Abstract
Background
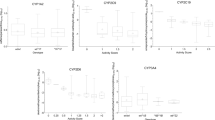
CYP2C19 is an important drug-metabolizing enzyme, responsible for metabolism of approximately 10% of the drugs on the market. Large inter-individual differences exist in metabolic activities, which are primarily attributed to genetic polymorphism of CYP2C19 gene. Conflicting results have been published about the role of CYP2C19 polymorphisms in metabolism of CYP2C19 substrates and clinical outcomes; thus, we aimed to investigate CYP2C19 genotype-phenotype associations, and we sought to elicit potential causes of discrepancies in the genotype-based prediction by incorporating the liver donors’ demographic data, drug administration events and pathological conditions.
Methods
(S)-Mephenytoin was used to assess CYP2C19 activities in human liver microsomes derived from 114 Hungarian organ donors. CYP2C19 genotype was determined by SNP genotyping for CYP2C19*2, CYP2C19*3, CYP2C19*4 and CYP2C19*17 variants, and CYP2C19 mRNA levels were measured by qPCR method. Clinical data of the donors were considered in the genotype-based phenotype prediction.
Results
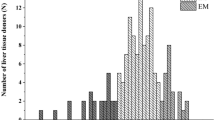
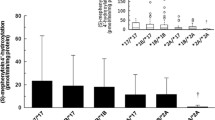
CYP2C19 phenotype of 40% of the donors was well-predicted from the genotype data, whereas the phenotype of 13% was underestimated displaying higher activity, and of 47% was overestimated displaying lower activity than predicted from CYP2C19 genotype. Among the donors with overestimated phenotype, one was treated with CYP2C19 substrate/inhibitor, 9 were on amoxicillin-clavulanic acid therapy, 7 were chronic alcohol consumers and 9 had disease with inflammatory processes.
Conclusions
CYP2C19 genotype only partially determines the CYP2C19 phenotypic appearance; co-medication, diseases with inflammatory processes and aspecific factors, such as chronic alcohol consumption and amoxicillin-clavulanic acid therapy (or any drug therapy resulting in liver injury) seem to be potential phenotype-modifying factors.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- CAR:
-
constitutive androstane receptor
- CI:
-
confidence interval
- CYP:
-
cytochrome P450
- EM:
-
extensive metabolizer
- GAPDH:
-
glyceraldehyde 3-phosphate dehydrogenase
- GR:
-
glucocorticoid receptor
- IM:
-
intermediate metabolizer
- PCR:
-
polymerase chain reaction
- PM:
-
poor metabolizer
- PXR:
-
pregnane X receptor
- SNP:
-
single nucleotide polymorphism
- UM:
-
ultra-rapid metabolizer
References
Bahar M.A., Setiawan D, Hak E, Wilffert B. Pharmacogenetics of drug-drug interaction and drug-drug-gene interaction: a systematic review on CYP2C9, CYP2C19 and CYP2D6. Pharmacogenomics 2017;18(7):701–39.
Ohtsuki S, Schaefer O, Kawakami H, Inoue T, Liehner S, Saito A, et al. Simultaneous absolute protein quantification of transporters, cytochromes P450, and UDP-glucuronosyltransferases as a novel approach for the characterization of individual human liver: comparison with mRNA levels and activities. Drug Metab Dispos 2012;40(1):83–92.
Zanger UM, Schwab M. Cytochrome P450 enzymes in drug metabolism: regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacol Ther 2013;138(1):103–41.
Hulot JS, Bura A, Villard E, Azizi M, Remones V, Goyenvalle C, et al. Cytochrome P450 2C19 loss-of-function polymorphism is a major determinant of clopidogrel responsiveness in healthy subjects. Blood 2006;108(7):2244–7.
Li-Wan-Po A, Girard T, Farndon P, Cooley C, Lithgow J. Pharmacogenetics of CYP2C19: functional and clinical implications of a new variant CYP2C19*17. Br J Clin Pharmacol 2010;69(3):222–30.
