Abstract
Mitochondrial disorders are a group of pathologies characterized by impairment of mitochondrial function mainly due to defects of the respiratory chain and consequent organellar energetics. This affects organs and tissues that require an efficient energy supply, such as brain and skeletal muscle. They are caused by mutations in both nuclear- and mitochondrial DNA (mtDNA)-encoded genes and their clinical manifestations show a great heterogeneity in terms of age of onset and severity, suggesting that patient-specific features are key determinants of the pathogenic process. In order to correlate the genetic defect to the clinical phenotype, we used a cell culture model consisting of fibroblasts derived from patients with different mutations in the mtDNA-encoded ND5 complex I subunit and with different severities of the illness. Interestingly, we found that cells from patients with the 13514A>G mutation, who manifested a relatively late onset and slower progression of the disease, display an increased autophagic flux when compared with fibroblasts from other patients or healthy donors. We characterized their mitochondrial phenotype by investigating organelle turnover, morphology, membrane potential and Ca2+ homeostasis, demonstrating that mitochondrial quality control through mitophagy is upregulated in 13514A>G cells. This is due to a specific downregulation of mitochondrial Ca2+ uptake that causes the stimulation of the autophagic machinery through the AMPK signaling axis. Genetic and pharmacological manipulation of mitochondrial Ca2+ homeostasis can revert this phenotype, but concurrently decreases cell viability. This indicates that the higher mitochondrial turnover in complex I deficient cells with this specific mutation is a pro-survival compensatory mechanism that could contribute to the mild clinical phenotype of this patient.
Similar content being viewed by others
Main
Mitochondrial disorders include a wide range of pathological conditions characterized by defects in organelle homoeostasis and energy metabolism, in particular in the electron transport chain (ETC) complexes. They are mostly caused by mutations in nuclear- or mtDNA-encoded genes of the respiratory chain complexes leading to a variety of clinical manifestations, ranging from lesions in specific tissues, such as in Leber’s hereditary optic neuropathy, to complex multisystem syndromes, such as myoclonic epilepsy with ragged-red fibers, Leigh syndrome or the mitochondrial encephalomyopathy, lactic acidosis and stroke-like episodes syndrome (MELAS).1, 2 Despite the detailed knowledge of the molecular defects in these diseases, their pathogenesis remains poorly understood. The heterogeneity of signs and symptoms depends on the diversity of the genetic background and on patient-specific compensatory mechanisms. Several studies investigated the consequences of nuclear DNA mutations on intracellular organelle physiology and Ca2+ homeostasis.3, 4 Here we analyzed a cohort of patients with mutations in the mtDNA-encoded ND5 subunit of NADH dehydrogenase in order to correlate the clinical phenotype with relevant intracellular parameters involved in mitochondrial physiology, such as the rate of autophagy and mitophagy fluxes, mitochondrial Ca2+ dynamics, mitochondrial membrane potential and their functional relationship.
Mitochondrial Ca2+ is a key regulator of organelle physiology, and impairment of cation homeostasis is a general feature of many pathological conditions, including mitochondrial diseases.5 In addition, Ca2+ uptake in this organelle has recently been demonstrated to be a fundamental regulator of autophagy.6, 7 Autophagy is involved in physiological organelle turnover and in the removal of damaged or non-functional mitochondria by autophagy (called ‘mitophagy’)8, 9, 10, 11 and is critical for organelle quality control. Given the pivotal role of mitochondrial Ca2+ in the adaptation of adenosine triphosphate (ATP) production to cellular energy demand, the recent identification of the channel responsible for Ca2+ entry into the organelle, the mitochondrial Ca2+ uniporter (MCU), is instrumental for the understanding of the regulation of mitochondrial Ca2+ transport in both physiological and pathological conditions. MCU was identified in 2011,12, 13 and in the following years, molecular insight on its complex regulatory mechanism was obtained. The pore region is composed of MCU, its isoform MCUb14 and essential MCU regulator (EMRE).15 The channel is gated by the Ca2+-sensitive proteins mitochondrial Ca2+ uptake 1 (MICU1) and MICU216, 17, 18, 19 and further regulated by the SLC25A23 protein.20 As to its cellular function, mitochondrial Ca2+ has been shown to stimulate ATP production by positive regulation of three key dehydrogenases of the tricarboxylic acid cycle21 and of the ETC.22 In parallel, unregulated and sustained organelle Ca2+ overload can also lead to the opening of the mitochondrial permeability transition pore,23, 24 with consequent dissipation of mitochondrial membrane potential (ΔΨmt), release of caspase cofactors and activation of the apoptotic cascade.5 Despite the significant molecular understanding of all these cellular processes, their role in the pathogenesis of mitochondrial diseases is still poorly understood. Here we investigated the interplay of these pathways and the possibility of their contribution to determine the severity of the pathology in a cellular model consisting of fibroblasts from patients carrying mutations in the mitochondrial ND5 gene.
Results
The autophagic flux is increased in cells with 13514A>G mutation
Given the wide variability in the clinical manifestations of complex I deficiency, we investigated whether a correlation between the severity of the clinical phenotype and mutation-specific cellular features could exist. We used a cellular model consisting of primary human skin fibroblasts derived from a cohort of four individuals (Pat#1 (patient 2 in Corona et al.25), Pat#2 (patient 8 in Bugiani et al.26), Pat#3 (patient 4 in Bugiani et al.26), Pat#4 (patient 6 in Malfatti et al.27)) bearing mutations in the ND5 gene and from three independent healthy individuals as controls (Ctrl#1, Ctrl#2 and Ctrl#3). Pat#1 and Pat#2 are two unrelated patients with the same 13514A>G mutation, displaying atypical MELAS and Leigh syndrome, respectively, Pat#3 is a Leigh patient with a different mutation (13513G>A) and Pat#4 has the 13063G>A mutation and is affected by typical MELAS. Interestingly, the atypical phenotype of Pat#1 and Pat#2 is characterized by late onset, relatively slow progression of the neurological signs, normal plasma lactate and muscle morphology.
