Abstract
In addition to promoting cell death and senescence, p53 also has important cellular survival functions. A mutant p53, lacking a proline-rich domain (p53ΔP), that is deficient in controlling both cell death and cell cycle arrest, was employed to determine the biological means by which p53 mediates survival upon DNA damage. While p53ΔP and p53−/− cells were equally resistant to many DNA damaging agents, p53ΔP cells showed an exquisite resistance to high doses of the alkylating agent Diazald (N-Methyl-N-(p-tolylsulfonyl)nitrosamide), as compared to cells completely deficient for p53 function. We determined that p53ΔP was capable of transcribing the repair gene, MGMT (O6-methylguanine-DNA methyltransferase) after irradiation or alkylation damage, resulting in DNA repair and cell survival. Consistent with these observations, p53ΔP mice show enhanced survival after IR relative to p53−/− mice. Suppression or deletion of MGMT expression in p53ΔP cells inhibited DNA repair and survival after alkylation damage, whereas MGMT overexpression in p53-deficient cells facilitated DNA repair and conferred survival advantage. This study shows that when cell death and cell cycle arrest pathways are inhibited, p53 can still mediate MGMT-dependent repair, to promote cell survival upon DNA damage.
Similar content being viewed by others
Main
P53 is one of the most frequently mutated genes in human tumors, and its proper activity and function is crucial for the maintenance of genomic stability and tumor suppression.1 Although it remains undisputed that p53 is a potent suppressor of tumorigenesis, the biological means by which p53 prevents cellular transformation remains unclear.2 To date, no ablation of a p53 target gene fully recapitulates the cancer predisposition of p53 null mice.3, 4, 5 The long-held hypothesis in the field predicted that upon oncogene-induced cellular stress, cell cycle arrest or apoptosis was induced, thus allowing the cell time for efficient repair of the damage, or the elimination of cells that acquired irreparable damage.6, 7 A genetic approach in which three p53 target genes required for cell cycle arrest and apoptosis were deleted, failed to phenocopy the spontaneous tumor incidence seen in p53−/− mice.8 However, given the requirement for p53 in the transcriptional regulation of hundreds of genes, it seems unlikely that the deletion of individual p53-mediated genes will produce a loss of tumor suppressor activity.8, 9, 10 We used a genetic model in which the proline-rich domain (PRD) of p53 (p53ΔP) was deleted, resulting in an altered transcription function.11 This mutant allowed us to evaluate the biological role of p53 that a majority of its target functions are ablated.
The PRD, defined by residues 58–98 and residues 62–91 of human and mouse p53 respectively, promotes p53-dependent transcription by an unclear mechanism.12 Some studies suggest that deletion of this region changes the stability of p53 protein, as a consequence of diminished Pin1 binding resulting in increased susceptibility of p53 to MDM2-dependent degradation. Others suggest that the PRD instead mediates binding to cofactors important for efficient transcription.13, 14, 15, 16, 17, 18 Nevertheless, deletion of the PRD of p53 (p53ΔP (75–91); and mΔpro (58–88)) resulted in a loss of cell cycle arrest and apoptosis; yet mice lacking this region rarely develop spontaneous tumors seen in p53−/− mice.11, 19, 20, 21, 22, 23
The repair of DNA lesions is crucial to maintain genomic stability and p53 mediates the expression of numerous repair genes involved in nucleotide excision repair, base excision repair, mismatch repair, and non-homologous end joining.24, 25 Mice with mutations or deletions in one of several DNA repair genes are prone to developing tumors.26 Methylating agents, such as chemotherapies can modify DNA at many different sites, with the most mutagenic lesion being the alkylation of O6 position of guanine (O6MeG).27, 28 These lesions are highly mutagenic unless repaired, and fail to block DNA replication.29, 30, 31 As a consequence, following two rounds of DNA replication these lesions manifest as G:C to A:T point mutations and represent a DNA adduct with the potential for tumor formation.32 In some cases, A:T transitions have been shown to arise in the KRAS oncogene and p53 tumor suppressor gene, thus promoting tumorigenesis.33, 34, 35 P53 induces the transcription of O6-methylguanine-DNA methyltransferase (MGMT), an enzyme that repairs O6MeG, via a one-step suicide reaction, involving the irreversible transfer of the methyl groups on the DNA lesion, to an internal cysteine residue in the active center of the alkyltransferase.36, 37, 38 This reaction is stoichiometric involving one molecule of MGMT for each methyl group removed.39 Following repair, MGMT is tagged by ubiquitin and degraded via the proteasome.40, 41 While MGMT-deficient mice do not develop spontaneous tumors, they are highly vulnerable to oncogenesis induced by treatment with alkylating agents.42, 43, 44, 45, 46 Conversely, mice with elevated MGMT expression levels exhibit resistance to alkylating agents and tumor formation.47, 48, 49 This suggests that MGMT is important in tumor suppression.
We determined that p53 lacking the PRD failed to mediate the expression of key genes involved in apoptosis or cell cycle arrest. While cells harboring p53ΔP were equally resistant to cell death induced by several DNA damaging agents as compared to p53−/− cells, they surprisingly showed a greater resistance to high doses of the alkylating agent Diazald, than did p53−/− cells. We determined that p53ΔP was capable of transcribing the repair gene, MGMT and could thereby induce repair alkylating DNA damage, thus promoting p53ΔP cell survival after alkylation damage.
Results
DNA damage-induced apoptosis and cell cycle arrest requires p53 with a functional proline-rich region
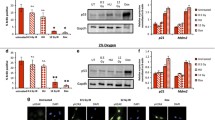
Since the proline-rich region of p53 is described to be essential for the efficient transcription function of p53,12, 50 we first determined whether common target genes involved in apoptosis and cell cycle arrest were induced upon DNA damage in cells expressing p53 lacking a proline-rich region (p53ΔP (75–91)). H1299 cells, that lack the p53 gene, were reconstituted with wild-type p53 (p53WT) or p53 lacking the proline-rich region (p53ΔP) fused to a modified estrogen receptor steroid-binding domain (ERTam) that binds 4-Hydroxytamoxifen (4OHT) specifically. Upon the addition of 4OHT for 5 h, both p21 and PUMA gene expression levels and protein levels were induced in cells expressing p53WT, but not p53ΔP (Figures 1a and c). P21 and PUMA gene expression levels were also induced after 6 h in B cells treated with 5 Gy of irradiation isolated from p53WT mice, but not those from p53ΔP or p53−/− mice (Figure 1b). E1A/RAS transformed mouse embryonic fibroblasts (MEF) from p53WT, p53ΔP, and p53−/− mice were treated with a panel of DNA damage agents and assessed for Annexin V externalization after 18 h by flow cytometry. P53WT-expressing MEF were sensitive to all DNA damaging agents tested, but p53ΔP-expressing cells and p53−/− cells were equally resistant to these death-inducing stimuli (Figure 1d). In addition, p53ΔP-expressing, and p53−/− cells exhibited reduced cell cycle arrest upon 5Gy irradiation (IR) compared to p53WT-expressing cells (Figure 1e). Even without irradiation, a greater number of p53ΔP and p53−/− cells were observed to be in S-phase as compared to p53WT-expressing cells, and exhibited more rapid growth than p53WT cells (Figure 1f). Irreversible cell cycle arrest was also compromised in p53ΔP-expressing cells, as evidenced by an inability to undergo replicative- and oncogene-induced senescence (Figures 1g and h respectively).
