Abstract
Mycoplasma hyopneumoniae (M. hyopneumoniae) is a primary agent of porcine enzootic pneumonia, a disease that causes significant economic losses to pig farming worldwide. Commercial vaccines induce partial protection, evidencing the need for a new vaccine against M. hyopneumoniae. In our work, three chimeric proteins were constructed, composed of potentially immunogenic domains from M. hyopneumoniae proteins. We designed three chimeric proteins (Q1, Q2, and Q3) based on bioinformatics analysis that identified five potential proteins with immunogenic potential (MHP418, MHP372, MHP199, P97, and MHP0461). The chimeric proteins were inoculated in the murine model to evaluate the immune response. The mice vaccinated with the chimeras presented IgG and IgG1 against proteins of M. hyopneumoniae. There was induction of IgG in mice immunized with Q3 starting from 30 days post-vaccination, and groups Q1 and Q2 showed induction at 45 days. Mice of the group immunized with Q3 showed the production of IgA. In addition, the mice inoculated with chimeric proteins showed a proinflammatory cytokine response; Q1 demonstrated higher levels of TNF, IL-6, IL2, and IL-17. In contrast, animals immunized with Q2 showed an increase in the concentrations of TNF, IL-6, and IL-4, whereas those immunized with Q3 exhibited an increase in the concentrations of TNF, IL-6, IL-10, and IL-4. The results of the present study indicate that these three chimeric proteins can be used in future vaccine trials with swine because of the promising antigenicity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mycoplasma hyopneumoniae (M. hyopneumoniae) is the primary agent of enzootic pneumonia (EP) in swine, a chronic respiratory worldwide disease, and one of the primary agents involved in the porcine respiratory disease complex [1]. It is mainly transmitted through nose-to-nose contact and indirectly via aerosols among susceptible pigs or airborne transmission among farms [2].
The control of M. hyopneumoniae infections in pig farms can be accomplished in many ways, namely by optimizing management and biosecurity practices and housing conditions, vaccination, and antimicrobial medication [3]. Vaccination is a helpful strategy to control M. hyopneumoniae and widely used worldwide [1]. The commercial vaccines commonly used are composed of inactivated bacterial whole cells, but they do not stimulate a potent immune response against M. hyopneumoniae infection [4, 5] and do not prevent colonization by M. hyopneumoniae [3].
In vitro M. hyopneumoniae cultivation is fastidious and laborious and requires an enriched medium [6]. Reverse vaccinology is being used to predict vaccine targets, especially for microorganisms that are difficult to cultivate in the laboratory [7]. Data produced using the complete genome of seven M. hyopneumoniae strains (7448, 7442, 232, 168, 168-L, J, and KM014) [8,9,10,11,12] allowed the analysis of coding regions associated with pathogenesis. Some of these proteins are adhesins (P97, P102, and MHP0461), lipoprotein (MHP372), membrane protein (P46), and a hypothetical protein (MHP418) [13,14,15].
The use of chimeric proteins constructed from immunogenic regions of antigens has become an attractive strategy in the construction of vaccine candidates, mainly due to the possibility of having several immunodominant epitopes from a single protein [16, 17]. Also, appropriate adjuvants help to improve the immunogenicity of these proteins. The B subunit of the thermolabile enterotoxin (LTB) of Escherichia coli stimulates both mucosal and systemic immunity, mainly by increasing the antigen presentation by dendritic cells to T cells [18].
The chimeric proteins used in this work were designed based on previous results from our group that investigated the genetic characteristics of Brazilian field M. hyopneumoniae strains [19, 20]. In addition, we also used the antigens identified from previous works [13, 15, 18, 21,22,23,24,25,26,27,28] to design three chimeric proteins to access their immunogenicity in a mouse model as a preclinical evaluation.
Material and methods
Design and expression of chimeric proteins
Three chimeric proteins (Q1, Q2, and Q3) were designed using five potential M. hyopneumoniae immunogenic proteins: MHP418, MHP372, MHP199, P97, and MHP0461. These proteins' surface or immunogenic protein domains were grouped in tandem through the rigid protein linker (EAAAKEAAAK). A domain representing the B subunit of thermolabile enterotoxin (LBT) of E. coli was added to each chimera. For Q1 construction, the proteins MHP418, MHP372, and MHP 199 were evaluated for most surface regions, using the Miyazawa hydrophobicity scale [29]. For Q2 and Q3 construction, antigenic regions of the proteins P46 and MHP0461 were selected using structural models of these proteins. Thus, proteins P46 and MHP0461 had their structural models produced through homology modeling using the I-TASSER software [30,31,32], with subsequent stereochemical and energetic evaluation through the Rampage [33] and ProSa programs [34]. Both models had their antigenic regions evaluated; for Q2, we used the ELLIPro program [35], and for Q3, we employed the Discotope [36] and Epitopia programs [37]. In this study, the gene sequences from the complete genome of M. hyopneumoniae strain 7448 (GenBank: NC 007332.1) and the protein sequence of LTB (GenBank: ABI14553.1) were used for all bioinformatics analyses and synthetic gene design. A gene was designed for the synthesis of the LTB protein (reLTB) to function as a cross-reactivity control of the experiments since this protein is present in the structure of E. coli bacteria, which mainly occur in the intestinal tract of endotherms [18]. The proteins' theoretical molecular weight and the isoelectric point were calculated according to a previous report [38].
