Abstract
Shiga toxin producing Escherichia coli (STEC) and enteropathogenic E. coli (EPEC) are pathovars that affect mainly infants’ health. Cattle are the main reservoir of STEC. Uremic hemolytic syndrome and diarrheas can be found at high rates in Tierra del Fuego (TDF). This study aimed to establish the prevalence of STEC and EPEC in cattle at slaughterhouses in TDF and to analyze the isolated strains. Out of 194 samples from two slaughterhouses, STEC prevalence was 15%, and EPEC prevalence was 5%. Twenty-seven STEC strains and one EPEC were isolated. The most prevalent STEC serotypes were O185:H19 (7), O185:H7 (6), and O178:H19 (5). There were no STEC eae + strains (AE-STEC) or serogroup O157 detected in this study. The prevalent genotype was stx2c (10/27) followed by stx1a/stx2hb (4/27). Fourteen percent of the strains presented at least one stx non-typeable subtype (4/27). Shiga toxin production was detected in 25/27 STEC strains. The prevalent module for the Locus of Adhesion and Autoaggregation (LAA) island was module III (7/27). EPEC strain was categorized as atypical and with the ability to cause A/E lesion. The ehxA gene was present in 16/28 strains, 12 of which were capable of producing hemolysis. No hybrid strains were detected in this work. Antimicrobial susceptibility tests showed that all strains were resistant to ampicillin and 20/28 were resistant to aminoglycosides. No statistical differences could be seen in the detection of STEC or EPEC either by slaughterhouse location or by production system (extensive grass or feedlot). The rate of STEC detection was lower than the one reported for the rest of Argentina. STEC/EPEC relation was 3 to 1. This is the first study on cattle from TDF as reservoir for strains that are potentially pathogenic to humans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Shiga toxin producing Escherichia coli (STEC) and Enteropathogenic E. coli (EPEC) are microorganisms known for causing severe gastroenteritis in infants [1]. STEC can cause a wide range of clinical disease patterns, including watery diarrhea, acute bloody diarrhea, and hemolytic uremic syndrome (HUS), which can lead to death by producing at least one of two cytotoxins called Shiga toxins (Stx), which are capable of inhibiting protein synthesis [2]. There are two types of Stx (Stx1 and Stx2), as well as several different subtypes [3, 4]. Non-typeable subtypes (NT) have also been reported [5]. There are 4 Stx1 subtypes (a, c–e) and 12 Stx2 subtypes (a–l) so far reported [5, 6]. EPEC causes diarrhea, affecting mainly children under 1 year of age, with high morbidity and mortality in developing countries [7]. It produces a lesion in the intestine epithelia known as “attaching and effacing” (A/E), produced by factors encoded by genes located in a pathogenicity island (PAI) called Locus of Enterocyte Effacement (LEE). The information needed to synthesize the protein intimin and a type 3 secretion system, along with different effector proteins, lies within LEE [8]. EPEC strains can be considered typical (tEPEC) or atypical (aEPEC) based on the presence of a type IV fimbriae called bundle forming pili (BFP). Atypical strains have been found in different animal species, and even some of these aEPEC are related to human gastroenteritis [7].
Cattle are the main reservoir for STEC and can eliminate the agent present in their intestinal microbiota through feces [9, 10]. Regarding EPEC, serogroups implicated in human diseases have been isolated from cattle suggesting that these animals could represent a reservoir for the pathogen [11, 12]. STEC causes a zoonotic disease, which can affect humans by oral ingestion of fecal-contaminated products [10]. The serotype O157:H7 is the most prevalent serotype and is responsible for the most severe disease patterns. However, more than 400 serotypes of E. coli can produce Stx [13].
Some STEC strains harbor LEE and can produce the A/E lesion. Fakih et al. [14] denominated these strains as AE-STEC. Patients who suffer HUS caused by AE-STEC strains can develop immune responsiveness to LEE-coded factors [15]. Yet in spite of the fact that AE-STEC is linked to human diseases in up to 80% of cases, LEE is not essential for successful infection (Rivas et al., 2016) [16]. Strains that do not have the LEE PAI are known as LEE negatives [17]. Montero et al. [18] reported the existence of another PAI that may contribute to intestinal colonization called Locus of Adhesion and Autoaggregation (LAA), associated with LEE negative strains. LAA can be present as a complete or incomplete structure, with four modules described that can be analyzed by gene markers as hes, iha, lesP, pagC, tpsA, tpsB, and agn4 [18].
E. coli can also share genetic markers from different pathovars, generating hybrid strains like the Stx-producing EAEC O104:H4 strain, which has the ability to produce more complex clinical patterns [16, 19].
HUS is an endemic disease in Argentina, with reported cases increasing during the summer [20, 21]. The higher rates of HUS are located in the southern regions of the country, particularly in Chubut and Tierra del Fuego (TDF). TDF reports also high rates of acute diarrhea [22].