We first monitored the activation of autophagy, a key pro-survival process induced by a variety of stress conditions. We challenged patient and control fibroblasts with nutrient deprivation (Starvation) and monitored the appearance of autophagosomes by immunofluorescence of endogenous microtubule-associated protein 1A/1B light chain 3B (LC3). In this experimental setup, the lipidated form of LC3, LC3-II, redistributes to autophagosomes, thus appearing as fluorescent dots inside the cells.28 Interestingly, fibroblasts derived from both patients carrying the 13514A>G ND5 mutation (Pat#1 and Pat#2) clearly show, already in the fed condition, an increase in the number and size of the LC3-positive dots compared with controls, Pat#3 and Pat#4 cells (Figures 1a and b). Thus the higher cellular autophagy correlates with the milder clinical phenotype, suggesting that this process could be protective during pathogenesis.
Autophagic flux is enhanced in 13514A>G-mutant fibroblasts and cybrids. (a) Representative images of healthy donor (Ctrl)- and patient-derived (Pat) fibroblasts, before (Fed) or after 4 h of KRB (Starved). Cells were fixed and immunocytochemistry was performed with an antibody against LC3 followed by incubation with Alexa 488-conjugated secondary antibody. The scale bars represent 10 μm. (b) Quantification of autophagosome number of cells treated as in panel (a). (c) Quantification of the autophagosome number of selected healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts, before or after 4 h of starvation, with or without chloroquine (50 μM for 1 h). (d) mRNA was extracted from healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts, and quantitative real-time PCR for SQSTM1/p62 transcript was performed as detailed in Materials and Methods section. Data are normalized on Rpl32 mRNA level and presented as mean±S.E. (e) Representative images of wt and mutated (13514A>G) cybrids, before (Fed) or after 4 h of KRB (Starved), with or without chloroquine (50 μM for 1 h). Cells were fixed and immunocytochemistry was performed with an antibody against LC3 followed by incubation with Alexa 488-conjugated secondary antibody. The scale bars represent 10 μm. (f) Quantification of the autophagosome number of cells treated as in panel (e). Data are presented as mean±S.E. Descriptive statistics can be found in Supplementary Table S1. ***P<0.0001 compared with control. ###P<0.0001 between the indicated groups
The induction of autophagosome formation observed in Pat#1 and Pat#2 cells could either be due to the cellular response to the pathological mutation or to a dysfunction in autophagy. In order to exclude a block in the recycling of the autophagic cargo, we evaluated the rate of the autophagic flux by incubating cells with the inhibitor of lysosomal acidification, chloroquine.28 When looking at the fed condition, chloroquine induced a significant increase of autophagosome number and volume in both control and Pat#1 fibroblasts (Figure 1c and Supplementary Figures S1a and b). This indicates that the enhanced autophagy displayed by 13514A>G cells is not due to a decrease of autophagosome clearance. This was further confirmed by immunofluorescence staining of a different marker of autophagy sequestosome1 (SQSTM1/p62). This protein is incorporated into mature autophagosomes and degraded in autolysosomes, serving as a readout of the autophagic pathway.29 The distribution of SQSTM1/p62 puncta perfectly matches that of LC3 puncta (Supplementary Figures S1c and d). Its transcriptional upregulation in Pat#1 fibroblasts further demonstrates that 13514A>G cells have an increased autophagosome formation rather than a block in their degradation (Figure 1d).
Additionally, we further confirmed that the increased autophagy is the direct consequence of the 13514A>G mutation by developing osteosarcoma cells with homoplasmic 13514A>G mutant mtDNA (13514A>G cybrids) or wt mtDNA (control cybrids). Our results clearly demonstrate that 13514A>G cybrids also have a basal increased number of autophagosomes compared with controls. In this cellular model, this is due to enhanced autophagy induction and not to a block in autophagosome clearance, as revealed by the chloroquine treatment (Figures 1e and f and Supplementary Figure S1e).
Mitochondrial turnover through autophagy is higher in 13514A>G fibroblasts
In addition to its role in global cellular homeostasis, autophagy has an important role in removing damaged organelles. Mitochondria can be specific targets of this process through a selective process called mitophagy, which is critical for organelle quality control.30, 31 We hypothesized that increased autophagy could represent an adaptive and/or compensatory mechanism aimed at eliminating dysfunctional and damaged organelles in 13514A>G fibroblasts, thus mitigating the effect of the pathological mutation. To test this hypothesis, we monitored the rate of mitochondria degradation via autophagy by implementing the recently described technique based on the protein Keima. Keima is a fluorescent protein characterized by a pH-dependent shift of its excitation peak (479 nm at neutral and 570 nm at acidic pH),32 which is instrumental to monitor autophagic cargo delivery to lysosomes. Transfection of the mitochondria-targeted version of Keima in patient fibroblasts revealed a significant increase of the ‘mitophagy index’ (calculated as the ratio between images acquired at 570 and 479 nm excitation) selectively in 13514A>G cells (Figures 2a and b). This points to an increased elimination of mitochondria through lysosomal degradation in Pat#1 and Pat#2. This was also confirmed by the enhanced translocation of Parkin to mitochondria33, 34 in Pat#1 cells compared with the control cells (Supplementary Figure S2a).