P53ΔP cells do not express p21 or PUMA and are resistant to apoptosis, cell cycle arrest, and senescence. p21 and PUMA gene expression levels in (a) H1299 cells expressing p53WTERTam or p53ΔPERTam constructs 5 h after 4OHT treatment, and (b) primary B cells 6 h post irradiation. (c) p21 and PUMA protein levels in H1299 cells expressing p53WTERTam or p53ΔPERTam constructs 8 h post 4OHT. (d) Cell death at 18 h as measured by flow cytometric analysis of Annexin V staining in p53WT, p53ΔP, and p53−/− MEF cells treated with a variety of apoptotic stimuli. (e) Brdu incorporation into MEFs 18 h after 4 Gy IR. Growth of (f) E1A/RAS transformed MEF cells over 4 days and (g) primary MEF cells over time. (h) Colony-forming assay looking at survival of primary MEF cells transduced with RAS oncogene
MGMT mediates repair and long-term survival in response to alkylating damage in cells expressing p53ΔP
We next used clonogenic assays to examine the long-term survival and proliferation of cells treated with DNA damaging agents. We suspected that since p53ΔP and p53−/− cells are resistant to the apoptotic effects of DNA damage in short-term assays, they would likely exhibit enhanced long-term survival and proliferation, as compared to p53WT cells. Indeed, p53ΔP-expressing and p53−/− cells exhibited better survival than their p53WT-expressing counterparts when treated with intermediate doses of UV (5 mJ/cm2) and STS (3.5 μM) (Figure 2a, Supplementary Figure 1A). P53ΔP-expressing cells exhibited a slightly enhanced sensitivity to intermediate doses of IR (12 Gy) as compared with p53−/− cells, but higher doses (14 Gy) resulted in equally diminished survival in both cell types (Figure 2a, Supplementary Figure 1A). P53 status was not important for long-term survival after treatment with Doxorubicin (0.75 μM and 1.0 μM) and Etoposide (25 μM and 50 μM).
P53ΔP cells exhibit a differential long-term survival response to alkylating agent Diazald compared with other damaging agents. (a) Colony-forming assay examining the survival of p53WT, p53ΔP and p53−/− MEF cells treated with intermediate or high doses of death-inducing agents. Red circle highlights the increase in survival of p53ΔP cells after Diazald (250 μM) treatment as compared to p53−/− cells. (b) Colony-forming assays examining survival in p53WT, p53ΔP, and p53−/− MEF cells after 250 μM Diazald treatment for increasing time intervals
Significantly however, p53ΔP-expressing cells were highly resistant to the high dose of alkylating agent, Diazald (250 μM), surviving and proliferating better than either p53WT-expressing or p53−/− cells (Figure 2a, Supplementary Figure 1A) (highlighted by red circle). To evaluate this observation further, we performed a time-course of high dose (250 μM) Diazald-treatment and analyzed proliferation and survival by clonogenic assay (Figure 2b). Compared with p53WT and p53−/− cells, p53ΔP-expressing cells showed a superior capacity to survive, even when treated for 8 h with a high dose of Diazald (Figure 2b). Only after a 24 h exposure to 250 μM Diazald, did cells expressing p53ΔP succumb to DNA damage and fail to survive (Figure 2b).
Alkylating damage induces specific types of DNA adducts at the N- and O-atoms in DNA bases. Although O6-methylguanine (O6MeG) adducts are less common, they are stable and will persist unless specifically repaired by the DNA repair protein MGMT.32 We determined that MGMT gene expression was elevated following either IR (5 Gy) or Diazald-treatment of p53WT and p53ΔP immune cells, but not in p53−/− cells (Figure 3a). P53ΔP cells expressed higher levels of MGMT after IR at 6–8 h in vitro and ex vivo compared with p53WT cells. Diazald treatment of thymocytes for 18 h showed approximately equal expression of MGMT in p53WT and p53ΔP cells (Figure 3a). P53-dependent transcription of MGMT is consistent with the identification of a consensus binding sequence for p53 within the MGMT promoter. CHIP assays confirmed direct binding of p53.51, 52 We tested whether the difference in the levels of MGMT observed in p53WT and p53ΔP after IR may be due to a difference in the regulation and stability of p53ΔP protein after damage. However, less p53ΔP is stabilized after damage, compared with p53WT, with a concomitant decrease in its negative regulator, MDM2, suggesting that there is not an increased turnover of p53ΔP protein (Supplementary Figure 2A). The difference in MGMT expression levels after IR between p53WT- and p53ΔP-cells may be due to the differences in cell survival between the cells.
The repair gene MGMT is induced in p53ΔP cells upon alkylating damage and mediates DNA repair and cell survival in cells. (a) MGMT expression levels 6–8 h IR (5 Gy) in irradiated immune cells (in vitro) or immune cells isolated from irradiated mice (ex vivo). MGMT expression levels in thymocytes treated with 50 μM Diazald for 18 h. (b) Intracellular pH2AX levels measured by flow cytometry in p53WT, p53ΔP and p53−/− MEF cell treated with 100 μM Diazald at 3 h to assess damage, or at 24 h to assess repair
Since the PRD is crucial for the efficient transcription function of p53, we were interested to determine if additional repair genes could also be induced upon IR treatment in p53ΔP cells. No repair genes analyzed were induced in a p53ΔP-manner (Supplementary Figure 1B). To assess repair of Diazald-mediated damage by MGMT, levels of phosphorylated H2AX (γH2AX) were assessed at 3 h and 24 h after Diazald treatment by intracellular staining and subsequent flow cytometry. Briefly, cells were treated for 3 h with Diazald after which they were stained for γH2AX, or washed, and re-suspended in fresh media for an additional 24 h before staining for γH2AX. Compared with untreated cells, Diazald treatment for 3 h induced an equivalent shift in fluorescence (based on mean fluorescence intensity), indicating damaged DNA, in p53WT, p53ΔP and p53−/− cells (Figure 3b). Twenty-four hours after Diazald treatment, both p53WT- and p53ΔP-expressing cells showed a decrease in the levels of detectable γH2AX staining, comparable to those measured in untreated cells, indicating that repair of Diazald-mediated damage had occurred (Figure 3b). By contrast, p53−/− cells, which are unable to induce MGMT expression, maintained high levels of γH2AX staining after 24 h, consistent with an inability to repair Diazald-imposed DNA damage (Figure 3b).