Hicks JK, Swen JJ, Thorn CF, Sangkuhl K, Kharasch ED, Ellingrod VL, et al. Clinical Pharmacogenetics Implementation Consortium guideline for CYP2D6 and CYP2C19 genotypes and dosing of tricyclic antidepressants. Clin Pharmacol Ther 2013;93(5):402–8.
Fricke-Galindo I, Céspedes-Garro C, Rodrigues-Soares F, Naranjo ME, Delgado À, de Andrés F, et al. Interethnic variation of CYP2C19 alleles, ‘predicted’ phenotypes and ‘measured’ metabolic phenotypes across world populations. Pharmacogenomics J 2016;16(2):113–23.
Zhou Y, Ingelman-Sundberg M, Lauschke VM. Worldwide distribution of cytochrome P450 alleles: a meta-analysis of population-scale sequencing projects. Clin Pharmacol Ther 2017;102(4):688–700.
Helsby NA, Burns KJE. Molecular mechanisms of genetic variation and transcriptional regulation of CYP2C19. Front Genet 2012;3:206.
Chen Y, Goldstein JA. The transcriptional regulation of the human CYP2C genes. Curr Drug Metab 2009;10(6):567–78.
Harada S, Zhou Y, Duncan S, Armstead AR, Coshatt GM, Dillon C, et al. Precision medicine at the University of Alabama at Birmingham: laying the foundational processes through implementation of genotype-guided antiplatelet therapy. Clin Pharmacol Ther 2017;102(3):493–501.
Sofi F, Giusti B, Marcucci R, Gori AM, Abbate R, Gensini GF. Cytochrome P450 2C19*2 polymorphism and cardiovascular recurrences in patients taking clopidogrel: a meta-analysis. Pharmacogenomics J 2011;11(3):199–206.
Furuta T, Shirai N, Sugimoto M, Ohashi K, Ishizaki T. Pharmacogenomics of proton pump inhibitors. Pharmacogenomics 2004;5(2):181–202.
Uckun Z, Baskak B, Ozel-Kizil ET, Ozdemir H, Devrimci Ozguven H, Suzen HS. The impact of CYP2C19 polymorphisms on citalopram metabolism in patients with major depressive disorder. J Clin Pharm Ther 2015;40(6):672–9.
Steimer W, Zöpf K, von Amelunxen S, Pfeiffer H, Bachofer J, Popp J, et al. Amitriptyline or not, that is the question: pharmacogenetic testing of CYP2D6 and CYP2C19 identifies patients with low or high risk for side effects in amitriptyline therapy. Clin Chem 2005;51(2):376–85.
Gao N, Tian X, Fang Y, Zhou J, Zhang H, Wen Q, et al. Gene polymorphisms and contents of cytochrome P450s have only limited effects on metabolic activities in human liver microsomes. Eur J Pharm Sci 2016;92:86–97.
Lewis JP, Stephens SH, Horenstein RB, O’Connell JR, Ryan K, Peer CJ, et al. The CYP2C19*17 variant is not independently associated with clopidogrel response. J Thromb Haemost 2013;11(9):1640–6.
Shirasaka Y, Chaudhry AS, McDonald M, Prasad B, Wong T, Calamia JC, et al. Interindividual variability of CYP2C19-catalyzed drug metabolism due to differences in gene diplotypes and cytochrome P450 oxidoreductase content. Pharmacogenomics J 2016;16(4):375–87.
Sanford JC, Guo Y, Sadee W, Wang D. Regulatory polymorphisms in CYP2C19 affecting hepatic expression. Drug Metabol Drug Interact 2013;28(1):23–30.
Coller JK, Somogyi AA, Bochner F. Comparison of (S)-mephenytoin and proguanil oxidation in vitro: contribution of several CYP isoforms. Br J Clin Pharmacol 1999;48(2):158–67.
Goldstein JA. Clinical relevance of genetic polymorphisms in the human CYP2C subfamily. Br J Clin Pharmacol 2001;52(4):349–55.
van der Hoeven TA, Coon MJ. Preparation and properties of partially purified cytochrome P-450 and reduced nicotinamide adenine dinucleotide phosphate-cytochrome P-450 reductase from rabbit liver microsomes. J Biol Chem 1974;249(19):6302–10.