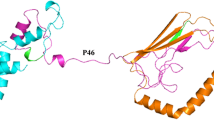
The protein domains of the chimeras are represented in Fig. 1. The synthetic genes were synthesized and cloned into the pET29a plasmid (Genscript, NJ, USA), and the N-terminus 6 × histidine-tagged chimeric proteins were expressed in Escherichia coli strain BL21-CodonPlus (DE3)—RIL (Agilent Technologies). Protein purification was performed by affinity chromatography using a HisTrap crude FF (GE), according to the manufacturer's recommendations in an FPLC system (AKTA TM purifier). The chimeric proteins were checked using a 12% SDS-PAGE gel and Western blot (WB) analysis. For Western blot assays, we used 6 × His Tag antibodies (Sigma) at 1:2,000 and HRP conjugated anti mouse IgG (Sigma) at 1:5,000.
Linear sequence and recombinant protein domains obtained by the modeling program I Tasser. A and B represent Chimera 1. C and D represent chimera 2. E chimera 3. In parentheses, the amino acid sequences of native proteins used to form chimeric proteins are shown. #(MHP046172-135), ##(MHP0461286-312). Modeling program I Tasser [30,31,32]. The Q3 protein could not solve the structure by the modeling program
Chimeric protein immunogenicity assessment
Four protein formulations were produced (Table 1). As a positive control, proteins from whole bacterial cell lysates of M. hyopneumoniae strain J were used (MhJ, GenBank: NC 007295.1). The MhJ was cultured in Friis medium as previously described [39]. The protein concentrations were quantified using the Bradford protein assay (BioRad, USA). Phosphate-buffered saline (PBS) and reLTB protein were used as negative and adjuvant controls, respectively.
The immunization assay was performed using male BALB/c mice aged 4–6 weeks, divided into six groups (Table 1). The animals were subcutaneously inoculated with three doses at 15-day intervals. Blood samples were collected from the tail at 0, 15, 30, and 45 days after the first immunization (d.p.i.). On day 45, the mice were euthanized, and the spleen was aseptically removed to perform the cytokine assay. The animals were obtained from the local breeding facility (Viçosa, MG, Brazil) and maintained under controlled environmental conditions. All experiments were performed according to the local Ethics Committee on Animal Experimentation (Protocol Number 54/2015).
Detection of IgA, IgG, IgG1, and IgG2a antibodies against M. hyopneumoniae proteins
A 96-well Maxisorp microtiter plate (Nunc) was coated with 1 ug/well of MhJ protein or recombinant proteins in 100 uL of carbonate-bicarbonate buffer and incubated on a 96-well ELISA microplate at 4 °C for 14 h. After incubation for 14 h at 4 °C, the wells were washed and incubated with 5% BSA in PBS. Subsequently, the plates were washed three times with PBS-Tween 0.05%, and 100 μL of diluted serum samples at 1:50 was added and incubated at 37 °C for 1 h. The plates were washed three times with PBS-T, followed by incubation for 1 h at 37 °C with 100 uL diluted anti-mouse IgG antibody (1:6,000), anti-mouse IgG1 (1:3,000), anti-mouse IgG2 (1:3,000), or anti-mouse IgA (1:2,000) from Sigma. Color development was achieved by adding a solution containing H2O2, OPD, and 0.1 M citrate buffer (pH 5.0). The readout was performed at a wavelength of 492 nm. Both positive and negative controls were added to all plates, and each sample was tested in triplicate.
Evaluation of antibody induction using Western blot
Serum samples 45 d.p.i were used to analyze the reactivity against the proteins from whole M. hyopneumoniae cell lysates and those of Mycoplasma hyorhinis by Western blot (WB). The proteins were analyzed in SDS-PAGE and electrophoretically transferred onto a nitrocellulose membrane. The membrane was then blocked with PBS-BSA for 14 h at room temperature. After washing with PBS-T twice, the membrane was incubated with serum at 1:100 for 2 h at room temperature. After five washes, it was incubated with anti-mouse IgG (Sigma, 1:6,000), and after another five washes, the membrane was incubated with a DAB solution. To control for the cross-reactivity of antibodies present in mouse sera, an isolated sample of pig lung was use. This sample tested positive for Mycoplasma hyorhinis and negative for M. hyopneumoniae.
Cytokines quantification
Spleens were aseptically removed, macerated with 10 mL of RPMI medium, and centrifuged at 250 g at 4ºC for 5 min. The pellet was resuspended in 5 mL of lysis buffer (0.17 M Tris–HCl, 0.16 M NH2Cl) and incubated for 5 min at room temperature. Cells were washed with RPMI medium supplemented with 10% fetal bovine serum, centrifuged, and then adjusted to 5 × 105 cells/well. These cells were stimulated with chimeric proteins (Q1, Q2, and Q3), reLTB, PBS, and MhJ for 48 h at 37 °C in a 5% CO2 incubator. Cytokine concentrations were evaluated in the supernatants using the BD cytometric bead array (CBA) mouse Th1/Th2/Th17 Cytokine Kit and BD FACSVerse flow cytometer.
Statistical analysis
Statistical analyses were performed by comparing the responses of antibodies and cytokines between the groups inoculated with chimeras in relation to the group vaccinated with reLTB. Additionally, the group immunized with MhJ was compared with that inoculated with PBS. One-way analysis of variance (ANOVA) followed by Tukey's multiple comparisons test was performed using GraphPad Prism 8.4.3 (GraphPad Software, San Diego, California USA, www.graphpad.com).