DNA damage engages the DNA damage response pathway to engage p53, and elucidate an appropriate biological response to prevent the perpetuation of potentially tumorigenic mutations.25 We therefore suggest that following DNA damage p53ΔP can induce expression of the enzyme MGMT to mediate DNA repair and promote survival.
The role of MGMT in tumorigenesis following alkylation damage has already been characterized in p53-sufficient and -deficient mice.53 To further explore this idea, we asked whether DNA damage-mediated MGMT induction promotes survival of mice after exposure to 4 Gy IR at p7. Survival after IR was examined in p53WT, p53ΔP and p53−/− mice, and as expected, p53−/− mice all died at roughly 3 months of age as a consequence of tumor formation (Figure 4a). However, mice lacking the PRD (p53ΔP) showed significant survival after IR relative to p53−/− mice, although with a slightly diminished survival when compared with p53WT mice (Figure 4a). Without MMR or apoptosis induction in p53ΔP cells, any un-repaired Diazald-mediated damage would be propagated to daughter cells, resulting in genomic instability and aneuploidy.54 We therefore examined the incidence of aneuploidy in p53WT, p53ΔP and p53−/− MEF 72 h after 4 Gy IR, as assessed by flow cytometric analysis of >4n cell cycle distribution after propidium iodide staining (Figure 4b). Compared with p53WT cells, p53ΔP and p53−/− cells displayed more aneuploidy and increased genomic instability. We observed a further increase in the incidence of aneuploidy in p53ΔP MEF lacking MGMT at 72 h following exposure to IR. This suggests that MGMT mediates some control of genomic stability in p53ΔP cells. This however was not reflected in vivo, since there was no difference in the survival of mice lacking MGMT after IR at p7 (Figure 4a).
MGMT-dependent repair does not promote survival in mice treated with IR. (a) Survival curve of mice irradiated with 4 Gy on day 7. Gehan–Breslow–Wilcoxon test used for statistical analysis. (b) Quantification of the percentage of MEF cells with aneuploidy (> 4n chromosomes) 72 h after 4 Gy IR as assessed by flow cytometric analysis of cell cycle distribution after propidium iodide staining
MGMT silencing results in diminished repair and long-term survival in response to alkylating damage in cells expressing p53ΔP
We hypothesized that cells expressing p53ΔP show superior survival after Diazald treatment because MGMT-mediated DNA repair is present and functional, despite the fact that these cells are unable to undergo cell cycle arrest and apoptosis after DNA damage (Supplementary Figure 2B). Since p53ΔP cells and p53WT cells exhibit no differential expression of MGMT protein, either basally or following DNA damage, the enhanced survival observed in p53ΔP cells is most likely due to their defect in apoptosis and/or cell cycle arrest (Figure 5a). Therefore p53WT-expressing cells show reduced survival as compared to p53ΔP cells, because they are able to undergo cell cycle arrest and apoptosis in response to alkylating damage (Supplementary Figure 2B). A lack of p53 ensures that apoptosis, cell cycle arrest, and MGMT-mediated repair are not engaged upon alkylating DNA damage. Although in short-term assays p53−/− cells are protected from apoptosis (Figure 1d), their inability to survive and proliferate long term suggests that alternative non-apoptotic cell death mechanisms are likely engaged (Figure 2b). It is possible that necrosis or mitotic catastrophe and subsequent death are engaged in these cells upon high dose Diazald treatment. We therefore hypothesized that silencing MGMT in p53ΔP-expressing cells results in less survival after Diazald treatment, similar to that seen with p53−/− cells. A lentiviral expression system was used to silence MGMT in p53WT, p53ΔP and p53−/− transformed MEF and the efficiency of silencing was confirmed in all cell lines (Figure 5a,Supplementary Figure 2C). P53ΔP and p53−/− cells remained resistant to apoptosis induced by UV, IR and Diazald as compared to p53WT-expressing cells, regardless of whether MGMT was silenced or not (Figure 5b). However, when MGMT was silenced in p53ΔP-expressing cells, Diazald-mediated damage could no longer be repaired (Figure 6a). Importantly, we also saw that these cells could no longer survive and proliferate as compared to MGMT-expressing cells when treated with high dose Diazald (Figure 6e). We also confirmed this result in MEFs isolated from p53ΔP,MGMT−/− mice (Figure 6e). In fact, these MGMT-silenced cells showed similar levels of survival and proliferation as compared to p53−/− cells (Figure 6e). Since p53−/− cells do not express MGMT, it was not surprising that no difference in repair or survival after Diazald treatment was detected when MGMT was silenced or not (Figures 6a and e). Diazald-treatment of p53WT cells in which MGMT was silenced showed less survival and proliferation in long-term assays (Figure 6e). We noted that Diazald-induced damage appeared to be repaired by 24 h in p53WT cells in which MGMT was silenced (Figure 6a). Since p53WT-expressing cells can undergo cell cycle arrest and apoptosis following the failure to repair damaged DNA (Supplementary Figure 2B), we examined cell death in parallel to repair at the 24-h endpoint. Briefly, cells were treated for 3 h with Diazald, washed and cultured for an additional 18 h. We observed twice as much apoptosis in p53WT cells lacking MGMT than any other cell line (Figure 6b). We suggest that the apparent repair of DNA damage in p53WT cells lacking MGMT is attributed to a small percentage of cells that failed to efficiently silence MGMT, maintained their repair function, and therefore escaped deletion by apoptosis or cell cycle arrest. To test this possibility, we overexpressed the anti-apoptotic protein BCLxL in all cell lines sufficient or deficient for MGMT (Supplementary Figure 3B). Diazald-induced cell death was completely inhibited in p53WT cells sufficient or deficient for MGMT in which BCLxL was overexpressed (Figure 6c). When Diazald-mediated apoptosis was inhibited by BCLxL expression, a population of cells clearly unable to repair alkylating damage was now visible in p53WT cells lacking MGMT (Figure 6d). P53−/− cells were unable to repair damage in an MGMT-mediated manner, whereas p53ΔP-expressing cells did so, unless MGMT was silenced (Figure 6a). Interestingly, despite BCLxL protection from Diazald-mediated cell death in p53WT-expressing cells, this did not confer significant long-term survival (Supplementary Figure 3C). This may be due to cells undergoing permanent cell cycle arrest and/or alternative forms of cell death. To test this we silenced p21 levels in p53WT cells overexpressing BCLxL, rendering these cells unable to engage cell cycle arrest or apoptosis (Supplementary Figure 3B). These cells showed slightly better survival relative to p53WT cells after alkylation, but did not phenocopy the survival seen in p53ΔP cells (Supplementary Figure 3C). This suggests that the response in p53ΔP differs from that of p53WT in additional ways to mediate survival after alkylation damage (Supplementary Figure 2B).
