Abstract
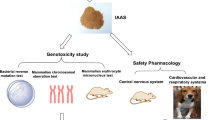
The leaves, stems, and fruits of Annona atemoya (A. atemoya; AA), a fruit-bearing plant of the family Annonaceae, exhibit anti-angiogenic, anti-oxidative, anti-inflammatory, and neuroprotective activities. However, the safety of AA has not been comprehensively elucidated. In this study, we evaluated the potential genotoxicity of an AA leaf (AAL) ethanol extract using a standard three-test battery constituting in vitro mammalian chromosomal aberration, in vivo micronucleus, and bacterial reverse mutation (also known as the Ames test) tests, as recommended by the Ministry of Food and Drug Safety of Korea. In vitro chromosomal aberration assay revealed that AAL extract did not induce structural or numerical aberrations, with or without metabolic activation (S9). In vivo micronucleus assay revealed that the number of micronucleated polychromatic erythrocytes (PCEs) and the PCE/normochromatic erythrocyte ratio after AAL extract treatment were not substantially different from those in the negative control. Changes in body weight and mortality were not observed. However, AAL extract partially induced mutagenic activity in all three bacterial strains in the bacterial reverse mutation assay, indicating that it could potentially aid in determining the genotoxic safety of AAL. QuantSeq 3′ mRNA sequencing analysis to elucidate the genotoxicity mechanisms of AAL extract using TK6 cells revealed that the genotoxic effects of AAL may be associated with cellular morphology-associated (cell development and keratinization), nucleotide metabolism, and electron transport chain functions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants are used as food or traditional medicine. They are increasingly being investigated as therapeutic agents or functional foods [1, 2], as replacements for chemical drugs to reduce various drug side effects, and for their pharmacological effects against various diseases [1]. Considering the biological effects of natural products, information on the risks of medicinal plants is essential to the identification of food supplements from plants [1, 3]. Notably, the overutilization of natural materials or compounds could exert side effects, such as mutagenicity and carcinogenicity [4], which necessitate the careful consideration of the efficacy and possible risks of natural herbal products [5, 6].
Annona atemoya (AA) is a commercially available fruiting plant of the family Annonaceae. It grows in tropical and subtropical climates, including Jeju Island in South Korea, the USA, and Africa [7, 8]. The AA leaves (AAL) grow to an average length of 15 cm, are leathery, and are relatively hairy [8]. Although several parts of AA, including its fruits, leaves, and seeds, are beneficial, only the fruits of AA are edible and safe [8,9,10]. According to previous in vivo and in vitro studies, AA extracts exhibit pharmacological effects, such as anti-angiogenic [11, 12], anti-bacterial, anti-oxidative [13], anti-inflammatory [14], and neuroprotective effects [15, 16].
Recently, an increasing demand for intellectual scrutiny has emerged owing to concerns surrounding the potential harm or toxicity attributed to products derived from natural herbs, their interactions with other pharmaceutical drugs, or their inherent toxic properties [6, 17]. Genotoxic substances hold notable significance owing to their ability to induce congenital disabilities or function as carcinogens, thereby presenting considerable risks to human health [18]. Genotoxicity tests, which involve assessment of direct damage to the DNA or chromosomes caused by a test substance to determine the mechanisms underlying morphological changes or functional abnormalities, are also used to determine the genotoxicity of drugs, pesticides, and various chemicals. However, given the diverse array of mechanisms that cause toxicity, a precise assessment of the genotoxic potential of a test substance requires a comprehensive approach involving multiple genotoxicity tests rather than relying solely on a single test. Currently, the standard battery of genotoxicity assessment comprises three primary assays: the in vitro chromosomal aberration assay, in vivo micronucleus assay, and bacterial reverse mutation test.
We recently demonstrated that AAL has neuroprotective effects in scopolamine-induced memory impairment and Alzheimer’s disease-like animal models [15, 16]. Despite the scientifically reported efficacy of AAL, its genetic toxicity has not been comprehensively assessed. Therefore, we aimed to evaluate the genotoxicity of AAL ethanol extract using the standard three-battery test in this non-clinical study, in accordance with the guidelines of the Korean Ministry of Food and Drug Safety (Guidebook No. 2017-0290-02).
Materials and methods
Chemicals and reagents
Chinese hamster lung (CHL) cells and TK-6 cells (ATCC CRL-1935, CRL-8015, American Type Culture Collection, Rockville, MD, USA), minimum essential medium (MEM, Cat. No. 11095080, Gibco Life Technologies Corp, Carlsbad, CA, USA), RPMI-1640 medium (Welgene, Gyeongsan, Korea), fetal bovine serum (FBS) (Cat. No. 10099141, Gibco Life Technologies), penicillin, streptomycin (Gibco Life Technologies), mitomycin C (MMC, Cas. No. 50-07-7, Sigma-Aldrich Co., Burlington, MA, USA), cyclophosphamide monohydrate (CPA, Cas. No. 6055-19-2, Sigma-Aldrich), furylfuramide (AF-2, Cas. No. 3688-53-7, Wako Pure Chemical Industries, Ltd., Tyoko, Japan), sodium azide (NaN3), 2-aminoanthracene (2-AA), 9-aminoacridine (9-AA), 2-(2-furyl)-3-(5-nitro-2-furyl) acrylamide (AF-2), benzo(a)pyrene (Cas. No. 26628-22-8, 613-13-8, 90-45-9, 50-32-8, Sigma-Aldrich), Samonella typhimurium TA 100, TA1535, TA98, TA1537, Escherichia coli WP2uvrA, S9 MIX (Molecular Toxicology Inc., Boone, NC, USA), Bacto-agar (Becton Dickinson and Company, Franklin Lake, New Jersey, USA), sodium chloride (Sigma-Aldrich), Vogel-bonner medium (× 10, Sigma-Aldrich), trypsin- ethylenediaminetetraacetic acid (EDTA; Life Technologies Corp), acridine orange (cas no 65-61-2, Sigma-Aldrich), and dimethyl sulfoxide (DMSO; Wako Pure Chemical Industries).
