Abstract
Wheat stripe/yellow rust, caused by Puccinia striiformis f. sp. tritici (Pst), is one of the serious diseases of wheat. The application of resistance cultivars underpins control of wheat stripe rust. The Chinese wheat landrace Dabaimai displayed resistance to most of the Chinese yellow rust races. To decipher the genetic bases of resistance in Dabaimai, a total of 202 recombinant inbred lines F6 of Taichung 29/Dabaimai were tested under field conditions with a predominant Pst race CYR32. Based on SSR markers and field resistance data, two resistance QTLs were detected and named QYr.caas-1AS and QYr.caas-4BS, respectively. QYr.caas-1AS, located on chromosome 1AS between markers Xgwm136 and Xcfd15, can explain 45.97% and 26.83% of the phenotypic variance for infection type (IT) and disease severity (DS), respectively. QYr.caas-4BS, located on chromosome 4BS between markers Xwmc652 and Xgpw4388, can explain 43.38% and 21.30% of the phenotypic variance for IT and DS, respectively. The adult plant stripe rust resistance loci and linked SSR markers will be valuable in combining with other effective genes for combating wheat stripe rust.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat stripe rust, caused by Puccinia striiformis f. sp. tritici (Pst), is an economically important disease around the world. Stripe rust is damaging wheat acreage in more than 60 countries and has become one of the major diseases threatening global wheat production (Chen 2005). In China, four important stripe rust epidemics had occurred during 1950, 1964, 1990 and 2002, respectively (Wan et al. 2007). Stripe rust can reduce wheat yield by 20–30%, and it can exceed 60% if epidemic was severe (Kang et al. 2015). Rational development and deployment of stripe rust-resistant wheat cultivars are the most feasible, environmentally friendly and effective way of stripe rust management (Line 2002). Due to unwise large-scale deployment of single gene cultivars and the evolution and occurrence of new virulent pathotypes of Pst, the stripe rust resistance is easily compromised. Therefore, it is highly essential to decipher and utilize new resistant genes.
Two major types of resistances are known in wheat against stripe rust: One is known as all-stage resistance (ASR) or race-specific resistance which remains active through plant life, while the other is described as adult plant resistance (APR) or non-race-specific resistance which becomes active during later stages of plant development (Chen 2005). Race-specific resistance is easy to be used in wheat breeding. However, race-specific resistance can be frequently overcome by the emergence of new virulent races as a result of strong selection pressure (Chen et al. 1995). On the other hand, non-race-specific resistance is quantitatively inherited and considered durable (Singh et al. 2005). But non-race-specific resistance is hard to prove that can be effective to all races of Pst. Many stripe rust resistance genes have been documented globally (Wang and Chen 2017). There are more than 80 stripe rust resistance genes which were permanently designated using the Yr symbol (https://shigen.nig.ac.jp/wheat/komugi/genes/symbolClassList.jsp). Of the Yr genes so far, some were from common wheat and others were from alien sources. ASR is high proportion, and APR is lower proportion in the resistance genes. Both ASR and APR are important to achieve genetic diversity for resistance in cultivars. Except Yr genes, there is also quantitative trait locus (QTL) which has been reported for resistance to wheat stripe rust. QTLs are distributed on every chromosome with the most on chromosome 2B (Chen and Kang 2017). Some QTLs contribute major effects, while others with small effects can contribute collectively to obtain high levels of resistance to wheat stripe rust. In addition, some QTLs and Yr genes are located in the same region. Compared to QTLs, Yr genes are relatively easier to transfer and obtain resistant cultivars. Since major genes have been widely used in breeding programs, some genes have been overcome by new virulent races. To date, stripe rust resistance genes including Yr5, Yr10, Yr15, Yr18 and Yr36 are still effective against prevalent Chinese races. Though these known genes have facilitated wheat stripe rust breeding in China, identification of new resistance sources is very important for increasing the level of stripe rust resistance and diversifying the resistance genes. Resistant resources landraces or wheat-related species are a good choice as they were regarded as an important source of wheat stripe rust resistance (Kertho et al. 2015).
