Abstract
Cryptococcus neoformans is primarily responsible for cases of cryptococcal meningitis in individuals with HIV/AIDS. This study evaluated the susceptibility of C. neoformans obtained from individuals with cryptococcal meningitis associated with HIV/AIDS in Manaus, Amazonas, Brazil, against the action of the antifungals amphotericin B, flucytosine, fluconazole, itraconazole and posaconazole and analyzed it using Multilocus Sequence Typing (MLST) in order to identify the Sequence Types (STs). We analyzed 30 isolates of C. neoformans, from 24 HIV/AIDS patients diagnosed with cryptococcosis from 2017 to 2019 in a reference hospital, in addition to 3 environmental isolates: 1 isolate obtained in the home of a patient and 2 isolates obtained from neighboring homes of patients. 86.6% (n = 26/30) of the clinical isolates were identified as C. neoformans VNI ST93, 6.6% (n = 2/30) as C. neoformans VNI ST5, 3.3% (n = 1/30) as C. neoformans VNI ST32 and 3.3% (n = 1/30) as C. neoformans VNB ST232. The environmental isolates were identified as C. neoformans VNI ST93 (n = 3/3). 96.6% (n = 29/30) isolates were sensitive to amphotericin B, though there was variation in the MIC. 60% (n = 18/30) presented a MIC above the proposed epidemiological cutoff values for one or more antifungals. All environmental isolates were sensitive to the tested antifungals. The MLST showed that there is an important relationship between C. neoformans VNI ST93 and individuals with HIV/AIDS, including in the environmental isolates analyzed. C. neoformans VNB ST232 was observed for the first time in Amazonas. Amphotericin B was proven to be the best drug, but fluconazole and posaconazole also showed relevant action.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cryptococcosis is a global invasive fungal infection with considerable morbidity and mortality [1], and cryptococcal meningitis is one of the opportunistic infections that most commonly affects individuals with HIV/AIDS [2]. In Latin American countries, C. neoformans VNI is the main agent of cryptococcosis associated with HIV/AIDS [3].
In Brazil, although the public health system provides free antiretroviral therapy (ART), the mortality rate for cryptococcal meningitis caused by C. neoformans still remains high [4]. According to the international standard induction regimen, the treatment of cryptococcal meningitis is performed with amphotericin B and flucytosine, followed by fluconazole [5]. However, in Latin America, flucytosine is not available and the induction of combined therapy, of amphotericin with fluconazole, is not frequent [6]. Adequate therapy associated with early diagnosis of the causative agent is essential in order to reduce deaths and sequelae of cryptococcal meningitis; however, the emergence of strains resistant to the antifungals used for the treatment of cryptococcosis is a factor that further hinders effective antifungal therapy [7].
In the state of Amazonas, northern Brazil, cases of meningitis with various etymologies have been observed since 1976 [8]. AIDS-associated cryptococcosis has been linked primarily to C. neoformans VNI with ST93 [9]. Alves et al. [10] reported a high mortality rate from cryptococcal meningitis in HIV/AIDS patients in Manaus, Brazil, which reflects the importance of cryptococcosis as an AIDS-defining disease and an important public health problem in the region, in addition to indicating that the home environment is a potential source of infection/reinfection by C. neoformans VNI. It is evident that epidemiological surveillance of the antifungal resistance of Cryptococcus in the Amazon region is very important in view of the high lethality of cryptococcal meningitis 7.
The present study aimed to determine the susceptibility of Cryptococcus neoformans isolates obtained from individuals with HIV/AIDS and diagnosed with cryptococcal meningitis in a reference hospital for AIDS cases in Manaus, Brazil, and identify the sequence types (STs) responsible for infections in patients via multilocus sequence typing (MLST).
Methods
Acquisition of the isolates and ethical approval
During the period from 2017 to 2019, 30 clinical isolates of C. neoformans were obtained from 24 patients with HIV/AIDS and diagnosed with cryptococcosis at the Fundação de Medicina Tropical Doutor Vieira Dourado (FMT-HVD). To verify the possibility of new infection or reinfection by different cryptococcosis agents, all isolates from the same patient obtained during the study period were analyzed. The isolates were maintained on Sabouraud dextrose agar at 4 °C in the Mycology Laboratory at FMT, and were given to us for the experiments that were carried out in the Mycology Laboratory of the Instituto Leônidas e Maria Deane (ILMD/FIOCRUZ).