Preparation of extract
AAL were provided by the Sunny Farm (Jeju, Korea). A voucher specimen (no. A111-2019-2) was preserved at the KM Convergence Research Division of the Korea Institute of Oriental Medicine. The AAL extract was prepared following the good manufacturing practice (GMP) standards described by Huons Foodience Co., Ltd. (Chuncheon, Korea). Dried AAL (31.5 kg) were extracted twice with 70% aqueous ethanol (630 L) at 80 °C for 3 h. The extracted solution was filtered using Whatman filter paper, then concentrated under vacuum at 60 °C using a rotary evaporator and freeze-dried to obtain the powdered extract. The extraction powder (yield = 7.8 kg, 24.76%) was stored at − 20 °C until use for toxicological evaluation. The extracts were dissolved in distilled water for all toxicity tests.
Mammalian chromosomal aberration test in CHL cells
The mammalian chromosomal aberration test was performed at the Korea Testing & Research Institute (Gwacheon, Korea) per the Organization for Economic Co-operation and Development (OECD) guidelines for the testing of chemicals, Section 4, Test No. 473 in vitro mammalian chromosomal aberration test [19, 20] and Principles of Good Laboratory Practice (OECD 1998) [21]. CHL cells were used to measure the structural abnormalities in chromosomes caused by AAL extract. A 5 mL cell suspension prepared with a density of 4 × 103 cells/mL was incubated in a 25 cm culture flask and cultured for 3 days. After removing the MEM, the test solution, S9 mix, and MEM were added to the culture flask in the subsequent composition. For the negative control group, the same procedure was performed using the solvent. The cells were treated for 6 h using the short-term treatment and 24 h using the continuous treatment. After 6 h of short-term treatment, the cell surface was washed 2–3 times with PBS, and 5 mL of fresh MEM was added, followed by incubation for 18 h. The cells were treated with AAL extract, and the positive controls, CPA and MMC, were dissolved in distilled water. To measure the relative cell number, the cell surfaces were washed with PBS and then treated with 0.25% trypsin—EDTA at 37 °C for 5 min. MEM was added, and the cells were detached by pipetting. Live cells were counted using a cytometer (Beckman Coulter, Brea, CA, USA). The increase in relative cell numbers yielded a cytotoxic concentration of 55 ± 5% in relative increase in cell counts (RICC). RICC55 was calculated as a straight line for two concentrations, with a cell proliferation rate of 55 ± 5%. Based on the concentration determination test results, the concentration exceeding RICC55 was set as the highest concentration under each treatment condition, and the concentrations were calculated. The RICC55, a cytotoxicity indicator, was 155.25 µg/mL in the absence of the S9 mix, 880.51 µg/mL in the presence of the S9 mix in 6 h, and 51.71 µg/mL after treatment for 24 h. The concentration range was based on cytotoxicity, with or without S9. Metaphase cells were added to each culture flask to a final concentration of 0.2 μg/mL 2 h before the end of the treatment. After treatment, the cell surface was washed with PBS, trypsinized, and centrifuged at 1000 rpm for 5 min at 37 °C to collect the cells. After removing the supernatant, KCl solution (0.075 mol/L) was added, incubated at 37 °C for 15 min, and then fixed with a cool fixative (methanol: glacial acetic acid mixture [3:1, v/v]). The suspended cells were dropped onto a glass slide at 2–3 places to dry and then treated with 5% (v/v) Giemsa staining solution. Three hundred metaphase cells per dose were observed using a light microscope (Olympus, Tokyo, Japan). The results of the in vitro mammalian chromosomal aberration test were evaluated in accordance with the criteria reported by Honma et al. [20]. In the case of the negative control group, the frequency of appearance of structurally abnormal cells and numerically abnormal cells was set to < 5%, and in the case of the positive control group, the frequency of appearance of structurally abnormal cells was determined to be > 10%.
In vivo micronucleus test
The micronucleus test was performed at the Korea Testing & Research Institute (TGK-2019-000268, Gwacheon, Korea) following the OECD guidelines for the testing of chemicals, Section 4, Test No. 474 Micronucleus test (OECD 1997) [22] and OECD Principles of Good Laboratory Practice (OECD 1997) [21]. This method was used to detect chromosomal or mitotic apparatus damage caused by the bone marrow or peripheral blood cells of mice treated with AAL extract. Five male CrlOri:CD1 (ICR) mice aged 7 weeks were treated with AAL extract at oral doses of 500, 1000, and 2000 mg/kg/day twice daily at 24-h-intervals or intraperitoneal injection of CPA (70 mg/kg/day) as the positive control. Mice were treated with AAL extract two or three times, usually 24 h apart, in accordance with OECD guidelines. We conducted a dose range-finding test before performing the main test. The highest dose was determined based on this test, and 2000 mg/kg, which was the limit dose for treatment up to 14 days according to the OECD guidelines, was selected as the maximum dose. All the mice were observed daily for clinical signs. To prepare mouse bone marrow cells, mice were euthanized via inhalation anesthesia, and bone marrow was extracted according to the method reported by Schmid [23]. The bone marrow cells were washed with 0.5 mL of FBS, and the cell suspension was collected and centrifuged at 1000 rpm for 5 min. After removing the supernatant, bone marrow cells were smeared on a glass slide and dried at room temperature. The methanol-fixed specimen was dripped with an acridine orange solution (40 μg/mL), followed by fluorescence staining by covering it with a cover glass. In total, 4000 polychromatic erythrocytes (PCEs) per animal were scored to determine the frequency of micronucleus polychromatic erythrocytes (MNPCEs). To evaluate the myelotoxic effects, 500 erythrocytes were scored per animal to determine the ratio of PCEs to the total number of erythrocytes. Slides were observed using a blinded method but using a fluorescence microscope at 400 × magnification or a microscope at 1000 × magnification. PCEs were differentiated by red fluorescence without a nucleus, and chromatic erythrocytes were distinguished according to the size of the PCEs, exhibiting no fluorescence at all, and only according to shade. The largest criterion for determining the size of micronuclei was half the diameter of the red blood cells, and the smallest micronucleus was identifiable. The nuclei were primarily circular, donut-shaped, or semicircular. They were determined to have green fluorescence, similar to the nucleus of the adjacent nucleated cell.