Landraces are farmer-developed and maintained as traditional cultivars. They have formed through long-term natural selection and artificial selection with the most extensive adaptability to local conditions. In addition, wheat landraces possess complex genetic diversity and valuable resistance genes (Bariana and Bansal 2017). Many genes in wheat landraces conferring resistance to stripe rust were identified, such as Yr45 (Li et al. 2011), Yr53 (Xu et al. 2013), Yr64 and Yr65 (Cheng et al. 2014) for all-stage resistance and Yr52 (Ren et al. 2012), Yr59 (Zhou et al. 2014), Yr62 (Lu et al. 2014), Yr81 (Gessese et al. 2019) and Yr82 (Pakeerathan et al. 2019) for adult plant resistance. Genetic studies on stripe rust resistance of Chinese landraces have been reported. Yr1 were identified from the wheat landraces Chinese 166. Lankao 5, from Henan Province, demonstrated high level of resistance to stripe rust and a different number of genes were found to be involved when inheritance studies were carried out against race CYR30, Sun11-11, CYR31, CYR32, Sun11-4 and CYR29. The stripe rust resistance gene YrLk in Lankao 5 was mapped on chromosome 7B (Yao et al. 2017). Similarly, Baidatou from Gansu Province displayed good APR to Pst races CYR25, CYR27, CY29, CYR30, CYR31, CYR32, CYR33 and Sun11-4. Stripe rust resistance gene YrBai in Baidatou was mapped on chromosome 6DS (Ma et al. 2015). Pingyuan 50, another landrace from China, demonstrated APR against race CYR32 and three QTLs were detected using composite interval mapping approach on chromosomes 2BS, 5AL and 6BS, respectively (Lan et al. 2010).
China has a large number of wheat landrace resource (Liu et al. 2000). Based on the success and achievement of utilization of wheat landraces, it is essential to further explore and exploit the stripe rust resistance gene in Chinese wheat landraces to achieve gradual diversification of resistant genes in wheat cultivars and upcoming germplasm in the country. Keeping this in background, a Chinese wheat landrace “Dabaimai” was chosen due to its strong resistance to stripe rust in adult plant stage and poorly understood the genetic basis of its APR. This paper reports findings of the genetic study of resistance and detection of stripe rust resistance loci in Dabaimai.
Materials and methods
Source of host, pathogen and plant population
Dabaimai, a Chinese wheat landrace, was used as the male parent and crossed with the susceptible spring wheat cultivar Taichung 29 (T29), which is highly susceptible to most Chinese Pst races. The recombinant inbred lines (RILs) population was advanced to F6 generation through selfing. A total of 202 F6 RILs were evaluated for phenotyping and genotyping analysis. Chinese Spring (CS) and its nulli-tetrasomic lines were used to determine the chromosomal location of resistance loci. To determine the relationship of the resistance loci in Dabaimai with Yr genes previously reported on chromosome 4B, PI 192252 (Yr62), CH223 (Yr50) and Daws (YrDa1), which are the known genes carrier materials, were used to test the resistant loci.
The predominant Chinese Pst race CYR32 was used for seedling and adult plant tests which was virulent to all Chinese stripe rust differentials except Zhong 4, Triticum spelta album, Guinong22 and all world differentials except Moro. The stripe rust race was multiplied on Mingxian 169, and spores were maintained after harvesting at the Institute of Plant Protection (IPP), Chinese Academy of Agricultural Sciences (CAAS), China.
Seedling tests
The resistant parent Dabaimai, the known genes carrier materials PI 192252, CH223 and the susceptible parent Taichung 29 were evaluated at the seedling stage in greenhouse. Plants were sown in 9 × 9 cm pots. Each of three replicates consisted of 15–20 plants of each cultivar and was grown in a rust-free condition. After 10 days at two-leaf stage, plants were inoculated at the two-leaf stage by urediniospores of pathotype. CYR32 suspended in 0.05% Tween using manual sprayer. Inoculated seedlings were shifted to a humid chamber for 18–24 h maintained at 10–12 °C. Seedlings were then moved to greenhouse maintained at 15–18 °C/10–14 °C (day/night) with 16 h photoperiod and 5000-6000 Lx. Infection types (IT) were recorded 15–18 days after inoculation using 0–9 scale (Line and Qayoum 1992).