Results
Characterization of chimeras
Chimeric proteins Q1, Q2, and Q3 showed theoretical molecular weights of 104.18, 88.41, and 83.79 kDa, respectively. The most antigenic regions of proteins with immunogenic potential against M. hyopneumoniae were selected according to the bioinformatics analysis. The proteins MHP418, MHP372, and MHP199 (P102) showed the residues 101–305, 500–749, and 500–749, respectively, of amino acids most likely to be exposed to the solvent, according to the hydrophobicity profile of the amino acids. Using the ELLIPro server, the proteins P46 and MHP0461 showed the regions of excellent flexibility and solvent exposure among the residues 90–300 and 90–348 of amino acids, respectively. Using the Discotope and Epitopia server, the protein P46 showed the residues 80–135 and 286–312 of amino acids, and the protein MHP0461 showed the residues 72–135 and 286–312 of amino acids. The C-terminal region of the protein P97 was used in the chimera composition, considering that this region has already been described with a robust immunogenic capacity [40].
Chimeras 1 and 2 were detected in the insoluble fraction, whereas chimera 3 and reLTB were detected in both soluble and insoluble fractions. The expression parameters were evaluated by varying the IPTG concentration, temperature, and induction time. The optimal induction time for all three proteins was 4 h, and the optimal IPTG concentrations were 0.75 mM for Q1 and Q2, 0.25 mM for Q3, and 0.5 mM for reLTB. Figure 2 shows the results of the purification of the chimeras and WB.
Recombinant chimera protein expression, purification, and characterization. A SDS-PAGE 12%: Q3 (chimera 3 purified), Q2 (chimera 2 purified), Q1 (chimera 1 purified) and MM (protein ladder). The arrow indicates the chimeras. B Western blot (WB): Q3 (chimera 3 purified), Q2 (chimera 2 purified), Q1 (chimera 1 purified), and MM (protein ladder)
Induction of antibodies against recombinant proteins
The immunogenicity of the recombinant proteins was evaluated through the detection of antibodies specific to each chimera at 45 d.p.i. The animals in the groups Q1 and MhJ showed IgG, IgG1, and IgG2a induction against Q1 in comparison to reLTB and PBS groups, respectively (Suppl. Fig. S1A-C). The group immunized with Q2 showed increased IgG, IgG1, and IgG2a binding to Q2 compared to the reLTB-immunized group (Suppl. Fig. S1D-F). The animals immunized with Q3 only showed increased IgG2a binding to Q3 compared to the reLTB group (Suppl. Fig. S1I). The animals that were vaccinated with whole-cell proteins of M. hyopneumoniae did not present significant seroconversion specific to Q3 compared to the PBS group (Suppl. Fig. S1G-I). The animals immunized with M. hyopneumoniae whole-cell proteins showed IgG, IgG1, and IgG2a binding to Q1 and IgG1 binding to Q2 compared to those of the PBS group (Suppl. Fig. S1A-C and S1E).
Induction of antibodies against whole-cell proteins of M. hyopneumoniae
Next, we evaluated the antibodies specific to M. hyopneumoniae proteins in the serum of mice at different groups and time points. As expected, all evaluated immunoglobulins were significantly higher in MhJ-immunized mice (positive control group) compared to mice inoculated only with PBS (negative control group) after 15 d.p.i. (Fig. 3).
IgG and IgA antibodies against MhJ proteins. Mice were immunized with recombinant proteins, and immunoglobulin levels (Ig) were evaluated at 0, 15, 30, and 45 d.p.i. IgG (A) IgG1 (B) and IgG2a (C) against whole-cell proteins as ELISA antigens were evaluated in the six immunized groups. IgA antibodies against Mhp proteins (D). Mice were immunized with recombinant proteins, and serum IgA levels against MhJ proteins were evaluated at 45 d.p.i. in the six immunized groups. Results are expressed as S/P ratio: (OD sample – OD negative control) / (OD positive control – OD negative control) with mean ± standard error of the mean (SEM) for individual values obtained in each experimental group. *p < 0.05; ** < 0.01; ***p < 0.001; ****p < 0.0001
Specific IgG antibodies against MhJ in the Q1, Q2, and Q3 groups were observed at later time points (30 and 45 d.p.i.). Both IgG and IgG1 levels increased in the Q2 and Q3 groups at day 30 p.i. and in all Q1, Q2, and Q3 groups at day 45 p.i., compared to the reLTB group (Fig. 3A and B). The IgG2a level was significantly higher in the Q1, Q2, and Q3 groups at day 30 p.i. but only in the Q3 group at day 45 p.i. (Fig. 3C).
In addition to IgG antibodies, anti-MhJ IgA was evaluated in serum samples at 45 d.p.i. The IgA specific to MhJ was significantly higher in the Q3 group than in the reLTB group (Fig. 3D).
Western blot analysis showed distinct patterns among the groups against M. hyopneumoniae whole-cell proteins from strain J (Fig. 4A). Sera from the Q1 group exhibited markings on proteins within the range of 35–48 and 48–63 kDa. Sera from the Q2 and Q3 groups displayed markings on proteins within the range of 25–35, 35–48, and 75–100 kDa. The positive control exhibited markings on several proteins with several molecular weights, whereas the negative control group presented only weak nonspecific markings. There was no reaction from the sera of the animals from the Q1, Q2, and Q3 groups with M. hyorhinis proteins (Fig. 4B).
Induction of IgG 45 d.p.i against whole bacterial cell lysates from M. hyorhinis and MhJ assessed by Western blot. A Serum reactivity against whole-cell protein of MhJ. B Serum reactivity against whole proteins of M. hyorhinis. MM (protein ladder), Q1 (serum from Q1 group), Q2 (serum from Q2 group), Q3 (serum from Q3 group), PBS (serum from PBS group), LTB (serum from reLTB group), MhJ (serum from Mh strain J group)
Evaluation of the cytokine profile
To gain insights into the immune response induced by the chimeras, cytokine quantification was performed in supernatants from splenocyte cultures derived from immunized animals. Animals immunized with M. hyopneumoniae whole-cell proteins from strain J (MhJ) showed increased production of TNF, IL-6, IL-2, and IL-4 after in vitro stimulation with MhJ (Suppl. Fig. S2). Additionally, within this group, an elevated IL-10 level was observed following stimulation with all chimeras, as compared to reLTB (Suppl. Fig. 2C). Among the cytokines evaluated in the different stimuli, the TNF presented the highest levels (Suppl. Fig. 2H).