Impairment of mitochondrial turnover and mitophagy in 13514A>G-mutant fibroblasts. (a) Representative images of healthy donor- (Ctrl) and patient-derived (Pat) fibroblasts transfected with mt-Keima. After 24 h, cells were imaged as detailed in Materials and Methods section. Images are presented as merged pictures of mt-Keima excited at 479 nm (green) and at 570 nm (red). The scale bars represent 10 μm. (b) Quantification of mitophagy index, measured as the image ratio (570/479 nm) of cells treated as in panel (a). (c) Healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts were harvested after treatment with 10 μM cycloheximide for the indicated time, and total proteins were extracted and subjected to western blotting analysis with antibodies against Tom20, MCU and β-Tubulin. Data are presented as mean±S.E. Descriptive statistics can be found in Supplementary Table S1. **P<0.001; ***P<0.0001 compared with control
We also monitored the degradation of selected mitochondrial proteins in our system by blocking protein synthesis with cycloheximide. Despite no differences could be detected in the overall level of mitochondrial proteins in both resting conditions and after nutrient deprivation (Supplementary Figure S2b), MCU (in the inner mitochondrial membrane) and Tom20 (in the outer mitochondrial membrane (OMM)) show a much faster degradation in Pat#1 compared with control fibroblasts after blocking protein synthesis with cycloheximide, indicating a higher organelle turnover (Figure 2c).33, 34
Defective mitochondrial Ca2+ homeostasis in 13514A>G fibroblasts
As previously reported, an impairment of mitochondrial Ca2+ handling is associated with mitochondrial diseases.35 Recently, Cardenas et al.6 reported that constitutively Ca2+ release from the ER controls cellular bioenergetics and autophagy. Based on these observations, we investigated cellular Ca2+ homeostasis in fibroblasts with different ND5 mutations. [Ca2+] measurements were carried out with aequorin-based cytosolic, mitochondrial and ER Ca2+ probes (cytAEQ,36 mtAEQ37 and erAEQ,38 respectively). Although control and mutant cells showed no differences in Ca2+ handling in the cytoplasm, a significant reduction of the [Ca2+]mt peak evoked by agonist stimulation was observed in Pat#1 and Pat#2 fibroblasts (Figures 3a and b and Supplementary Figures S3a and b), indicating selective alteration of mitochondrial Ca2+ homeostasis in 13514A>G cells. This was confirmed in experiments with erAEQ (Supplementary Figure S3d) and in permeabilized cells, where the contribution of plasma membrane or ER channels to the mitochondrial Ca2+ response is absent (Supplementary Figure S3c). In parallel, we performed [Ca2+] measurements in wt and mutant cybrids. Even in this experimental setup, the 13514A>G mutation causes a specific decrease of mitochondrial Ca2+ uptake (Supplementary Figures S3e–g), further confirming the presence of a correlation between altered autophagy and impaired mitochondrial Ca2+ dynamics associated with the 13514A>G mutation.
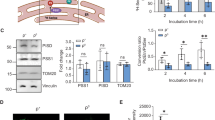
Impairment of mitochondrial Ca2+ uptake and MCU complex components' expression in complex I-deficient fibroblasts. (a) [Ca2+]cyt measurements in intact healthy donor (Ctrl)- and patient-derived (Pat) fibroblasts, challenged with maximal histamine stimulation. (b) [Ca2+]mt measurements in intact healthy donor (Ctrl)- and patient-derived (Pat) fibroblasts, challenged with maximal histamine stimulation. (c) Healthy donor (Ctrl)- and patient-derived (Pat) fibroblasts before (Fed) or after 4 h of KRB (Starved) were harvested, and total proteins were extracted and subjected to western blotting analysis with antibodies against phosphorylated AMPK on Ser172 (p-AMPK), total AMPK and β-Tubulin. (d) Healthy donor (Ctrl)- and patient-derived (Pat) fibroblasts were harvested, and total proteins were extracted and subjected to western blotting analysis with antibodies against MCU, MICU1, MICU2, EMRE and β-Tubulin. Data are presented as mean±S.E. Descriptive statistics can be found in Supplementary Table S1. *P<0.05; **P<0.001; ***P<0.0001 compared with control
We next analyzed the signaling pathways linking mitochondrial Ca2+ to autophagy by monitoring AMP-activated protein kinase (AMPK), the major metabolic sensor of AMP/ATP ratio.6, 39, 40 Western blotting analysis of AMPK phosphorylation on Ser172 (p-AMPK)40 revealed low p-AMPK level in resting conditions in control cells. Conversely, all patient-derived fibroblasts displayed a slight increase in p-AMPK levels, as expected in cells with deficiency of the respiratory chain complexes, but in Pat#1 this increase was dramatic. AMPK phosphorylation was induced after nutrient depletion in all the cells, except for Pat#1 fibroblasts, where phosphorylation level was already maximal (Figure 3c). Other pathways involved in autophagy such as Akt, mammalian target of rapamycin and its target S6 were only modestly affected in 13514A>G fibroblasts (Supplementary Figure S4). Thus the pattern of AMPK activation was directly correlated with the level of autophagy and inversely associated with the amplitude of the [Ca2+]mt rise in our system.
Impairment of the MCU complex in 13514A>G cells
To further dissect the molecular mechanisms underlying the impairment of mitochondrial Ca2+ handling, we evaluated the major mitochondrial parameters involved in the control of organellar cation homeostasis, that is, (i) expression of the MCU complex components, (ii) ER–mitochondria contact sites, (iii) morphology and (iv) membrane potential.
The expression level of the MCU complex components showed small but significant differences. In particular, in Pat#1 and, to a lesser extent, in Pat#2, the positive regulators of MCU, MICU1 and EMRE were downregulated, while the negative regulator MICU2 was slightly increased (Figure 3d). Importantly, this MCU complex remodeling was not present (except for EMRE) in Pat#3 and Pat#4, in line with the [Ca2+]mt homeostasis data. In addition, we recently reported that MICU1 homodimer and MICU1/MICU2 heterodimer regulates MCU sensitivity to extramitochondrial [Ca2+] and exert opposite effects on the activity of the channel (stimulatory for MICU1 and inhibitory for MICU2).19 The analysis of the ratio between MICU1/MICU1 and MICU1/MICU2 dimers in non-reducing conditions showed that two bands were present in control fibroblasts, corresponding to the MICU1/MICU2 heterodimer and the MICU1 homodimer. Conversely, Pat#1 cells lacked the MICU1 homodimer (Supplementary Figure S3h), which is responsible for the rapid and efficient mitochondrial Ca2+ uptake. This remodeling of the MCU complex can, at least in part, explain the reduced organelle Ca2+ transients observed in these cells.
