Abstract
Screening of wheat genotypes containing suitable candidate genes for salt tolerance is an essential step in stress-breeding. This study aimed to determine genetic variation, and screen salt-tolerant genotypes using morphological and molecular markers as selection criteria. In this study, 44 genotypes of wheat were screened at the seedling stage by hydroponic culture. The treatments included a control (1 dSm−1), and low (9 dSm−1), moderate (12 dSm−1) and high (15 dSm−1) salinity stresses. Moreover, these genotypes were subjected to marker-assisted selection using 15 salt tolerance related gene-linked simple sequence repeats (SSRs). The total salt stress response indices (TSSRI) categorized wheat genotypes into four groups; tolerant, moderately tolerant, moderately susceptible, and susceptible. TSSRI and principal component analysis (PCA) revealed that Akbar was the most salt-tolerant wheat genotype, followed by Barigom-20, Barigom-22, BW-1284, BW-1262, BW-1237, and Barigom-24. It was evident that the growth of most of the genotypes was significantly inhibited by salt treatment. The tolerant genotypes, however, had to face a lower reduction in morphological parameters. The highest value (0.7127) of polymorphism information content (PIC) was shown by the marker Xwmc17. UPGMA dendrogram showed a clear differentiation of 43 wheat genotypes into seven major clusters considering genetic similarity. Cluster-I consisted of twelve genotypes, seven of which were found salt-tolerant, cluster-VI was composed of seven genotypes, of which four emerged as salt-tolerant, and Cluster-VII accommodated three genotypes all of which were tolerant based on TSSRI. On the contrary, seven genotypes in Cluster-II, and five in Cluster-III were found salt-susceptible according to phenotypic assessment at the seedling stage. In conclusion, considering the morphological characterization and molecular markers response, Akbar, Barigom-20, Barigom-22, BW-1284, and BW-1262 are identified as tolerant genotypes. Therefore, these screened genotypes could be utilized as valuable genetic resources in wheat breeding for salinity tolerance research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat (Triticum aestivum L.) is a staple cereal crop of the Poaceae family (Shewry 2009), which is utilized as food for about 35% world population (Hussain et al. 2015) and cultivated in Bangladesh as the second most economically valuable food crop. The population in the globe is predicted to be extended by 9.6 billion in the next 30 years (DESA 2015), therefore, the world's food supply needs to be enhanced by at least 70% by this time to ensure food security. Bangladesh has been able to produce only 1.0 million tons (Mt) of wheat from 0.40 million hectares (Mha) of land against the national demand of 3.0–3.5 Mt annually (BARI 2010). Moreover, global warming-induced climate change has increased the saline-prone regimes in Bangladesh and severely affected the yield of even the high-yielding wheat cultivars (Siddiqui et al. 2017
Salinity caused an agro-economic impact by affecting vast amounts of arable land and reducing crop growth and productivity (Mbarki et al. 2018; Ahmad et al. 2019). Munns and Tester (2008) and Osman et al. (2020) previously reported that more than 800 Mha of global soils are seriously affected by salinity. Furthermore, over 45 Mha of irrigated lands are negatively influenced by salt stress-related problems worldwide and approximately 50% of farming land could be salt-affected by 2050 (Shrivastava and Kumar 2015). Hence, soil salinity is a major obstacle to crop cultivation and sustainable agriculture. About 1.51 Mha of agricultural fields in Bangladesh are severely affected by varying degrees of salinity which continuously threatens the present cropping ecosystem (SRDI 2010). These saline-prone coastal areas of Bangladesh could be a potential source of cultivable lands which remain fallow during most of the part of the year. Thus, it is essential to develop salt-tolerant and high-yielding varieties that can maintain agricultural sustainability in satisfying the endless yearning for food (Al-Ashkar et al. 2020).
Response of wheat to salt stress is a multiplex phenomenon that closely involves changes in several morpho-physiological characteristics and different biochemical pathways. Wheat plants are generally salt-sensitive and extremely vulnerable to salinity throughout all developmental stages, especially at seedling stages (Munns 2002; Badridze et al. 2009; Ahmad et al. 2013). Furthermore, salinity stress severely hampers plant growth by creating water deficiency and Na+ ion toxicity (Ashraf and Foolad 2007; Acosta-Motos et al. 2017; Ismail and Horie 2017). The plant tries to prevent the salt-induced adverse effects by antioxidants production, accumulation of organic osmolytes, ion exclusion, and changes in mineral and nutrients uptake (Ismail and Horie 2017; Rahneshan et al. 2018).
Our understanding of morpho-physiological mechanisms of plant response to salt-induced stress to successfully isolate the salt-tolerant cultivars is limited and needs further deepened for the development of salt-tolerant cultivars. Several phenotypic traits have been previously used as selection criteria to determine the most appropriate candidate genotypes for the identification of salt-tolerant varieties (Negrão et al. 2017). Field level screening is extremely difficult and laborious due to soil heterogeneity and environmental factors that may interfere with the normal physiological processes. However, screening through hydroponic culture conditions in the laboratory is considered to be more advantageous than screening in the field (Munns et al. 2006). Many researchers examined large-scale wheat genotypes for salt tolerance in a hydroponic medium at the seedling stage (Shahzad et al. 2012; Ahmad et al. 2013; Hussain et al. 2015; Haque et al. 2020).
Besides morpho-physiological study, molecular marker application is fruitful for different genetic studies, molecular marker-assisted selection, quantitative trait loci (QTL) mapping, assessing genetic variability, and screening of candidate genotypes for stress tolerance. Simple sequence repeat (SSR) markers are the most important for the genomic study and are extensively used because of their significant advantages as they are multi-allelic, highly polymorphic, accurate, informative and impartial. Moreover, SSR markers are highly abundant and locus-specific and speedy, cost-friendly than other molecular markers; redundant and distributed widely across the genome; co-dominant and high-throughput in nature (Powell et al. 1996; Chen et al. 2012; Singh et al. 2018). Therefore, SSR markers are applied widely for diversity analysis for selecting stress-tolerant lines and varieties. SSRs are tools of choice in genetics and molecular breeding applications and biotechnological research.
It has been used in several previous studies and reported to be very informative and highly capable to distinguish between wheat genotypes for salinity tolerance (Shahzad et al. 2012; Ahmad et al. 2013; Elshafei et al. 2019; Al-Ashkar et al. 2020; Haque et al. 2020). Since precise phenotyping is a prerequisite for determining the potential candidate salt-tolerant wheat, a combined approach of phonemics and genomics is essential to ensure the development of salinity-tolerant wheat varieties. The current investigation screened 44 wheat genotypes at seedling stages for salt tolerance by a combination of morpho-physiological traits and SSR marker-based characterization.
Materials and methods
Phenotyping wheat seedlings
There were 44 wheat genotypes used in the current investigation of which 5 genotypes were landraces, 9 genotypes were high yielding varieties and the remaining 30 genotypes were advanced line (Table S1). The current experiment used a randomized complete block design replicated thrice. The wheat genotypes are tested under four different salinity levels (1, 9, 12, and 15 dSm−1). The hydroponic culture conditions were constructed following a protocol developed by IRRI (Gregoria et al. 1997).