Loss of MGMT in cells does not change their apoptotic response to death inducing stimuli regardless of p53 status. (a) MGMT and p53 protein levels in p53WT, p53ΔP and p53−/− MEF cells, deficient or sufficient for MGMT, 18 h after IR (5 Gy) or Diazald (75 μM) treatment. (b) Cell death as measured flow cytometric analysis of Annexin V staining in MEF cells with or without MGMT expression treated with different doses of UV, IR or Diazald for 18 h
Loss of MGMT in cells inhibits repair and survival of p53ΔP cells. Intracellular pH2AX levels measured by flow cytometry in p53WT, p53ΔP and p53−/− MEF cells (a) sufficient or deficient for MGMT or, (d) expressing BCLxL, treated with 100 μM Diazald for 3 h to assess damage or 24 h to assess repair. (b) Cell death as measured flow cytometric analysis of Annexin V staining in MEF cells treated for 3 h with 100 μM Diazald, washed and assessed for Annexin V staining 24 h later. (c) Cell death as measured flow cytometric analysis of Annexin V staining in MEF cells expressing BCLxL and treated with 100 μM Diazald for 24 h. (e) Colony-forming assays examining survival in Left: p53WT, p53ΔP and p53−/− MEF cells sufficient or deficient for MGMT; Right: MEF cells from p53ΔP, MGMT−/− mice after 250 μM Diazald treatment for increasing time intervals
MGMT expression in p53−/− cells results in repair and survival in response to alkylating DNA damage
MGMT was overexpressed in p53WT, p53ΔP and p53−/− cells (Supplementary Fig 3A), which were then subjected to Diazald treatment. We confirmed that MGMT expression did not alter sensitivity to Diazald-induced cell death in any of the cell lines (Figure 7a). Importantly however, p53−/− cells expressing MGMT were now capable of repairing Diazald-induced DNA damage to a similar extent as that seen in p53ΔP-expressing cells (Figure 7b). Clonogenic assays confirmed that p53−/− cells expressing MGMT showed increased survival and proliferation in response to Diazald treatment (Figures 6c and d). Overexpression did not fully mimic the survival seen in p53ΔP cells, probably because MGMT-mediated repair is stoichiometric such that survival is limited to the absolute amount of MGMT molecules per cell.39
MGMT overexpression facilitates repair and some survival advantage in p53−/− cells. (a) Cell death at 18 h as measured by flow cytometric analysis of Annexin V staining in MEF cells overexpressing MGMT and treated with 100 μM Diazald. (b) Intracellular pH2AX levels measured by flow cytometry in MEF cells overexpressing MGMT and treated with 100 μM Diazald for 3 h to assess damage, or at 24 h to assess repair. (c) Colony-forming assays examining survival in p53WT, p53ΔP or p53−/− MEF cells with or without MGMT overexpression after 250 μM Diazald for increasing time intervals. (d) Quantification of colony-forming assay in p53WT, p53ΔP, or p53−/− MEF cells
Discussion
While some studies utilizing p53ΔP mice suggested that cell cycle arrest is constrained but apoptosis remains functional, others suggest that the opposite is true.11, 22 In the study described herein, we found that p53ΔP cannot induce the expression of p21 and PUMA, and cells lacking the PRD are correspondingly deficient for apoptosis, cell cycle arrest, and senescence in response to DNA damage to a similar extent as seen in cells fully deficient for p53 function. Importantly however, unlike p53−/− mice that develop spontaneous tumors, p53ΔP mice rarely do, although some eventually succumbed to tumors with a median survival of approximately 400 days.22, 55 We propose that biological functions of p53 are maintained in p53ΔP, and must be engaged for tumor suppression.
We observed that p53ΔP cells exhibited a differential long-term survival response to the alkylating agent Diazald, as compared to other DNA damaging agents. We determined that MGMT levels were induced in a p53-dependent manner in response to IR and Diazald, resulting in repair and survival of cells exposed to DNA damage. Importantly, our results corroborated those using genetic mouse models in which p53 showed an altered biological function.8, 52 Cells from mice deficient for p21, PUMA, and NOXA displayed γ-irradiation-induced MGMT expression (among other repair genes), and could clear DNA lesions more rapidly than that observed in p53-deficient cells.8 Additionally, gene expression profiles of oncogene-transduced HRAS-transduced MEF from the TAD mutant knockin (p53L25Q, W26S) mouse showed the induction of repair genes, including MGMT, and identified a p53 consensus-binding site in the MGMT promoter.52 These studies suggested that MGMT is induced in a p53-dependent manner after DNA-damage or in response to oncogenes. Our studies show that in the absence of apoptosis, cell cycle arrest, and senescence, p53ΔP can mediate cell survival through MGMT-mediated DNA repair.
P53−/− mice have an increased incidence of MNU-induced thymic lymphomas compared to MNU-treated wild-type mice, and overexpression of MGMT in the thymus of p53+/− mice significantly reduced this lymphoma incidence.53 We found that whole body ionizing radiation (4 Gy) induced p53-mediated MGMT expression, facilitating DNA repair. While p53−/− mice succumbed to tumors by 3 months, this was dramatically reduced in p53ΔP mice after whole body irradiation. Other studies have shown some differences in the survival between p53WT and p53ΔP mice after whole body irradiation, which we also observed.22, 55 It is feasible that the accumulating aneuploidy seen in p53ΔP cells after irradiation may contribute to the difference in survival seen in p53ΔP mice compared with p53WT mice. Indeed, MEF lacking MGMT show increased aneuploidy after irradiation.56 Both MEF cells and mice lacking the PRD, exhibit elevated aneuploidy in comparison to WT cells, under basal and irradiated conditions.21, 55
Although MGMT-deficient mice fail to develop spontaneous tumors, they do exhibit an increased sensitivity to the genotoxic effects of methylating agents, and are highly vulnerable to tumors induced by alkylating agents as compared to WT mice.42, 43, 44, 45, 46 We saw that contrary to our expectation, irradiated p53ΔP mice lacking MGMT had a median survival of 250 days, failing to phenocopy p53−/− mice. As p53ΔP, MGMT−/− MEF showed higher levels of aneuploidy after irradiation than with p53ΔP MEF, the extent to which aneuploidy may contribute to tumor induction remains unclear. Mice with defective mitotic checkpoint genes exhibit chromosomal changes in the absence of structural abnormalities, and despite the presence of aneuploidy, many of these mice are not predisposed to tumor formation.57 A p53 mutant mouse (p533K KI) exhibits aneuploidy and genomic instability without spontaneous tumor formation.58, 59 Therefore, chromosomal aneuploidy may be a consequence rather than a cause of deregulated cell growth.