In addition to the clinical isolates, three environmental isolates of C. neoformans obtained from the homes of patients and their neighbors were also used. Environmental samples of household dust, soil, bird excreta and atmospheric air were collected from 51 households (17 from patients and 34 from neighboring households) in order to verify the presence of the fungus in the homes of patients and their neighbors [10]. Of these, the three isolates of C. neoformans used in this study were obtained: one isolate from soil from a patient’s home and two isolates from household dust from the homes of neighbors of patients.
The study was conducted according to the guidelines of the Declaration of Helsinki and approved by the Ethics Committee of the Fundação de Medicina Tropical Doutor Vieira Dourado (protocol code CAAE No. 82715917.4.0000.0005).
Multilocus sequence typing of C. neoformans
The isolates of C. neoformans were processed individually for the extraction of total DNA, according to the instructions of the QIAamp Tissue and Blood extraction kit (Qiagen, Hilden, Germany) with modifications in the pre-phase of mechanical maceration with glass beads.
Molecular typing using PCR–RFLP was performed with amplification of the primers URA5 (5´ATGTCCTCCCAAGCCCTCGACTCCG 3´) and SJ01 (5´ TTAAGACCTCTGAACACCGTACTC 3´), followed by enzymatic digestion with HhaI and Sau961 (ThermoScientific) according to Meyer et al. [11]. C. neoformans isolates were submitted to MLST via amplification of CAP59, GPD1, LAC1, PLB1, SOD1, IGS1 and URA5 genes according to the conditions proposed by Meyer et al. [12].
Amplicons were purified and sequenced according to the manufacturer’s instructions with the BigDye® Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems). Capillary electrophoresis was performed in genetic analyzer sequencer (ABI 3130). The sequences were manually edited using Sequencher 5.3 software (Gene Codes Corporation, Ann Arbor, MI, USA) and the contigs were aligned in the MEGA v6.06 program. The sequences were analyzed in the MLST database (http://mlst.mycologylab.org) for determination of alleles and ST. To verify the genetic relationship of the isolates, a phylogenetic tree was constructed based on the neighbor-joining (NJ) model, with bootstrap analysis using 1,000 replicates, and using related STs.
Antifungal susceptibility test
ETEST® (bioMérieux) tapes were used with the antifungals: amphotericin B, flucytosine, fluconazole, itraconazole and posaconazole. The inoculum was prepared from Cryptococcus colonies grown on Sabouraud agar medium and 0.85% sodium chloride (NaCl) solution, homogenized to a 1.0 McFarland turbidity and then placed in a Petri dish containing RPMI 1640 (Himedia Laboratorios Pvt. Ltd., Mumbai), supplemented with 1.5% agar and 2% glucose, and buffered to pH 7.00 ± 0.01 with 3N-morpholino-propanesulfonic acid – MOPS (Sigma Aldrich) with a thickness of 0.4 mm ± 0.1. Epidemiological cutoff value (ECV) were evaluated according to Spinel-Ingroff et al. [13, 14]. Whonet software (version) 5.6 was used to analyze the determination of the geometric mean. To calculate the minimum inhibitory concentration 90 (MIC90) and geometric mean, the value corresponding to 100% inhibition of fungal growth was used for amphotericin, 95% for flucytosine, and 80% for azole antifungals, according to the ETEST reading protocol. A sample of Candida parapsilosis ATCC 22019 was used as control.
Results
MLST of the clinical and environmental isolates of C. neoformans
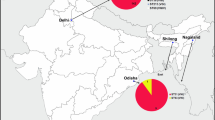
A total of 33 isolates (30 clinical and 3 environmental) were analyzed. Among the clinical isolates, 86.6% (n = 26/30) were identified as C. neoformans VNI with ST93, 6.6% (n = 2/30) as C. neoformans VNI with ST5, 3.3% (n = 1/30) as C. neoformans VNI with ST32, and 3.3% (n = 1/30) as C. neoformans VNI with ST 232. The environmental isolates were identified as C. neoformans VNI with ST93 (n = 3/3) (Table 1).
Phylogenetic analysis demonstrates a relationship of great identity in the grouping of all ST93 isolates, both between environmental strains and clinical strains found in the infection acquired by patients. The C. neoformans VNB isolate with ST 232 and the reference sequences formed a separate cluster (Fig. 1).