The morphological screening was done in glasshouse condition with 30/20ºC day/night temperatures, and relative humidity of 50 ± 10%. For breaking seed dormancy, wheat seeds were incubated in an oven at 50 °C for one day by following the procedure of Tabassum et al. (2021). Then the seeds were kept under distilled water for 1 day, transferred to Petri dishes (100 seeds/petri dish) and incubated at 32 °C temperature for 2 days. The seeds, after germination (10 of each genotype) were placed on a styrofoam sheet floating in plastic trays of 12L (36 × 27 × 34 cm) containing only tap water and kept for 3 days. Afterward, water-soluble Peters fertilizer (N:P:K = 20:20:20, 1 g L−1) and ferrous sulfate (FeSO4.7H2O, 200 mg L−1) were added as a source of nutrients to 6-days old pre-seedlings and kept for 4 days. The pH was checked daily by pH meter (Hanna HI 2211, Nasfalau, Romania) to maintain at 5.1–5.2. Ten-day-old wheat seedlings (at 2/3 leaf stage) were exposed to salinity treatment with the addition of crude salt which was unrefined seashore salt mainly containing NaCl and some trace elements. To detect the salt toxicity symptoms, we used the modified standard evaluation system (SES) of IRRI twice once at 14 days and again at 21 days. (Ali et al. 2014) previously concluded that the growth characteristics of rice at the seedling stage were not significantly affected by a low salinity treatment at 6 dSm−1. We, therefore, used control (1 dSm−1), low (9 dSm−1), moderate (12 dSm−1) and (high) 15 dSm−1 salinity treatments for wheat. The electrical conductivity (EC) was measured using an EC meter (WTW 2FD45C, Weilheim, Germany) and maintained with necessary adjustments. Data on shoot length (SL), root length (RL), shoot fresh weight (SFW), and root fresh weight (RFW) was measured after 21 days of salinization. Moreover, the fresh roots and shoots of each seedling were dried for two days at 80 °C to determine the shoot dry weight (SDW) and root dry weight (RDW).
SSR marker-based genotyping
Twenty-one days old young green leaves of wheat were stored at −20 °C immediately after collection. DNA was extracted from leaf samples following the modified Cetyl Trimethyl Ammonium Bromide (CTAB) method (Zidani et al. 2005). DNA quality was checked by using a Nanodrop spectrophotometer (Nanodrop- ND One UV–vis Spectrophotometer, Thermo Scientific, Waltham, USA). The concentration of DNA was measured using OD values at 260/280 nm and 260/230 nm and diluted to approximately 50 ng μL−1 by adding nuclease-free sterile water. Initially, fifteen SSR markers were used for the polymorphism survey (Moghaieb et al. 2011; Shahzad et al. 2012; Ahmad et al. 2013; Vaja et al. 2016; Singh et al. 2018), however, eleven markers showed clear polymorphism out of fifteen (Table 2; Fig. S1-11). These primers located on chromosomes 1–7 and in locus A, B, and D, and associated with salt tolerance related genes were used for molecular screening of 44 wheat genotypes.
The locus names of primers, sequences, corresponding annealing temperatures, and allele size are enlisted in Table S2. The polymerase chain reaction (PCR) cocktail including 1 μL−1 genomic DNA sample had a total volume of 10 μL per reaction mixture for SSR analysis. PCR cocktail composed of 5 μL GoTaq® Green Master Mix that includes Taq DNA polymerase, MgCl2, dNTPs, dNTPs, and reaction buffers (Promega Corporation, USA), 0.5 μL (10 pM) each of forwarding and reverses primers, 3 μL nuclease-free sterile water. The PCR was performed by preheating at 94 °C for 3 min followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 55–65 °C (each primer has its annealing temperature) for 45 s, elongation at 72 °C for 2 min and final extension at 72 °C for 7 min using Biometra Tone Thermocycler, Göttingen, Germany. The amplified PCR products were separated by vertical electrophoresis accomplished on an 8% polyacrylamide gel in 10 × TBE buffer. The gel was stained with ethidium bromide (1 μg mL−1) for 30 min. The resolved bands were documented using UVP® 97–0165-01 BioDoc-It™ 210 UV Imaging System (Jena, Germany). The amplified fragments’ size was estimated by distinguishing their migration distance compared with the molecular weight of 50 and 100 bp DNA ladders.
Calculations and statistical analysis
Firstly, the recorded value of six morphological parameters was divided by the value of those particular parameters at control (C) conditions and summed up all the estimated values of each parameter to calculate cumulative salt stress response indices (CSSRI). CSSRI at low salinity (LS) level was calculated as:
\(CSSRI\left( {LS} \right) \, = \frac{SL(LS)}{{SL(C)}} + \frac{RL(LS)}{{RL(C)}} + \frac{ SFW(LS)}{{RFW(C)}} + \frac{ RFW(LS)}{{RFW(C)}} + \frac{SDW(LS)}{{SDW(C)}} + \frac{ RDW (LS)}{{RDW (C)}}\)
CSSRIs at moderate salinity (MS), and high salinity (HS) were calculated following the CSSRI (LS). Finally, the total salt stress response indices (TSSRIs) were calculated by adding the CSSRI values of the respective salinity level.
The principal component analysis (PCA) was performed using R 3.6.3 with ‘FactoMineR’, ‘factoextra’ and ‘ggplot2’ packages. The 'corplot' was used to visualize the correlation coefficient and scatterplot matrix. The molecular weights of amplified fragments were estimated using Alpha-Ease FC 4 software (Alpha Innotech, USA). The allele’s number, major allele frequency, genetic diversity, and the polymorphism information content (PIC) were estimated through the software, POWER MARKER 3.23 software (Liu and Muse, 2005). The marker index (MI) for individual markers is calculated as the value of major allele frequencies multiplied by the value of allele number (Powell et al. 1996). A dendrogram was constructed employing the resultant similarity matrice and using Sequential Agglomerative Hierarchical Nesting (SAHN)-based unweighted pair group method of arithmetic means (UPGMA) following NTSYSPC 2.1 software (Rohlf, 2000).
Results
Phenotypic performance of wheat genotypes at different salinity levels
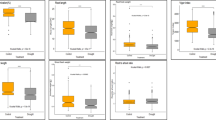
Box and whisker plots were used to display descriptive statistics and natural variation within phenotypic features of wheat seedlings (Fig. 1). The values of all morphological characteristics of seedlings had a significant decrease due to salinity stress and a significant genetic variation was observed among the salinity treatments for all growth-related parameters (Fig. 1).