We found that suppression or deletion of MGMT expression in p53ΔP cells inhibited DNA repair and survival after alkylation damage and that its overexpression in p53-deficient cells facilitated DNA repair and conferred a survival advantage to the cells.
While p53-induced MGMT repair of DNA damage may be beneficial to prevent genomic instability and potential transformation of cells, its expression in established tumors may be detrimental to patient treatment and survival. Interestingly, many different types of cancers show promoter hypermethylation of MGMT, resulting in diminished MGMT expression.60 The epigenetic silencing of the MGMT gene compromises DNA repair mechanisms and increases chemosensitivity, and has been shown to be a strong predictor of survival.60, 61, 62 While MGMT mediated repair of DNA damage is crucial for survival of normal cells, a lack of MGMT induction in established tumors lacking p53 may actually be advantageous for overall patient survival.
Materials and methods
Plasmids and retroviral transduction
The following previously published plasmids were used in this study to generate stable cell lines: pBabePuro.p53ERtam and pBabePuro.p53ΔPPERtam63 and LZRS-HA-BclxL.64 Mouse Mgmt cDNA (Thermo Scientific, Waltham, MA, USA) was cloned into the retroviral expression vector, LZRS-Zeo, and fully sequenced. To generate stable cell lines, Phoenix virus producer cells were transfected with the appropriate plasmid by using Lipofectamine 2000 for 48 h. Target cells were infected with filtered virus containing the culture medium from packaging cells supplemented with 5 μg/ml polybrene. Stable transductants were selected after adding 200 μg/ml Zeocin (pLZRS vectors) (Invitrogen, Waltham, MA, USA), or 1 μg/ml puromycin (pBabe vectors) (Sigma-Aldrich, St. Louis, MO, USA). To generate knockdown of MGMT in MEF cells, we used SMARTchoice mouse MGMT-GFP shRNA (Thermo Scientific), and selected for GFP-positive clones. We used SMARTpool ON-TARGETplus Cdkn1a siRNA (Dharmacon, Lafayette, LA, USA), to silence p21.
Cell lines and cell culture
All cell lines were maintained at 37 °C/5% v/v CO2 in a humidified incubator. H1299 cells ATCC expressing pBabePuro.p53ERtam or pBabePuro.p53ΔPPERtam were cultured in Dulbecco’s modified essential medium (DMEM) (GIBCO, Waltham, MA, USA) supplemented with 10% dextran/charcoal-treated fetal bovine serum (dcFBS), L-glutamine, and penicillin. Primary MEF cells were transformed with H-RasG12V and 12S E1A (pWZL-Hygro-12S-E1A and pBabe-Puro-H-RasG12V) and cultured in DMEM supplemented with 10% FBS, 2 mM L-glutamine (GIBCO), and 100 U/ml penicillin–streptomycin (GIBCO), 1 mM sodium pyruvate (GIBCO), nonessential amino acids (GIBCO), and 55 μM β-mercaptoethanol (GIBCO).
Flow cytometry
For Annexin V staining, detached and adherent cells were harvested, washed, and re-suspended in Annexin V binding buffer (10 mM HEPES, 150 mM NaCl, 5 mM KCl, 1 mM MgCl2, and 1.8 mM CaCl2) with Annexin V-APC or Annexin V-FITC (1/200; Caltag Laboratories, Waltham, MA, USA). For intracellular pH2AX staining, cells were treated with 100 μM Diazald (Sigma-Aldrich) for 3 h to assess damage, or treated with 100 μM Diazald for 3 h, washed twice with PBS and re-suspended in cell culture media (DMEM (GIBCO) supplemented with 10% FBS, 2 mM L-glutamine (GIBCO), to assess repair. Cells at 3 and 24 h were fixed and permeabilized using a cytofix/cytoperm kit according to the manufacturer’s instructions (BD Biosciences, San Jose, CA, USA), and stained with Alexa Fluor-647 or Alexa Fluor-488 mouse anti-H2AX (pS139) antibody (1/200; BD Biosciences). Experiments were quantitated by flow cytometry, using FACScan and FACsCalibur systems (BD Biosciences) and FlowJo Collectors’ Edition software (Tree Star, Ashland, USA). Flow cytometry data are represented as mean±S.D. of three independent experiments, unless otherwise stated.
Cell growth, senescence and cell cycle analysis
Transformed MEF cells were plated at 5e4 and growth rate determined by viable cell count through trypan-blue dye exclusion. Cell cycle analysis was performed 24 h after 4 Gy y-irradiation using the BD Pharmingen BrdU Flow Kit staining procedure, and analyzed by flow cytometry and cell cycle analysis. Replicative senescence was examined in primary MEF after serial passaging. All primary MEF were plated at 2e5 and the number of days taken to reach 90% confluence was noted. Cells were subsequently split at a ratio of 1:4, which represents a population doubling of 2. Total cell number was then calculated based on the number of days taken to reach 90% confluence. Oncogene-induced senescence was examined in primary MEF cells plated at 1e3 and transduced with H-RasG12V. Puromycin-resistant clones were stained with methylene blue. To determine aneuplodidy, cells were treated with 4 Gy y-irradiation and 72 h later cells were lysed and stained using a hypotonic buffer (0.1% Triton-100, 0.1% sodium citrate, 50 μg/ml PI) for 30 min on ice to determine the fraction of nuclei with >4N DNA content by flow cytometry and cell cycle analysis.