Splenocytes derived from Q1-immunized animals stimulated with whole-cell proteins of M. hyopneumoniae (MhJ) presented higher concentrations of TNF, IL-6, IL-10, IL-2, IFN-γ, IL-17A, and IL-4 when compared to the negative control (PBS) (Fig. 5A-G). Upon stimulation with Q1, an increase in IL-6, IL-2, and IL-17 was observed compared to reLTB (Fig. 5B, D, and F). Regarding the stimulus represented by whole bacterial cell lysates from M. hyopneumoniae and Q1 protein in splenocytes, the results showed an increase in the levels of TNF, IL-6, IL2, and IL-17 (Fig. 5).
Cytokine profile induced in the Q1 group. A TNF level; B IL-6 level; C IL-10 level; D IL-2 level; E INF-γ level; F IL-17A level; G IL-4 level. reLTB, Q1, MHJ (whole-cell protein), and PBS are the stimuli used in the culture of splenocytes from mice inoculated with Q1 chimera protein. Results are expressed as the mean ± standard deviation (SD) of the individual values obtained for each experimental group. H Cytokine array heat map. # represents a significant difference in TNF compared to other cytokines evaluated in the same group. & represents a significant difference between IL-6 and IL-10 compared to other cytokines evaluated in the same group. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001
Splenocytes derived from the mice vaccinated with Q2 presented higher concentrations of TNF, IL-6, and IL-4 when stimulated with whole-cell proteins of M. hyopneumoniae in comparison to those stimulated with PBS (Fig. 6A, B, and G). Stimuli with Q2 induced elevated levels of TNF, IL-6, IL-2, IFN-γ, IL-17A, and IL-4 compared to reLTB (Fig. 6). As TNF was the major cytokine produced by Q2-immunized mice (Fig. 6H), an increase in TNF, IL-6, and IL-4 levels was observed when splenocytes from mice vaccinated with Q2 were stimulated with either the MPH whole-cell proteins or Q2 stimuli.
Cytokine profile induced in the Q2 group. A TNF level; B IL-6 level; C IL-10 level; D IL-2 level; E INF-γ level; F IL-17A level; G IL-4 level. reLTB, Q2, MhJ (whole-cell proteins of MhJ), and PBS are the stimuli used in the culture of splenocytes from mice inoculated with Q2 chimera. Results are expressed as the mean ± standard deviation (SD) of the individual values obtained for each experimental group. H Cytokine array heat map. # represents a significant difference in TNF compared to other cytokines evaluated in the same group. & represents a significant difference in IL-6 in comparison to other cytokines evaluated in the same group. $ represents a significant difference in IFN-γ in comparison to other cytokines evaluated in the same group. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001
The Q3-immunized animals presented splenocytes secreting higher concentrations of all assessed cytokines (Fig. 7A-G). Upon stimulation with whole-cell proteins of M. hyopneumoniae, the splenocytes secreted higher levels of TNF, IL-6, IL-10, and IL-4 (Fig. 7A-C, G). Thus, an increase in TNF, IL-6, IL-10, and IL-4 was detected when splenocytes from mice immunized with Q3 were stimulated with either the MPH or Q3 stimuli (Fig. 7).
Cytokine profile induced in the Q3 group. A TNF level; B IL-6 level; C IL-10 level; D IL-2 level; E INF-γ level; F IL-17A level; G IL-4 level. reLTB, Q3, MhJ (whole-cell proteins of MhJ), and PBS are the stimuli used in the culture of splenocytes. Results were expressed as the mean ± standard deviation (SD) of the individual values obtained for each experimental group. H Cytokine array heat map. # represents a significant difference in TNF compared to other cytokines evaluated in the same group. & represents a significant difference in IL-6 in comparison to other cytokines evaluated in the same group. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001
Discussion
Despite the worldwide use of bacterins in controlling M. hyopneumoniae, they provide only partial protection, highlighting the need for more efficient vaccines [6]. In this sense, chimeric vaccines may represent a better alternative to control M. hyopneumoniae. Drawing from the outcomes of advanced reverse vaccinology, immunogenicity analyses of molecules and specific vaccine regions can be conducted via in silico analysis, facilitating an initial selection of potential vaccine candidates. In this context and intending to produce an immunogenic vaccine against M. hyopneumoniae, in this work, we developed and evaluated the immunogenicity of three protein chimeras designed from M. hyopneumoniae proteins. These proteins were subsequently expressed in a bacterial system. When vaccinated in the murine model, the three chimeras showed antigenicity and could elicit specific IgG antibodies. Also, the sera from animals immunized with whole-cell proteins of M. hyopneumoniae exhibited reactivity against Q1 and Q3. Galli and collaborators [26] showed that the proteins present in Q1 (MHP418, MHP372, MHP199, or P102) are antigenic and react with convalescent pig sera collected from a commercial herd chronically infected by M. hyopneumoniae. Upon analyzing the antibody production against the whole-cell protein of M. hyopneumoniae, we observed IgG production in the Q1, Q2, and Q3 groups. Additionally, when MHP418 and MHP372 were inoculated in mice, they stimulated the production of antibodies against M. hyopneumoniae [26]. These findings underscore the capability of the antibodies produced by the three chimeras developed in this study to interact with M. hyopneumoniae proteins.