Box and whisker plots showing the descriptive statistics and natural variation of seedlings’ morphological traits. (A) shoot length, cm, (B) root length, cm, (C) shoot fresh weight, g, (D) root fresh weight, g, (E) Shoot dry weight, g, (F) root dry weight, g under control (C), low salinity (LS), moderate salinity (LS), high salinity (HS) stress conditions. The Whisker, lower limit of the box represents the Q1 (first quartile/25th percentile) which displays the first 25% of data distribution whereas the whisker upper limit of the box represents the Q3 (third quartile/75th percentile) which displays the last 25% of data distribution and the length of the box represents IQR (interquartile range/25th to 75th percentile), which displays 25 to 75% of the data distribution. The horizontal line and red circle within the box indicate the median value for that particular trait. Descriptive line(s) on the boxes indicate a significant difference at P< 0.05 by Games-Howell pairwise test
Salt-treated plants showed a substantial decrease in SL, and RL, however, comparatively a higher decrease in RL were observed than the SL under all salinity treatment conditions (Fig. 1a, b). Contrarily, a greater reduction in SFW was recorded in comparison with RFW (Fig. 1c, d). At the LS level, SL, SFW, and SDW were decreased by 16.34, 53.57, and 30.23%, respectively, over control (Fig. 1a, c, e), whereas 22.68, 31.58, and 33.33% decreased were recorded in RL, RFW, RDW, respectively (Fig. 1). Moreover, all these shoot and root-related parameters followed the same trend in response to MS, and HS stresses (Fig. 1). SL and RL gradually declined under LS, MS, and HS treatment conditions in comparison with the control treatment (Fig. 1a, b).
At MS stress, maximum SL, and RL was recorded in BW-1147, however, SL was drastically reduced for all wheat genotypes at HS condition and ranged between 25.08 (BW-1147) to 13.09 cm (EP-6), and had an average value of 19.36 cm (Table S3; Fig. 1a, b). Moreover, in the HS condition, RL had a sudden decline and varied from 12 (BW-1293) to 3.40 cm (EP-46), giving an average value of 7.06 cm (Table S3; Fig. 1b). The highest values of SFW, RFW, SDW, and RDW were found in the control condition and the values of these traits decreased significantly with the gradual increase of the salinity levels (Fig. 1c–f). At MS and HS stress, the maximum SFW, and RFW were determined in BW-1293 and EP-3, respectively (Table S3; Fig. 1c, d), however, SFW and RFW were reduced significantly in all wheat genotypes, ranging from 0.47 g (BW-1293) and 0.44 g (EP-3) to 0.079 g (Barigom-32) and 0.1 g (EP-44, and EP-48), with an average of 0.22 g and 0.19 g at HS stress (Table S3; Fig. C, D). The mean value of SDW dropped from 0.13 to 0.09 g as the salinity stress changed from LS to MS (Fig. 1f). Moreover, under HS conditions, SDW was significantly decreased for all wheat genotypes and ranged from 0.096 g (BW-1293) to 0.04 g (Barigom-26), with an average of 0.06 g (Table S3; Fig. 1d). The mean value of RDW at MS was recorded at 0.03, however, no significant difference was found in the case of RDW mean value at HS conditions (Fig. 1f).
Correlation analysis among the morphological traits and treatments
The degree of association among the traits was determined by their correlation coefficients. Pearson correlation analysis revealed a significant relationship among observed seedling traits (Fig. 2). Overall, shoot and root-related traits showed positive and significant correlations among all traits at the P<0.001 level. However, at LS and HS treatments, SL exhibited a positively significant correlation with all morphological traits except RFW whereas RL was significantly correlated with SFW, SDW, and RDW under all salinity levels (P<0.001). Moreover, a significant (P<0.001) correlation was found between SFW with RFW, SDW, and RDW under C and HS conditions. Furthermore, RFW was significantly correlated with SDW and RDW at a P<0.001 level in response to C and HS stress. SDW was significantly (P<0.05) correlated with RDW at all four salinity conditions. Interestingly, no significant correlation was found between SL and RFW under all salinity conditions except control, whereas at MS salinity conditions non-signification correlation showed by RL with SFW (Fig. 2).
Scatter plot, a correlation matrix of the studied morphological traits of all studied wheat genotypes grown under control and different salinity stress conditions. In the upper panel, red and blue boxes indicate positive and negative correlations, respectively, with increasing color intensity reflecting a higher coefficient. The diagonal panel indicates the distribution histogram of correlated traits. The lower panel indicates a scatter plot and trend-line of the correlated traits. *, ** and *** indicate significance at P<0.05, P<0.01 and P<0.001. Shoot length (SL), root length (RL), shoot fresh weight (SFW), root fresh weight (RFW), shoot dry weight (SDW), and root dry weight (RDW)
TSSRI and PCA-based screening for salt-tolerance
All the examined genotypes were classified into four response groups following TSSRI in which 11 genotypes were identified as susceptible, 12 genotypes were found as moderately susceptible, 11 genotypes appeared to be moderately tolerant, and 10 genotypes were considered tolerant (Table 1).
TSSRI values ranged from 6.85 for genotype Barigom-30 which was identified as the most salt-susceptible to 17.55 for Akbar, the most salt-tolerant genotype. The ten genotypes belonging to the tolerant group are Akbar, Pavon-76, EP-11, Barigom-20, Barigom-22, BW-1284, EP-37, BW-1262, BW-1237, and Barigom-24. Besides, the moderately tolerant eleven genotypes are EP-46, Triticale, EP-13, EP-3, EP-6, EP-31, EP-33, EP-15, EP-27, EP-35, and EP-44. On the contrary, the moderately susceptible genotypes are EP-17, EP-29, Sonalika, EP-20, BW-1295, Binagom-1, EP-41, EP-39, BW-1194, EP-22, Barigom-26, and BW-1274. Finally, EP-48, BW-1147, Barigom-25, EP-25, Barigom-28, BW-1278, Kanchan, EP-9, EP-32, BW-1293, and Barigom-30 genotypes are included in the susceptible group.
The PCA was used to discriminate the potential salt-tolerant genotypes by estimating the principal components of recorded morphological characters in all the salt-exposed genotypes (Fig. 3). The distinguished morphological loadings in principal component one (PC1) and principal component two (PC2) are compared to investigate the contributors of the principal components. The PCA individual plot was used to dissect all genotypes into four response categories: tolerant, moderately tolerant, moderately susceptible, and susceptible based on the distribution of the genotypes into two principal components, PC1 and PC2 that defined the majority of data variability (Fig. 3).
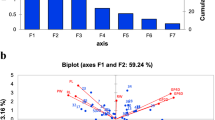
Principal component analysis (PCA) for the first two principal components (PC1 and PC2) describing the grouping of 44 wheat genotypes into four different responses based on measured phenotypic traits after 21 days of treatments at (A) low salinity, (B) moderate salinity, (C) high salinity stress. Tolerant (T), moderate tolerant (MT), moderate susceptible (MS), susceptible (S)
Under LS level, the PC1 and PC2 accounted for 16.1 and 67% of data variability, respectively as they comprised 83.1% of data variability (Fig. 3a). Here, nine genotypes, Barigom-24, Barigom-22, Triticale, BW-1147, Barigom-20, Akbar, Barigom-25, BW-1237, and BW-1194 are considered salt-tolerant; eight genotypes as moderate salt-tolerant; fourteen genotypes appeared as moderate salt-susceptible; thirteen genotypes as salt-susceptible, respectively (Fig. 3a). At the MS level, the PC1 and PC2 represented 58.9% (PC1) and 14.4% (PC2) and collectively 73.3% data variation (Fig. 3b). At this stress condition, eight genotypes, EP-3, EP-6, BW-1293, Barigom-24, Triticale, Akbar, BW-1295, and BW-1284 clustered together as salt-tolerant; eleven genotypes as moderate salt-tolerant; eleven genotypes as moderate susceptible; fourteen genotypes as salt-susceptible (Fig. 3b).