The province of Tierra del Fuego, Antártida e Islas del Atlántico Sur, is an island located in the southernmost part of Argentina and has different climate conditions than the rest of the country. Argentina’s legislation restricts the movement of cattle to TDF, so meat production on the island is mainly locally produced. Therefore, E. coli strains in TDF may have characteristics only locally found. Also, it is interesting to assess antimicrobial resistance, given the emergence of global antimicrobial resistance, and the importance of cattle industries in Argentina. Prack McCormick et al. [23] found that in E. coli, multidrug resistance is associated with both animal species and animal production, being more prevalent in intensive productions. Information regarding antimicrobial resistance in beef cattle is still missing.
It is important to stress that there are no previous studies of STEC and EPEC in cattle in TDF. The correlation between STEC’s prevalence and other E. coli pathovars in reservoirs and sources of infections has not been yet studied.
The aim of this study is to determine the prevalence of STEC (STEC LEE-negative / AE-STEC) and EPEC in cattle at slaughterhouses in TDF, as well as to evaluate the virulence profile of the strains and analyze the correlation between these pathovars.
Methods
Samples
We carried out a cross-sectional epidemiological study in June 2016 in order to determine the prevalence of STEC (STEC LEE-negative/AE-STEC) and EPEC in cattle from TDF.
The sample size was 126 animals from each of the two slaughterhouses, one located in Ushuaia and the other in Rio Grande. We considered a standard normal distribution (Z = 1.96), precision at 5%, 95% confidence level, and an estimated 9% of STEC and EPEC prevalence based on data [24]. The cattle were swabbed at the rectum, and each sample was sent to the lab maintained in Stuart Transport Media. The study was approved by the Institutional Committee on Animal Care and Use of Experimental Animals (CICUAL; Nº 2017/8) at the Facultad de Ciencias Veterinarias, Universidad de Buenos Aires.
Detection and isolation of strains
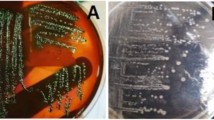
In order to detect non-O157 EPEC and STEC strains, each swab sample was enriched in 5 mL trypticase soy broth (TSB), incubated at 37 °C for 6 h, and then streaked onto Mac Conkey agar (MAC) for further incubation at 37 °C another 18 h.
STEC O157 detection was carried out by enriching each sample in 5 mL of TSB plus tellurite cefixime and incubated at 37 °C for 6 h. Then, we screened the serogroup by immunochromatography test (Reveal® E. coli O157) following the supplier’s instructions. The positive samples were then subjected to Immunomagnetic separation and streaked onto sorbitol Mac Conkey agar (SMAC) [25, 26].
We obtained a loop from the confluence culture zone in order to extract DNA and conducted multiplex PCR for STEC markers stx1, stx2, and rfbO157 [27] and single PCR for EPEC marker eae [28]. Primers for detection of genetic markers are referred in Table S1.
We then analyzed up to 50 colonies by PCR to find those markers identified in the confluence culture zones [25, 26].
For an AE-STEC strain to be classified as such, it has to carry both eae and at least one of the stx. According to the results, the samples were thus classified as negative, positive by screening, and positive by isolation. We characterized the isolates phenotypically (morphology, Gram stain, and motility) and confirmed them as E. coli through biochemical tests [29].
STEC and EPEC characterization
Genotypical characterization
In STEC strains, we analyzed the Stx subtypes: Stx1 subtypes (stx1a, stx1c, stx1d) and Stx2 subtypes (stx2a, stx2b, stx2c, stx2d, stx2e, stx2g), for identification by conventional PCR amplification [4]. Non-typeable strains were characterized by RFLP-PCR [3]. In addition, we evaluated the presence of eae gene. In STEC LEE-negative strains, we searched for the presence of the four modules (I-IV) of the LAA Island [18]. In EPEC isolates, we searched for the gene bfpA [30].
In all strains, we analyzed by PCR the presence of an enterohaemolysin (ehxA) and STEC autoagglutinating adhesin (saa) [31]. The presence of hybrid strains was determined searching for genetic markers of other E. coli pathovars: aaiC [32] and aagR [33] for enteroaggregative E. coli strains (EAEC); elt and estA for enterotoxigenic E. coli strains (ETEC) [34]; daaE for diffuse aggregation E. coli strains (DAEC) [35]; and invE for enteroinvasive E. coli strains (EIEC) [34]. Primers for detection of genetic markers are referred in Table S1.
Phenotypic characterization
The production of enterohaemolysin was tested in a TS-based agar using washed blood and non-washed blood ovine red blood cells [36].
To identify the production of Stx, we used the SHIGA TOXIN QUIK CHEK, which is a rapid membrane enzyme immunoassay test. For that purpose, we followed the supplier’s recommendations.