Apoptosis and survival assays
Cells were treated for 18–24 h with the following DNA damaging agents: Actinomycin D (ActD; Sigma-Aldrich), Staurosporine (STS; Sigma-Aldrich), Etoposide (Sigma-Aldrich), Diazald (Sigma-Aldrich), y-irradiation, and UV irradiation (using a Stratagene UV cross-linker). Clonogenic survival was assessed after methylene blue staining of cells treated with y-irradiation, and UV irradiation, or after treatment with very high doses of DNA damaging agents; Staurosporine (3.5 and 10 μM), Etoposide (25 and 50 μM), Doxorubicin (Sigma-Aldrich, 0.75 and 1 μM) and Diazald (150 and 250 μM) for 3 h, washed and suspended in media.
Irradiation tumor model
Mice were treated with 4 Gy γ-irradiation at p7 and monitored for clinical symptoms. All experiments were done in accordance with the Guide for the Care and Use of Laboratory Animals, and the St. Jude Institutional Animal Care and Use Committee approved all animal procedures.
Western blotting
Cells were lysed in cell lysis buffer (50 mM Tris-Cl pH 7.4, 150 mM NaCl, complete protease inhibitors cocktail (Roche, Indianapolis, IN, USA), and 0.5% Nonidet P-40). Protein concentration in cell lysates was measured by the BCA assay (Pierce, Waltham, MA, USA) and systematically normalized before western blotting. Lysates were resolved by SDS-PAGE. The proteins were transferred to supported Hybond C nitrocellulose (Amersham Bioscience, Pittsburgh, PA, USA) and immunodetected using appropriate primary and peroxidase-coupled secondary antibodies (Amersham Bioscience). Proteins were visualized by enhanced chemiluminescence (Amersham Bioscience).
The following antibodies were used for western blotting: anti-hp53 (Do7 clone, BD Pharmingen, San Jose, CA, USA) anti-mp53 (1C12, Cell Signaling, Danvers, MA, USA), anti-p21 (clone SXM30, BD Pharmingen), anti-PUMA/bbc3 (N-terminal, Sigma-Aldrich), anti-MGMT (Clone 300008, RnD systems, Minneapolis, MN, USA), anti-BCLxLs/L (D3, Santa Cruz, Dallas, TX, USA) and anti-actin (clone C4, Santa Cruz, St. Louis, MO, USA).
qRT-PCR
Cells were treated with 4 Gy γ-irradiation or with 100 μM 4-Hydroxytamoxifen (Sigma-Aldrich) and RNA for qRT-PCR was extracted using the RNeasy Mini Kit (Qiagen, Germantown, MD, USA). Reverse-transcription reactions were preformed with M-MLV reverse transcriptase (Invitrogen) following the manufacturer’s protocol and using random hexamers. Real-time PCR was performed with SYBR Green and a 7500 Fast Real-Time PCR System (Applied Biosystems, Waltham, MA, USA). The following primers were used: m18S-S 5′-ATGGTAGTCGCCGTGCCTAC-3′, m18S-AS 5′-CCGGAATCGAACCCTGATT-3′; mp21-S 5′-CCGTTGTCTCTTCGGTCCC-3′, mp21-AS 5′-CATGAGCGCATCGCAATC-3′; mPUMA-S 5′-AGCAGCACTTAGAGTCGCC-3′, mPUMA-AS 5′-CCTGGGTAAGGGGAGGAGT-3′; mMGMT-S 5′-CTGCATGGGATACGGTTGCT-3′, mMGMT-AS 5′-GTTCACGGAAATAGGCTTCCAG-3′; hGAPDH-S 5′-TCATTTCCTGGTATGACAACG-3′, hGAPDH-AS 5′-ATGTGGGCCATGAGGT-3′; hp21-S 5′-GCGATGGAACTTCGACTTTG-3, hp21-AS 5′-CAGGTCCACATGGTCTTCCT-3′; hPUMA-S 5′-CTCAACGCACAGTACGAG-3′; hPUMA-AS 5′-GTCCCATGATGAGATTGTACAG-3′; mGADD45a-S 5′-CCGAAAGGATGGACACGGTG-3′, mGADD45a-AS 5′-TTATCGGGGTCTACGTTGAGC-3′; mMLH1-S 5′-GTTTTACTCCATTCGGAAGCAGT-3′, mMLH1-AS 5′-TGTGAGCGGAAGGCTTTATAGAT-3′; mPOLK-S 5′-AGCTCAAATTACCAGCCAGCA-3′, mPOLK-AS 5′-GGTTGTCCCTCATTTCCACAG-3′; mXPC-S 5′-TCCAGGGGACCCCACAAAT-3′, mXPC-AS 5′-GCTTTTTGGGTGTTTCTTTGCC-3′; mPMS2-S 5′-GAGCAAACCGAAGGCGTGA-3′, mPMS2-AS 5′-GCGGTGCTTAAACTGAGTACC-3′; mMSH2-S 5′-GTGCAGCCTAAGGAGACGC-3′, mMSH2-AS 5′-CTGGGTCTTGAACACCTCGC-3′; mRRM2B-S 5′-GAGCCACTCCTAAGAAAGAGTTC-3′, mRRM2B-AS 5′-GAGGGAGGTCCTTTGACAAGT-3′.
References
Lane DP . Cancer. p53, guardian of the genome. Nature 1992; 358: 15–16.
Bieging KT, Mello SS, Attardi LD . Unravelling mechanisms of p53-mediated tumour suppression. Nat Rev Cancer 2014; 14: 359–370.
Harvey M, McArthur MJ, Montgomery CA Jr., Butel JS, Bradley A, Donehower LA . Spontaneous and carcinogen-induced tumorigenesis in p53-deficient mice. Nat Genet 1993; 5: 225–229.
Donehower LA, Harvey M, Slagle BL, McArthur MJ, Montgomery CA Jr., Butel JS et al. Mice deficient for p53 are developmentally normal but susceptible to spontaneous tumours. Nature 1992; 356: 215–221.
Jacks T, Remington L, Williams BO, Schmitt EM, Halachmi S, Bronson RT et al. Tumor spectrum analysis in p53-mutant mice. Curr Biol 1994; 4: 1–7.
Levine AJ . p53, the cellular gatekeeper for growth and division. Cell 1997; 88: 323–331.
Jeggo PA, Pearl LH, Carr AM . DNA repair, genome stability and cancer: a historical perspective. Nat Rev Cancer 2016; 16: 35–42.
Valente LJ, Gray DH, Michalak EM, Pinon-Hofbauer J, Egle A, Scott CL et al. p53 efficiently suppresses tumor development in the complete absence of its cell-cycle inhibitory and proapoptotic effectors p21, Puma, and Noxa. Cell Rep 2013; 3: 1339–1345.
Valente LJ, Grabow S, Vandenberg CJ, Strasser A, Janic A . Combined loss of PUMA and p21 accelerates c-MYC-driven lymphoma development considerably less than loss of one allele of p53. Oncogene 2015; 35: 3866–3871.