However, at HS conditions, cumulatively 77.2% of data variability was accounted for by PC1 and PC2 (Fig. 3c). Here, six genotypes, EP-3, BW-1262, Akbar, BW-1274, Triticale, and BW-1278 are separated as salt-tolerant, thirteen genotypes as moderate salt-tolerant, nine genotypes as moderate salt-susceptible, and sixteen genotypes as salt-susceptible (Fig. 3c). As assumed, the discrimination by PCA revealed a decrease in the number of tolerant genotypes with the increase of salinity levels from LS to MS and HS (Fig. 3a–c). At the LS level, the genotypes e.g., Barigom-24, Barigom-22, Triticale, Barigom-20, Akbar, and BW-1237 emerged as salt-tolerant considering both PCA and TSSRI results together (Fig. 3a; Table 1). At the MS level, the results of PCA and TSSRI indicated that EP-3, EP-6, Barigom-24, Triticale, Akbar, and BW-1284 appeared as salt-tolerant. Considering PCA and TSSRI results together, the genotypes EP-3, BW-1262, Akbar, and Triticale appeared to have the highest salt tolerance at the HS level (Fig. 3c; Table 1). Overall results of PCA and TSSRI revealed that Akbar, Barigom-20, Barigom-22, Barigom-24, BW-1237, BW-1262, and BW-1284 were identified to be the salt-tolerant genotypes. Morphological findings also supplemented that the genotypes identified as tolerant faced the least degrees of reduction in root and shoot growth parameters when imposed with salinity stress under a hydroponic culture (Table 1; Fig. 3). Barigom-20 and BW-1262, among the identified tolerant genotypes (Table 1; Fig. 3), had higher values of all phenotypic parameters; Akbar and Barigom-24 had higher values of all measured morphological traits except SL; and Barigom-22 and BW-1284 had higher values of SL, RL, SFW, and SFW at LS stress in comparison with control. At MS conditions, Akbar, Barigom-20, Barigom-24, BW-1237, and BW-1284 showed greater RL as compared to the control (Table S3). Furthermore, at HS stress, Barigom-20, BW-1237, and BW-1284 showed higher SL as compared to control conditions whereas Barigom-22 and BW-1262 had greater RFW (Table S3).
Molecular screening for salt-tolerant wheat genotypes
Genetic diversity of SSR molecular markers
The genotypic data were evaluated to observe the genetic diversity among the studied wheat genotypes and identify the potential salt-tolerant candidate genotypes. The proper polymorphic amplicons generated by 11 SSR markers (among 15 SSR markers) showed differences among the tested genotypes. The marker, Xgwm455, amplified the amplicons of the lowest size (120 bp), and the highest amplicon size(458 bp) was found with the marker, Xgmc661. Alleles produced by eleven tested SSRs efficiently produced consistent, well-resolved, and reproducible alleles that had very clear allelic patterns in 44 studied wheat genotypes (Fig. S1-11). A total of 45 alleles were produced by eleven polymorphic SSR markers and the number of microsatellite alleles ranged from 2.00 for Xbarc 45 to 7.00 for Xgwm296 where the average alleles per locus were found at 4.09 (Table 2). The investigated primers produced an average of 1.27 unusual alleles, while the tested primers produced an average of 1.09 null alleles (Table 2).
The major allelic frequencies of the eleven marker loci were found to range from 0.2955 for Xwmc17 to 0.8409 for Xbarc45 (Table 2). The highest (0.7562) gene diversity, among the loci, was found in locus Xwmc17 and the lowest (0.2676) was in loci Xbarc45 having a mean diversity of 0.5604. It appeared that markers that detected the lower number of alleles had concomitant lower gene diversity than the markers detecting the more alleles revealing higher gene diversity. The PIC was estimated for measuring genetic diversity. The loci polymorphism was considered high at the PIC value above 0.5 (Vaiman et al. 1994). PIC values of the markers in this study ranged from 0.2318 to 0.7127 giving an average of 0.4967 (Table 2). Xwmc17 gave the highest PIC, and the lowest PIC value was found in Xbarc45 (Table 2). The PIC value of 5 markers (Xgwm296, Xwmc44, Xgwm249, Xwmc17, and Xwmc149) was above 0.5 (Table 2). The examined markers displayed an average relatively high PIC value. This fact proves that the markers used were highly polymorphic. The MI was highest for the marker Xgwm296 and Xwmc149 (3.18), and lowest for the marker Xwmc17 (1.48). The higher values of PIC and MI together suggest that the primers applied in this study are efficient in distinguishing the genotypes for salt tolerance.
UPGMA-based genetic similarity and cluster analysis
Nei’s (1973) genetic distance-based UPGMA dendrogram showed a clear differentiation among forty-three wheat genotypes and grouped the genotypes into seven major clusters (Cluster-I, -II, -III, -IV, -V, -VI, and -VII) and a single isolated genotype (Fig. 4).
Cluster-I consisted of twelve genotypes, seven (Akbar, Barigom-20, Barigom-22, BW-1284, EP-6, EP-15, and EP-33) of which were found salt-tolerant in the morphological study based on TSSRI (Fig. 4; Table 1). Similarly, cluster-IV included two genotypes and cluster-VII accommodated three genotypes of which Barigom-24, Pavon-76, BW-1237, and EP-3 are salt-tolerant based on TSSRI (Fig. 4; Table 1). Moreover, cluster-VI was composed of seven genotypes of which four (EP-11, EP-35, EP-37, and EP-44) emerged as salt-tolerant by TSSRI (Fig. 4; Table 1). On the contrary, out of eight genotypes in both Cluster-II and cluster-III, seven were determined to be salt-susceptible in Cluster-II (Barigom-25, Barigom-28, Barigom-30, Sonalika, Binagom-1, BW-1194, and BW-1293) and five in Cluster-III (EP-17, EP-22, EP-25, EP-29, and EP-39) according to phenotypic assessment at the seedling stage (Fig. 4; Table 1). Moreover, three genotypes were grouped under cluster-IV of which Kanchan and EP-9 emerged as salt-susceptible (Fig. 4; Table 1). Interestingly, the dendrogram revealed with few exceptions that the released varieties and advanced inbred lines of BARI are grouped in clusters-I, -II, -IV, and -VII (Fig. 4) while the accessions obtained from CIMMYT are grouped in cluster-II, -V, and -VI (Fig. 4). Again, the tolerant genotypes grouped in clusters -I, -IV, -VI and -VII while the susceptible genotypes were in the rest of the clusters.
Discussion
Salinity inhibits the plants’ cell-cycle machinery under salt-stressed conditions, and consequences to the cellular dysfunction in cell division and differentiation finally retards the plant growth, including wheat (De Veylder et al. 2007; Al-Ashkar et al. 2020). Even though numerous earlier findings on wheat salinity tolerance have been published around the world, studies in Bangladesh are rarely mentioned. Thus, it is critical to identify genetic variation among genotypes and to identify promising salt-tolerant genotypes that can be used as breeding candidates. The current is aimed to discern wheat genotypes that are potential salt-tolerant and their genetic diversity analysis based on morphological traits and molecular marker characterization.