EPEC strains were tested for adherence pattern to HeLa cells [37, 38]. We incubated the HeLa cells with 20 µL of an overnight culture of the strains (3- and 6-h assays). Then, the cultures were washed with PBS, fixated with methanol, stained with Giemsa, and finally analyzed under the microscope. We also examined the ability of the strains to cause A/E lesion using fluorescent actin stain (FAS). For this assay, HeLa cells and strains cultures were mixed, incubated for 3 h, and then fixated with formaldehyde 3%. The fixed cells were treated with 0.1% Triton X-100, washed with PBS, and then stained with fluorescein isothiocyanate-phalloidin (5 μg/mL). We kept the cells in a dark and humid environment for 30 min; we washed the cells again with PBS and analyzed the cells under the fluorescence microscope. The existence of fluorescent areas in adhesion zones was considered a positive FAS test [39].
Finally, all isolates were serotyped by agglutination assays with rabbit antisera: 187 sera were used against somatic antigens (O), and 53 sera were used against flagellar antigens (H) at the Facultad de Medicina, Universidad Nacional Autónoma de México (UNAM) [40, 41].
Antimicrobial susceptibility testing
Antimicrobial susceptibility was conducted in all STEC and EPEC isolated strains, against the antibiotics enlisted by the reference center INEI-ANLIS: amikacin, ampicillin, ciprofloxacin, colistin, chloramphenicol, gentamicin, norfloxacin, nalidixic acid, tetracycline, streptomycin, and trimethoprim-sulfamethoxazole [36]. We used the Kirby-Bauer Disk Diffusion Susceptibility Test Protocol according to CLSI recommendations [42]. To evaluate colistin resistance, we used the recommended method COLISTIN AGAR SPOT [43]. In the strains resistant to ampicillin, we evaluated the presence of broad-spectrum beta-lactamase (BLEE, AmpC, and KPC), performing the Kirby-Bauer test with strategic colocation of the following discs alongside boronic acid: amoxicillin clavulanic, cefotaxime, cefotaxime clavulanic, ceftazidime, meropenem, and imipenem [44].
Statistical analysis
Statistical analysis of the obtained data was carried out using the test of differences between proportions (InfoStat 2016e), considering the following variables: production system, farm, and slaughterhouse location.
Results
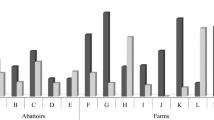
From a total of 194 samples, 124 were obtained in Rio Grande (RG) and 70 in Ushuaia (U). The samples in RG were taken at 3 different farms, 2 of them with extensive grass cattle and the other one feedlot. The samples in U were obtained from a single extensive grass breeding facility.
STEC prevalence at the screening stage was 15% (30/194), while EPEC was detected in 5% of the samples (10/194). No sample revealed the genetic markers for both pathovars at the same time. We isolated 27 LEE-negative STEC strains and one EPEC strain from the positive samples. AE-STEC strains were not detected. All the isolated strains were identified as E.coli.
Regarding the type of stx, 59.2% of STEC strains carried stx2 alone (16/27), 7.4% carried stx1 (2/27), and 33.3% of STEC strains carried both stx1/stx2 (9/27). Considering subtypes, the most prevalent was stx2c (10/27), followed by stx1a/stx2hb (4/27), and 15% of the strains (4/27) carried at least one non-typeable stx (stxNT). A total of 25/27 of the isolated STEC strains were able to produce Stx. None of the strains carried the gene saa; 15/27 carried the gene ehxA, 12 of which were able to produce the expected hemolysis. In search for the LAA island, we found that 11/27 strains where positive for at least one module, and module III was prevalent (7/27). None of the strains carried module IV, and no strain carried the 4 modules of the LAA PAI at the same time. Results are referred in Table 1.
The isolated EPEC strain was bfpA negative; therefore, we classified it as atypical EPEC (aEPEC). In regard to additional virulence factors, we characterized the strain as saa—ehxA + , without the capability to produce hemolysis (Table 1). HeLa essays did not reveal a defined adhesion pattern. However, the latter did not affect the strain’s capability to produce A/E lesion.
No hybrid strains were detected in this study.
The 27 isolated STEC strains belonged to 12 different serotypes, the most prevalent O185:H19 (26%), O185:H7 (22,2%), and O178:H19 (18,5%). The remaining 9 serotypes (33,3%) were O1:H21, O6:H34, O113:H21, O130:H11, O130:H-, O171:H2, O174:H28, O179:H8, and O187:H7, with one strain for each serotype. The EPEC strain belonged to the O152:H25 serotype.