Bacterial reverse mutation test (Ames test)
The in vitro Ames test was performed at Korea Testing & Research Institute (Gwacheon, Korea) in accordance with the OECD guidelines for the testing of chemicals, Section 4; Test No. 471: Bacterial reverse mutation test (OECD 1997) [24] and OECD Principles of Good Laboratory Practice (OECD 1997) [21]. Preliminary tests for range-finding and the main tests to determine the dose range of AAL extract in the absence and presence of the S9 mix were conducted using three plates per dose and five strains: TA98, TA100, TA1535, TA1537, and WP2uvrA. Test strains were purchased from a commercial manufacturer. S9 mix, the metabolic activation system, was prepared and immediately placed on ice. The preliminary range-finding test was performed at concentrations of 5000, 1500, 500, 150, and 50 μg/plate according to the aforementioned guidelines (Supplementary Table S1). As a positive control, NaN3 (5 μg/mL) was dissolved in sterile distilled water, and AF-2 (0.01 or 0.1 µg/mL), 9-AA (40 µg/mL), 2-AA (2 or 10 μg/mL), and benzo(a)pyrene (2.5 μg/mL) were dissolved in DMSO. Ames tests were performed using the pre-incubation method with and without the S9 mix. Precipitation and turbidity were observed with the naked eye. Bacterial growth inhibition was observed under a microscope after 48 h of incubation, and the colonies were counted.
Statistical analysis for the toxicity tests
Data were processed for statistical analysis using SPSS (Ver. 19, IBM, San Francisco, CA, USA). Statistical analyses of micronuclei were performed using the Kruskal–Wallis H-test and Mann–Whitney U-test for micronucleus induction frequency and comparison of the negative and positive controls. For PCE/(PCE + NCE), the negative control and vehicle (positive control) were analyzed using ANOVA and Dunnett’s tests. Chromosomal aberration test data with a frequency of appearance of abnormal cells below 5% were not statistically analyzed. The negative and positive controls were compared using Student’s t-test. Results with p < 0.05 were considered statistically significant. All values were presented as mean ± standard deviation (SD).
Cytotoxicity test
The TK6 cell line (CRL-8015) was maintained in RPMI1640 medium (Welgene) supplemented with 10% heat-inactivated FBS (Gibco), penicillin (100 U/mL)/streptomycin (100 μg/mL) (Gibco) in a humidified incubator with 5% CO2 at 37 °C. Cell Counting Kit-8 (CCK; Dojindo Molecular Technologies Inc., Kumamoto, Japan) assay was performed to measure the cytotoxicity of AAL extract against TK6 cells. Cells were plated onto 96-well microplates at 1.5 × 104 cells/well and treated with various concentrations of AAL extracts (0, 0.78, 1.57, 3.12, 6.25, 12.5, 25, 50, 100, 200, or 400 μg/mL) for 24 h. Following incubation with CCK reagent for 3 h, optical density (OD) at 450 nm was read using a Benchmark plus microplate reader (Bio-Rad Laboratories Inc., Hercules, CA, USA). Cell viability was determined using the following equation: Cell viability (%) = (mean ODAAL-treated cells/mean ODuntreated control) × 100.
RNA isolation
Total RNA was extracted from AAL extract-treated TK6 cells using TRIzol reagent, according to the manufacturer’s instructions (Invitrogen, Carlsbad, CA, USA). RNA quality was assessed using the Agilent TapeStation 4000 system (Agilent Technologies, Amstelveen, Netherlands). RNA was quantified using a NanoDrop-2000 Spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). Only samples with RNA integrity > 7.0 were included in the QuantSeq analysis.
Library preparation and sequencing for QuantSeq analysis
The construction of library was performed using QuantSeq 3′ mRNA-Seq Library Prep Kit (Lexogen, Inc., Vienna, Austria) according to the manufacturer’s instructions. In brief, each total RNA was prepared and an oligo-dT primer containing an Illumina-compatible sequence at its 5′ end was hybridized to the RNA, and reverse transcription was performed. After degradation of the RNA template, second strand synthesis was initiated by a random primer containing an Illumina compatible linker sequences at its 5′ end. The double-stranded library was purified using magnetic beads to remove all reaction components. The library was amplified to add the complete adapter sequences required for cluster generation. The completed library was purified from PCR components. High-throughput sequencing was performed as single-end 75 sequencing using NextSeq 550 (Illumina, Inc., San Diego, CA, USA).
QuantSeq data analysis
QuantSeq 3′ mRNA-Seq reads were aligned with the human genome database (GRCh38) using STAR [25]. The alignment file was used for assembling transcripts, estimating their abundances and detecting differential expression of genes. The read count data were processed based on the trimmed mean of M-values (TMM) + counts per million (CPM) normalization method using EdgeR within R (R development Core Team, 2020) using Bioconductor [26].
Isolation of genes and gene ontology
Genes responding to AAL treatment were defined as genes that exhibited more than 1.5-fold changes in expression by AAL treatment compared to genes in the normal control. We then clustered these genes based on similarity in expression levels using Gene Cluster 3.0 and visualized them using Java TreeView [27]. Gene ontology terms associated with AAL-responsive genes were obtained from the DAVID site (https://david.ncifcrf.gov), where modified Fisher’s exact test was used to measure statistical significance [28]. Gene ontology networks were measured using the ClueGO (version 2.5.10) Cytoscape plugin program, with default parameter settings, and GO-terms fusion check [29].
Results
Effect of AAL extract on chromosomal aberration
A chromosomal aberration test was conducted using cultured mammalian cells to evaluate the potential inducibility of chromosomal aberrations by the test substance, AAL extract. The concentration range was determined based on cytotoxicity, with or without the S9 mix, as observed in the preliminary dose-range test. Results presented in Table 1 demonstrate that following short-duration treatments (6–18 h), the observed frequency levels of chromosomal structure abnormalities were 0.0%, 0.3%, 0.0%, and 0.3% at concentrations of 0, 50, 100, and 150 μg/mL in the absence of S9 mix, respectively. Correspondingly, the frequency of numerical abnormalities was 0.0%, 0.0%, 0.0%, and 0.0%. Additionally, frequencies of chromosomal aberrant cells were 0.3%, 0.0%, 0.3%, 0.0%, and 0.0%, and the frequencies of numerical abnormal cells were 0.0%, 0.0%, 0.0%, 0.0%, and 0.0% at concentrations of 0, 225, 450, 900, and 1000 μg/mL in the presence of + S9 mix. Moreover, 24 h treatment yielded frequency levels of chromosomal aberrations and abnormalities of 0.0%, 0.0%, 0.0%, and 0.3% at concentrations of 0, 12.5, 25, and 50 μg/mL, respectively. However, AAL extract without S9 mix exhibited cytotoxicity, determined by a cytotoxicity indicator (RICC55), at 200 μg/mL under treatment for a short duration or 100 μg/mL under continuous treatment in the preliminary dose-range test, precluding the assessment of chromosomal aberrations. In the negative control group, the frequency of appearance of structural and numerical abnormal cells was confirmed to be less than 5%. In contrast, the positive control group exhibited a frequency of appearance of structurally abnormal cells exceeding 10%.