Screening for salt-tolerance in wheat genotypes
A significant reduction of all tested morphological parameters of what seedling was observed under different salinity conditions (Fig. 1). The damage to the wheat seedling’s growth might be due to the excess accumulation of Na+ at different plant parts that might have created a disruption of water absorption, oxidative injury, and mineral imbalance (Castillo et al. 2007; Mekawy et al. 2015; Zhang et al. 2018).
This reduction of morphological parameters had an increase with the increment of salinity stress as the LS stress reduced the morphological characteristics of wheat seedlings at the early stage to a lesser degree than MS and HS stresses (Fig. 1). Similar findings were reported in some crop plants such as rice (Oryza sativa) (Ahmed et al. 2019; Tabassum et al. 2021), wheat (Triticum aestivum) (Ahmad et al. 2013), maize (Zea mays) (Masuda et al. 2021) and barley (Hodeum vulgare) (Allel et al. 2016) under salt stress. Our findings showed that RL was affected to a higher degree by salinity than the SL (Fig. 1a, b; Table S3). It might be because high salinity repressed the growth of wheat roots by suppressing the division and elongation of root cells (Duan et al. 2015). This result is supported by (Nakamura et al. 2021) who earlier observed that roots were more sensitive than shoots to salinity during the seedling stage of wheat plants. It remained unclear if the main target tissues of salt stress lie in roots and/or shoots/leaves. Nevertheless, many findings showed that shoots and leaves are more sensitive than roots (Munns and Tester 2008; Sharma 2015; Tao et al. 2021), higher sensitivity of roots than shoots (Bilkis et al. 2016; Soares et al. 2018; Bacu et al. 2020) or both the roots and shoots are equally sensitive in wheat (Hussain et al. 2015; Hasan et al. 2017).
Most genotypes' root and shoot growth-related parameters were slightly lower and/or sometimes higher under LS stress than those of the control condition (Table S3). These findings indicate that the studied genotypes were affected marginally at 9 dSm−1 salinity level and may be capable of reducing the salt-induced toxicity in this concentration. Moreover, this observation suggests that the tested wheat plants are tolerant to 9 dSm−1 salt stress at the seedling stage which was strongly supported by SESbased visual scoring as they score 3 in a 1–9 scoring system. A 3 score means the seedlings showed almost normal growth with few leaf tips or leaves whitish and rolled). Although wheat is a moderately tolerant crop (Maas and Hoffman 1977), its yield could be significantly reduced at 10 dSm−1 saline soil conditions, however, the rice plants could die in the same salinity treatment conditions (Munns et al. 2006, 2008).
The correlation studies can define the nature and degree of association between any pairs of morphological traits grouped by treatments. All the morphological traits used in this investigation showed highly significant correlations among them (Fig. 2), indicating that a change in any one of the used traits significantly changed the other traits. These findings revealed that phenotypic features played a significant role in determining the salt stress response under salinity conditions. Therefore, these seedling traits can be used as a selection criterion for identifying salt-tolerant wheat genotypes. These findings are supported by many previous reports described by (Shahzad et al. 2012; Ahmad et al. 2013; Haque et al. 2020) as they found similar kinds of correlation among the tested genotypes under salinity stress.
Different salt tolerance indices were previously used to discriminate between tolerant and sensitive wheat genotypes (Genc et al. 2007; Shahzad et al. 2012). We used TSSRI results and revealed that Akbar, Pavon-76, EP-11, Barigom-20, Barigom-22, BW-1284, BW-1262, BW-1237, and Barigom-24 maintained relatively higher values in all studied root and shoot traits under salinity conditions (Table 1; Table S3). The maintenance of higher root and shoot traits under salinity conditions is probably due to the ability of these genotypes to cope with salt stress and show tolerance by the adoption of some morpho-physiological mechanisms (Islam et al. 2009). PCA was previously used to categorize rice (Rasel et al. 2020a), canola (Singh et al. 2008), and corn (Wijewardana et al. 2016) for salinity tolerance. It can be used to group the genotypes based on the similarity and dissimilarity in their responses (Kim et al., 2013; Tuhina-Khatun et al. 2015). PCA can explain and indicate the trait-related crucial components responsible for salt tolerance between genotypes and treatments (Tabassum et al. 2021). PCA findings showed that with the increase in salt stress level the morphological parameters of wheat seedlings were affected resulting in a lesser number of tolerant genotypes at MS and HS (Fig. 1, 3). The results obtained from TSSRI are strongly supported by PCA findings, as the tolerant and susceptible genotypes were separated, and potential salt-tolerant genotypes could be successfully detected (Table 1; Fig. 3). Kakar et al. (2019) conducted an experiment on rice where they proved that the results TSSRI results agree with PCA method based findings at seedling stage phenotyping for salt tolerance. The findings of phenotypic observation suggested that the tolerant genotypes may adapt some tolerance mechanism to improve root and shoot morphology, which can help the wheat plant survive in saline conditions. According to Kakar et al. (2019), increased root biomass in rice (e.g., RFW and RDW) contributes to vigorous shoot growth in salt-tolerant genotypes, possibly through salt exclusion, controlling the excess amount of toxic Na+ uptake in the shoots (Negro et al. 2017), and thus lessening salinity injury symptoms in leaves.
Molecular screening for salt-tolerant wheat genotypes
SSR marker-based investigation was performed to confirm the presence of desirable candidate genes in the selected candidates. The molecular study showed eleven SSR markers and 44 alleles in the 44 genotypes. The higher (4.09) average alleles per locus indicate a greater genetic diversity among the studied genotypes (Ram et al. 2007). Ali et al. (2014) detected 58 alleles with ten SSR primers among 33 rice genotypes and (Ram et al. 2007) previously reported 4.86 alleles per locus in 33 rice genotypes by performing 25 SSR markers, which was similar to the result of the current investigation.
The examined primers produced an average 1.09 null allele, which failed to amplify during PCR, most likely due to polymorphism at one or both of the primers' hybridization sites (Dakin and Avise 2004). Genetic diversity is valuable as raw material in breeding for genetic improvement, increasing the yield potential and improving desired traits related to environmental stress tolerance (Gepts 2006). The genetic diversity was measured in the term of PIC. The PIC is extensively used as the measurement of each marker’s usefulness to distinguish one individual from another It is considered that the loci polymorphism is high, medium or low when PIC values are greater than 0.5, between 0.5 and 0.25, and less than 0.25, respectively (Vaiman et al. 1994). The PIC values of the investigated markers varied from 0.2318 to 0.7127, with a mean value of 0.4967. The mean PIC values observed in the current experiment are near 0.5 and half of the investigated markers showed the PIC values of more than 0.05. It was evident that the SSR markers we used were highly polymorphic. The PIC value of the present molecular study is higher than the PIC values described in the investigations of (Vignesh et al. 2021) and lower than the observation of (Sardouie-Nasab et al. 2013). Though it can be argued that the use of abundant SSR markers could be more effective to characterize and describe genotypes, our results revealed that the use of a lesser number of markers would be also efficient for identifying salt-tolerant genotypes.