The affinity of the antibodies induced by the chimeras against whole bacterial cell lysates from M. hyopneumoniae was also assessed using WB. The antibodies induced by Q1 showed strong reactivity against one protein in the range of 35–48 kDa, probably protein P102 in its cleaved fraction P42, and one protein in the range of 48–63 kDa, probably protein P60, another cleaved fraction of P102 [41]. The sera from Q2 and Q3 groups showed the same patterns of antibody reactivity. They reacted against a protein between 25 and 35 kDa, probably protein P97 in its fraction P28, a protein between 35 and 48 kDa, possibly protein P42, and a protein between 75 and 100 kDa, probably protein P97 [42]. These results indicate that the chimeras possess B lymphocyte epitopes common to M. hyopneumoniae. We suggest that the induction of antibodies by Q1, Q2, and Q3 is related to their regions corresponding to the antigen determinant of M. hyopneumoniae.
Our work quantified the concentration of cytokines associated with Th1, Th2, and Th17 responses using CBA kit analyses. Animals inoculated with the chimeras had heterogeneous levels of increased cytokines in each group when the splenocytes were stimulated with whole-cell proteins of M. hyopneumoniae or chimeras. We therefore highlight the significative response of cytokines from splenocytes of groups inoculated with Q1, Q2, or Q3, where a statistical difference was noted between the stimulation by whole-cell proteins of M. hyopneumoniae and purified chimera versus control groups (PBS and reLTB, respectively).
A common characteristic of the vaccinated Q1, Q2, and Q3 mice was the increased production of TNF. Generally, TNF is mainly produced by activated macrophages, T lymphocytes, and natural killer (NK) cells [43]. Proinflammatory cytokines, such as TNF, play a pivotal role in the pathogenesis of enzootic pneumonia [44]. Recently, a study showed that vaccination using bacterins induced a strong TNF-α+ in pigs, and there was a relation with immune protection [45]. The IL-6 showed another cytokine increase in the three experimental groups (Q1, Q2, and Q3); it is a proinflammatory cytokine with a central role in the integration of immune defense against infections and functions in both innate and adaptive immune responses against pathogens [46, 47]. Various cell types, such as mast cells, macrophages, dendritic cells, and T and B cells, can express this cytokine [47]. Both IL-6 and TNF were cytokines that showed the highest concentrations in chimera groups after immunization (Figs. 5H, 6H, and 7H). In addition, IL-4 was increased in the Q2 and Q3 groups. The IL-4 is a cytokine that plays a role in developing Th2 responses and promotes the growth and activation of B cells to produce IgG1 [47]. Figure 4B shows an increase in IgG1 levels at 30 days post-vaccination in the Q2 and Q3 groups, and at 45 d.p.i, the mice in all three experimental groups displayed high levels of IgG1. Yang and collaborators [47] showed that the IL-4 and IL-6 fused gene as an adjuvant could significantly improve the protection efficacy of vaccination against M. hyopneumoniae.
The splenocytes from the Q1 group exhibited increased levels of IL2 and IL-17. The species M. hyopneumoniae is a mucosal bacterial species that attaches to the epithelial cilia in the lower airways [48], and studies have suggested that the Th17 response plays a role in the early stage of M. hyopneumoniae infection [49, 50]. In addition, the mice from the Q1 group showed an IL-2 increase. The IL-2 may be related to protection against M. hyopneumoniae infection in vaccinated and infected pigs [51].
Splenocytes obtained from animals vaccinated with Q3 exhibited IL-10 expression upon stimulation. The IL-10 is produced by various cell types, including macrophages, CD8 + and CD4 + T cells, γδ-T cells, NK cells, B cells, and dendritic cells. Macrophages are the major source of IL-10, and this cytokine is a crucial anti-inflammatory cytokine that can inhibit proinflammatory responses [52, 53]. Pathogenic bacteria induce IL-10 expression as a strategy to evade the host immune response [53]. This suggests that evading or delaying the host's immune response is a potential mechanism to facilitate persistent infection [54].
In vaccinated pigs, there was a higher concentration of IL-10-producing cells in the bronchial lymph nodes. Vaccination-induced IL-10 secretion may also lead to a reduced influx of macrophages in the bronchoalveolar lymphoid tissue compared to non-vaccinated pigs after experimental infection with M. hyopneumoniae [22]. Further, IL-10 has an anti-proliferative effect, preventing the pathological effects of inflammatory cytokines [21]. This mechanism may also contribute to the reduction of lung tissue damage observed in vaccinated animals following infection [15, 20].
Some studies performed vaccination assays using chimera proteins against M. hyopneumoniae. Oliveira et al. [25] describe the construction of a multi-antigen chimera composed of four antigens, namely P97R1, P46, P95, and P42. This protein was tested as a vaccine in a murine model. Conceição and collaborators [28] tested a recombinant chimera as a vaccine composed of the R1 repeat region of P97 adhesin and evaluated the immune response in animal models. The studies cited above showed positive results as a vaccine candidate against M. hyopneumoniae in mouse models or pigs. However, there was no significant protection against M. hyopneumoniae infection in pigs when testing a chimeric protein composed of the C-terminal portion of P97, heat shock protein P42, and NrdF as vaccine candidates [18]. Here, we employed different designs of chimeric proteins as vaccine candidates that had not been tested previously.