Phylogenetic tree of STs obtained in the state of Amazonas. Phylogeny reconstruction was performed by the neighbor-joining algorithm (bootstrap with 1,000 replicates) with the concatenated sequences of the CAP59, GPD1, LAC1, PLB1, SOD1, URA5 and IGS genes of the isolates obtained in the present study and sequences acquired from the MLST database
Susceptibility to antifungals
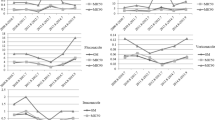
The isolates of C. neoformans (environmental and clinical) showed a large variation in the MIC for all the antifungal agents tested, though the greatest variation was observed for flucytosine (0.032 to > 32). Among the isolates, 40% (n = 12/30) were sensitive to all antifungals tested. 60% (n = 18/30) of the clinical isolates presented a MIC above the proposed ECVs for one or more antifungals. The isolate P33-515 presented high MICs for amphotericin, fluconazole and itraconazole; only this isolate was resistant to amphotericin B, and was sensitive only to flucytosine and posaconazole. For fluconazole susceptibility, 10% (n = 3/30) of clinical isolates demonstrated a MIC above what is established.
Isolates from the same patient, obtained at an interval of 15 days, showed different results against antifungals. While isolate P24-430/02 showed no elevated MIC for any antifungal, isolate P24-606/02 presented a MIC higher than the proposed ECVs for itraconazole and posaconazole. Isolate P39-1008 presented a MIC higher than the ECVs proposed for fluconazole and itraconazole, whereas isolate P39-1052 presented a higher MIC only for itraconazole. None of the environmental isolates presented a MIC higher than the proposed ECVs (Table 1). The geometric mean and MIC90 can be seen in Table 2.
Discussion
According to Ficarative, Meyer & Castañeda [3], in Latin America, the population of C. neoformans has less diversity in its STs than the population of C. gattii. In Brazil, the five molecular types VNI, VNB, VNII, VNIII and VNIV have already been identified. C. neoformans VNI with ST93 being the most frequent, followed by ST77, ST2, ST5 and ST23. ST93 also predominates among environmental isolates. C. neoformans VNII with ST40 and ST41 has been observed causing infection in humans and animals [15,16,17], and C. neoformans VNIV with ST11 and ST160 causing infection in humans [15]. C. neoformans VNIII is rare in Brazil with only one environmental isolate recorded [18].
Ferreira-Paim et al. [4], in southeastern Brazil, reported a higher frequency of ST93 when analyzing both clinical isolates of C. neoformans obtained from patients with HIV/AIDS and environmental isolates. In northern Brazil, ST93 is also the main subgenotype of VNI strains that causes cryptococcosis in individuals with HIV/AIDS [9].
Rocha et al. [9], in a study conducted from 2014 to 2016 (in the three years preceding this study), identified only ST93 in isolates of C. neoformans VNI, as the cause of cryptococcosis in 25 patients with HIV/AIDS, in Manaus, Brazil; while, in the present study, isolates from 24 patients were analyzed and, in addition to ST93, three other STs were also identified, ST5 and ST32 (C. neoformans VNI), in addition to ST232 (C. neoformans VNB), which via PCR–RFLP genotyping presented a molecular pattern identical to the molecular type VNII [10]. The VNB population may be underestimated due to the close relationship between VNB and VNII, especially when only RFLP PCR is used to identify the molecular type [15].
Ferreira-Paim et al. [4] also identified ST32 in clinical isolates from patients with cryptococcosis in Minas Gerais, southeastern Brazil. ST32 is rarely found in South America, being more observed in South Africa where it appears to present worse patient survival after the first year of infection [19]. As in our study, ST93, ST5 and ST32 were related to patients with HIV/AIDS. C. neoformans with ST93, ST5 and ST32 has also been associated with AIDS in Asia [20, 21]. In East Asia, ST5 is predominant, being one of the most virulent STs [22], but in Brazil the isolation of this subgenotype is considered rare [4, 15].
The VNB genotype shows high virulence in Africa and in countries outside the African continent is considered rare [23]. Previously, in Brazil, VNB with ST504 and ST527 was identified in environmental isolates in the southeastern region [15], and VNB with ST 232 was identified in two clinical isolates of HIV/AIDS patients in Rio de Janeiro [16] and, more recently, in our study. According to Rhodes et al. [24], the formerly “African” VNB lineage occurs naturally in the South American environment, suggesting a migration of the VNB lineage between Africa and South America before the diversification of this lineage, whose nature is quite dispersive. The geographic niche of the VNB lineage could be separated when the continents split, in Pangea [25].