Effect of AAL extract on body weight
Oral toxicity was monitored using the micronucleus test for AAL extract. Doses of 500, 1000, or 2000 mg/kg of AAL extract or 70 mg/kg of positive control CPA per day were administered to the animals for 2 days (Table 2). No signs of toxicity, such as changes in body weight or mortality, were observed in any experimental group.
Effect of AAL extract on micronucleus test
Based on the observation results of > 4000 polychromatic red blood cells per individual presented in Table 3, the frequencies of red blood cells with micronuclei among polychromatic red blood cells were 0.10 ± 0.02%, 0.08 ± 0.02%, 0.06 ± 0.03%, and 0.06 ± 0.01% at 0, 500, 1000, and 2000 mg/kg, respectively. The frequency of the positive group was 6.75 ± 0.52%. In all test substance-administered groups, the frequency of red blood cells with micronuclei among polychromatic red blood cells did not increase compared with that in the negative control group. In contrast, the frequency of micronuclei in the positive control group was significantly higher than that in the negative control group (p < 0.05). The ratio [PCE/(PCE + NCE)], an indicator of cytotoxicity, was averaged to be 51.24 ± 1.21%, 51.20 ± 1.18%, 51.13 ± 2.03%, and 51.71 ± 0.75% at 0, 500, 1000, and 2000 mg/kg of AAL extract, respectively, and 45.79 ± 0.68% in the positive control group; no significant difference was observed between the AAL extract-treated groups and the negative control group. In contrast, the [PCE/(PCE + NCE)] ratio of the positive control group was significantly different from that of the negative control group (p < 0.05).
Effect of AAL extract in Ames test
To evaluate whether AAL extract induced bacterial reverse mutation, the Ames test was conducted using four types of S. typhimurium histidine (TA98, TA100, TA1535, and TA1537) and a type of E. coli tryptophan (WP2uvrA) in the absence or presence of the S9 mix. In the preliminary dose range-finding test of the TA100 strain without the S9 mix, and the TA98, TA100, TA1537 and WP2uvr A strains with the S9 mix, the number of colonies was increased in a concentration-dependent manner compared with that in the negative control group (0 μg/plate). Accordingly, the main test was conducted at the following concentrations, with or without the S9 mix: 313, 625, 1250, 2500, or 5000 μg/plate (TA98, TA100, TA1535, TA1537, and WP2uvrA). In the main test, an increase in colony number compared to that in the negative control was observed in the TA100 strain without the S9 mix and in the TA98, TA100, and TA1537 strains with the S9 mix. No precipitation or turbidity was observed on the plate in any AAL extract-treated groups, regardless of whether a metabolic activation system was applied. These findings suggest that AAL is mutagenic (positive), as evidenced by the results of the Ames test under the aforementioned conditions (Table 4).
Genotoxicity mechanisms of AAL extract
QuantSeq 3′ mRNA sequencing analysis was conducted to elucidate the genotoxicity mechanisms of AAL using a TK6 thymidine kinase heterozygote cell line. The cytotoxicity of AAL extract was determined using the CCK assay in TK6 cells (Fig. 1). Based on the results of cytotoxicity test, cells were treated with 50 μg/mL for 24 h and used for 3′ mRNA sequencing. Transcriptome analysis was performed to measure overall changes in gene expression. A total of 439 genes were identified as AAL-responsive genes based on the aforementioned criterion. Figure 2a shows the expression pattern of AAL-responsive genes. The list of the representative top-ranked down- and up-regulated genes is shown in Table 5. These AAL-responsive genes are associated with diverse biological functions. Especially, cellular morphology-associated functions (cell development and keratinization), nucleotide metabolism functions, and electron transport functions were significantly enriched with AAL-responsive genes (Fig. 2b). Network analysis also showed the relationship among these functional gene ontology terms (Fig. 2c).
Cytotoxicity of AAL extract against TK6 cells. Cells were plated onto 96-well microplates and treated with various concentrations of AAL extract ranging from 0 to 400 μg/mL for 24 h. Cell viability (% of control) was assessed using the Cell Counting Kit-8 assay. The values are expressed as the mean ± standard error of four independent experiments
Genotoxicity mechanisms of AAL extract in TK6 cells. QuantSeq 3′ mRNA sequencing analysis was performed using TK6 cells treated with AAL for 24 h. a AAL-responsive genes that exhibited more than 1.5-fold changes in expression by AAL treatment compared to the normal control were clustered based on similarity in expression levels. As shown in the scale bar, red and blue represent high and low levels, respectively. b A list of enriched gene ontology terms (p < 0.001) associated with AAL-responsive genes is shown. Red, blue, and green represent morphology-associated functions, nucleotide metabolism functions, and electron transport functions, respectively. c These enriched ontology terms are interconnected in the network. Representative ontology terms are indicated
Discussion
Advancement in research on the pharmacological efficacy of AAL extract and its potential as an alternative medicine or food necessitates a comprehensive evaluation of its safety. In this study, we systematically evaluated the genotoxicity of AAL, a natural resource for food products, using a three-battery assay test. Each non-clinical test was conducted in compliance with the OECD guidelines [19, 22, 24] and GLP regulations of the test substance registration [21], aligning with the guidelines of the Korean Ministry of Food and Drug Safety (Guideline No. 2017-0290-02) [30]. Our findings demonstrated that AAL extract exhibited minimal inducibility of chromosomal aberrations in the mammalian cell-based test, with negligible cytotoxic effects. Furthermore, the micronucleus and Ames tests did not reveal substantial chromosomal damage or mutagenic potential. Notably, the Ames test revealed that the AAL possessed mutagenic properties, presenting a significant aspect in the assessment of genotoxicity.