The mouse serves as the primary model for studying mammalian diseases due to its small size, which makes it a cost-effective choice, high breeding capacity, and immunological reagents available [55]. There are some similarities in the immune response between pigs and mice because they are both mammals. Recently, Zhou et al. [56] showed the susceptibility of mice to Mycoplasma hyopneumoniae infection. However, there are genetic variations that can result in slight differences in their immune systems. Whilst these chimeric proteins were able to induce strong immune responses in mice, there is no assurance that the same immune response can be elicited in pigs when using these chimeric proteins as a vaccine.
Three novel chimeric proteins were successfully designed based on regions of M. hyopneumoniae proteins. These recombinant chimeric proteins induced an immune response in a murine model, able to induce specific antibodies against M. hyopneumoniae proteins and cytokines. Moreover, the chimeric proteins could induce proinflammatory cytokines such as TNF and IL-6. Further studies using different combinations with the three chimeric proteins are necessary, and clinical tests in pigs may be assayed using chimeric proteins.
References
Maes D, Boyen F, Haesebrouck F, Gautier-Bouchardon AV (2020) Antimicrobial treatment of Mycoplasma hyopneumoniae infections. Vet J 259:259–105474. https://doi.org/10.1016/j.tvjl.2020.105474
Villarreal I, Meyns T, Dewulf J et al (2011) The effect of vaccination on the transmission of Mycoplasma hyopneumoniae in pigs under field conditions. Vet J 188(1):48–52. https://doi.org/10.1016/j.tvjl.2010.04.024
Maes D, Segales J, Meyns T, Sibila M, Pieters M, Haesebrouck F (2008) Control of Mycoplasma hyopneumoniae infections in pigs. Vet Microbiol 126(4):297–309. https://doi.org/10.1016/j.vetmic.2007.09.008
Tao Y, Shu J, Chen J, Wu Y, He Y (2019) A concise review of vaccines against Mycoplasma hyopneumoniae. Res Vet Sci 123:144–152. https://doi.org/10.1016/j.rvsc.2019.01.007
Fisch A, Marchioro SB, Gomes CK, Galli V, Oliveira NR, Simionatto S, Dellagostin OA, Mendonça M, Moreira AN, Conceição FR (2016) Commercial bacterins did not induce detectable levels of antibodies in mice against Mycoplasma hyopneumoniae antigens strongly recognized by swine immune system. Trials Vaccinol 5:32–37. https://doi.org/10.1016/j.trivac.2016.01.001
Simionatto S, Marchioro SB, Maes D, Dellagostin OA (2013) Mycoplasma hyopneumoniae: from disease to vaccine development. Vet Microbiol 165(3–4):234–242. https://doi.org/10.1016/j.vetmic.2013.04.019
Rodrigues TCV, Jaiswal AK, De Sarom A et al (2019) Reverse vaccinology and subtractive genomics reveal new therapeutic targets against Mycoplasma pneumoniae: a causative agent of pneumonia. R Soc Open Sci 6(7). https://doi.org/10.1098/rsos.190907
Liu W, Feng Z, Fang L et al (2011) Complete genome sequence of Mycoplasma hyopneumoniae strain 168. J Bacteriol 193(4):1016–1017. https://doi.org/10.1128/JB.01305-10
Minion FC, Lefkowitz EJ, Madsen ML, Cleary BJ, Swartzell SM, Mahairas GG (2004) The genome sequence of Mycoplasma hyopneumoniae strain 232, the agent of swine mycoplasmosis. J Bacteriol 186(21):7123–7133. https://doi.org/10.1128/JB.186.21.7123-7133.2004
Siqueira FM, Thompson CE, Virginio VG et al (2013) New insights on the biology of swine respiratory tract mycoplasmas from a comparative genome analysis. BMC Genomics 14(1):1–17. https://doi.org/10.1186/1471-2164-14-175
Vasconcelos ATR, Ferreira HB, Bizarro CV et al (2005) Swine and poultry pathogens: the complete genome sequences of two strains of Mycoplasma hyopneumoniae and a strain of Mycoplasma synoviae. J Bacteriol 187(16):5568–5577. https://doi.org/10.1128/JB.187.16.5568-5577.2005
Han J, Park B-S, Shin D-J, Song S-Y, Jeong Y-J, Lee N (2017) Complete genome sequence of the mycoplasma hyopneumoniae strain KM014, a clinical isolate from South Korea. Genome Announc 5:4–5. https://doi.org/10.1128/genomeA.01012-17
Galli V, Simionatto S, Marchioro SB et al (2012) Immunisation of mice with Mycoplasma hyopneumoniae antigens P37, P42, P46 and P95 delivered as recombinant subunit or DNA vaccines. Vaccine 31(1):135–140. https://doi.org/10.1016/j.vaccine.2012.10.088
Jarocki VM, Santos J, Tacchi JL et al (2015) MHJ-0461 is a multifunctional leucine aminopeptidase on the surface of Mycoplasma hyopneumoniae. Open Biol 5(1). https://doi.org/10.1098/rsob.140175
Simionatto S, Marchioro SB, Galli V et al (2012) Immunological characterization of Mycoplasma hyopneumoniae recombinant proteins. Comp Immunol Microbiol Infect Dis 35(2):209–216. https://doi.org/10.1016/j.cimid.2012.01.007
Batra SA, Shanthalingam S, Donofrio G, Srikumaran S (2016) A chimeric protein comprising the immunogenic domains of Mannheimia haemolytica leukotoxin and outer membrane protein PlpE induces antibodies against leukotoxin and PlpE. Vet Immunol Immunopathol 175:36–41. https://doi.org/10.1016/j.vetimm.2016.05.004