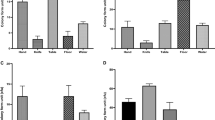
Antimicrobial susceptibility tests revealed resistance to ampicillin in all the analyzed strains. The assays did not confirm the presence of broad-spectrum beta-lactamases. Some strains showed resistance to streptomycin and amikacin; however, none of them were resistant to more than two families of antibiotics (Table S2).
No significant differences were observed (p > 0.05) between variables (production system, farm, and slaughterhouse location). STEC/EPEC detection showed a ratio 1 to 3 considering screening positive samples.
Discussion
We studied the prevalence of STEC (LEE-negative and AE-STEC) and EPEC in TDF ‘s cattle. We analyzed 194 rectal swabs samples from cattle in slaughterhouses. STEC prevalence at the screening stage was 15%, with all the strains being STEC LEE-negative. This result is lower than the prevalence reported for cattle in the rest of the country, which ranges between 22 and 67% [9, 45,46,47].
EPEC prevalence was 5%, similar to the one reported by Pizarro et al. [46] in Mendoza. As far as we know, there are no other studies regarding EPEC’s prevalence in the country.
In concordance with Masana et al. [48], STEC detection showed no significant statistical differences regarding production system (feedlot and grass cattle). On the other hand, Padola et al. [45] and Tanaro et al. [49] revealed in their studies higher rates of detection of STEC in feedlot samples.
In our study, we detected mainly strains stx2 + , similarly to the ones found in other studies from Argentina [9, 46, 50, 51]. Of the 27 STEC strains isolated in this work, 25 (92%) presented the gene stx2, and 11 strains carried the gene stx1 (40%). The most prevalent subtype for Stx2 was stx2c (11/25), whereas 10 isolated strains could not be sub-typified using conventional PCR [4]. Of the latter 10 strains, 7 were identified using PCR- RFLP as stx2hb [3], and the remaining 3 strains could not be subtyped (stx2NT). These 3 stx2NT were obtained as follows: 2 from grass cattle from Ushuaia and 1 from the feedlot in Rio Grande. The most prevalent subtype for Stx1 was stx1a (9/11). One of the Stx1 could not be identified with the methods described [3, 4] (Table 1).
Considering the genetic profiles, stx2c was the most prevalent gene (10/27), followed by stx1a/stx2hb (4/27), and 15% of the strains (4/27) carried at least one non-typeable Stx (Table 1). Other studies in mainland Argentina also found non-typeable (NT) strains yet in a lesser proportion [49,50,51,52,53]. These results may suggest that STEC strains from TDF’s cattle have characteristics of their own. In order to further investigate NT strains, it will be necessary to perform their sequencing. We found that 25/27 STEC strains were able to produce Stx using SHIGA TOXIN QUIK CHEK. A total of 3/25 of the Stx producing strains carried NT toxins (24fRGIII, 24fRGII, U41V) (Table 1). Considering that the sensitivity of the test may vary according to the Stx subtype, the characteristics of the strain, and the amount of Stx produced, the absence of Stx production in the remaining 2 strains should be confirmed through toxicity assays in Vero cells cultures.
In this work, we did not find AE-STEC. Orden et al. [24] neither found AE-STEC strains in healthy cattle in Spain, being the most prevalent profile stx2 + eae-. This is also true in previous reports from other studies [9, 46, 54, 55].
Etcheverría and Padola [56] analyzed STEC strains from different sources (minced beef, cattle, and environment), and they did find AE-STEC, yet in a lower proportion than STEC LEE-negative. Other studies in mainland Argentina revealed a higher degree of detection of AE-STEC O157 [48, 53, 57] and non-O157 AE-STEC [50, 51]. The differences in the results could be related to different sampling methods and detection protocols.
It is important to highlight that nowadays, the diagnostic pathways are better and include the search for non-O157 serogroups [58]. In 2008, Coombes et al. [59] reported a 60% increase in clinical HUS worldwide related to non-O157 serogroups, while the HUS cases related to O157 strains increased by only 13%. In 2015, Byrne et al. [60] carried out a study in England that revealed that non-O157 strains were related to highest rates of HUS hospitalization than O157 strains. Moreover, Valilis et al. [61] analyzed the pathogenic role of non-O157 strains, concluding that they are more related to acute diarrhea in humans than O157 strains.
The LEE-negative strains isolated in our work belonged to non-O157 serogroups: O1:H21; O6:H34; O113:H21; O130:H11; O130:H-; O171:H2; O174:H28; O178:H19; O179:H8; O185:H7; O185:H19; and O187:H7 (Table 1). Serotypes O113:H21, O130:H11, O174:H28, O178:H19, and O185:H7 have been previously detected in Argentina [50, 51, 56, 62]. Etcheverría and Padola [56] detected O8:H19, O26:H11, O91:H21, O113:H21, O117:H7, O130:H11, O145:H-, O157:H7, O171:H2, and O178:H19 as prevalent serotypes, some of them equally present in different sources.