The chromosomal aberration test involves the treatment of incubated mammalian cells with test substances in the absence or presence of an exogenous metabolic system (S9 mix). Double-strand breaks, a major cause of structural chromosomal abnormalities, can be induced directly or indirectly by repair errors or the replication of DNA lesions [31]. Various cultured cell lines, including mammalian CHL cells, can be used to test the chemical induction of DNA damage and are suitable for identifying possible mutagens and carcinogens [32]. Chromosomal aberration test results of AAL extract indicated chromosomal structure and numerical abnormalities < 5% for both short-term and continuous treatments. This test was validated, as the frequency of occurrence of chromosomally aberrant cells in the negative and positive control groups fell within the range of historical background data [19], suggesting that AAL did not induce chromosomal abnormalities in CHL/IU cells under these test conditions.
The micronucleus test is the most reliable method for quantifying chromosomal damage caused by test substances by enumerating erythrocytes collected from the bone marrow and peripheral blood lymphocytes of rodents [33]. An increase in the number of micronucleus-bearing PCEs can signify chromosomal damage or hematopoietic perturbations [34]. Our findings suggested that AAL extract-treated mice did not exhibit mortality- or toxicity-related clinical signs in general appearance or body weight changes up to 2000 mg/kg (Table 2). The frequency of micronucleus-bearing erythrocytes among PCE in all administration groups did not exhibit an increasing trend compared with that in the negative control group and indicated no statistical significance. Additionally, PCE/(PCE + NCE) ratios for all erythrocytes did not differ significantly from those of the negative control group. The mean values of the MNPCE occurrence frequency and PCE ratio in the negative control group were within the range of historical background data [35]. Furthermore, statistical significance was observed in the positive control group. These results suggest that AAL extract does not induce micronuclei formation in male ICR mouse bone marrow cells under our test conditions.
The Ames test, a pivotal non-clinical safety assessment, helps detect the mutagenic effects of dietary and plant extracts with simplicity and sensitivity [33, 36]. It employs S9 metabolic activation for detecting indirect mutagens or operates without it for direct mutagen detection, using various S. typhimurium and E. coli test strains [37]. Strains TA100, TA1535, and WP2uvrA harbor base-pair substitution mutations, whereas the TA98 and TA1537 strains harbor frameshift mutations [38]. Bacterial reverse mutation test results revealed concentration-dependent increases in the mutant colonies in the TA100 strain with metabolic activation, and in TA98, TA100 and TA1537 without metabolic activation, compared with those in the negative control. These results suggest potential direct or indirect mutagenic properties within the AAL extract. Reproducibility was confirmed using the preliminary tests for range-finding and the main tests to determine the dose range of AAL extract in the absence or presence of the S9 mix, and the negative and positive control values were within the appropriate range. Furthermore, the number of reversible mutations induced in the positive controls demonstrated significantly increased positive results for all test strains, with or without metabolic activity.
Considering the results of the Ames test, we elucidated the genotoxicity mechanisms of AAL extract. We utilized the TK6 thymidine kinase heterozygote cell line broadly used for toxicogenomic studies to understand genotoxicity mechanisms [39,40,41,42]. Transcriptome analysis showed that a total of 439 genes exhibited a 1.5-fold change in expression by AAL treatment. The top five most changed down- or up-regulated genes included NADH:ubiquinone oxidoreductase subunit B3, transcription factor B2, centromere protein A, apurinic/apyrimidinic endodeoxyribonuclease 1, heterogenous nuclear ribonucleoprotein A/B, keratin 6A, keratin 1, cystatin A, filaggrin, and loricrin cornified envelope precursor protein. These genes are mainly associated with cellular morphology-associated (cell development and keratinization), nucleotide metabolism, and electron transport chain functions. Several studies have described the crucial roles of these molecular functions in the regulation of genotoxicity. Genotoxicity directly or indirectly induces DNA damage [43]. DNA-damaged cells respond by controlling defense modes, such as cell cycle regulation, apoptotic cell death, and DNA repair system [44, 45]. Keratinization is the differentiation of epidermal cells to keratin-rich cells. Keratin structure can be damaged by ultraviolet (UV) radiation, an environmental genotoxic factor, and keratin 17A is a robust marker of UV-mediated stress in human keratinocytes [46, 47]. Disturbances in nucleotide metabolism involving nucleotide synthesis, degradation, and recycling influence genomic stability, making cells more susceptible to genotoxic insults and increasing the risk of mutagenesis and carcinogenesis [48, 49]. For instance, Jia et al. recently found a prominent and conserved role for the endonuclease ENDU-2 in regulating nucleotide metabolism in response to genotoxic stress, suggesting it as a potential target for cancer therapy [50]. The electron transport chain comprises a series of protein complexes that plays a role in the transfer of electrons from electron donors to acceptors, eventually producing ATP in the inner mitochondrial membrane. Park et al. reported that pendimethalin-induced genotoxicity causes loss of electron transport chain activity and mitochondrial dysfunction [51]. Gunaydin-Akyildiz et al. demonstrated that the genotoxic effect of favipiravir (T-705), an anti-influenza viral drug, causes ATP depletion by inhibiting the activities of electron transport chain enzymes [52]. Based on these evidences and our data from the toxicogenomic analysis, we propose that the genotoxicity of AAL may be induced by regulating the following molecular functions: cellular morphology-associated (cell development and keratinization), nucleotide metabolism, and electron transport chain functions.
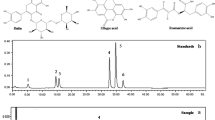
The clinical relevance of the Ames test for plant extracts ultimately lies in its ability to detect the capacity to protect human health. Identifying mutagenic compounds early in the development process can prevent their inclusion in products used by people, ultimately reducing the risk of adverse health effects like cancer. Typically, plant extracts are mixtures of multiple components with phytotherapeutic and phytonutritional properties [53, 54]. The Ames test is generally used to evaluate synthetic chemicals [55]. If a plant extract shows positive results in the Ames test, it can be suspected to contains mutagenic compounds and could be relevant in situations where there is concern about the safety of botanical extracts, which might contain mutagenic compounds that could pose a risk to human health. In our previous study, we used high-performance liquid chromatography (HPLC) analysis data for quality control (QC) of AAL extract using the six indicator substances: (β)-catechin, chlorogenic acid, epicatechin, rutin, isoquercitrin, and kaempferol-3-O-rutinoside [15, 16]. The components of AAL that induce mutagenic genotoxicity remain unknown based on the bacterial reverse mutation test. Noncarcinogenic substances frequently yield false-positive results in genotoxicity assays [32, 56]. Plant extracts will likely have positive mutagenic outcomes in the Ames test [57, 58]. Regulatory agencies, such as the U.S. Food and Drug Administration, require safety assessments for substances used in foods or medicinal drugs. If plant extracts are found to be mutagenic in the Ames test, the regulatory agencies may impose restrictions or necessitate further safety test before approving them for human consumption. Gomes da Silva Dantas et al. [59] reported in their systematic review that 18% of the species of medicinal plants were mutagenic, of which 54% were mutagenic in the presence or absence of S9. In particular, strains TA98 and TA100 showed a sensitivity of 93% in detecting plant extracts with mutagenic potential. They indicated that methodological issues should be considered to detect the mutagenicity of plant extracts, such as the use of maximum doses for plating and the lack of cytotoxicity information of tested plants. They finally suggested that the guidelines of the regulatory agencies are required to be updated to test the mutagenic potential of plant extracts, including the doses used in the study for the mixture and the most sensitive strains employed to detect the mutagenesis.