Adult plant tests
The RIL population along with parents was evaluated at adult plant stage for stripe rust response at Langfang Experimental Station of the Institute of Plant Protection, Chinese Academy of Agricultural Sciences in 2017. About 30 seeds of each line were sown in 1.5 m long row, and the distance between rows is 30 cm. The susceptible cultivar Mingxian 169 was raised as rust spreader to ensure adequate inoculum development around experimental field which was irrigated frequently during the growing season to promote disease development. The spreader rows were inoculated by spraying the predominant Pst pathotype CYR32 at the evening with windless and low temperature (12–15 °C). The plants were covered with plastic to obtain a relative humidity at saturation which contributed to the pathogen infection. The next day the plastic was uncovered. The essential nutrients (N, P and K) were regularly arranged. Rust data of RIL population and their parents were initiated when the susceptible cultivar Mingxian 169 reached 80–90% disease severity. Infection type (IT) was scored using 0–9 scale (Line and Qayoum 1992), while disease severity (DS) was scored using 13 scales (Peterson et al. 1948).
DNA extraction and PCR amplification
Genomic DNA of F6 plants along with parents was extracted from leaf tissues using CTAB method (Allen et al. 2006) and dissolved in 1 × TE buffer (10 mM Tris–HCl, 1 mM EDTA, pH 8.0). DNA quantity was evaluated using spectrophotometry and quality using 1.5% agarose gel electrophoresis. DNA samples were stored at − 20 °C and were diluted to a concentration of 50 ng/μl for PCR reactions. Ten RILs resistant plants DNA and 10 RILs susceptible plants DNA were separately bulked to make resistant and susceptible bulks, respectively. The DNA samples from Dabaimai, Taichung 29 and the two bulks were screened using 1124 SSR markers covering all chromosomes. SSR primers information was obtained from GrainGenes 2.0 (http://www.wheat.pw.usda.gov).
PCR was performed with volume of 10 µl comprising 2 µl (100 ng) template DNA, 5 µl 5 × TsingKe Master Mix, 0.5 µl SSR forward primer (10 µM), 0.5 µl SSR reverse primer (10 µM) and 2 µl sterilized ddH2O. PCR reaction was performed in T100 PCR, programmed for 5 min of initial denaturation followed by 35 cycles, each cycle consisting of 60 s at 95 °C, 45 s annealing temperature depending on primers and 60 s at 72 °C, followed by extension of 10 min at 72 °C. PCR products were added 6 µL of formamide loading buffer (98% formamide, 10 mM EDTA, 0.5% xylene cyanol and 0.5% bromophenol blue) and denatured at 94 °C for 5 min. A total of 5 µL mixtures of PCR product and loading buffer were loaded and analyzed by 6% polyacrylamide gel electrophoresis (PAGE). After electrophoresis, silver-stained method was used to visualize the genotype of the samples.
Statistical analysis
A genetic map based on molecular markers was constructed with JoinMap 4.0 using Kosambi mapping function. The genetic map was used for the QTL analysis which was performed with WinQTLCart 2.5 (Wang et al. 2012) using composite interval mapping (CIM) using both phenotype and genotype. A threshold (LOD) value of 2.5 was used to determine the presence of significant QTL.
Results
Resistance to stripe rust in Dabaimai
Dabaimai and Taichung 29 were challenged by Pst pathotype CYR32 in seedling (Fig. 1a) and adult plant stage (Fig. 1b). Result indicated that landrace Dabaimai displayed infection type 8, while susceptible parent Taichung 29 had infection type 9 at the seedling stage. During adult plant stage, Dabaimai demonstrated infection type 1 and susceptible parent Taichung 29 had infection type 9. The seedling and adult plant tests results indicated that resistance to CYR32 in landrace Dabaimai was of adult plant type.
Inheritance of resistance
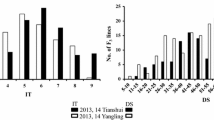
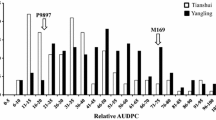
The RIL population and their parents were tested in field with CYR32 to decipher resistance and inheritance of resistance. Variability in infection types of RIL population was recorded against CYR32, which was resistant with necrotic flecks to susceptible having abundant sporulation (Fig. 1c). The RIL population exhibited a continuous distribution of infection type (IT) and disease severity (DS) (Fig. 2), indicating that not a single gene controlling this observed rust resistance.