The O174 serogroup has been detected in various sources (cattle, rodents, food, humans), and it has been related to HUS in Argentina [50, 52]. It was also described by Masana et al. [50] as one of the non-O157 serogroups responsible for 30% of clinical HUS in Argentina.
Previous reports in Argentina detected STEC O130:H11 and STEC O178:H19 as prevalent serotypes in dairy and feedlot cattle, abattoirs, and local markets [50, 62,63,64]. Both serotypes have also been associated to HUS in Argentina [63]. STEC O178:H19 has also been related to HUS worldwide [65,66,67,68,69]. A comparison between strains of STEC O178:H19 from different sources confirmed the absence of the eae gene. Its absence has no impact on STEC O178.H19 pathogenesis [70].
Qin et al. [71] isolated the O6:H34 serotype from clinical pediatric HUS cases, and Wang et al. [72] found O6:H34 in 10% of the isolated strains in a persistence study in cattle from Canada. Previously, Delannoy et al. [73] analyzed O1 and O2 STEC strains in human and cattle feces. Serogroups O113 and O179 have been also reported in bovine feces and carcasses in Argentina [50, 51], as well as in minced meat [68]. No information has been found about the O185:H19 serotype, yet O185:H9 and O185:NT have been previously reported in bovine feces in Germany [74].
As far as the serotype O187:H7 is concerned, there are not many findings yet. Bai et al. [75] found 4 human hybrid STEC/ETEC strains in Sweden belonging to rare serogroups, including O187. Hybrid strains represent a high risk for public health, and although they have been found in cattle before [76,77,78], we did not find any hybrid strain in our study.
Our aEPEC strain belonged to the O152:H25 serotype (Table 1). HeLa essays did not reveal a defined adhesion pattern. However, this did not affect the O152:H25 capability to produce A/E lesion. As far as we know, this serotype has not been associated with human diarrhea, yet clinical cases of EIEC O152 serogroup have been reported [79,80,81].
Regarding additional STEC virulence factors, the ehxA gene was found in 57% (16/28) of strains and 12/16 were able to produce the expected hemolysis (Table 1). Strain capability to produce this hemolysin has been associated with strains with clinical impact and has been used for screening detection [82]. The aEPEC strain (93fRG) was one of the carrier strains of gene ehxA without the capability to produce hemolysis, as has been also reported in previous studies [11, 83,84,85].
None of the analyzed strains carried either the saa gene or the four modules of the LAA island at the same time (Table 1). In their analysis of LEE negative strains from various sources (cattle, food, environment), Colello et al. [86] found the complete LAA sequence in 46% of the strains, without correlation between the detection of LAA and serogroup. Although the high variation of LEE negative strains makes it difficult to establish their virulence, Montero et al. [87] described the gene hes among the most prevalent virulence factors. In our study the hes gene was not prevalent (5/27) (Table 1). To establish the importance of LAA in our strains, it will be necessary to carry out further functional assays, as well as analyze the presence of other adhesins.
Finally, our susceptibility tests detected ampicillin resistance in all strains (Table S2). Other studies conducted in Latin America revealed similar results [88, 89]. In the analysis of feces from different animal sources, Pantozzi et al. [90] found ampicillin resistance in up to 4.4% of cattle strains. In 2017, SENASA (Servicio Nacional de Sanidad y Calidad Agroalimentaria) reported levels of ampicillin resistance related mainly to swine and poultry intensive production in Argentina. The resistance to this antibiotic in cattle was 6.5% [91]. Regarding other antibiotics, some strains in our work detected resistance or intermediate resistance to aminoglycosides, specifically streptomycin (71%), and amikacin (21%) (Table S2). Pantozzi et al. [90] also found strain resistance in cattle to streptomycin and amikacin, yet in lower proportions (2.2% and 9%, respectively). None of our strains revealed phenotypic resistance to colistin, none was resistant to more than two families of antibiotics, and there was no beta-lactamase extended spectrum resistance.
There are not many studies that allow the comparison between two E. coli pathovars in cattle feces samples [14, 23, 46, 92]. The analysis of 194 samples from local TDF cattle detected 30/194 positives to STEC at the screening stage and 10/194 positives to EPEC. None of the strains was classified as AE-STEC. The correlation between STEC/EPEC was 3 to 1, so it is three times more likely (considering the same source of infection) for TDF infants to be exposed to STEC, with all the health risks it implies. Considering the gene homology between these two pathovars in LEE, Calderon Toledo et al. [93] suggested crossed immunity between EPEC and AE-STEC strains. They inoculated BALB/c mice with these strains and reported that the infection from EPEC could provide immunity to AE-STEC infection. However, all the STEC strains isolated in our work were LEE-negative. In this sense, the immunity provided by EPEC would not be effective for a consequent STEC LEE-negative infection. This could contribute to the high HUS rates in TDF.