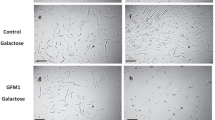
We next investigated the ER–mitochondria contact sites, a key determinant of mitochondrial Ca2+ uptake in intact cells, by monitoring the expression of the components of the tethering complex in mutant and control fibroblasts. The protein levels of the best characterized ER-mitochondria tether, Mitofusin 2 (Mfn2),41 as well as other proteins located at the interface between the organelles, such as SigmaR1, showed a significant and specific decrease in 13514A>G cells (Figure 4a). In addition, the downregulation of Mfn2 transcript in Pat#1 cells suggests that this effect occurs at the transcriptional level (Supplementary Figure S5a). Following this, we directly measured the formation of ER–mitochondria contact sites in Pat#1 and control cells by using a novel technique based on the split-green fluorescent protein (GFP) technology. An OMM-anchored GFP lacking the essential beta strand 11 (GFP 1–10) and an ER-anchored beta strand 11, both facing the cytosolic side, were expressed in our fibroblasts. These constructs are per se non-fluorescent, but they reconstitute a functional fluorescent GFP as they come in close proximity (i.e., at ER–mitochondria contact sites) (Calì et al., submitted manuscript). Consequently, the fluorescence intensity correlates with the extent of ER–mitochondria contact sites. As shown in Figures 4b and c, contacts between the two organelles are reduced in Pat#1 fibroblasts, in agreement with the decrease of mitochondrial Ca2+ uptake detected in these cells.
Effect of 13514A>G mutation on ER–mitochondria tethering components' expression, ΔΨmt and mitochondrial morphology. (a) Healthy donor (Ctrl)- and patient-derived (Pat) fibroblasts were harvested, and total proteins were extracted and subjected to western blotting analysis with antibodies against Mfn2, Mfn1, grp75, SigmaR1 and β-Tubulin. (b) Healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts were co-transfected with an OMM-anchored GFP lacking an essential beta strand (GFP 1–10) and an ER-anchored beta strand 11 required for GFP1–10 fluorescence reconstitution upon complementation. After 24 h, cells were fixed and imaged as detailed in Materials and Methods section. The scale bars represent 10 μm. (c) Quantification of contact sites between ER and mitochondria of cells processed as in panel (b). (d) Healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts were fixed and immunocytochemistry was performed with Tom20 antibody followed by incubation with Alexa 488-conjugated secondary antibody (green). The scale bars represent 10 μm. (e) Quantification of the number of objects per cell (top panel) and mean volume of objects (bottom panel) of cells immunostained with antibody against Tom20. (f and g) Tetra-methyl-rhodamine-methyl ester fluorescence analysis of healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts. Where indicated, 4 μM rotenone (R), 5 μM oligomycin (O) and 4 μM FCCP (F) were added. Data report mean of three independent experiments±S.E., expressed as percentage of the initial fluorescent value. Data are presented as mean±S.E. Descriptive statistics can be found in Supplementary Table S1. *P<0.05; **P<0.001; ***P<0.0001 compared with control
Mitochondrial fragmentation was also assessed in Pat#1 cells by immunostaining of the mitochondrial marker Tom20. No major alterations of organelle morphology could be observed (Figure 4d). Indeed, morphometric analysis of mitochondrial network showed no significant differences in mitochondrial volume and number (Figure 4e).
Finally, ΔΨmt, the main driving force for Ca2+ entry into mitochondria, was monitored in control and Pat#1 fibroblasts using the lipophilic dye tetra-methyl-rhodamine-methyl ester. No differences were observed after treatment with the ATP synthase inhibitor oligomycin or the complex I inhibitor rotenone. After rotenone addition, both cell lines maintained their ΔΨmt through the reverse activity of ATP synthase, as demonstrated by the collapse of ΔΨmt induced by oligomycin (Figure 4f). On the other hand, when applied first, oligomycin caused a slight ΔΨmt increase, indicating that both control and Pat#1 cells possess a functional ETC and generate ΔΨmt that is partly consumed by ATP synthesis through the mitochondrial ATP synthase (Figure 4g). In addition, a slight increase in complex III and V could be noticed in Pat#1 fibroblasts (Supplementary Figure S5b), which may at least in part explain the normal steady-state ΔΨmt (Supplementary Figure S5c).
Together these results suggest that both the remodeling of the MCU complex composition and the rearrangement of ER–mitochondria tethering contribute to the decrease in mitochondrial Ca2+ uptake in 13514A>G cells.
Genetic and pharmacological manipulation of mitochondrial Ca2+ uptake restores the normal autophagic flux in 13514A>G fibroblasts
In order to test the possibility that the lowering of mitochondrial Ca2+ uptake is upstream of the increased autophagy in 13514A>G fibroblasts, we corrected the Ca2+ signaling alteration by molecular (i.e., MCU overexpression) or pharmacological (i.e., treatment with compounds that enhance mitochondrial Ca2+ uptake) approaches, and experimentally verified whether the normal autophagic flux was restored. MCU was overexpressed in both Ctrl#1 and Pat#1 fibroblasts leading to an increase of [Ca2+]mt transients in resting as well as starved cells, restoring organellar Ca2+ signals of 13514A>G cells to control levels (Figure 5a). When we monitored the translocation of the autophagy marker LC3 in the same conditions, we found that induction of the autophagic flux inversely correlates with the amplitude of mitochondrial Ca2+ signals (Figures 5b and c). Most importantly, MCU overexpression in Pat#1 cells was able to restore AMPK phosphorylation to levels undistinguishable from control (Figure 5d). To confirm this effect, we treated control and patient cells with two structurally different MCU activators, Kaempferol (a plant-derived antioxidant flavonoid)42 and SB202190 (also an inhibitor of p38 MAPK).43 Both compounds enhanced mitochondrial Ca2+ uptake in Pat#1 fibroblasts to values similar to control cells (Figure 6a). Interestingly, Kaempferol and SB202190 treatments in patient cells restored autophagy to the level of resting control fibroblasts (Figures 6b and c).