Construction of genetic linkage map and screening resistant linkage markers
A total of 1124 SSR markers covering 21 chromosomes were used. Among these markers, 73 on chromosome 1A and 97 on chromosome 4B were tested. A total of 8 SSR markers were found to be polymorphic between the two parents and bulks based on field IT data. Xcfa2153, Xcfd15, Xgpw7072 and Xgwm136 were located on chromosome 1A. Xgwm4388, Xgwm375, Xwmc652 and Xwmc349 were located on chromosome 4B. By JoinMap 4.0, two linkage maps for chromosomes 1A and 4B were constructed.
QTL analysis
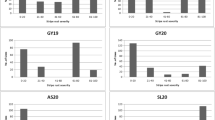
Based on SSR markers and resistant identification, two resistance QTLs on chromosomes 1AS and 4BS, named QYr.caas-1AS and QYr.caas-4BS, were detected using composite interval mapping method by WinQTLCart 2.5 (Table 1; Fig. 3). For infection type, QYr.caas-1AS was located between marker Xgwm136 and Xcfd15 and the genetic distance from the marker Xgwm136 was 1.00 cM. QYr.caas-1AS explained 45.97% phenotypic variance. QYr.caas-4BS was located between the marker Xwmc652 and Xgpw4388, and the genetic distance from the marker Xgpw4388 was 1.30 cM. QYr.caas-4BS explained 43.38% phenotypic variance. For disease severity, QYr.caas-1AS was situated on the same locus with marker Xgwm136 and explained 26.83% of the phenotypic variance. QYr.caas-4BS was mapped between marker Xwmc652 and Xgpw4388, and the genetic distance from the marker Xgpw4388 was 1.30 cM. QYr.caas-4BS explained 21.30% of the phenotypic variance.
To determine the location of two resistant loci, the linkage markers were tested on 21 nulli-tetrasomic lines of Chinese Spring. SSR marker Xgwm136 amplified fragments about 300 bp in CS except N1AT1D. SSR marker Xgwm4388 amplified fragments of 247 bp in CS except N4BT4D. The results showed that QYr.caas-1AS and QYr.caas-4BS were located on chromosomes 1AS and 4BS, respectively.
Discussion
With continually evolving new Pst pathogen races, which can be capable of overcoming deployed resistance genes, and the limited availability of resistance genes in wheat cultivars, there is an ongoing need to discover and utilize novel rust resistance sources for wheat breeding. It is of high significance and great practical value to characterize, validate and utilize rust-resistant resources for wheat production. Rational deployment of resistance genes can avoid the epidemics of wheat stripe rust. In this study, we identified two new stripe rust adult plant resistance loci located on chromosomes 1AS and 4BS in wheat landrace Dabaimai.
QYr.caas-1AS was a locus of APR on chromosome 1AS. There was no officially named resistance gene on chromosome 1A, while the temporarily named resistance genes (YrHA and YrZhong12-2) were not located on 1AS chromosome. Only a single resistance gene, YrDa1, was reported to be located on chromosome 1A (Chen et al. 1995) without specific location. The linkage marker Xcfd15 screened in this study was used to test Daws, carrier of known gene YrDa1, and Dabaimai by polypropylene gel electrophoresis detection (Fig. 4a), suggesting that the resistance loci in Daws and Dabaimai were different. QYr.cim-1AS from cultivar Sujata (Lan et al. 2015) and QYrid.ui-1A from IDO444 (Chen et al. 2012) are in distal region of short arm of 1A chromosome; QYr.sig-1A.1 from Kariega (Prins et al. 2011), QYr.tam-1AS from TAM111 (Basnet et al. 2014) and QYr.sun-1A from Janz (Bariana et al. 2010) were reported near the centromere of the 1A chromosome. It was therefore unlikely that the loci in Sujata, IDO444, Kariega, TAM111 and Janz were same that in Dabaimai. QYr.caas-1AS was a new resistant locus.