The three environmental isolates from patient’s homes/patient’s neighbors’ homes are C. neoformans with ST93, which corroborates other environmental studies [4, 26]. Brito-Santos et al. [27] revealed an endemic pattern in domestic microenvironments in the Negro River microregion of the Brazilian Amazon and observed the presence of C. neoformans VNI with ST93 and also C. neoformans VNI with ST5.
ETEST is considered an excellent method for distinguishing yeasts that are resistant to amphotericin B [28, 29], besides being reproducible and simpler than dilution in broth [29]. There are no well-established clinical cutoff points in the revised CLSI M27-A3 and CLSI M27-S3 documents to determine the susceptibility of C. neoformans /C. gattii isolates in tests with antifungals, so the epidemiological cutoff values (ECVs) established by Espinel-Ingroff et al. [13, 14], which proposes the adoption of species-specific values, were used. According to Lockhart et al. [30], the ECV can predict whether an isolate, for which there is not enough data to establish cutoff points, has acquired mechanisms of resistance to an antifungal agent with already known activity.
Nishikawa et al. [7], when analyzing the antifungal susceptibility of C. neoformans isolates from the northern region of Brazil, especially from the state of Pará, observed higher voriconazole activity among the azoles tested and lower fluconazole activity and high frequency of C. neoformans VNI isolates with MICs higher than the ECVs proposed for amphotericin B. Unlike the study by Nishikawa et al. [7], the present study used posaconazole and not voriconazole, and this, together with fluconazole, showed good results against the isolates analyzed. In addition, only one isolate presented MICs higher than the ECVs proposed for amphotericin B.
Rocha et al. [9] evaluated isolates of C. neoformans VNI with ST93 from patients with HIV/AIDS, in Manaus, Amazonas, Brazil, and via the broth microdilution technique, observed the susceptibility of all isolates to amphotericin B, fluconazole and itraconazole. This same observation is made in Silva et al. [31] and in other studies [5, 32,33,34]. In the present study, 60% (n = 18/30) of the clinical isolates presented MICs higher than the ECVs proposed for one or more antifungals, with variation in MICs even among ST93 isolates, and 96.6% (n = 29/30) of the clinical isolates tested were sensitive to amphotericin B.
The C. neoformans VNB isolate (P5-25) did not show an elevated MIC for any of the antifungals tested. Naicker et al. [35] observed low MIC values for the antifungal Fluconazole when analyzing 15 VNB isolates and no difference between the STs, including ST232.
In patients with HIV/AIDS and cryptococcal meningitis, induction therapy with amphotericin B associated with flucytosine and followed up by fluconazole for up to 10 weeks is the treatment of choice. After this period, fluconazole is used with a reduced dose according to the patient’s clinical status [5]. In Amazonas state, fluconazole and amphotericin B are the drugs of choice for the treatment of patients with cryptococcosis associated with HIV/AIDS [9, 10]. Our patients were treated with these two antifungals. When evaluating the in vitro antifungal activity of Amphotericin B and Fluconazole, was observed a variation in MIC for the different isolates of C. neoformans VNI with ST93 and 50% (n = 10/20) of patients infected with these isolates died. The patient from whom the only strain that presented a high MIC for Amphotericin B was isolated died. This same strain also showed a high MIC for Fluconazole. The remaining patients who died were infected with isolates with MIC below the ECV for Amphotericin B and Fluconazole, but all from ST93.
The two VNI isolates of C. neoformans with ST5 showed high MIC for 5-FC and Itraconazole, which may demonstrate a possible resistance of this molecular type to these antifungals, but none of the patients used these antifungals and none died.
Matos et al. [36], when analyzing isolates from patients with cryptococcal meningitis in Bahia, northeastern Brazil, observed isolates resistant to amphotericin B, fluconazole and flucytosine. In our study, isolate P33-515 had elevated MICs for amphotericin B, fluconazole and itraconazole, with lower MICs for flucytosine and posaconazole. It is important to highlight that patient P33-515 used liposomal amphotericin B and died seven days after the diagnosis of cryptococcosis and initiation of treatment with this antifungal [10], which is the induction treatment in cases of cryptococcosis, administered with 2 to 4 weeks. The other isolates studied were susceptible to Amphotericin B, but P36-878 and P39-1008 showed high MIC for Fluconazole, which is used for consolidation therapy in the treatment of cryptococcosis.