Riley T, Sontag E, Chen P, Levine A . Transcriptional control of human p53-regulated genes. Nat Rev Mol Cell Biol 2008; 9: 402–412.
Toledo F, Krummel KA, Lee CJ, Liu CW, Rodewald LW, Tang M et al. A mouse p53 mutant lacking the proline-rich domain rescues Mdm4 deficiency and provides insight into the Mdm2-Mdm4-p53 regulatory network. Cancer Cell 2006; 9: 273–285.
Walker KK, Levine AJ . Identification of a novel p53 functional domain that is necessary for efficient growth suppression. Proc Natl Acad Sci USA 1996; 93: 15335–15340.
Dornan D, Shimizu H, Burch L, Smith AJ, Hupp TR . The proline repeat domain of p53 binds directly to the transcriptional coactivator p300 and allosterically controls DNA-dependent acetylation of p53. Mol Cell Biol 2003; 23: 8846–8861.
Berger M, Vogt Sionov R, Levine AJ, Haupt Y . A role for the polyproline domain of p53 in its regulation by Mdm2. J Biol Chem 2001; 276: 3785–3790.
Wulf GM, Liou YC, Ryo A, Lee SW, Lu KP . Role of Pin1 in the regulation of p53 stability and p21 transactivation, and cell cycle checkpoints in response to DNA damage. J Biol Chem 2002; 277: 47976–47979.
Zheng H, You H, Zhou XZ, Murray SA, Uchida T, Wulf G et al. The prolyl isomerase Pin1 is a regulator of p53 in genotoxic response. Nature 2002; 419: 849–853.
Zilfou JT, Hoffman WH, Sank M, George DL, Murphy M . The corepressor mSin3a interacts with the proline-rich domain of p53 and protects p53 from proteasome-mediated degradation. Mol Cell Biol 2001; 21: 3974–3985.
Berger M, Stahl N, Del Sal G, Haupt Y . Mutations in proline 82 of p53 impair its activation by Pin1 and Chk2 in response to DNA damage. Mol Cell Biol 2005; 25: 5380–5388.
Toledo F, Lee CJ, Krummel KA, Rodewald LW, Liu CW, Wahl GM . Mouse mutants reveal that putative protein interaction sites in the p53 proline-rich domain are dispensable for tumor suppression. Mol Cell Biol 2007; 27: 1425–1432.
Edwards SJ, Hananeia L, Eccles MR, Zhang YF, Braithwaite AW . The proline-rich region of mouse p53 influences transactivation and apoptosis but is largely dispensable for these functions. Oncogene 2003; 22: 4517–4523.
Campbell HG, Mehta R, Neumann AA, Rubio C, Baird M, Slatter TL et al. Activation of p53 following ionizing radiation, but not other stressors, is dependent on the proline-rich domain (PRD). Oncogene 2013; 32: 827–836.
Slatter TL, Ganesan P, Holzhauer C, Mehta R, Rubio C, Williams G et al. p53-mediated apoptosis prevents the accumulation of progenitor B cells and B-cell tumors. Cell Death Differ 2010; 17: 540–550.
Sakamuro D, Sabbatini P, White E, Prendergast GC . The polyproline region of p53 is required to activate apoptosis but not growth arrest. Oncogene 1997; 15: 887–898.
Hoeijmakers JH . Genome maintenance mechanisms for preventing cancer. Nature 2001; 411: 366–374.
Williams AB, Schumacher B . p53 in the DNA-damage-repair process. Cold Spring Harb Perspect Med 2016; 6: 1–16.
Bernstein C, Bernstein H, Payne CM, Garewal H . DNA repair/pro-apoptotic dual-role proteins in five major DNA repair pathways: fail-safe protection against carcinogenesis. Mutat Res 2002; 511: 145–178.
Beranek DT . Distribution of methyl and ethyl adducts following alkylation with monofunctional alkylating agents. Mutat Res 1990; 231: 11–30.
Goth-Goldstein R . Inability of Chinese hamster ovary cells to excise O6-alkylguanine. Cancer Res 1980; 40: 2623–2624.
Toorchen D, Topal MD . Mechanisms of chemical mutagenesis and carcinogenesis: effects on DNA replication of methylation at the O6-guanine position of dGTP. Carcinogenesis 1983; 4: 1591–1597.
Larson K, Sahm J, Shenkar R, Strauss B . Methylation-induced blocks to in vitro DNA replication. Mutat Res 1985; 150: 77–84.
Snow ET, Foote RS, Mitra S . Base-pairing properties of O6-methylguanine in template DNA during in vitro DNA replication. J Biol Chem 1984; 259: 8095–8100.
Kaina B, Christmann M, Naumann S, Roos WP . MGMT: key node in the battle against genotoxicity, carcinogenicity and apoptosis induced by alkylating agents. DNA Repair (Amst) 2007; 6: 1079–1099.
Nagasaka T, Goel A, Notohara K, Takahata T, Sasamoto H, Uchida T et al. Methylation pattern of the O6-methylguanine-DNA methyltransferase gene in colon during progressive colorectal tumorigenesis. Int J Cancer 2008; 122: 2429–2436.
Esteller M, Risques RA, Toyota M, Capella G, Moreno V, Peinado MA et al. Promoter hypermethylation of the DNA repair gene O(6)-methylguanine-DNA methyltransferase is associated with the presence of G:C to A:T transition mutations in p53 in human colorectal tumorigenesis. Cancer Res 2001; 61: 4689–4692.
Zaidi NH, Pretlow TP, O'Riordan MA, Dumenco LL, Allay E, Gerson SL . Transgenic expression of human MGMT protects against azoxymethane-induced aberrant crypt foci and G to A mutations in the K-ras oncogene of mouse colon. Carcinogenesis 1995; 16: 451–456.
Rafferty JA, Clarke AR, Sellappan D, Koref MS, Frayling IM, Margison GP . Induction of murine O6-alkylguanine-DNA-alkyltransferase in response to ionising radiation is p53 gene dose dependent. Oncogene 1996; 12: 693–697.
Grombacher T, Eichhorn U, Kaina B . p53 is involved in regulation of the DNA repair gene O6-methylguanine-DNA methyltransferase (MGMT) by DNA damaging agents. Oncogene 1998; 17: 845–851.
Pegg AE . Mammalian O6-alkylguanine-DNA alkyltransferase: regulation and importance in response to alkylating carcinogenic and therapeutic agents. Cancer Res 1990; 50: 6119–6129.