There are six genes reported on the 4B chromosome. Officially named three resistance genes were Yr50 (Liu et al. 2013), Yr62 (Lu et al. 2014) and Yr68, and temporarily named three resistance genes were YrZH22 (Wang et al. 2017), YrXY (Jing et al. 2007) and YrElm2 (Yang et al. 2008). To compare the response to Pst, CH223 (Yr50), PI 192252 (Yr62), Dabaimai (QYr.caas-4BS) and susceptible variety Taichung 29 were tested with races CYR32 in the seedling stage. The three varieties displayed different infection types: CH223 (Yr50) with a resistant reaction (IT 1), PI 192252 (Yr62) and Dabaimai with a susceptible reaction (Fig. 5), indicating the resistance gene in them was different. In addition, the linkage marker Xgpw4388 was used to test CH223 (Yr50), PI 192252 (Yr62) and Dabaimai by polypropylene gel electrophoresis (Fig. 4b). The polymorphic bands were different indicating QYr.caas-4BS were different from Yr50 and Yr62. Furthermore, Yr62 was flanked between Xgwm251 and Xgwm192, whose distance was 0.60 cM to Yr62. QYr.caas-4BS has interval of 28.50 cM to Xgwm192, so it can confirm that QYr.caas-4BS and Yr62 were different. YrElm2 was located 7.60 cM to Xgwm495, but QYr.caas-4BS was 18.40 cM to Xgwm495, indicating they have different resistant loci. YrZH22 was mapped 7.24 cM to Xwmc47, which was away QYr.caas-4BS about 16.90 cM, so they have different genes. YrXY has a distance of 7.08 cM to Xgwmc107, but the relative position of QYr.caas-4BS and YrXY was different (Fig. 6). QYr.caas-4BL (Lu et al. 2009) was identified between Xgwm165 and Xgwm149, QYr.ufs-4BL (Agenbag et al. 2012) was situated between Xgwm165 and Xgwm495, QYr.sun-4BL (Zwart et al. 2010) was located between Xwmc238 and Xgwm368, and Qpst.jic-4BL (Jagger et al. 2011) was linked to Xwmc692. These four resistant loci were different to QYr.caas-4BS by comparing relative position (Fig. 6). QYr.caas-4BS was a new resistant locus.
Dabaimai is a Chinese wheat landrace, which was the artificial and natural selection results during long-term production practice. Dabaimai had provided stripe rust resistance to CYR32 in adult plant stage. QYr.caas-1AS and QYr.caas-4BS were two adult plant resistance loci, and their phenotypic variation rates can reach 26.83–45.97% and 21.30–43.38%. So it is of great practical value to be exploited and utilized in combination with other resistance genes for high-level wheat resistance breeding. Also, the polymorphic markers can be used in marker-assisted selection for incorporating resistance genes into commercial cultivars. The selection process can be accelerated, and the selection efficiency can be improved.
References
Agenbag GM, Pretorius ZA, Boyd LA, Bender CM, Prins R (2012) Identification of adult plant resistance to stripe rust in the wheat cultivar Cappelle–Desprez. Theor Appl Genet 125:109–120
Allen GC, Floresvergara MA, Krasynanski S, Kumar S, Thompson WF (2006) A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat Protoc 1:2320–2325
Bariana HS, Bansal UK (2017) Breeding for disease resistance A2-Thomas, Brian. In: Murray BG, Murphy DJ (eds) Encyclopedia of applied plant sciences, 2nd edn. Academic Press, Oxford, pp 69–76
Bariana HS, Bansal UK, Schmidt A, Lehmensiek A, Kaur J, Miah H, Howes N, McIntyre CL (2010) Molecular mapping of adult plant stripe rust resistance in wheat and identification of pyramided QTL genotypes. Euphytica 176:251–260
Basnet BR, Ibrahim AMH, Chen XM, Singh RP, Mason ER, Bowden RL, Liu SY, Hays DB, Devkota RN, Subramanian NK, Rudd JV (2014) Molecular mapping of stripe rust resistance in hard red winter wheat TAM 111 adapted to the U.S. high plains. Crop Sci 54:1361–1373