This is the first study of STEC and EPEC prevalence in cattle in TDF. High HUS rates in TDF highlight the need for further research, in particular serial sampling.
We need to know if the proportion of serotypes, the relationship between E. coli detection and cattle production facility, and the prevalence obtained are stable over time. It is of importance as well the study of other sources of infection and reservoirs.
References
Trabulsi LR, Keller R, Gomez TAT (2002) Typical and atypical enteropathogenic Escherichia coli. Emerg Infect Dis 8:508–513. https://doi.org/10.3201/eid0805.010385
Guth BEC, Prado V, Rivas M (2010) Shiga toxin-producing Escherichia coli. En Pathogenic Escherichia coli in Latin America, Torres, A.G. (Ed). Bentham Science Publishers U.S.A, pp 25–47 https://doi.org/10.2174/97816080519221100101
Tyler SD, Johnson WM, Lior H, Wang G, Rozee KR (1991) Identification of verotoxin type 2 variant B subunit genes in Escherichia coli by the polymerase chain reaction and restriction fragment length polymorphism analysis. J Clin Microbiol 29(7):1339–431343. https://doi.org/10.1128/jcm.29.7.1339-1343.1991
Scheutz F, Teel LD, Beutin L, Pierard D, Buvens G, Karch H, Mellmann A, Caprioli A, Tozzoli R, Morabito S, Strockbine NA, Melton-Celsa AR, Sanchez M, Persson S, O’Brien AD (2012) Multicenter evaluation of a sequence-based protocol for subtyping Shiga toxins and standardizing Stx nomenclature. J Clin Microbiol 50:2951–63. https://doi.org/10.1128/JCM.00860-12
Werber D, Scheutz F (2019) The importance of integrating genetic strain information for managing cases of Shiga toxin-producing E. coli infection. Epidemiol Infect 147:e264. https://doi.org/10.1017/S0950268819001602
EFSA BIOHAZ Panel, Koutsoumanis K, Allende A, Alvarez-Ordoñez A, Bover-Cid S, Chemaly M, Davies R, De Cesare A, Herman L, Hilbert F, Lindqvist R, Nauta M, Peixe L, Ru G, Simmons M, Skandamis P, Suffredini E, Jenkins C, Monteiro Pires S, Morabito S, Niskanen T, Scheutz F, da Silva Felício MT, Messens W, Bolton D (2020) Pathogenicity assessment of Shiga toxin-producing Escherichia coli (STEC) and the public health risk posed bycontamination of food with STEC. Efsa J Sci Opin. https://doi.org/10.2903/j.efsa.2020.5967
Gomes TAT, González-Pedrajo B, Enteropathogenic Escherichia coli (EPEC) (2010). En Pathogenic Escherichia coli in Latin America, Torres AG (ed). Bentham Science Publishers U.S.A, pp 25–47. https://doi.org/10.2174/97816080519221100101
Nataro JP, Kaper JB (1998) Diarrheagenic Escherichia coli. Clin Microbiol Rev 11(1):142–201. https://doi.org/10.1128/CMR.11.1.142
Parma AE, Sanz ME, Blanco JE, Blanco J, Viñas MR, Blanco M, Padola NL, Etcheverría AI (2000) Virulence genotypes and serotypes of verotoxigenic Escherichia coli isolated from cattle and foods in Argentina. Eur J Epidemiol 2000(16):757–762. https://doi.org/10.1023/a:1026746016896
Piérard D, De Greve H, Haesebrouck F, Mainil J (2012) O157:H7 and O104:H4 Vero/Shiga toxin-producing Escherichia coli outbreaks: respective role of cattle and humans. Vet Res 43(1):13. https://doi.org/10.1186/1297-9716-43-13
Hernandes RT, Elias WP, Vieira MAM, Gomes TAT (2009) An overview of atypical enteropathogenic Escherichia coli. FEMS Microbiol Lett 297(2):137–149. https://doi.org/10.1111/j.1574-6968.2009.01664.x
Kolenda R, Burdukiewicz M, Schierack, (2015) A systematic review and meta-analysis of the epidemiology of pathogenic Escherichia coli of calves and the role of calves as reservoirs for human pathogenic E. coli. Front Cell Infect Microbiol. https://doi.org/10.3389/fcimb.2015.002
Acuña P, Rojas N, Florentin M, Rodriguez F (2019) Estandarización de una técnica de PCR múltiple para la detección de los serogrupos O157, O104 y “big six” de Escherichia coli productora de la toxina Shiga (STEC). Memorias del Instituto de Investigaciones en Ciencias de la Salud 17(2):71–76. https://doi.org/10.18004/mem.iics/1812-9528/2019.017.02.71-076