The Ames test is used for the comprehensive evaluation of a compound’s safety and provides valuable information about the mutagenic potential of substances. Other toxicity tests and studies are typically conducted to fully assess the potential impact on human health. If the Ames test yields positive results, chromosomal aberration and comet testing (single-cell gel electrophoresis assay) can serve as alternatives [60, 61]. The in vivo comet assay using plant extract can help validate in vitro genotoxicity findings [56, 62]. Therefore, further studies, including a comet assay, are warranted to clarify the mechanisms by which AAL causes these mutations and validate the genotoxicity profile of AAL. In addition, to comprehensively assess AAL extract’s safety and risk–benefit profile, future studies might integrate broader assessments considering factors like metabolism, individual/sex-based variation, and long-term exposure scenarios.
Collectively, our results suggested that AAL could induce mutations, such as substitution and frameshift mutations, evident from reverse mutations identified in the bacterial strains TA98, TA100, and TA1537; however, mutations in WP2uvrA and TA1535, identified in the bacterial reverse mutation test but not micronucleus and chromosomal aberration tests, were excluded. Nonetheless, a positive Ames test result does not inherently signify that AAL is inherently carcinogenic [63]. Therefore, despite the Ames test being an accessible screening tool for setting priorities, further extensive testing is required to determine whether AAL extract can cause cancer. Furthermore, the disparity between in vitro and in vivo results in this study highlights a need for further investigation to bridge this gap and ensure translatability to human health implications. Although no chromosomal aberrations or micronucleus toxicity of AAL was observed in this study, further research using the comet assay is recommended to elucidate the mutagenic risks of AAL.
Data availability
The data used to support the findings of this study are accessible from the corresponding author upon request.
References
Sen T, Samanta SK (2015) Medicinal plants, human health and biodiversity: a broad review. Adv Biochem Eng Biotechnol 147:59–110. https://doi.org/10.1007/10_2014_273
Akkol EK, Çankaya IT, Karatoprak GŞ et al (2021) Natural compounds as medical strategies in the prevention and treatment of psychiatric disorders seen in neurological diseases. Front Pharmacol 12:669638. https://doi.org/10.3389/fphar.2021.669638
Lim MK, Kim JY, Jeong J et al (2021) Evaluation of subchronic Toxicity and genotoxicity of ethanolic extract of Aster glehni leaves and stems. Evid Based Complement Alternat Med 2021:1018101. https://doi.org/10.1155/2021/1018101
Roberts SM, James RC, Williams PL (2015) Principles of toxicology: environmental and industrial applications. Wiley, Hoboken
Posadzki P, Watson LK, Ernst E (2013) Adverse effects of herbal medicines: an overview of systematic reviews. Clin Med 13:7–12. https://doi.org/10.7861/clinmedicine.13-1-7
Zhou J, Ouedraogo M, Qu F, Duez P (2013) Potential genotoxicity of traditional Chinese medicinal plants and phytochemicals: an overview. Phytother Res 27:1745–1755. https://doi.org/10.1002/ptr.4942
Morton JF, Dowling CF (1987) Fruits of warm climates. Distributed by Creative Resources System, Miami
Kazman BSMA, Harnett JE, Hanrahan JR (2020) The phytochemical constituents and pharmacological activities of Annona atemoya: a systematic review. Pharmaceuticals (Basel) 13:269. https://doi.org/10.3390/ph13100269
Tiangda CH, Gritsanapan W, Sookvanichsilp N, Limchalearn A (2000) Anti-headlice activity of a preparation of Annona squamosa seed extract. Southeast Asian J Trop Med Public Health 31(Suppl 1):174–177
Yi J-M, Park J-S, Lee J et al (2014) Anti-angiogenic potential of an ethanol extract of Annona atemoya seeds in vitro and in vivo. BMC Complement Altern Med 14:353. https://doi.org/10.1186/1472-6882-14-353
Nugraha AS, Damayanti YD, Wangchuk P, Keller PA (2019) Anti-infective and anti-cancer properties of the Annona species: their ethnomedicinal uses, alkaloid diversity, and pharmacological activities. Molecules 24:4419. https://doi.org/10.3390/molecules24234419
Mannino G, Gentile C, Porcu A et al (2020) Chemical profile and biological activity of Cherimoya (Annona cherimola Mill.) and Atemoya (Annona atemoya) leaves. Molecules 25:2612. https://doi.org/10.3390/molecules25112612
Rabêlo SV, Costa M, Libório RC (2014) Almeida JRGdS: antioxidant and antimicrobial activity of extracts from atemoia (Annona cherimola Mill. × A. squamosa L.). Rev Bras Frutic 36:265–271. https://doi.org/10.3390/ph13100269
Do Hallison NS, Suzana VR, Tamara CD et al (2017) Antinociceptive and anti-inflammatory activities of ethanolic extract from atemoya (Annona cherimola Mill × Annona squamosa L.). Afr J Pharm Pharmacol 11:224–232. https://doi.org/10.5897/ajpp2017.4778
Sohn E, Lim H-S, Kim YJ et al (2019) Annona atemoya leaf extract improves scopolamine-induced memory impairment by preventing hippocampal cholinergic dysfunction and neuronal cell death. Int J Mol Sci 20:3538. https://doi.org/10.3390/ijms20143538
Lim H-S, Kim YJ, Sohn E et al (2019) Annona atemoyaleaf extract ameliorates cognitive impairment in amyloid-β injected Alzheimer’s disease-like mouse model. Exp Biol Med (Maywood) 244:1665–1679. https://doi.org/10.1177/1535370219886269