Chen XM (2005) Epidemiology and control of stripe rust (Puccinia striiformis f. sp. tritici) on wheat. Can J Plant Pathol 27:314–337
Chen X, Kang Z (2017) Stripe rust. Springer, Dordrecht, p 444
Chen XM, Line RF, Jones SS (1995) Chromosomal location of genes for resistance to Puccinia striiformis in winter wheat cultivars Heines VII, Clement, Moro, Tyee, Tres, and Daws. Phytopathology 85:1362–1367
Chen JL, Chu C, Souza EJ, Guttieri MJ, Chen XM, Xu S, Hole D, Zemetra R (2012) Genome-wide identification of QTL conferring high-temperature adult-plant (HTAP) resistance to stripe rust (Puccinia striiformis f. sp. tritici) in wheat. Mol Breed 29:791–800
Cheng P, Xu LS, Wang MN, See D, Chen XM (2014) Molecular mapping of genes Yr64 and Yr65 for stripe rust resistance in hexaploid derivatives of durum wheat accession PI 331260 and PI 480016. Theor Appl Genet 127:2267–2277
Gessese M, Bariana H, Wong D, Hayden M, Bansal U (2019) Molecular mapping of stripe rust resistance gene Yr81 in a common wheat landrace Aus27430. Plant Dis. https://doi.org/10.1094/PDIS-06-18-1055-RE
Jagger LJ, Newell C, Berry ST, Maccormack R, Boyd LA (2011) The genetic characterisation of stripe rust resistance in the German wheat cultivar Alcedo. Theor Appl Genet 122:723–733
Jing JX, Xu ZB, Wang DB, Wang MN, Yao QY, Shang HS, Li ZQ (2007) Genetic analysis of gene conferring resistance to stripe rust in Xiaoyan6. Sci Agric Sin 40:499–504
Kang ZS, Wang XJ, Zhao J, Tang CL, Huang LL (2015) Advances in research of pathogenicity and virulence variation of the wheat stripe rust fungus Puccinia striiformis f. sp. tritici. Sci Agric Sin 48:3439–3453
Kertho A, Mamidi S, Bonman JM, McClean PE, Acevedo M (2015) Genome-wide association mapping for resistance to leaf rust and stripe rust in winter-habit hexaploid wheat landraces. PLoS ONE 10(6):e0129580. https://doi.org/10.1371/journal.pone.0129580
Lan CX, Liang SS, Zhou XC, Zhou G, Lu QL, Xia XC, He ZH (2010) Identification of genomic regions controlling adult-plant stripe rust resistance in Chinese landrace Pingyuan 50 through bulked segregant analysis. Phytopathology 100:313–318
Lan CX, Zhang YL, Herrera-Foessel SA, Basnet BR, Huerta-Espino J, Lagudah ES, Singh RP (2015) Identification and characterization of pleiotropic and co-located resistance loci to leaf rust and stripe rust in bread wheat cultivar Sujata. Theor Appl Genet 128:549–561
Li Q, Chen XM, Wang MN, Jing JX (2011) Yr45, a new wheat gene for stripe rust resistance on the long arm of chromosome 3D. Theor Appl Genet 122:189–197
Line RF (2002) Stripe rust of wheat and barley in North America, a retrospective historical review 1. Annu Rev Phytopathol 40(1):75–118
Line RF, Qayoum A (1992) Virulence, aggressiveness, evolution and distribution of races of Puccinia striiformis (the cause of stripe rust of wheat) in North America, 1968–1987. Technical Bulletin Number 1788, USDA-ARS, Washington, DC
Liu SC, Zheng DS, Cao YS, Song CH, Chen MY (2000) Genetic diversity of landrace and bred varieties of wheat in China. Sci Agric Sin 33:20–24
Liu J, Chang ZJ, Zhang XJ, Yang ZJ, Li X, Jia JQ, Zhan HX, Guo HJ, Wang JM (2013) Putative Thinopyrum intermedium-derived stripe rust resistance gene Yr50 maps on wheat chromosome arm 4BL. Theor Appl Genet 126:265–274
Lu YM, Lan CX, Liang SS, Zhou XC, Liu D, Zhou G, Lu QL, Jing JX, Wang MN, Xia XC, He ZG (2009) QTL mapping for adult-plant resistance to stripe rust in Italian common wheat cultivars Libellula and Strampelli. Theor Appl Genet 119:1349–1359
Lu Y, Wang MN, Chen XM, See D, Chao SM, Jing JX (2014) Mapping of Yr62 and a small-effect QTL for high-temperature adult-plant resistance to stripe rust in spring wheat PI 192252. Theor Appl Genet 127:1449–1459
