events in osteoblasts. Bone Res 2014; 2: pii 14011.
Paz S, Krainer AR, Caputi M . HIV-1 transcription is regulated by splicing factor SRSF1. Nucleic Acids Res 2014; 42: 13812–13823.
Walsh CM, Suchanek AL, Cyphert TJ, Kohan AB, Szeszel-Fedorowicz W, Salati LM . Serine arginine splicing factor 3 is involved in enhanced splicing of glucose-6-phosphate dehydrogenase RNA in response to nutrients and hormones in liver. J Biol Chem 2013; 288: 2816–2828.
Krasteva V, Buscarlet M, Diaz-Tellez A, Bernard MA, Crabtree GR, Lessard JA . The BAF53a subunit of SWI/SNF-like BAF complexes is essential for hemopoietic stem cell function. Blood 2012; 120: 4720–4732.
Costessi L, Porro F, Iaconcig A, Muro AF . TDP-43 regulates β-adducin (Add2) transcript stability. RNA Biol 2014; 11: 1280–1290.
Camps J, Erdos MR, Ried T . The role of lamin B1 for the maintenance of nuclear structure and function. Nucleus 2015; 6: 8–14.
Shi Z, Li Z, Li ZJ, Cheng K, Du Y, Fu H et al. Cables1 controls p21/Cip1 protein stability by antagonizing proteasome subunit alpha type 3. Oncogene 2015; 34: 2538–2545.
Huang YW, Surka MC, Reynaud D, Pace-Asciak C, Trimble WS . GTP binding and hydrolysis kinetics of human septin 2. FEBS J 2006; 273: 3248–3260; Erratum in: FEBS J 2006; 273: 4577.
Seshacharyulu P, Pandey P, Datta K, Batra SK . Phosphatase: PP2A structural importance, regulation and its aberrant expression in cancer. Cancer Lett 2013; 335: 9–18.
Acknowledgements
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2013R1A1A2058019 and NRF-2011-0003111). In addition, this research was supported by a grant from the Institute of Health Sciences, Gyeongsang National University (IHS GNU-2013-05).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Experimental & Molecular Medicine website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Jung, E., Chung, K., Bae, DW. et al. Proteomic analysis of novel targets associated with the enhancement of TrkA-induced SK-N-MC cancer cell death caused by NGF. Exp Mol Med 48, e235 (2016). https://doi.org/10.1038/emm.2016.33
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/emm.2016.33
- Springer Nature Limited