Lin X, Xiao G, Luo D et al (2016) Chimeric epitope vaccine against Leptospira interrogans infection and induced specific immunity in Guinea pigs. BMC Microbiol 16(1):1–9. https://doi.org/10.1186/s12866-016-0852-y
Marchioro SB, Fisch A, Gomes CK et al (2014) Local and systemic immune responses induced by a recombinant chimeric protein containing Mycoplasma hyopneumoniae antigens fused to the B subunit of Escherichia coli heat-labile enterotoxin LTB. Vet Microbiol 173(1–2):166–171. https://doi.org/10.1016/j.vetmic.2014.07.009
Assao VS, Scatamburlo TM, Araujo EN et al (2019) Correction to: Genetic variation of Mycoplasma hyopneumoniae from Brazilian field samples. BMC Microbiol 19:234. https://doi.org/10.1186/s12866-019-1637-x
Gonzaga NF, de Souza LFL, Santos MR et al (2020) Antimicrobial susceptibility and genetic profile of Mycoplasma hyopneumoniae isolates from Brazil. Brazilian J Microbiol 51(1):377–384. https://doi.org/10.1007/s42770-019-00185-0
Marchioro SB, Simionatto S, Dellagostin O (2016) Development of Mycoplasma hyopneumoniae recombinant vaccines. Methods Mol Biol 1404:39–50. https://doi.org/10.1007/978-1-4939-3389-1_2
Marchioro SB, Maes D, Flahou B et al (2013) Local and systemic immune responses in pigs intramuscularly injected with an inactivated Mycoplasma hyopneumoniae vaccine. Vaccine 31(9):1305–1311. https://doi.org/10.1016/j.vaccine.2012.12.068
Jorge S, de Oliveira NR, Marchioro SB et al (2014) The Mycoplasma hyopneumoniae recombinant heat shock protein P42 induces an immune response in pigs under field conditions. Comp Immunol Microbiol Infect Dis 37(4):229–236. https://doi.org/10.1016/j.cimid.2014.07.001
Simionatto S, Marchioro SB, Galli V et al (2010) Cloning and purification of recombinant proteins of Mycoplasma hyopneumoniae expressed in Escherichia coli. Protein Expr Purif 69(2):132–136. https://doi.org/10.1016/j.pep.2009.09.001
de Oliveira NR, Jorge S, Gomes CK et al (2017) A novel chimeric protein composed of recombinant Mycoplasma hyopneumoniae antigens as a vaccine candidate evaluated in mice. Vet Microbiol 201:146–153. https://doi.org/10.1016/j.vetmic.2017.01.023
Galli V, Simionatto S, Marchioro SB, Klabunde GHF, Conceição FR, Dellagostin AO (2013) Recombinant secreted antigens from Mycoplasma hyopneumoniae delivered as a cocktail vaccine enhance the immune response of mice. Clin Vaccine Immunol 20(9):1370–1376. https://doi.org/10.1128/CVI.00140-13
Marchioro SB, Sácristan RDP, Michiels A et al (2014) Immune responses of a chimaeric protein vaccine containing Mycoplasma hyopneumoniae antigens and LTB against experimental M. hyopneumoniae infection in pigs. Vaccine 32(36):4689–4694. https://doi.org/10.1016/j.vaccine.2014.05.072
Conceição FR, Moreira ÂN, Dellagostin AO (2006) A recombinant chimera composed of R1 repeat region of Mycoplasma hyopneumoniae P97 adhesin with Escherichia coli heat-labile enterotoxin B subunit elicits immune response in mice. Vaccine 24(29–30):5734–5743. https://doi.org/10.1016/j.vaccine.2006.04.036
Miyazawa S, Jernigan LR (1985) Estimation of effective interresidue contact energies from protein crystal structures: quasi-chemical approximation. Macromolecules 18:534–552. https://doi.org/10.1021/ma00145a039
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER suite: protein structure and function prediction. Nat Methods 12(1):7–8. https://doi.org/10.1038/nmeth.3213
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5(4):725–738. https://doi.org/10.1038/nprot.2010.5
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9:40. https://doi.org/10.1186/1471-2105-9-40
Lovell SC, Davis IW, Arendall WB et al (2003) Structure validation by Cα geometry: phi, psi and Cβ deviation. Proteins 50(3):437–450. https://doi.org/10.1002/prot.10286
Wiederstein M, Sippl MJ (2007) Interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 2007:407–410. https://doi.org/10.1093/nar/gkm290
Ponomarenko J, Bui HH, Li W et al (2008) ElliPro: a new structure-based tool for the prediction of antibody epitopes. BMC Bioinformatics 9:1–8. https://doi.org/10.1186/1471-2105-9-514
Haste Andersen P, Nielsen M, Lund O (2006) Prediction of residues in discontinuous B-cell epitopes using protein 3D structures. Protein Sci 15(11):2558–2567. https://doi.org/10.1110/ps.062405906
Rubinstein ND, Mayrose I, Martz E, Pupko T (2009) Epitopia: a web-server for predicting B-cell epitopes. BMC Bioinformatics 10:287. https://doi.org/10.1186/1471-2105-10-287
Walker JM (2005) The proteomics protocols handbook. Humana Press, New York
Souza LFL, Gonzaga N, Santos MR, Assao VS, Rycroft A, Deeney AS, Fietto JLR, Bressan GC, Moreira MAS, Silva-Júnior A (2022) Identification of extracellular vesicles from J strain and wild isolate of mycoplasma hyopneumoniae. Brazil J Microbiol 53(2):1081–1084. https://doi.org/10.1007/s42770-022-00726-0