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ. Protein measurement with the Folin phenol reagent. J Biol Chem 1951;193(1):265–75.
Srivastava PK, Yun CH, Beaune PH, Ged C, Guengerich FP. Separation of human liver microsomal tolbutamide hydroxylase and (S)-mephenytoin 4′-hydroxylase cytochrome P-450 enzymes. Mol Pharmacol 1991;40(1):69–79.
Pedersen RS, Brasch-Andersen C, Sim SC, Bergmann TK, Halling J, Petersen MS, et al. Linkage disequilibrium between the CYP2C19*17 allele and wildtype CYP2C8 and CYP2C9 alleles: identification of CYP2C haplotypes in healthy Nordic populations. Eur J Clin Pharmacol 2010;66(12):1199–205.
Shah RR, Smith RL. Inflammation-induced phenoconversion of polymorphic drug metabolizing enzymes: hypothesis with implications for personalized medicine. Drug Metab Dispos 2015;43(3):400–10.
Rideg O, Háber A, Botz L, Szücs F, Várnai R, Miseta A, et al. Pilot study for the characterization of pharmacogenetically relevant CYP2D6, CYP2C19 and ABCB1 gene polymorphisms in the Hungarian population. Cell Biochem Funct 2011;29(7):562–8.
Rost KL, Brockmöller J, Esdorn F, Roots I. Phenocopies of poor metabolizers of omeprazole caused by liver disease and drug treatment. J Hepatol 1995;23(3):268–77.
Yucel E, Sancar M, Yucel A, Okuyan B. Adverse drug reactions due to drug-drug interactions with proton pump inhibitors: assessment of systematic reviews with AMSTAR method. Expert Opin Drug Saf 2016;15(2):223–36.
Chung WG, Park CS, Roh HK, Lee WK, Cha YN. Oxidation of ranitidine by isozymes of flavin-containing monooxygenase and cytochrome P450. Jpn J Pharmacol 2000;84(2):213–20.
Liu A, Wang C, Hehir M, Zhou T, Yang J. In vivo induction of CYP in mice by carbamazepine is independent on PXR. Pharmacol Rep 2015;67(2):299–304.
Lakehal F, Wurden CJ, Kalhorn TF, Levy RH. Carbamazepine and oxcarbazepine decrease phenytoin metabolism through inhibition of CYP2C19. Epilepsy Res 2002;52(2):79–83.
Fontana RJ, Shakil AO, Greenson JK, Boyd I, Lee WM. Acute liver failure due to amoxicillin and amoxicillin/clavulanate. Dig Dis Sci 2005;50(10):1785–90.
Helsby NA, Lo WY, Sharples K, Riley G, Murray M, Spells K, et al. CYP2C19 pharmacogenetics in advanced cancer: compromised function independent of genotype. Br J Cancer 2008;99(8):1251–5.
Frye RF, Schneider VM, Frye CS, Feldman AM. Plasma levels of TNF-alpha and IL-6 are inversely related to cytochrome P450-dependent drug metabolism in patients with congestive heart failure. J Card Fail 2002;8(5):315–9.
Tóth K, Bűdi T, Kiss À, Temesvári M, Háfra E, Nagy A, et al. Phenoconversion of CYP2C9 in epilepsy limits the predictive value of CYP2C9 genotype in optimizing valproate therapy. Per Med 2015;12(3):199–207.
Frye RF, Zgheib NK, Matzke GR, Chaves-Gnecco D, Rabinovitz M, Shaikh OS, et al. Liver disease selectively modulates cytochrome P450-mediated metabolism. Clin Pharmacol Ther 2006;80(3):235–45.
Aitken AE, Morgan ET. Gene-specific effects of inflammatory cytokines on cytochrome P450 2C, 2B6 and 3A4 mRNA levels in human hepatocytes. Drug Metab Dispos 2007;35(9):1687–93.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kiss, Á.F., Vaskó, D., Déri, M.T. et al. Combination of CYP2C19 genotype with non-genetic factors evoking phenoconversion improves phenotype prediction. Pharmacol. Rep 70, 525–532 (2018). https://doi.org/10.1016/j.pharep.2017.12.001
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1016/j.pharep.2017.12.001