Overexpression of MCU restores the normal autophagic flux in 13514A>G-mutant fibroblasts. (a) [Ca2+]mt measurements in intact healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts before (Fed) or after 4 h of KRB (Starved), transfected with pcDNA or MCU-encoding plasmid, challenged with maximal histamine stimulation. (b) Healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts, treated as in panel (a), were fixed and immunocytochemistry was performed with an antibody against LC3 followed by incubation with Alexa 488-conjugated secondary antibody. The scale bars represent 10 μm. (c) Quantification of the autophagosome number of cells processed as in panel (b). (d) Healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts, before (Fed) or after 4 h of KRB (Starved), transfected with pcDNA or MCU-encoding plasmid, were harvested, and total proteins were extracted and subjected to western blotting analysis with antibodies against phosphorylated AMPK on Ser172 (p-AMPK), AMPK, MCU and β-Tubulin. Data are presented as mean±S.E. Descriptive statistics can be found in Supplementary Table S1. *P<0.05; **P<0.001; ***P<0.0001 compared with control. ###P<0.0001 between the indicated groups
Pharmacological modulation of mitochondrial Ca2+ uptake restores the normal autophagic flux in 13514A>G-mutant fibroblasts. (a) [Ca2+]mt measurements in intact healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts, untreated or treated with Kaempferol (20 μM) or SB-202190 (20 μM) for 24 h and challenged with maximal histamine stimulation. (b) Healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts, processed as in panel (a), were fixed and immunocytochemistry was performed with an antibody against LC3 followed by incubation with Alexa 488-conjugated secondary antibody. The scale bars represent 10 μm. (c) Quantification of the autophagosome number of cells processed as in panel (c). (e) Healthy donor (Ctrl#1)- and 13514A>G patient-derived (Pat#1) fibroblasts, untreated or treated with Kaempferol (20 μM), SB-202190 (20 μM) or infected with MCU-encoding adenovirus for 24 or 48 h as indicated. At the end of treatments, the cell viability was determined using MTT assay. Cell viability is expressed as the percentage of the untreated control. Data are presented as mean±S.E. Descriptive statistics can be found in Supplementary Table S1. *P<0.05; **P<0.001; ***P<0.0001 compared with control. ###P<0.0001 between the indicated groups
Finally, we tested cell viability in fibroblasts treated with the two compounds or overexpressing MCU. As expected, both of these conditions lowered cell viability in control cells, confirming the notion that sustained [Ca2+]mt overload enhances cell death.44, 45 More importantly, the same manipulations cause a much higher decrease of cell viability in Pat#1 cells (Figure 6d). This indicates that the decrease of mitochondrial Ca2+ uptake in 13514A>G fibroblasts is a pro-survival compensatory mechanism in these cells.
Discussion
Mitochondrial disorders caused by mutations in mtDNA affect approximately 1 in every 10 000 adults.46, 47 The broad heterogeneity of clinical manifestations, in terms of severity, age of onset and prognosis, relies on the specific molecular defect, the complexity of mitochondrial genetics48 and the individual genotypic differences. The large diversity, however, could also depend on the cellular mechanisms that allow to compensate for the respiratory deficiency or promote the turnover of the damaged mitochondria. Recently, autophagy has been recognized as an important response to energetic defects,49 as well as the leading mechanism for eliminating damaged organelles.30, 31 In addition, mitochondrial Ca2+ signaling has a role in the regulation of autophagy.6 This complex relationship could allow, in the context of a mitochondrial disorder, the triggering of a cellular response that compensate, at least in part, the energetic defect, dampening the clinical phenotype. In this work, we investigated autophagy and mitochondrial Ca2+ signaling in human fibroblasts from control subjects and from a set of patients carrying mutations in the mtDNA-encoded ND5 subunit of the ETC complex I.25, 26, 27 Despite the impaired complex I activity25, 26 and the increased ROS production (data not shown), these cells do not show any major defects in resting ΔΨmt (Supplementary Figure S5c) or in organelle morphology (Figures 4d and e). However, we observed a marked increase of the autophagic flux in fibroblasts carrying the 13514A>G mutation (Figures 1a–d and Supplementary Figures S1a–d). Interestingly, both the patients with this specific mutation display atypical MELAS and Leigh syndromes, characterized by late onset, slow progression, normal plasma lactate level and muscle morphology.
We thus focused our study on the molecular mechanism underlying the exaggerated autophagic flux in 13514A>G cells and tested whether this could justify the milder phenotype associated with this mutation. Notably, a similar increase in autophagy was also observed in cybrids homoplasmic for the same mtDNA mutation. Even in this system, this was clearly due to an enhanced formation of autophagosomes rather than a decrease of their clearance (Figures 1e and f and Supplementary Figure S1e). Importantly, the accelerated autophagy underlies a faster mitochondrial turnover in Pat#1 compared with Ctrl#1 cells, as revealed by the shorter lifespan of mitochondrial proteins (Figure 2c) and by the increase of mitochondria delivered to the lysosome (Figures 2a and b). However, no difference at the steady state could be observed in the overall mitochondrial mass (Figures 4d and e) or in the total amount of mitochondrial proteins (Supplementary Figure S2b), suggesting that a compensatory upregulation of mitochondrial biogenesis matches the rapid elimination of mitochondria.
We then investigated the molecular basis of the enhanced autophagy in 13514A>G cells by combining the observations that mitochondrial Ca2+ signals inhibits autophagy6 with the previously reported defects of mitochondrial Ca2+ homeostasis in mitochondrial disorders.35 The analysis of Ca2+ homeostasis indicates a selective impairment of mitochondrial Ca2+ uptake in 13514A>G cells (Figures 3a and b and Supplementary Figures S3a–d) and in homoplasmic cybrids (Supplementary Figures S3e–g). Accordingly, the p-AMPK, and consequently its activity, are higher in 13514A>G cells compared with control (Figure 3c). This impairment of mitochondrial Ca2+ uptake was due to both a decrease in interorganellar contact sites (Figures 4a–c) as well as to differences in the supramolecular organization of the MCU complex (Figure 3d). However, as the defect in mitochondrial Ca2+ accumulation in Pat#1 cells is also evident in permeabilized cells (Supplementary Figure S3c), we conclude that the assembly of the MCU complex has a major role in lowering mitochondrial Ca2+ uptake. In particular, Pat#1 and Pat#2 (but not Pat#3 and Pat#4) cells show a significant decrease of MICU1 and EMRE levels, known as positive MCU regulators,15, 16, 17, 18 and an increase in the expression of MICU2. MICU2 is a negative MCU modulator as it forms the MICU1/MICU2 heterodimer, thus sequestering MICU1 and preventing the MICU1 homodimer stimulatory activity on MCU, as we recently demonstrated.19 Indeed, we detected the presence of MICU1 homodimers in control but not in Pat#1 fibroblasts (Supplementary Figure S3h). Such an imbalance of MCU regulators could explain the reduced efficiency of mitochondrial Ca2+ uptake in 13514A>G cells.