Ma DF, Li Q, Tang MS, Chao KX, Li JC, Wang BT, Jing JX (2015) Mapping of gene conferring adult-plant resistance to stripe rust in Chinese wheat landrace Baidatou. Mol Breed 35:157–165
Pakeerathan K, Bariana H, Qureshi N, Wong D, Hayden M, Bansal U (2019) Identification of a new source of stripe rust resistance Yr82 in wheat. Theor Appl Genet 132:3169–3176
Peterson RF, Campbell AB, Hannah AE (1948) A diagrammatic scale for estimating rust intensity on leaves and stems of cereals. Can J Res 26:496–500
Prins R, Pretorius ZA, Bender CM, Lehmensiek A (2011) QTL mapping of stripe, leaf and stem rust resistance genes in a Kariega × Avocet S doubled haploid wheat population. Mol Breed 27:259–270
Ren RS, Wang MN, Chen XM, Zhang ZJ (2012) Characterization and molecular mapping of Yr52 for high-temperature adult-plant resistance to stripe rust in spring wheat germplasm PI 183527. Theor Appl Genet 125:847–857
Singh RP, Huerto-Espino J, William HM (2005) Genetics and breeding for durable resistance to leaf and stripe rusts in wheat. Turk J Agric For 29:121–127
Somers D, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Wan AM, Chen XM, He ZH (2007) Wheat stripe rust in China. Aust J Agric Res 58:605–619
Wang MN, Chen XM (2017) Stripe rust resistance. In: Chen XM, Kang ZS (eds) Stripe rust. Springer, Dordrecht, pp 353–558
Wang S, Basten CJ, Zeng ZB (2012) Windows QTL Cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh
Wang Y, Xie JZ, Zhang HZ, Guo BM, Ning SZ, Chen YX, Lu P, Wu QH, Li MM, Zhang DY, Guo GH, Zhang Y, Liu DC, Zou SK, Tang GW, Zhao H, Wang XC, Li J, Yang WY, Cao TJ, Yin GH, Liu ZY (2017) Mapping stripe rust resistance gene YrZH22, in Chinese wheat cultivar Zhoumai 22 by bulked segregant RNA-Seq (BSR-Seq) and comparative genomics analyses. Theor Appl Genet 130:2191–2201
Xu LS, Wang MN, Cheng P, Kang ZS, Hulbert S, Chen XM (2013) Molecular mapping of Yr53, a new gene for stripe rust resistance in durum wheat accession PI480148 and its transfer to common wheat. Theor Appl Genet 126:523–533
Yang MN, Jing J, Liu P (2008) Genetic analysis and microsatellite makers of a resistance gene YrElm2 in wheat from Elymus mollis (Trin.) Hara. J Northwest Agric For Univ (Nat Sci Ed) 4:187–192
Yao Q, He MM, Hou L, Yan JH, Guo QY, Jing JX, Kang ZS (2017) Genetic analysis and molecular mapping of stripe rust resistance genes in Chinese native wheat (Triticum aestivum) Lankao 5. Australas Plant Pathol 46:213–221
Zhou XL, Wang MN, Chen XM, Lu Y, Kang ZS, Jing JX (2014) Identification of Yr59 conferring high temperature adult plant resistance to stripe rust in wheat germplasm PI 178759. Theor Appl Genet 127:935–945
Zwart RS, Thompson JP, Milgate AW, Bansal UK, Williamson PM, Raman H, Bariana HS (2010) QTL mapping of multiple foliar disease and root-lesion nematode resistances in wheat. Mol Breed 26:107–124
Acknowledgements
This research was supported by the National Natural Science Foundation of China (No. 31871923) and the National Key Research and Development Program of China (Nos. 2016YFD0300705 and 2018YFD0200500).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no financial and associative relationships with other people or organizations that represent a conflict of interest in connection with the work submitted.
Additional information
Communicated by K. Posta.
Rights and permissions
About this article
Cite this article
Wang, M.Y., Xu, M.R., Wang, F.T. et al. Characterization and validation of QTLs for adult plant stripe rust resistance in Chinese wheat landrace Dabaimai. CEREAL RESEARCH COMMUNICATIONS 49, 91–98 (2021). https://doi.org/10.1007/s42976-020-00071-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42976-020-00071-8