Pegg AE . Properties of mammalian O6-alkylguanine-DNA transferases. Mutat Res 1990; 233: 165–175.
Srivenugopal KS, Yuan XH, Friedman HS, Ali-Osman F . Ubiquitination-dependent proteolysis of O6-methylguanine-DNA methyltransferase in human and murine tumor cells following inactivation with O6-benzylguanine or 1,3-bis(2-chloroethyl)-1-nitrosourea. Biochemistry 1996; 35: 1328–1334.
Xu-Welliver M, Pegg AE . Degradation of the alkylated form of the DNA repair protein, O(6)-alkylguanine-DNA alkyltransferase. Carcinogenesis 2002; 23: 823–830.
Sakumi K, Shiraishi A, Shimizu S, Tsuzuki T, Ishikawa T, Sekiguchi M . Methylnitrosourea-induced tumorigenesis in MGMT gene knockout mice. Cancer Res 1997; 57: 2415–2418.
Iwakuma T, Sakumi K, Nakatsuru Y, Kawate H, Igarashi H, Shiraishi A et al. High incidence of nitrosamine-induced tumorigenesis in mice lacking DNA repair methyltransferase. Carcinogenesis 1997; 18: 1631–1635.
Glassner BJ, Weeda G, Allan JM, Broekhof JL, Carls NH, Donker I et al. DNA repair methyltransferase (Mgmt) knockout mice are sensitive to the lethal effects of chemotherapeutic alkylating agents. Mutagenesis 1999; 14: 339–347.
Shiraishi A, Sakumi K, Sekiguchi M . Increased susceptibility to chemotherapeutic alkylating agents of mice deficient in DNA repair methyltransferase. Carcinogenesis 2000; 21: 1879–1883.
Becker K, Gregel C, Fricke C, Komitowski D, Dosch J, Kaina B . DNA repair protein MGMT protects against N-methyl-N-nitrosourea-induced conversion of benign into malignant tumors. Carcinogenesis 2003; 24: 541–546.
Dumenco LL, Allay E, Norton K, Gerson SL . The prevention of thymic lymphomas in transgenic mice by human O6-alkylguanine-DNA alkyltransferase. Science 1993; 259: 219–222.
Nakatsuru Y, Matsukuma S, Nemoto N, Sugano H, Sekiguchi M, Ishikawa T . O6-methylguanine-DNA methyltransferase protects against nitrosamine-induced hepatocarcinogenesis. Proc Natl Acad Sci USA 1993; 90: 6468–6472.
Becker K, Dosch J, Gregel CM, Martin BA, Kaina B . Targeted expression of human O(6)-methylguanine-DNA methyltransferase (MGMT) in transgenic mice protects against tumor initiation in two-stage skin carcinogenesis. Cancer Res 1996; 56: 3244–3249.
Zhu J, Jiang J, Zhou W, Zhu K, Chen X . Differential regulation of cellular target genes by p53 devoid of the PXXP motifs with impaired apoptotic activity. Oncogene 1999; 18: 2149–2155.
Fritz G, Tano K, Mitra S, Kaina B . Inducibility of the DNA repair gene encoding O6-methylguanine-DNA methyltransferase in mammalian cells by DNA-damaging treatments. Mol Cell Biol 1991; 11: 4660–4668.
Brady CA, Jiang D, Mello SS, Johnson TM, Jarvis LA, Kozak MM et al. Distinct p53 transcriptional programs dictate acute DNA-damage responses and tumor suppression. Cell 2011; 145: 571–583.
Reese JS, Allay E, Gerson SL . Overexpression of human O6-alkylguanine DNA alkyltransferase (AGT) prevents MNU induced lymphomas in heterozygous p53 deficient mice. Oncogene 2001; 20: 5258–5263.
Karran P, Bignami M . DNA damage tolerance, mismatch repair and genome instability. Bioessays 1994; 16: 833–839.
Adams CJ, Yu JS, Mao JH, Jen KY, Costes SV, Wade M et al. The Trp53 delta proline (Trp53DeltaP) mouse exhibits increased genome instability and susceptibility to radiation-induced, but not spontaneous, tumor development. Mol Carcinog 2015; 55: 1387–1396.
Kaina B . Mechanisms and consequences of methylating agent-induced SCEs and chromosomal aberrations: a long road traveled and still a far way to go. Cytogenet Genome Res 2004; 104: 77–86.
Ricke RM, van Ree JH, van Deursen JM . Whole chromosome instability and cancer: a complex relationship. Trends Genet 2008; 24: 457–466.
Li T, Kon N, Jiang L, Tan M, Ludwig T, Zhao Y et al. Tumor suppression in the absence of p53-mediated cell-cycle arrest, apoptosis, and senescence. Cell 2012; 149: 1269–1283.
Li T, Liu X, Jiang L, Manfredi J, Zha S, Gu W . Loss of p53-mediated cell-cycle arrest, senescence and apoptosis promotes genomic instability and premature aging. Oncotarget 2016; 7: 11838–11849.
Jacinto FV, Esteller M . MGMT hypermethylation: a prognostic foe, a predictive friend. DNA Repair (Amst) 2007; 6: 1155–1160.
Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 2005; 352: 997–1003.
Wiewrodt D, Nagel G, Dreimuller N, Hundsberger T, Perneczky A, Kaina B . MGMT in primary and recurrent human glioblastomas after radiation and chemotherapy and comparison with p53 status and clinical outcome. Int J Cancer 2008; 122: 1391–1399.
Chipuk JE, Maurer U, Green DR, Schuler M . Pharmacologic activation of p53 elicits Bax-dependent apoptosis in the absence of transcription. Cancer Cell 2003; 4: 371–381.
Llambi F, Moldoveanu T, Tait SW, Bouchier-Hayes L, Temirov J, McCormick LL et al. A unified model of mammalian BCL-2 protein family interactions at the mitochondria. Mol Cell 2011; 44: 517–531.
Acknowledgements
The authors thank H. Beere for manuscript editing. National Institute of Health (NIH 096208) and ALSAC supported this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Melino
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Rights and permissions
About this article
Cite this article
Baran, K., Yang, M., Dillon, C. et al. The proline rich domain of p53 is dispensable for MGMT-dependent DNA repair and cell survival following alkylation damage. Cell Death Differ 24, 1925–1936 (2017). https://doi.org/10.1038/cdd.2017.116
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2017.116
- Springer Nature Limited
This article is cited by
-
Elucidating the clonal relationship of esophageal second primary tumors in patients with laryngeal squamous cell carcinoma
Infectious Agents and Cancer (2023)
-
Cell death in cancer in the era of precision medicine
Genes & Immunity (2019)