The actual relevance of the correlation between autophagy and mitochondrial Ca2+ homeostasis was corroborated by the demonstration that genetically or pharmacologically enhanced mitochondrial Ca2+ uptake slows down the autophagic flux in 13514A>G cells (Figures 5 and 6). Notably, long term treatment with these compounds as well as MCU overexpression decreased the cells' viability to a greater extent in 13514A>G cells compared with controls (Figure 6d). This supports the hypothesis that the lowering of mitochondrial Ca2+ uptake is a compensatory, potentially beneficial cellular mechanism that, on the one hand, activates organelle turnover via autophagy and, on the other hand, reduces the efficiency of death pathways, to which cells harboring mtDNA defects are more vulnerable.
Overall, our data demonstrate that, in a model of a mitochondrial disorder displaying a relatively mild phenotype, organelle dysfunction is relieved by increasing mitochondrial turnover through mitophagy. This effect is triggered by decreasing mitochondrial Ca2+ uptake, leading to upregulation of cellular catabolic processes through an AMPK-dependent mechanism. Consequently, selective stimulation of mitochondrial quality control could be a potentially promising approach to ameliorate the clinical manifestation of the most severe forms of mitochondrial diseases.
Materials and Methods
Chemicals, cell culture and transfection
All the experiments were performed on primary skin fibroblasts derived from MELAS or Leigh syndrome patients carrying different point mutations in the ND5 subunit of complex I (Pat#1 (patient 2 in Corona et al.25), Pat#2 (patient 8 in Bugiani et al.26), Pat#3 (patient 4 in Bugiani et al.26), Pat#4 (patient 6 in Malfatti et al.27)) and three independent control fibroblast cultures from healthy individuals (Ctrl#1, Ctrl#2 and Ctrl#3). Primary fibroblasts were cultured in Dulbecco’s modified Eagle’s medium (DMEM), supplemented with 20% fetal bovine serum (FBS), penicillin (100 U/ml) and streptomycin (100 μg/ml) and transfected with Lipofectamine 2000 (Life Technologies, Waltham, MA, USA), according to the manufacturer’s instructions. The experiments were carried out 24 h after transfection. Some experiments were performed on transmitochondrial cybrids. These cybrids were obtained by polyethylene glycol fusion of fibroblast-derived cytoplasts from Pat#1 and a 143B rho-zero cell line, as previously described.50 After selection, two different clones containing 13514A>G wild-type mtDNA (wt) and 100% of 13514A>G mutant mtDNA (13514A>G) were chosen. Cybrids were cultured in DMEM (Life Technologies), supplemented with 10% FBS (Life Technologies) and uridine (50 μg/ml), penicillin (200 U/ml) and streptomycin (200 μg/ml) and transfected with Lipofectamine 2000 (Life Technologies), according to the manufacturer’s instructions. The experiments were carried out 24 h after transfection.
For glucose- and serum-deprivation experiments (Starved condition), fibroblasts and cybrids were washed with phosphate-buffered saline (PBS) and then were incubated for 4 h in sterile modified Krebs–Ringer buffer (KRB; 135 mM NaCl, 5 mM KCl, 1 mM MgSO4, 0.4 mM K2HPO4, 20 mM HEPES, pH=7.4, supplemented with 1 mM CaCl2). Where indicated, chloroquine (50 μM) was added to the same medium for 1 h.
All chemicals were purchased from Sigma-Aldrich (St. Louis, MO, USA), unless specified.
The MCU-flag/pcDNA expression construct is described in De Stefani et al.12
The Parkin full-length expression construct is described in Cali et al.51
The MCU-flag adenovirus production is described in Raffaello et al.14
Immunofluorescence
Control and mutated fibroblasts or wt and mutated cybrids were starved (KRB for 4 h), treated with chloroquine (50 μM for 1 h), Kaempferol (20 μM for 24 or 48 h) or SB202190 (20 μM for 24 or 48 h) and transfected with Parkin, MCU or empty vector as control, as indicated. Cells were fixed and permeabilized in cold MeOH/Acetone 1 : 1, blocked in PBS containing 1% BSA, 5% goat serum and 0.3% triton X-100. Cells were then incubated with the indicated antibodies (LC3 from Cell Signaling (Danvers, MA, USA); SQSTM1/p62 from Sigma-Aldrich; Parkin and Tom20 from Santa Cruz Biotechnology (Dallas, TX, USA)) overnight at +4 °C. The appropriate isotype-matched AlexaFluor-conjugated secondary antibodies (Life Technologies) were used, and coverslips were mounted with ProLong Gold Antifade reagent (Life Technologies).
AEQ Ca2+ measurements
Control and mutated primary fibroblasts or wt and mutated cybrids were transfected with plasmid coding for the cytAEQ,36 mtAEQ37 or erAEQ,38 together with the indicated constructs, using Lipofectamine 2000. Mutated or wt mtAEQ37 were implemented for primary fibroblasts or cybrids, respectively. After 24 h of transfection, cells were processed for the [Ca2+] measurements as described in detail in Granatiero et al.52
All of the results are expressed as means±S.E., and Student's t test was used for the statistical analysis. Representative traces and histograms are shown in the figures, whereas the full data set is included in the Supplementary Table S1.
Full methods are available in the Supplementary Material.
Abbreviations
- AEQ:
-
aequorin
- AMPK:
-
AMP-activated protein kinase
- ATP:
-
adenosine triphosphate
- ΔΨmt:
-
mitochondrial membrane potential
- DMEM:
-
Dulbecco’s modified Eagle’s medium
- EMRE:
-
essential MCU regulator
- ETC:
-
electron transport chain
- GFP:
-
green fluorescent protein
- KRB:
-
Krebs–Ringer buffer
- LC3:
-
microtubule-associated protein 1 A/1B light chain 3B
- MCU:
-
mitochondrial Ca2+ uniporter
- MELAS:
-
mitochondrial encephalomyopathy, lactic acidosis and stroke-like episodes syndrome
- Mfn2:
-
Mitofusin2
- MICU1:
-
mitochondrial Ca2+ uptake 1
- mtDNA:
-
mitochondrial DNA
- OMM:
-
outer mitochondrial membrane
- PBS:
-
phosphate-buffered saline
- SQSTM1/p62:
-
sequestosome1
References
Zeviani M, Di Donato S . Mitochondrial disorders. Brain 2004; 127: 2153–2172.