The markers used in this study were salt tolerance-related SSRs. Hence, clustering based on genetic similarity might indicate the genetic potentiality of the selected genotypes for salt tolerance. Finally, the UPGMA analysis by Pairwise similarity coefficient values of SSR marker data classified 44 tested wheat genotypes into seven major clusters and one isolated single genotype considering their genetic similarity (Fig. 4).
Our molecular data are supported by the total salinity tolerance index (TSSRI) results to determine the candidate salt-tolerant and susceptible genotypes. Exceptionally, however, some tolerant genotypes were grouped into tolerant clusters and some susceptible genotypes were also found in the tolerant groups. These few exceptions might be because the markers we used might not be enough to cover the genomic regions where the salt-tolerant genes are probably located to be able to explore the potential salt-tolerant genotypes (Seetharam et al. 2009; Singh et al. 2018; Haque et al. 2020). Furthermore, it may be attributable to the molecular diversity not reflecting the diversity at the morphological level (Ali et al. 2014). Many previous studies have reported SSR marker-based clustering for the description of genetic diversity and characterization based on genetic similarity in crop plants including wheat under salt stress (Ali et al. 2014; Rubel et al. 2014; Kordrostami et al. 2017; Elshafei et al. 2019; Al-Ashkar et al. 2020; Huong et al. 2020; Rasel et al. 2020b).
In conclusion, we evaluate the overall growth and genetic diversity in wheat seedlings under different salinity treatments by assessing morphological parameters and SSR markers’ responses. The morphological findings showed that salinity interfered with the growth and growth contributing traits and caused severe damage to the growth of the seedlings. According to SSRI and PCA, Akbar was the most salt-tolerant genotype followed by Barigom-20, Barigom-22, BW-1284, BW-1262, BW-1237, and Barigom-24. These results evidence that the tolerant genotypes showed the lowest reduction in growth attributing traits. The highest PIC value was observed by the marker, Xwmc17 (0.7127). SSR marker-based characterization exhibited Akbar, Barigom-20, Barigom-22, BW-1284, EP-6, EP-15, and EP-33, Barigom-24, Pavon-76, BW-1237, EP-3, and BW-1262 as salt-tolerant genotypes taking into consideration the genetic similarity in the dendrogram and TSSRI results. These screened salt-tolerant wheat genotypes could be further utilized in wheat breeding to develop salt-tolerant varieties.
Data availability
The data of the current investigation are available from the corresponding author upon reasonable request.
References
Acosta-Motos JR, Ortuño MF, Bernal-Vicente A, Diaz-Vivancos P, Sanchez-Blanco MJ, Hernandez JA (2017) Plant responses to salt stress: Adaptive mechanisms. Agron 7:18. https://doi.org/10.3390/agronomy7010018
Ahmad M, Shahzad A, Iqbal M, Asif M, Hirani AH (2013) Morphological and molecular genetic variation in wheat for salinity tolerance at germination and early seedling stage. Aust J Crop Sci 7:66–74. https://doi.org/10.3316/informit.142656583967655
Ahmad R, Hussain S, Anjum MA, Khalid MF, Saqib M, Zakir I, Hassan A, Fahad S, Ahmad S (2019) Oxidative stress and antioxidant defense mechanisms in plants under salt stress. In Plant Abiotic Stress Tolerance, pp. 191–205. Springer. https://doi.org/10.1007/978-3-030-06118-0_8
Ahmed S, Anik TR, Islam A, Uddin I Haque MS (2019) Screening of Some Rice (Oryza sativa L.) Genotypes for Salinity Tolerance using Morphological and Molecular Markers. Biosci Biotechnol Res Asia, 16: 377–390. https://doi.org/10.13005/bbra/2753
Al-Ashkar I, Alderfasi A, Ben Romdhane W, Seleiman MF, El-Said RA, Al-Doss A (2020) Morphological and genetic diversity within salt tolerance detection in eighteen wheat genotypes. Plants 9:287. https://doi.org/10.3390/plants9030287
Ali MN, Yeasmin L, Gantait S, Goswami R, Chakraborty S (2014) Screening of rice landraces for salinity tolerance at seedling stage through morphological and molecular markers. Physiol Mol Bio Plants 20:411–423. https://doi.org/10.1007/s12298-014-0250-6
Allel D, Ben-Amar A, Badri M, Abdelly C (2016) Salt tolerance in barley originating from harsh environment of North Africa. Aust J Crop Sci 10:438–451. https://search.informit.org/doi/https://doi.org/10.3316/informit.135292448531387
Ashraf M, Foolad MR (2007) Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ Exp Bot 59:206–216. https://doi.org/10.1016/j.envexpbot.2005.12.006
Bacu A, Ibro V, Nushi M (2020) Compared salt tolerance of five local wheat (Triticum aestivum L.) cultivars of Albania based on morphology, pigment synthesis and glutathione content. E EuroBiotech J 4:42–52. https://doi.org/10.2478/ebtj-2020-0006
Badridze G, Weidner A, Asch F, Börner A (2009) Variation in salt tolerance within a Georgian wheat germplasm collection. Genet Resour Crop Evol 56:1125–1130. https://doi.org/10.1007/s10722-009-9436-0
BARI (2010) BARI, 2010 Wheat Production in Bangladesh: A success story. Retrieved from http://www.bari.gov.bd/index.php? (No. 2). Article 2.