Fu B, Wang N, Tan H-Y et al (2018) Multi-component herbal products in the prevention and treatment of chemotherapy-associated toxicity and side effects: a review on experimental and clinical evidences. Front Pharmacol 9:1394. https://doi.org/10.3389/fphar.2018.01394
Singh N, Manshian B, Jenkins GJS et al (2009) NanoGenotoxicology: the DNA damaging potential of engineered nanomaterials. Biomaterials 30:3891–3914. https://doi.org/10.1016/j.biomaterials.2009.04.009
Registre M, Proudlock R (2016) The in vitro chromosome aberration test. In: Genetic toxicology testing. Academic Press, Cambridge, pp 207–267
Honma M (1999) Evaluation of the mouse lymphoma TK assay (microwell method) as an alternative to the in vitro chromosomal aberration test. Mutagenesis 14:5–22. https://doi.org/10.1093/mutage/14.1.5
OECD (2019) OECD principles of good laboratory practice. Series on principles of good laboratory practice (GLP) and compliance monitoring, No 20. OECDiLibrary, Paris. https://doi.org/10.1787/2077785x
OECD (2016) Test no. 474: mammalian erythrocyte micronucleus test. OECDiLibrary, Paris. https://doi.org/10.1787/97892642647621_en
Schmid W (1975) The micronucleus test. Mutat Res Environ Mutag Related Subj 31:9–15. https://doi.org/10.1016/0165-1161(75)90058-8
OECD (2020) Test no. 471: bacterial reverse mutation test. OECDiLibrary, Paris. https://doi.org/10.1787/9789264071247_en
Dobin A, Davis CA, Schlesinger F et al (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29:15–21. https://doi.org/10.1093/bioinformatics/bts635
Gentleman RC, Carey VJ, Bates DM et al (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80. https://doi.org/10.1186/gb-2004-5-10-r80
De Hoon MJ, Imoto S, Nolan J, Miyano S (2004) Open source clustering software. Bioinformatics 20:1453–1454. https://doi.org/10.1093/bioinformatics/bth078
Dennis G, Sherman BT, Hosack DA et al (2003) DAVID: database for annotation, visualization, and integrated discovery. Genome Biol 4:3. https://doi.org/10.1186/gb-2003-4-9-r60
Bindea G, Mlecnik B, Hackl H et al (2009) ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25:1091–1093. https://doi.org/10.1093/bioinformatics/btp101
Korean Ministry of Food and Drug Safety Web (2019) Guidelines for nonclinical testing of herbal product. http://www.nifds.go.kr/brd/0290-02. Accessed 2 Apr 2019
Clare G (2012) The in vitro mammalian chromosome aberration test. Methods Mol Biol 817:69–91. https://doi.org/10.1007/978-1-61779-421-6_5
Lee S-B, Lee J-S, Wang J-H et al (2021) Genotoxicity of water extract from bark-removed Rhus verniciflua Stokes. Molecules 26:896. https://doi.org/10.3390/molecules26040896
Kim NS, Shin S, Shin G-G, Bang O-S (2019) Genotoxicity evaluation of a Phragmitis rhizoma extract using a standard battery of in vitro and in vivo assays. J Ethnopharmacol 241:112025. https://doi.org/10.1016/j.jep.2019.112025
Lee MY, Park Y-C, Jin M et al (2018) Genotoxicity evaluation of So-ochim-tang-gamibang (SOCG), a herbal medicine. BMC Complement Altern Med 18:47. https://doi.org/10.1186/s12906-018-2111-2
Lovell DP, Fellows M, Marchetti F et al (2018) Analysis of negative historical control group data from the in vitro micronucleus assay using TK6 cells. Mutat Res Genet Toxicol Environ Mutagen 825:40–50. https://doi.org/10.1016/j.mrgentox.2017.10.006
Lewis DFV, Ioannides C, Parke DV (1993) Validation of a novel molecular orbital approach (COMPACT) for the prospective safety evaluation of chemicals, by comparison with rodent carcinogenicity and Salmonella mutagenicity data evaluated by the U.S. NCI/NTP Mutat Res-Environ Mutag Related Subj 291:61–77. https://doi.org/10.1016/0165-1161(93)90018-u
Hong C-E, Lyu S-Y (2013) Evaluation of the mutagenic properties of two lignans from Acanthopanax koreanum Nakai. Toxicol Res 29:279–283. https://doi.org/10.5487/tr.2013.29.4.279
Oliveira NdMS, Resende MR, Morales DA, De Ragão UG, Boriollo MFG (2016) In vitro mutagenicity assay (Ames test) and phytochemical characterization of seeds oil of Helianthus annuus Linné (sunflower). Toxicol Rep 3:733–739. https://doi.org/10.1016/j.toxrep.2016.09.006
Platel A, Gervais V, Sajot N et al (2010) Study of gene expression profiles in TK6 human cells exposed to DNA-oxidizing agents. Mutat Res 689:21–49. https://doi.org/10.1016/j.mrfmmm.2010.04.004
Kuehner S, Holzmann K, Speit G (2013) Characterization of formaldehyde’s genotoxic mode of action by gene expression analysis in TK6 cells. Arch Toxicol 87:1999–2012. https://doi.org/10.1007/s00204-013-1060-2
Suh SK, Kim TG, Kim HJ et al (2007) Gene expression in profiling of genotoxicity induced by MNNG in TK6 cells. Mol Cell Toxicol 3:98–106. https://doi.org/10.1016/j.mrfmmm.2010.04.004
Li HH, Hyduke DR, Chen R et al (2015) Development of a toxicogenomics signature for genotoxicity using a dose-optimization and informatics strategy in human cells. Environ Mol Mutagen 56:505–519. https://doi.org/10.1002/em.21941
Phillips DH, Arlt VM (2009) Genotoxicity: damage to DNA and its consequences. Mol Toxicol 99:87–110. https://doi.org/10.1007/978-3-7643-8336-7_4
Wang Y, Su M, Chen Y et al (2023) Research progress on the role and mechanism of DNA damage repair in germ cell development. Front Endocrinol (Lausanne) 14:1234280. https://doi.org/10.3389/fendo.2023.1234280
Chen ACH, Peng Q, Fong SW et al (2021) DNA damage response and cell cycle regulation in pluripotent stem cells. Genes (Basel) 12:1548. https://doi.org/10.3390/genes12101548