Sproule DM, Kaufmann P . Mitochondrial encephalopathy, lactic acidosis, and strokelike episodes: basic concepts, clinical phenotype, and therapeutic management of MELAS syndrome. Ann NY Acad Sci 2008; 1142: 133–158.
Koopman WJ, Verkaart S, Visch HJ, van Emst-de Vries S, Nijtmans LG, Smeitink JA et al. Human NADH:ubiquinone oxidoreductase deficiency: radical changes in mitochondrial morphology? Am J Physiol Cell Physiol 2007; 293: C22–C29.
Willems PH, Valsecchi F, Distelmaier F, Verkaart S, Visch HJ, Smeitink JA et al. Mitochondrial Ca2+ homeostasis in human NADH:ubiquinone oxidoreductase deficiency. Cell Calcium 2008; 44: 123–133.
Rizzuto R, De Stefani D, Raffaello A, Mammucari C . Mitochondria as sensors and regulators of calcium signalling. Nat Rev Mol Cell Biol 2012; 13: 566–578.
Cardenas C, Miller RA, Smith I, Bui T, Molgo J, Muller M et al. Essential regulation of cell bioenergetics by constitutive InsP3 receptor Ca2+ transfer to mitochondria. Cell 2010; 142: 270–283.
Levine B, Kroemer G . Autophagy in the pathogenesis of disease. Cell 2008; 132: 27–42.
Suen DF, Narendra DP, Tanaka A, Manfredi G, Youle RJ . Parkin overexpression selects against a deleterious mtDNA mutation in heteroplasmic cybrid cells. Proc Natl Acad Sci USA 2010; 107: 11835–11840.
Geisler S, Holmstrom KM, Skujat D, Fiesel FC, Rothfuss OC, Kahle PJ et al. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat Cell Biol 2010; 12: 119–131.
Narendra D, Tanaka A, Suen DF, Youle RJ . Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol 2008; 183: 795–803.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005; 122: 927–939.
De Stefani D, Raffaello A, Teardo E, Szabo I, Rizzuto R . A forty-kilodalton protein of the inner membrane is the mitochondrial calcium uniporter. Nature 2011; 476: 336–340.
Baughman JM, Perocchi F, Girgis HS, Plovanich M, Belcher-Timme CA, Sancak Y et al. Integrative genomics identifies MCU as an essential component of the mitochondrial calcium uniporter. Nature 2011; 476: 341–345.
Raffaello A, De Stefani D, Sabbadin D, Teardo E, Merli G, Picard A et al. The mitochondrial calcium uniporter is a multimer that can include a dominant-negative pore-forming subunit. EMBO J 2013; 32: 2362–2376.
Sancak Y, Markhard AL, Kitami T, Kovacs-Bogdan E, Kamer KJ, Udeshi ND et al. EMRE is an essential component of the mitochondrial calcium uniporter complex. Science 2013; 342: 1379–1382.
Mallilankaraman K, Doonan P, Cardenas C, Chandramoorthy HC, Muller M, Miller R et al. MICU1 is an essential gatekeeper for MCU-mediated mitochondrial Ca(2+) uptake that regulates cell survival. Cell 2012; 151: 630–644.
Perocchi F, Gohil VM, Girgis HS, Bao XR, McCombs JE, Palmer AE et al. MICU1 encodes a mitochondrial EF hand protein required for Ca(2+) uptake. Nature 2010; 467: 291–296.
Csordas G, Golenar T, Seifert EL, Kamer KJ, Sancak Y, Perocchi F et al. MICU1 controls both the threshold and cooperative activation of the mitochondrial Ca(2)(+) uniporter. Cell Metab 2013; 17: 976–987.
Patron M, Checchetto V, Raffaello A, Teardo E, Vecellio Reane D, Mantoan M et al. MICU1 and MICU2 finely tune the mitochondrial Ca2+ uniporter by exerting opposite effects on MCU activity. Mol Cell 2014; 53: 726–737.
Hoffman NE, Chandramoorthy HC, Shanmughapriya S, Zhang XQ, Vallem S, Doonan PJ et al. SLC25A23 augments mitochondrial Ca(2)(+) uptake, interacts with MCU, and induces oxidative stress-mediated cell death. Mol Biol Cell 2014; 25: 936–947.
Denton RM . Regulation of mitochondrial dehydrogenases by calcium ions. Biochim Biophys Acta 2009; 1787: 1309–1316.
Jouaville LS, Pinton P, Bastianutto C, Rutter GA, Rizzuto R . Regulation of mitochondrial ATP synthesis by calcium: evidence for a long-term metabolic priming. Proc Natl Acad Sci USA 1999; 96: 13807–13812.
Bernardi P . The mitochondrial permeability transition pore: a mystery solved? Front Physiol 2013; 4: 95.
Rasola A, Bernardi P . Mitochondrial permeability transition in Ca(2+)-dependent apoptosis and necrosis. Cell Calcium 2011; 50: 222–233.
Corona P, Antozzi C, Carrara F, D'Incerti L, Lamantea E, Tiranti V et al. A novel mtDNA mutation in the ND5 subunit of complex I in two MELAS patients. Ann Neurol 2001; 49: 106–110.
Bugiani M, Invernizzi F, Alberio S, Briem E, Lamantea E, Carrara F et al. Clinical and molecular findings in children with complex I deficiency. Biochim Biophys Acta 2004; 1659: 136–147.
Malfatti E, Bugiani M, Invernizzi F, de Souza CF, Farina L, Carrara F et al. Novel mutations of ND genes in complex I deficiency associated with mitochondrial encephalopathy. Brain 2007; 130: 1894–1904.
Klionsky DJ, Abdalla FC, Abeliovich H, Abraham RT, Acevedo-Arozena A, Adeli K et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 2012; 8: 445–544.