Bilkis A, Islam M, Hafiz M, Hasan M (2016) Effect of NaCl induced salinity on some physiological and agronomic traits of wheat. Pak J Bot 48:455–460. http://www.pakbs.org/pjbot/PDFs/48(2)/05.pdf
Castillo EG, Tuong TP, Ismail AM, Inubushi K (2007) Response to salinity in rice: Comparative effects of osmotic and ionic stresses. Plant Prod Sci 10:159–170. https://doi.org/10.1626/pps.10.159
Chen X, Min D, Yasir A, Hu YG (2012) Genetic diversity, population structure and linkage disequilibrium in elite Chinese winter wheat investigated with SSR markers. PLoS ONE 7:e44510. https://doi.org/10.1371/journal.pone.0044510
Dakin E, Avise J (2004) Microsatellite null alleles in parentage analysis. Heredity 93:504–509. https://doi.org/10.1038/sj.hdy.6800545
De Veylder L, Beeckman T, Inzé D (2007) The ins and outs of the plant cell cycle. Nature Rev Mol Cell Biol 8:655–665. https://doi.org/10.1038/nrm2227
DESA U (2015) World population prospects: The 2015 revision, key findings and advance tables. United Nations Department of Economic and Social Affairs. Population Division Working Paper No. ESA/P/WP, 241. https://population.un.org/wpp/publications/files/key_findings_wpp_2015.pdf
Duan L, Sebastian J, Dinneny JR (2015) Salt-stress regulation of root system growth and architecture in Arabidopsis seedlings. Plant Cell Expansion. https://doi.org/10.1007/978-1-4939-1902-4_10
Elshafei AA, Afiah SAE-A, AlDoss AA, Ibrahim EI (2019) Morphological variability and genetic diversity of wheat genotypes grown on saline soil and identification of new promising molecular markers associated with salinity tolerance. J Plant Interactions 14:564–571. https://doi.org/10.1080/17429145.2019.1672815
Genc Y, Mcdonald GK, Tester M (2007) Reassessment of tissue Na+ concentration as a criterion for salinity tolerance in bread wheat. Plant Cell Environ 30:1486–1498. https://doi.org/10.1111/j.1365-3040.2007.01726.x
Gepts P (2006) Plant genetic resources conservation and utilization: The accomplishments and future of a societal insurance policy. Crop Sci 46:2278–2292. https://doi.org/10.2135/cropsci2006.03.0169gas
Gregoria, G. B, Senadhira, D, Mendoza, RD (1997) Screening rice for salinity tolerance. https://EconPapers.repec.org/RePEc:ags:irridp:287589
Hannan MA, Saha NR, Roy SK, Woo S-H, Haque MS (2021). Genetic Diversity Analysis and Molecular Screening for Salinity Tolerance in Wheat Germplasm. https://doi.org/10.9787/pbb.2021.9.3.185
Haque MS, Saha NR, Islam MT, Islam MM, Kwon S-J, Roy SK, Woo S-H (2020) Screening for drought tolerance in wheat genotypes by morphological and SSR markers. J Crop Sci Biotechnol. https://doi.org/10.1007/s12892-020-00036-7
Hasan M, Baque M, Habib M, Yeasmin M, Hakim M (2017) Screening of salt tolerance capability of wheat genotypes under salt stress condition. Universal Journal of Agricultural Research, 5:235–249. https://doi.org/10.13189/ujar.2017.050405
Huong CT, Anh TTT, Tran H-D, Duong VX, Trung NT, Dang Khanh T, Dang Xuan T (2020) Assessing Salinity Tolerance in Rice Mutants by Phenotypic Evaluation Alongside Simple Sequence Repeat Analysis. Agric 10:191. https://doi.org/10.3390/agriculture10060191
Hussain B, Khan AS, Ali Z (2015) Genetic variation in wheat germplasm for salinity tolerance atseedling stage: Improved statistical inference. Turkish J Agric Forestry 39:182–192. https://doi.org/10.3906/tar-1404-114
Islam MM, Hoque MA, Okuma E, Banu MNA, Shimoishi Y, Nakamura Y, Murata Y (2009) Exogenous proline and glycinebetaine increase antioxidant enzyme activities and confer tolerance to cadmium stress in cultured tobacco cells. J Plant Physiol 166:1587–1597. https://doi.org/10.1016/j.jplph.2009.04.002
Ismail AM, Horie T (2017) Genomics, physiology, and molecular breeding approaches for improving salt tolerance. Annual Review Plant Biol 68:405–434. https://doi.org/10.1146/annurev-arplant-042916-040936
Kim JK, Park S-Y, Lim S-H, Yeo Y, Cho HS, Ha S-H (2013) Comparative metabolic profiling of pigmented rice (Oryza sativa L.) cultivars reveals primary metabolites are correlated with secondary metabolites. J Cereal Sci 57:14–20. https://doi.org/10.1016/j.jcs.2012.09.012
Kordrostami M, Rabiei B, Kumleh HH (2017) Biochemical, physiological and molecular evaluation of rice cultivars differing in salt tolerance at the seedling stage. Physiol Mol Biol Plants 23:529–544. https://doi.org/10.1007/s12298-017-0440-0
Liu K, Muse SV (2005) PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 21:21282129. https://doi.org/10.1093/bioinformatics/bti282
Maas EV, Hoffman GJ (1977) Crop salt tolerance—Current assessment. J Irrig Drain Div 103:115–134. https://doi.org/10.1061/jrcea4.0001137
Masuda MS, Azad MAK, Hasanuzzaman M, Arifuzzaman M (2021) Evaluation of salt tolerance in maize (Zea mays L.) at seedling stage through morphological characters and salt tolerance index. Plant Physiol Reports. https://doi.org/10.1007/s40502-021-00611-2
Mbarki S, Sytar O, Cerda A, Zivcak M, Rastogi A, He X, Zoghlami A, Abdelly C, Brestic M (2018) Strategies to mitigate the salt stress effects on photosynthetic apparatus and productivity of crop plants. In Salinity Responses and Tolerance in Plants 1:85–136, Springer. https://doi.org/10.1007/978-3-319-75671-4_4
Mekawy AMM, Assaha DV, Yahagi H, Tada Y, Ueda A, Saneoka H (2015) Growth, physiological adaptation, and gene expression analysis of two Egyptian rice cultivars under salt stress. Plant Physiol Biochem 87:17–25. https://doi.org/10.1016/j.plaphy.2014.12.007
Moghaieb RE, Abdel-Hadi A-HA, Talaat NB (2011) Molecular markers associated with salt tolerance in Egyptian wheats. African J Biotechnol 10:18092–18103. https://doi.org/10.5897/ajb11.2576
Munns R (2002) Comparative physiology of salt and water stress. Plant Cell Environ 25:239–250. https://doi.org/10.1046/j.0016-8025.2001.00808.x
Munns R, James RA, Läuchli A (2006) Approaches to increasing the salt tolerance of wheat and other cereals. J Exp Bot 57:1025–1043. https://doi.org/10.1093/jxb/erj100
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. https://doi.org/10.1146/1nnurev.arplant.59.032607.092911
Nakamura C, Takenaka S, Nitta M, Yamamoto M, Kawazoe T, Ono S, Takenaka M, Inoue K, Takenaka S, Kawai S (2021) High sensitivity of roots to salt stress as revealed by novel tip bioassay in wheat seedlings. Biotech Biotechnol Equip 35:246–254. https://doi.org/10.1080/13102818.2020.1852890
Negrão S, Schmöckel S, Tester M (2017) Evaluating physiological responses of plants to salinity stress. Ann Bot 119:1–11. https://doi.org/10.1093/aob/mcw191
Osman MS, Badawy AA, Osman AI, Latef AAHA (2020) Ameliorative impact of an extract of the halophyte Arthrocnemum macrostachyum on growth and biochemical parameters of soybean under salinity stress. J Plant Growth Regul. https://doi.org/10.1007/s00344-020-10185-2
Powell W, Machray GC, Provan J (1996) Polymorphism revealed by simple sequence repeats. Trends in Plant Sci 1:215–222. https://doi.org/10.1016/s1360-1385(96)86898-0