In a study by Molloy et al. [37], the combination of amphotericin B plus flucytosine showed lower mortality of patients at ten weeks, but side effects were more frequent with two weeks of amphotericin B than with one week. However, as in all of Latin America, flucytosine is not available in Amazonas state. In countries where flucytosine is not available, the most frequent treatment is monotherapy with amphotericin B [6], or the induction of monotherapy with fluconazole is used; however, the rate of fungal clearance of fluconazole is lower when compared to amphotericin B, and mortality is 50 to 60% in 10 weeks, even with the use of high doses [33].
As for the environmental isolates, the MIC results were the same as those observed in most clinical isolates and susceptibility was similar to that reported in other environmental studies, though isolates resistant to the antifungals tested were not observed [38,39,40].
Conclusion
C. neoformans VNI with ST93 is mainly involved in infections that affect individuals with HIV/AIDS in Brazil and throughout Latin America. In Amazonas state, there is a predominance of this subgenotype, but there are other STs present that affect individuals with HIV/AIDS, namely ST5 and ST32, in addition to C. neoformans VNB with ST232 infections having also been observed. C. neoformans VNB, previously considered restricted to the African continent, has been observed in other parts of the world, including Brazil. In this study, C. neoformans VNB with ST232 was observed for the first time in a patient with HIV/AIDS in Manaus, Brazil. ST93, the ST that is most observed in clinical isolates, was also identified in the environmental isolates analyzed, thus demonstrating the importance of the home environment for possible infection/reinfection by the fungus. In all, 40% of the isolates were sensitive to all antifungals. Only one isolate presented a MIC above the proposed ECV for AMB, which demonstrates that AMB, despite being associated with side effects, is still the most recommended and effective drug for the treatment of cryptococcal meningitis. To reduce sequelae and mortality from cryptococcal meningitis, there is a need for early diagnosis and treatment of cryptococcosis, greater adherence to treatment with HAART and other effective therapeutic options.
References
Perfect JR, Dismukes WE, Dromer F, Goldman DL, Graybill JR, Hamils RJ et al (2010) Clinical practice guidelines for the management of cryptococcal disease: 2010 update by the infectious diseases society of America. Clin Infect Dis 50(3):291–322. https://doi.org/10.1086/649858
Park BJ, Wannemuehler KA, Marston NG, Pappas PG, Chiller TM (2009) Estimation of the current global burden of cryptococcal meningitis among persons living with HIV/AIDS. AIDS 23:525–530. https://doi.org/10.1097/QAD.0b013e328322ffac
Firacative F, Meyer M, Castañeda E (2021) Cryptococcus neoformans and Cryptococcus gattii species complexes in Latin America: a map of molecular types, genotypic diversity, and antifungal susceptibility as reported by the Latin American cryptococcal study group. J Fungi 7(4):282. https://doi.org/10.3390/jof7040282
Ferreira-Paim K, Andrade-Silva L, Fonseca FM, Ferreira TB, Mora DJ, Reis C et al (2017) MLST-Based Population Genetic Analysis in a Global Context Reveals Clonality amongst Cryptococcus neoformans var. grubii VNI Isolates from HIV Patients in Southeastern Brazil. PLOS Negl Trop Dis 18:1–29. https://doi.org/10.1371/journal.pntd.0005223
Saag MS, Graybill RJ, Larsen RA, Pappas PG, Perfect JR, Powderly WG (2000) Practice guidelines for the management of cryptococcal disease. CID 30:710–718. https://doi.org/10.1086/313757
Vidal JE, Oliveira ACP, Dauarc RF, Boulwared DR (2013) Strategies to reduce mortality and morbidity due to AIDS-related cryptococcal meningitis in Latin America. Braz J Infect Dis 17(3):353–362. https://doi.org/10.1016/j.bjid.2012.10.020