Fakih I, Thiry D, Duprez J-N, Saulmont M, Iguchi A, Piérard D, Jouant L, Daube G, Ogura Y, Hayashi T, Taminiau B, Mainil JG (2017) Identification of Shiga toxin-producing (STEC) and enteropathogenic (EPEC) Escherichia coli in diarrhoeic calves and comparative genomics of O5 bovine and human STEC. Vet Microbiol 202:16–22. https://doi.org/10.1016/j.vetmic.2016.02.017
Kaper JB, Nataro JP, Mobley HL (2004) Pathogenic Escherichia coli. Nat Rev Microbiol 2(2):123–140. https://doi.org/10.1038/nrmicro818
Rivas M, Chinen I, Guth BEC (2016) Enterohemorrhagic (Shiga toxin –producing) Escherichia coli. En Escherichia coli in the Americas, Torres AG (Ed) Springer International Publishing, U.S.A, 2016, 149–72. https://doi.org/10.2174/97816080519221100101
Newton HJ, Sloan J, Bulach DM, Seemann T, Allison CC, Tauschek M, Robins-Browne RM, Paton JC, Whittam TS, Paton AW, Hartland EL (2009) Shiga toxin–producing Escherichia coli strains negative for locus of enterocyte effacement. Emerg Infect Dis 15(3):372–380. https://doi.org/10.3201/eid1502.080631
Montero DA, Velasco J, Del Canto F, Puente JL, Padola NL, Rasko DA, Farfán M, Salazar JC, Vidal R (2017) Locus of Adhesion and Autoaggregation (LAA), a pathogenicity island present in emerging Shiga toxin-producing Escherichia coli strains. Sci Rep 7(1):7011. https://doi.org/10.1038/s41598-017-06999-y
Bielaszewska M, Mellmann A, Zhang W, Köck R, Fruth A, Bauwens A, Peters G, Karch H (2011) Characterisation of the Escherichia coli strain associated with an outbreak of haemolytic uraemic syndrome in Germany, 2011: a microbiological study. Lancet Infect Dis 11:671–76. https://doi.org/10.1016/S1473-3099(11)70165-7
Ministerio de Salud de la Nación. Boletín Integrado de Vigilancia N222-SE30 (2014) Informe Especial I: Síndrome urémico hemolítico (SUH) en Argentina, 2010 – 2013, p. 92 – 103. Retrieved from: https://www.argentina.gob.ar/salud/epidemiologia/boletines2014. Accessed June 2022
Amigo N, Mercado E, Bentancor A, Singh P, Vilte D, Gerhardt E, Zotta E, Ibarra C, Manning SD, Larzábal M, Cataldi A (2015) Clade 8 and clade 6 strains of Escherichia coli O157:H7 from cattle in Argentina have hypervirulent-like phenotypes. PLoS One 10(6):e0127710. https://doi.org/10.1371/journal.pone.0127710
Ministerio de Salud de la Nación. Boletín Integrado de Vigilancia N477-SE50 (2019). Retrieved from: https://www.argentina.gob.ar/salud/epidemiologia/. Accessed June 2022
Prack McCormick B, Quiroga MP, Álvarez VE, Centrón D, Tittonell P (2022) Diseminación de la resistencia a antimicrobianos asociada a prácticas de producción animal intensiva en Argentina: Una revisión sistemática y metaanálisis. Rev Argent Microbiol. https://doi.org/10.1016/j.ram.2022.07.001
Orden JA, Cid D, Ruiz-Santa-Quiteria JA, García S, Martínez S, de la Fuente R (2002) Verotoxin- producing Escherichia coli (VTEC), enteropathogenic E. coli (EPEC) and necrotoxigenic E. coli (NTEC) isolated from healthy cattle in Spain. J Appl Microbiol 93:29–35. https://doi.org/10.1046/j.1365-2672.2002.01649.x
Bentancor A, Rumi MV, Gentilini MV, Sardoy C, Irino K, Agostini A, Cataldi A (2007) Shiga toxin-producing and attaching and effacing Escherichia coli in cats and dogs in a high hemolytic uremic syndrome incidence region in Argentina. FEMS Microbiol Lett 267:251–256. https://doi.org/10.1111/j.1574-6968.2006.00569.x
Blanco Crivelli X, Bonino MP, Sanin M, Petrina JF, Disalvo V, Massa R, Miliwebsky E, Navarro A, Chinen I, Bentancor A (2021) Potential zoonotic pathovars of diarrheagenic Escherichia coli detected in lambs for human consumption from Tierra del Fuego. Argent Microorganisms 9(8):1710. https://doi.org/10.3390/microorganisms9081710