Rahneshan Z, Nasibi F, Moghadam AA (2018) Effects of salinity stress on some growth, physiological, biochemical parameters and nutrients in two pistachio (Pistacia vera L.) rootstocks. J Plant Interact 13:73–82. https://doi.org/10.1080/17429145.2018.1424355
Ram SG, Thiruvengadam V, Vinod KK (2007) Genetic diversity among cultivars, landraces and wild relatives of rice as revealed by microsatellite markers. J Appl Genet 48:337–345. https://doi.org/10.1007/bf03195230
Rasel M, Tahjib-Ul-Arif M, Hossain MA, Hassan L, Farzana S, Brestic M (2020a) Screening of Salt-Tolerant Rice Landraces by Seedling Stage Phenotyping and Dissecting Biochemical Determinants of Tolerance Mechanism. J Plant Growth Regul. https://doi.org/10.1007/s00344-020-10235-9
Rasel M, Tahjib-Ul-Arif M, Hossain MA, Sayed MA, Hassan L (2020b) Discerning of rice landraces (Oryza sativa L.) for morpho-physiological, antioxidant enzyme activity, and molecular markers’ responses to induced salt stress at the seedling stage. J Plant Growth Regul 39:41–59
Rohlf FJ (2000) NTSYS-pc: Numerical taxonomy and multivariate analysis system, version 2.1 Exeter Software. Setauket, New York, USA. Exeter Publishing. https://doi.org/10.4236/psych.2010.12018
Sardouie-Nasab S, Mohammadi-Nejad G, Zebarjadi A (2013) Haplotype analysis of QTLs attributed to salinity tolerance in wheat (Triticum aestivum). Mol Biol Reports 40:4661–4671. https://doi.org/10.1007/s11033-013-2561-z
Seetharam K, Thirumeni S Paramasivam K (2009) Estimation of genetic diversity in rice (Oryza sativa L.) genotypes using SSR markers and morphological characters. African J Biotechnol 8. https://www.ajol.info/index.php/ajb/article/view/60480
Shahzad A, Ahmad M, Iqbal M, Ahmed I, Ali G (2012) Evaluation of wheat landrace genotypes for salinity tolerance at vegetative stage by using morphological and molecular markers. Genet Mol Res 11:679–692. https://doi.org/10.4238/2012.march.19.2
Sharma R (2015) Genotypic response to salt stress: I–relative tolerance of certain wheat cultivars to salinity. Advances in Crop Science and Technology 3:192. https://doi.org/10.4172/2376-0354.1000158
Shewry PR (2009) Wheat Jexp Bot 60:1537–1553. https://doi.org/10.1093/jxb/erp058
Shrivastava P, Kumar R (2015) Soil salinity: A serious environmental issue and plant growth promoting bacteria as one of the tools for its alleviation. Saudi Journal of Biological Sciences 22(2):123–131. https://doi.org/10.1016/j.sjbs.2014.12.001
Siddiqui MN, Mostofa MG, Akter MM, Srivastava AK, Sayed MA, Hasan MS, Tran L-SP (2017) Impact of salt-induced toxicity on growth and yield-potential of local wheat cultivars: Oxidative stress and ion toxicity are among the major determinants of salt-tolerant capacity. Chemos 187:385–394. https://doi.org/10.1016/j.chemosphere.2017.08.078
Singh AK, Chaurasia S, Kumar S, Singh R, Kumari J, Yadav MC, Singh N, Gaba S, Jacob SR (2018) Identification, analysis and development of salt responsive candidate gene based SSR markers in wheat. BMC Plant Biol 18:1–15. https://doi.org/10.1186/s12870-018-1476-1
Singh S, Kakani V, Brand D, Baldwin B, Reddy K (2008) Assessment of cold and heat tolerance of winter-grown canola (Brassica napus L.) cultivars by pollen-based parameters. J Agron Crop Sci 194:225–236. https://doi.org/10.1111/j.1439-037x.2008.00309.x
Soares AL, Geilfus C-M, Carpentier SC (2018) Genotype-specific growth and proteomic responses of maize toward salt stress. Front Plant Sci 9:661. https://doi.org/10.3389/fpls.2018.00661
SRDI (2010) Soil Resources Development Institute (SRDI) 2010 Saline Soils of Bangladesh; SRDI, Ministry of Agriculture: Dhaka, Bangladesh. (No. 1). Article 1
Tabassum R, Tahjib-Ul-Arif M, Hasanuzzaman M, Sohag AAM, Islam MS, Shafi SSH, Islam MM, Hassan L (2021) Screening salt-tolerant rice at the seedling and reproductive stages: An effective and reliable approach. Environ Exp Bot. https://doi.org/10.1016/j.envexpbot.2021.104629
Tao R, Ding J, Li C, Zhu X, Guo W, Zhu M (2021) Evaluating and Screening of Agro-Physiological Indices for Salinity Stress Tolerance in Wheat at the Seedling Stage. Front Plant Sci 12:529. https://doi.org/10.3389/fpls.2021.646175
Tuhina-Khatun M, Hanafi MM, Rafii Yusop M, Wong M, Salleh FM, Ferdous J (2015) Genetic variation, heritability, and diversity analysis of upland rice (Oryza sativa L.) genotypes based on quantitative traits. BioMed Res Int. https://doi.org/10.1155/2015/290861
Vaiman D, Mercier D, Moazami-Goudarzi K, Eggen A, Ciampolini R, Lépingle A, Velmala R, Kaukinen J, Varvio S-L, Martin P (1994) A set of 99 cattle microsatellites: Characterization, synteny mapping, and polymorphism. Mamm Genome 5:288–297. https://doi.org/10.1007/bf00389543
Vaja KN, Gajera H, Katakpara ZA, Patel S, Golakiya B (2016) Microsatellite markers based genetic diversity analysis for salt tolerance in wheat genotypes. Indian J Agric Biochem 29:140–145. https://doi.org/10.5958/0974-4479.2016.00023.x
Vignesh M, Anbuselvam Y, Sharma A, Anandan R, Prakash M, Murugan S, Ramakrishnan M (2021) Haplotype and diversity analysis of indigenous rice for salinity tolerance in early-stage seedling using simple sequence repeat markers. Biotechnol Reports. https://doi.org/10.1016/j.btre.2021.e00666
Wijewardana C, Henry WB, Hock MW, Reddy KR (2016) Growth and physiological trait variation among corn hybrids for cold tolerance. Canadian J Plant Sci 96:639–656. https://doi.org/10.1139/cjps-2015-0286
Zhang Y, Fang J, Wu X, Dong L (2018) Na+/K+ balance and transport regulatory mechanisms in weedy and cultivated rice (Oryza sativa L.) under salt stress. BMC Plant Biol 18:1–14. https://doi.org/10.1186/s12870-018-1586-9
Zidani S, Ferchichi A, Chaieb M (2005) Genomic DNA extraction method from pearl millet (Pennisetum glaucum) leaves. African J Biotechnol 4:862–866. https://www.ajol.info/index.php/ajb/article/view/15198
Acknowledgements
We are grateful to the Ministry of Education and Ministry of Science and Technology Ministry, Government of the People’s Republic of Bangladesh, for financial support. We are thankful to BINA for its support during morphological experiments under the hydroponic system. Finally, we acknowledge the BAURES, BAU for the support throughout the entire period of the project.
Author information
Authors and Affiliations
Contributions
MSH, SY conceived the project and planned the experiment. MH, NRS, and SF performed the experimental work. MT-U-A, NRS, and SF analyzed the data. MH wrote the first draft of the paper. MSH and MT-U-A revised and edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
No potential conflict of interest was reported by the author(s).
Additional information
Communicated by M.A.R. Arif.
Rights and permissions
About this article
Cite this article
Hasanuzzaman, M., Saha, N.R., Farabi, S. et al. Screening of salt-tolerant wheat (Triticum aestivum L.) through morphological and molecular markers. CEREAL RESEARCH COMMUNICATIONS 51, 87–100 (2023). https://doi.org/10.1007/s42976-022-00278-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42976-022-00278-x