Nishikawa MM, Almeida-Paes R, Brito-Santos F, Nascimento CR, Fialho MM, Trilles L, Morales BP et al (2019) Comparative antifungal susceptibility analyses of Cryptococcus neoformans VNI and Cryptococcus gattii VGII from the Brazilian Amazon Region by the Etest, Vitek 2, and the clinical and laboratory standards institute broth microdilution methods. Med Mycol 57:864–873. https://doi.org/10.1093/mmy/myy150
Saraiva MGG, Santos ECS, Saraceni V, Rocha LLS, Monte RL, Albuquerque BC, Bastos MS et al (2015) Epidemiology of infectious meningitis in the State of Amazonas. Brazil Rev Soc Bras Med Trop 48(Suppl I):79–86. https://doi.org/10.1590/0037-8682-0116-2014
Rocha DFS, Cruz KS, Santos CSS, Menescal LSF, Neto JRS, Pinheiro SB et al (2018) MLST reveals a clonal population structure for Cryptococcus neoformans molecular type VNI isolates from clinical sources in Amazonas, Northern-Brazil. Plos-one 13(6):1–15. https://doi.org/10.1371/journal.pone.0197841
Alves MJ, do Nascimento IS, Cruz KS, Menescal VVF, Menescal LSF, Silva LSC, et al (2022) Cryptococcosis in HIV/AIDS patients in northern Brazil: clinical aspects, molecular types and isolation of agents from environmental samples associated with patients. Trop Med Int Health 00:1– 10.https://doi.org/10.1111/tmi.13737
Meyer W, Castañeda A, Jackson S, Huynh M, Castañeda E (2003) Molecular typing of Ibero American Cryptococcus neoformans isolates. Emerg Infect Dis 9:189–195. https://doi.org/10.3201/eid0902.020246
Meyer W, Aanensen DM, Boekhout T, Cogliati M, Diaz MR, Esposto MC et al (2009) Consensus multi-locus sequence typing scheme for Cryptococcus neoformans and Cryptococcus gattii. Med Mycol 47(6):561–570. https://doi.org/10.1080/13693780902953886
Espinel-Ingroff A, Chowdhary A, Cuenca-Estrella M et al (2012) Cryptococcus neoformans-Cryptococcus gattii species complex: an international stud of wild-type susceptibility endpoint distributions and epidemiological cutoff values for amphotericin B and flucytosine. Antimicrob Agents Chemother 56:3107–3113. https://doi.org/10.1128/AAC.06252-11
Espinel-Ingroff A, Aller AI, Canton E et al (2012) Cryptococcus neoformans-Cryptococcus gattii species complex: an international study of wildtype susceptibility endpoint distributions and epidemiological cutoff values for fluconazole, itraconazole, posaconazole, and voriconazol. Antimicrob Agents Chemother 56:5898–5906. https://doi.org/10.1128/aac.01115-12
Andrade-Silva LE, Ferreira-Paim K, Ferreira BT, Vilas-Boas A, Mora DJ, Manzato VM (2018) Genotypic analysis of clinical and environmental Cryptococcus neoformans isolates from Brazil reveals the presence of VNB isolates and a correlation with biological factors. PLoS ONE 5:1–21. https://doi.org/10.1371/journal.pone.0193237
Silva FA (2020) Genética populacional e epidemiologia molecular de isolados brasileiros clínicos e ambientais de Cryptococcus neoformans. 95 f. Dissertação (Mestrado em Pesquisa Clínica em Doenças Infecciosas) - Instituto Nacional de Infectologia Evandro Chagas, Fundação Oswaldo Cruz, Rio de Janeiro
Reis RS, Bonna ICF, Antonio IMS, Pereira AS, do Nascimento AS, Ferraris FK, et al (2021) Cryptococcus neoformans VNII as the main cause of cryptococcosis in domestic cats from Rio de Janeiro, Brazil. J Fungi 2021(7):980. https://doi.org/10.3390/jof711098
Trilles L, Lazera M, Wanke B, Theelen B, Boekhout T (2003) Genetic characterization of environmental isolates of the Cryptococcus neoformans species complex from Brazil. Med Mycol 41:383–390. https://doi.org/10.1080/1369378031000137206