Okamba FR, Arella M, Music N, Jia JJ, Gottschalk M, Gagnon CA (2010) Potential use of a recombinant replication-defective adenovirus vector carrying the C-terminal portion of the P97 adhesin protein as a vaccine against Mycoplasma hyopneumoniae in swine. Vaccine 28(30):4802–4809. https://doi.org/10.1016/j.vaccine.2010.04.089
Seymour LM, Deutscher AT, Jenkins C et al (2010) A processed multidomain Mycoplasma hyopneumoniae adhesin binds fibronectin, plasminogen, and swine respiratory cilia. J Biol Chem 285(44):33971–33978. https://doi.org/10.1074/jbc.M110.104463
Djordjevic SP, Cordwell SJ, Djordjevic MA, Wilton J, Minion FC (2004) Proteolytic processing of the Mycoplasma hyopneumoniae cilium adhesin. Infect Immun 72(5):2791–2802. https://doi.org/10.1128/IAI.72.5.2791-2802.2004
Christodoulides A, Gupta N, Yacoubian V, Maithel N, Parker J, Kelesidis T (2018) The role of lipoproteins in Mycoplasma-mediated immunomodulation. Front Microbiol 9:1682. https://doi.org/10.3389/fmicb.2018.01682
Almeida HMS, Mechler-Dreibi ML, Sonálio K, Ferraz MES, Storino GY, Barbosa FO, Maes D, Montassier HJ, de Oliveira LG (2020) Cytokine expression and Mycoplasma hyopneumoniae burden in the development of lung lesions in experimentally inoculated pigs. Vet Microbiol 244:108647. https://doi.org/10.1016/j.vetmic.2020.108647
Beuckelaere L, Haspeslagh M, Biebaut E et al (2022) Different local, innate and adaptive immune responses are induced by two commercial Mycoplasma hyopneumoniae bacterins and an adjuvant alone. Front Immunol 13:1–16. https://doi.org/10.3389/fimmu.2022.1015525
Murtaugh MP (1994) Porcine cytokines. Vet Immunol Immunopathol 43:37–44. https://doi.org/10.1016/0165-2427(94)90118-X
Yang X, Xiao YL, Chen JL, Chen C, Zhang H, Wan XP, Lv XB, Li JL, Wang ZZ, Gao R (2013) Potentiation of immunity of piglets to Mycoplasma hyopneumoniae vaccine with fused gene for pig IL-4 and IL-6 embodied in chitosan nanoparticles. Procedia Vaccinol 7:15–22. https://doi.org/10.1016/j.provac.2013.06.004
Mechler-Dreibi ML, Almeida HMS, Sonalio K et al (2021) Oral vaccination of piglets against Mycoplasma hyopneumoniae using silica SBA-15 as an adjuvant effectively reduced consolidation lung lesions at slaughter. Sci Rep 11(1):1–15. https://doi.org/10.1038/s41598-021-01883-2
Luo Y, Li C, Zhou Z, Gong Z, Zhu C, Lei A (2021) Biological functions of IL-17-producing cells in mycoplasma respiratory infection. Immunology 164(2):223–230. https://doi.org/10.1111/imm.13346
Xu L, Hao F, Wang J et al (2022) Th1 and Th17 mucosal immune responses elicited by nasally inoculation in mice with virulence factors of Mycoplasma hyopneumoniae. Microb Pathog 172:105779. https://doi.org/10.1016/j.micpath.2022.105779
Lorenzo H, Quesada Ó, Assunçao P, Castro A, Rodríguez F (2006) Cytokine expression in porcine lungs experimentally infected with Mycoplasma hyopneumoniae. Vet Immunol Immunopathol 109(3–4):199–207. https://doi.org/10.1016/j.vetimm.2005.07.021
Standiford TJ, Deng JC (2006) Interleukins: IL-10. In: Laurent GJ, Shapiro SD (eds) Encyclopedia of respiratory medicine. Academic Press, Cambridge, pp 373–377
Couper KN, Blount DG, Riley EM (2008) IL-10: the master regulator of immunity to infection. J Immunol 180(9):5771–5777. https://doi.org/10.4049/jimmunol.180.9.5771
Leal FMDA, Virginio VG, Martello CL, Paes JA, Borges TJ, Jaeger N, Bonorino C, Ferreira HB (2016) Mycoplasma hyopneumoniae and Mycoplasma flocculare differential domains from orthologous surface proteins induce distinct cellular immune responses in mice. Vet Microbiol 190:50–57. https://doi.org/10.1016/j.vetmic.2016.05.008
Phifer-Rixey M, Nachman MW (2015) The natural history of model organisms: Insights into mammalian biology from the wild house mouse Mus musculus. eLife 4:e05959. https://doi.org/10.7554/eLife.05959
Zhou G, Tian Y, Tian J, Ma Q, Huang S, Li Q, Wang S, Shi H (2022) Oral immunization with attenuated Salmonella choleraesuis expressing the P42 and P97 antigens protects mice against Mycoplasma hyopneumoniae challenge. Microbiol Spectr 10(6):e02361-e2422. https://doi.org/10.1128/spectrum.02361-22
Funding
This work was supported by the following Brazilian agencies: Minas Gerais State Agency for Research and Development (FAPEMIG, grant #APQ-01327–14), National Council for Scientific and Technological Development (CNPq, grant #304727/2016–4), Coordination for the Improvement of Higher Education Personnel (CAPES, grant #Finance code 001), and Foundation for Research Support of the State of Alagoas (FAPEAL, grant # APQ2022021000101).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Responsible Editor: David Germano Gonçalves Schwarz
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Santos, M.R., Toledo, L.T., Bassi, Ê.J. et al. Chimeric proteins of Mycoplasma hyopneumoniae as vaccine and preclinical model for immunological evaluation. Braz J Microbiol 55, 943–953 (2024). https://doi.org/10.1007/s42770-023-01240-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01240-7