Lombardo G, Melzi G, Indino S et al (2022) Keratin 17 as a marker of UVB-induced stress in human epidermis and modulation by vitis vinifera extract. Cells Tissues Organs 211:611–627. https://doi.org/10.1159/000520038
Dos Santos CP, Londero JEL, Dos Santos MB et al (2018) Sunlight-induced genotoxicity and damage in keratin structures decrease tadpole performance. J Photochem Photobiol B 181:134–142. https://doi.org/10.1016/j.jphotobiol.2018.03.013
Yoshioka KI, Kusumoto-Matsuo R, Matsuno Y, Ishiai M (2021) Genomic instability and cancer risk associated with erroneous DNA repair. Int J Mol Sci 22:12254. https://doi.org/10.3390/ijms222212254
Wu HL, Gong Y, Ji P, Xie YF et al (2022) Targeting nucleotide metabolism: a promising approach to enhance cancer immunotherapy. J Hematol Oncol 15:45. https://doi.org/10.1186/s13045-022-01263-x
Jia F, Chi C, Han M (2020) Regulation of nucleotide metabolism and germline proliferation in response to nucleotide imbalance and genotoxic stresses by EndoU nuclease. Cell Rep 30:1848-1861.e5. https://doi.org/10.1016/j.celrep.2020.01.050
Park H, Lee JY, Lim W, Song G (2021) Assessment of the in vivo genotoxicity of pendimethalin via mitochondrial bioenergetics and transcriptional profiles during embryogenesis in zebrafish: implication of electron transport chain activity and developmental defects. J Hazard Mater 411:125153. https://doi.org/10.1016/j.jhazmat.2021.125153
Gunaydin-Akyildiz A, Aksoy N, Boran T et al (2022) Favipiravir induces oxidative stress and genotoxicity in cardiac and skin cells. Toxicol Lett 371:9–16. https://doi.org/10.1016/j.toxlet.2022.09.011
Ananthi R, Chandra N, Santhiya ST, Ramesh A (2010) Genotoxic and antigenotoxic effects of Hemidesmus indicus R. Br. root extract in cultured lymphocytes. J Ethnopharmacol 127:558–560. https://doi.org/10.1016/j.jep.2009.10.034
Fateh AH, Mohamed Z, Chik Z et al (2019) Mutagenicity and genotoxicity effects of Verbena officinalis leaves extract in Sprague–Dawley rats. J Ethnopharmacol 235:88–99. https://doi.org/10.1016/j.jep.2019.02.007
Gollapudi BB, Krishna G (2000) Practical aspects of mutagenicity testing strategy: an industrial perspective. Mutat Res 455:21–28. https://doi.org/10.1016/s0027-5107(00)00114-7
Walmsley RM, Billinton N (2011) How accurate is in vitro prediction of carcinogenicity? Genotoxicity testing. Br J Pharmacol 162:1250–1258. https://doi.org/10.1111/j.1476-5381.2010.01131.x
Verschaeve L (2015) Genotoxicity and antigenotoxicity studies of traditional medicinal plants: How informative and accurate are the results? Nat Prod Commun 10:1934578X1501000. https://doi.org/10.1177/1934578x1501000843
Eren Y, Özata A (2014) Determination of mutagenic and cytotoxic effects of Limonium globuliferum aqueous extracts by Allium, Ames, and MTT tests. Rev Bras Farmacogn 24:51–59. https://doi.org/10.1590/0102-695x20142413322
da Silva Dantas FG, de Castilho PF, de Almeida-Apolonio AA et al (2020) Mutagenic potential of medicinal plants evaluated by the Ames Salmonella/microsome assay: a systematic review. Mutat Res Rev Mutat Res 786:108338. https://doi.org/10.1016/j.mrrev.2020.108338
Fei C, Zhang J, Lin Y et al (2015) Safety evaluation of a triazine compound nitromezuril by assessing bacterial reverse mutation, sperm abnormalities, micronucleus and chromosomal aberration. Regul Toxicol Pharmacol 71:585–589. https://doi.org/10.1016/j.yrtph.2015.01.011
Kirkland D, Reeve L, Gatehouse D, Vanparys P (2011) A core in vitro genotoxicity battery comprising the Ames test plus the in vitro micronucleus test is sufficient to detect rodent carcinogens and in vivo genotoxins. Mutat Res Genet Toxicol Environ Mutagen 721:27–73. https://doi.org/10.1016/j.mrgentox.2010.12.015
Langie SAS, Azqueta A, Collins AR (2015) The comet assay: past, present, and future. Front Genet 6:266. https://doi.org/10.3389/fgene.2015.00266
Wu M-F, Peng F-C, Chen Y-L et al (2011) Evaluation of genotoxicity of Antrodia cinnamomea in the Ames test and the in vitro chromosomal aberration test. In Vivo 25:419–423
Funding
This study was supported by the Korea Institute of Oriental Medicine (KIOM, Grant No. KSN1515293), and the National Research Foundation of Korea (NRF) grant (NRF-2020R1A2C2012917), funded by the Ministry of Science and ICT (MSIT), Republic of Korea.
Author information
Authors and Affiliations
Contributions
S-JJ: conceptualization, supervision, writing, review, and editing. ES: methodology, data curation, writing, original draft preparation, and investigation. B-YK: methodology, data curation, writing, and investigation. YJK: investigation and validation. All authors have reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Ethics approval
All experimental procedures were approved by the Institutional Animal Care and Use Committee of the Korea Testing and Research Institute of the Testing and Technical Consulting Institute (Approval No. TGK-2019-000268). The study was designed in compliance with the guidelines published by the Organization of Economic Cooperation and Development (OECD) as the Good Laboratory Practice Regulations for Nonclinical Laboratory Studies of the Korea Food and Drug Administration.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sohn, E., Kim, BY., Kim, Y.J. et al. Non-clinical safety assessment of Annona atemoya leaf extract: evaluation of genotoxicity. Toxicol Res. 40, 473–485 (2024). https://doi.org/10.1007/s43188-024-00241-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43188-024-00241-4