Beale MA, Sabiiti W, Robertson EJ, Fuentes-Cabrejo KM, O’Hanlon SJ, Jarvis JN, Loyse A, Meintjes G, Harrison TS, May RC, Fisher MC, Bicanic T (2015) Genotypic diversity is associated with clinical outcome and phenotype in cryptococcal meningitis across Southern Africa. PLoS Negl Trop Dis 9(6):e0003847. https://doi.org/10.1371/journal.pntd.0003847
Khayhan K, Hagen F, Pan W, Simwami S, Fisher MC, Wahyuningsih R, Chakrabarti A, Chowdhary A, Ikeda R, Taj-Aldeen SJ, Khan Z, Ip M, Imran D, Sjam R, Sriburee P, Liao W, Chaicumpar K, Vuddhakul V, Meyer W, Trilles L, van Iersel LJ, Meis JF, Klaassen CH, Boekhout T (2013) Geographically structured populations of Cryptococcus neoformans variety grubii in Asia correlate with HIV status and show a clonal population structure. PLoS One 8(9):e72222. https://doi.org/10.1371/journal.pone.0072222
Chen YH, Yu F, Bian ZY, Hong JM, Zhang N, Zhong (2018) Multilocus sequence typing reveals both shared and unique genotypes of Cryptococcus neoformans in Jiangxi Province China. Sci Rep 8:1495. https://doi.org/10.1038/s41598-018-20054-4
Thanh LT, PhanTH RS, Nguyen TM, Duong AV, Dacon C et al (2019) Multilocus sequence typing of Cryptococcus neoformans var. grubii from Laos in a regional and global contexto. Med Mycol 57:557–565. https://doi.org/10.1093/mmy/myy105
Ngamskulrungroj P, Gilgado F, Faganello J, Litvintseva AP, Leal AL, Tsui KM, Mitchell TG, Vainstein MH, Meyer W (2009) Genetic diversity of the Cryptococcus species complex suggests that Cryptococcus gattii deserves to have varieties. PLoS One 4(6):e5862. https://doi.org/10.1371/journal.pone.0005862. Erratum in: PLoS One 2009;4(7). https://doi.org/10.1371/annotation/3037bb69-1b8e-4d99-b169-afdf4b74ace2. Erratum in: PLoS One 2009;4(7). https://doi.org/10.1371/annotation/348c3375-3918-4e41-bb8c-27aa15d2bdc4
Rhodes J, Desjardins CA, Sykes SM, Beale MA, Vanhove M, Sakthikumar S et al (2017) Tracing genetic exchange and biogeography of Cryptococcus neoformans var. grubii at the global population level. Genetics 207:327–346. https://doi.org/10.1534/genetics.117.203836
Casadevall A, Freij JB, Hann-Soden C, Taylor J (2017) Continental drift and speciation of the Cryptococcus neoformans and Cryptococcus gattii species complexes. Msphere 2(2):e00103-e117. https://doi.org/10.1128/msphere.00103-17
Vélez N, Scandón P (2020) Multilocus sequence typing (MLST) of clinical and environmental isolates of Cryptococcus neoformans and Cryptococcus gattii in six departments of Colombia reveals high genetic diversity. Rev Soc Bras Med Trop 53:e20190422. https://doi.org/10.1590/0037-8682-0422-201
Brito-Santos F, Trilles L, Firacative C, Wanke B, Carvalho-Costa FA, Nishikawa MM (2020) Indoor dust as a source of virulent strains of the agents of cryptococcosis in the Rio Negro Micro-Region of the Brazilian Amazon. Microorganisms 8:682. https://doi.org/10.3390/microorganisms8050682.DOI:10.3390/microorganisms8050682
Lozano-Chiu M, Paetznick VL, Ghannoum MA, REX JH, (1998) Detection of resistance to amphotericin B among Cryptococcus neoformans clinical isolates: performances of three different media assessed by using E-Test and National Committee for Clinical Laboratory Standards M27-A methodologies. Jour clin micro 36(10):2817–2822. https://doi.org/10.1128/jcm.36.10.2817-2822.1998
Horta JÁ, Faganello J, Silva LKR, Oliveira LT, Santurio JM, Vainstein MH (2005) Susceptibility to heat and antifungal agents of Cryptococcus neoformans var. neoformans (serotype d) isolated from eucalyptus spp in Rio Grande do Sul, Brazil. Brazilian J Microbiol 36:1–6. https://doi.org/10.1590/S1517-83822005000100001
Lockart SR, Ghrannoum MA, Alexander BD (2017) Establishment and use of epidemiological cutoff values for molds and yeasts by use of the clinical and laboratory standards institute m57 standard. J Clin Microbiol 55(5):1262–1268. https://doi.org/10.1128/jcm.02416-16