Rogov V, Dotsch V, Johansen T, Kirkin V . Interactions between autophagy receptors and ubiquitin-like proteins form the molecular basis for selective autophagy. Mol Cell 2014; 53: 167–178.
Kim I, Rodriguez-Enriquez S, Lemasters JJ . Selective degradation of mitochondria by mitophagy. Arch Biochem Biophys 2007; 462: 245–253.
Zhang Y, Qi H, Taylor R, Xu W, Liu LF, Jin S . The role of autophagy in mitochondria maintenance: characterization of mitochondrial functions in autophagy-deficient S. cerevisiae strains. Autophagy 2007; 3: 337–346.
Katayama H, Kogure T, Mizushima N, Yoshimori T, Miyawaki A . A sensitive and quantitative technique for detecting autophagic events based on lysosomal delivery. Chem Biol 2011; 18: 1042–1052.
Narendra D, Walker JE, Youle R . Mitochondrial quality control mediated by PINK1 and Parkin: links to parkinsonism. Cold Spring Harb Perspect Biol 2012; 4: 11.
Winklhofer KF . Parkin and mitochondrial quality control: toward assembling the puzzle. Trends Cell Biol 2014; 24: 332–341.
Brini M, Pinton P, King MP, Davidson M, Schon EA, Rizzuto R . A calcium signaling defect in the pathogenesis of a mitochondrial DNA inherited oxidative phosphorylation deficiency. Nat Med 1999; 5: 951–954.
Brini M, Marsault R, Bastianutto C, Alvarez J, Pozzan T, Rizzuto R . Transfected aequorin in the measurement of cytosolic Ca2+ concentration ([Ca2+]c). A critical evaluation. J Biol Chem 1995; 270: 9896–9903.
Rizzuto R, Simpson AW, Brini M, Pozzan T . Rapid changes of mitochondrial Ca2+ revealed by specifically targeted recombinant aequorin. Nature 1992; 358: 325–327.
Montero M, Brini M, Marsault R, Alvarez J, Sitia R, Pozzan T et al. Monitoring dynamic changes in free Ca2+ concentration in the endoplasmic reticulum of intact cells. EMBO J 1995; 14: 5467–5475.
Hardie DG . AMP-activated/SNF1 protein kinases: conserved guardians of cellular energy. Nat Rev Mol Cell Biol 2007; 8: 774–785.
Andersen MN, Rasmussen HB . AMPK: A regulator of ion channels. Commun Integr Biol 2012; 5: 480–484.
de Brito OM, Scorrano L . Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature 2008; 456: 605–610.
Montero M, Lobaton CD, Hernandez-Sanmiguel E, Santodomingo J, Vay L, Moreno A et al. Direct activation of the mitochondrial calcium uniporter by natural plant flavonoids. Biochem J 2004; 384: 19–24.
Montero M, Lobaton CD, Moreno A, Alvarez J . A novel regulatory mechanism of the mitochondrial Ca2+ uniporter revealed by the p38 mitogen-activated protein kinase inhibitor SB202190. FASEB J 2002; 16: 1955–1957.
Pinton P, Giorgi C, Siviero R, Zecchini E, Rizzuto R . Calcium and apoptosis: ER-mitochondria Ca2+ transfer in the control of apoptosis. Oncogene 2008; 27: 6407–6418.
Celsi F, Pizzo P, Brini M, Leo S, Fotino C, Pinton P et al. Mitochondria, calcium and cell death: a deadly triad in neurodegeneration. Biochim Biophys Acta 2009; 1787: 335–344.
Rozwodowska M, Drewa G, Zbytniewski Z, Wozniak A, Krzyzynska-Malinowska E, Maciak R . Mitochondrial diseases. Med Sci Monit 2000; 6: 817–822.
Chinnery PF, Elliott HR, Hudson G, Samuels DC, Relton CL . Epigenetics, epidemiology and mitochondrial DNA diseases. Int J Epidemiol 2012; 41: 177–187.
Wong LJ . Molecular genetics of mitochondrial disorders. Dev Disabil Res Rev 2010; 16: 154–162.
Kroemer G, Marino G, Levine B . Autophagy and the integrated stress response. Mol Cell 2010; 40: 280–293.
King MP, Attardi G . Human cells lacking mtDNA: repopulation with exogenous mitochondria by complementation. Science 1989; 246: 500–503.
Cali T, Ottolini D, Negro A, Brini M . Enhanced parkin levels favor ER-mitochondria crosstalk and guarantee Ca(2+) transfer to sustain cell bioenergetics. Biochim Biophys Acta 2013; 1832: 495–508.
Granatiero V, Patron M, Tosatto A, Merli G, Rizzuto R . Using targeted variants of aequorin to measure Ca2+ levels in intracellular organelles. Cold Spring Harb Protoc 2014; 2014: 1.
Acknowledgements
We are grateful to Cristina Mammucari and Gyorgy Szabadkai for helpful discussion. This work was supported by grants from Telethon-Italy (GPP10005A), the Italian Ministries of Health (Ricerca Finalizzata) and of Education, University and Research (PRIN, FIRB), the European Union (ERC mitoCalcium, no. 294777), National Institutes of Health (Grant no. 1P01AG025532-01A1), Cariparo and Cariplo Foundations (Padua), the Italian Association for Cancer Research (AIRC). TC is supported by the University of Padova (Progetto Giovani GRIC128SP0, Bando 2012).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by N Chandel
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Granatiero, V., Giorgio, V., Calì, T. et al. Reduced mitochondrial Ca2+ transients stimulate autophagy in human fibroblasts carrying the 13514A>G mutation of the ND5 subunit of NADH dehydrogenase. Cell Death Differ 23, 231–241 (2016). https://doi.org/10.1038/cdd.2015.84
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2015.84
- Springer Nature Limited
This article is cited by
-
Mitochondrial calcium uniporter stabilization preserves energetic homeostasis during Complex I impairment
Nature Communications (2022)
-
Endoplasmic reticulum−mitochondria coupling increases during doxycycline-induced mitochondrial stress in HeLa cells
Cell Death & Disease (2021)
-
Implications of mitochondrial network organization in mitochondrial stress signalling in NARP cybrid and Rho0 cells
Scientific Reports (2017)