Silva BK, Freire AK, Bentes AD, Sampaio IL, Santos LO, Santos MS, Souza JVB (2012) Characterization of clinical isolates of the Cryptococcus neoformans-Cryptococcus gattii species complex from the Amazonas State in Brazil. Rev Iberoam Micol 29(1):40–43. https://doi.org/10.1016/j.riam.2011.05.003
Almeida AM, Matsumoto MT, Baeza LC, de Oliveira E, Silva RB, Kleiner AA, Melhem Mde S, Mendes Giannini MJ, Laboratory Group on Cryptococcosis (2007) Molecular typing and antifungal susceptibility of clinical sequential isolates of Cryptococcus neoformans from Sao Paulo State, Brazil. FEMS Yeast Res 7(1):152–164. https://doi.org/10.1111/j.1567-1364.2006.00128.x
Longley N, Muzoora C, Taseera K, Mwesigye J, Rwebembera J, Chakera A et al (2008) Dose response effect of high-dose fluconazole for HIV-associated cryptococcal meningitis in Southwestern Uganda. CID 47:1556–1561. https://doi.org/10.1086/593194
Lia M, Liao Y, Chena M, Pana W, Weng L (2012) Antifungal susceptibilities of Cryptococcus species complex isolates from AIDS and non-AIDS patients in Southeast China. Braz J Infect Dis 16(2):175–179. https://doi.org/10.1016/S1413-8670(12)70301-X
Naicker SD, Magobo RE, Maphanga TG, Firacative C, Schalkwyk EV, Monroy-Nieto J et al (2021) (2021) Genotype, antifungal susceptibility, and virulence of clinical South African Cryptococcus neoformans strains from national surveillance, 2005–2009. J Fungi 7:338. https://doi.org/10.3390/jof7050338
Matos CS, Andrade AS, Oliveira NS, Barros TF (2012) Microbiological characteristics of clinical isolates of Cryptococcus spp. in Bahia, Brazil: molecular types and antifungal susceptibilities. Eur J Clin Microbiol Infect Dis 31:1647–1652. https://doi.org/10.1007/s10096-011-1488-3
Molloy SF, Kanyama C, Heyderman RS, Loyse A, Kouanfack C, Chanda D et al (2018) Antifungal combinations for treatment of cryptococcal meningitis in Africa. N Engl J Med 2018:378. https://doi.org/10.1056/NEJMoa1710922
Souza LKH,1, Junior AHS, Costa CR, Faganello J, Vainstein MH, Chagas ALB, et al (2009) Molecular typing and antifungal susceptibility of clinical and environmental Cryptococcus neoformans species complex isolates in Goiania, Brazil. Mycoses 53:62–67.https://doi.org/10.1111/j.1439-0507.2008.01662.x
Cichon M, Vicente VA, Muro MD, Bordignon GPF, Queiroz-Telles F (2011) Isolation of Cryptococcus neoformans from environmental samples of Curitiba and metropolitan region (Paraná, Brazil), and susceptibility antifungal testing. RBAC 43(3):176–179
Alves GS, Freire AK, Bentes Ados S, Pinheiro JF, de Souza JV, Wanke B, Matsuura T, Jackisch-Matsuura AB (2016) Molecular typing of environmental Cryptococcus neoformans/C. gattii species complex isolates from Manaus, Amazonas, Brazil. Mycoses 59(8):509–515. https://doi.org/10.1111/myc.12499
Acknowledgements
The authors are grateful for the funding received from Fundação de Amparo à Pesquisa do Estado do Amazonas (Public call No. 002/2018—UNIVERSAL AMAZONAS/ FAPEAM) and PROEP/LABS/ILMD Fiocruz Amazônia LDMAIS. The authors also thank the “Laboratório Temático de Biologia Molecular – LTBM”, “Plataforma de genômica do Instituto de Pesquisa Leônidas e Maria Deane – ILMD/FIOCRUZ” and “Plataforma de genômica do Instituto Oswaldo Cruz- IOC Manguinhos/FIOCRUZ” for their technical support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest statement
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Communicated by Responsible Editor: Celia Maria de Almeida Soares.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Alves, M.J., Cruz, K.S., Alves, G.S.B. et al. Antifungal susceptibility and multilocus sequence typing (MLST) of Cryptococcus neoformans complex from HIV-associated cryptococcal meningitis patients in Manaus, Amazonas, Brazil. Braz J Microbiol 55, 2603–2611 (2024). https://doi.org/10.1007/s42770-024-01378-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-024-01378-y