Leotta G, Chinen I, Epszteyn S, Miliwebsky E, Melamed IC, Motter M, Ferrer M, Marey E, Rivas M (2005) Validación de una técnica de PCR múltiple para la detección de Escherichia coli productor de toxina Shiga. Revista Argentina de Microbiología 37:1–10. Retrieved from: https://www.redalyc.org/articulo.oa?id=213016778001
Blanco M, Blanco JE, Mora A, Dahbi G, Alonso MP, González EA, Bernárdez MI, Blanco J (2004) Serotypes, virulence genes, and intimin types of Shiga Toxin (Verotoxin)-producing Escherichia coli isolates from cattle in Spain and identification of a new intimin variant gene (eae). J Clin Microbiol 42:645–651. https://doi.org/10.1128/JCM.42.2.645-651.2004
MacFaddin JF (2003) Pruebas bioquímicas para la identificación de bacterias de importancia clínica, 3rd edn. Buenos Aires, Argentina, Médica Panamericana Ed, pp 54–421
Gunzburg ST, Tornieporth NG, Riley LW (1995) Identification of enteropathogenic Escherichia coli by PCR-based detection of the bundle-forming pilus gene. J Clin Microbiol 33:1375–1377. https://doi.org/10.1128/jcm.33.5.1375-1377.1995
Paton AW, Paton JC (2002) Direct detection and characterization of Shiga toxigenic Escherichia coli by multiplex PCR for stx1, stx2, eae, ehxA, and saa. J Clin Microbiol 40(1):271–274. https://doi.org/10.1128/JCM.40.1.271-274.2002
Boisen N, Scheutz F, Rasko DA, Redman JC, Persson S, Simon J, Kotloff KL, Levine MM, Sow S, Tamboura B, Toure A, Malle D, Panchalingam S, Krogfelt KA, Nataro JP (2012) Genomic characterization of enteroaggregative Escherichia coli from children in Mali. J Infect Dis 205(3):431–444. https://doi.org/10.1093/infdis/jir757
Wieler LH, Semmler T, Eichhorn I, Antao EM, Kinnemann B, Geue L, Karch H, Guenther S, Bethe A (2011) No evidence of the Shiga toxin-producing E. coli O104:H4 outbreak strain o enteroaggregative E. coli (EAEC) found in cattle faeces in northern Germany, the hostpot of the 2011. HUS outbreak area. Gut Pathogens 3:17. https://doi.org/10.1186/1757-4749-3-17
Itoh F, Ogino T, Itoh F, Watanabe H (1992) Differentiation and detection of pathogenic determinants among diarrheagenic Escherichia coli by polymerase chain reaction using mixed primers. Jpn J Clin Med 50:343–347
Vidal M, Kruger C, Duran R, Lagos R, Levine M, Prado V, Toro C, Vidal R (2005) Single PCR assay to identify simultaneously the six categories of diarrheagenic Escherichia coli associated with enteric infections. J Clin Microbiol 43(10):5362–5365. https://doi.org/10.1128/JCM.43.10.5362-5365.2005
Miliwebsky E, Deza N, Zolezzi G, Baschkier A, Carbonari CC, Manfredi E, D’Astek BA, Chinen I, Rivas M (2019) Manual de Procedimientos: Escherichia coli Productor de Toxina Shiga en el Marco de la Detección de E.coli diarreigénico. Retrieved from: http://sgc.anlis.gob.ar/handle/123456789/2307
Scaletsky IC, Silva ML, Trabulsi LR (1984) Distinctive patterns of adherence of Enteropathogenic Escherichia coli to HeLa cells. Infect Immun 45(2):534–536. https://doi.org/10.1128/iai.45.2.534-536.1984
Gomes TAT, Irino K, Girão DM, Girão VBC, Guth BEC, Vaz TMI, Moreira FC, Chinarelli SH, Vieira MAM (2004) Emerging enteropathogenic Escherichia coli strains? Emerg Infect Dis 10(10):1851–1855. https://doi.org/10.3201/eid1010.031093
Nakazato G, Gyles C, Ziebell K, Keller R, Trabulsi LR, Gomes TAT, Irino K, Da Silveira WD, Pestana De Castro AF (2004) Attaching and effacing Escherichia coli isolated from dogs in Brazil: characteristics and serotypic relationship to human enteropathogenic E. coli (EPEC). Vet Microbiol 101(4):269–77. https://doi.org/10.1016/j.vetmic.2004.04.009
Edwards PR, Ewing WH (1986) Edwards and Ewing’s identification of Enterobacteriaceae, 4th edn. Elsevier Science Publishing Inc, New York, USA
Orskov F, Orskov I (1984) Serotyping of Escherichia coli. Methods Microbiol 14:43–112. https://doi.org/10.1016/S0580-9517(08)70447-1
Performance Standards for Antimicrobial Susceptibility Testing (2018) 28th Edition. Weinstein, MP, Patel JB. Clinical and Laboratory Standards Institut. Retrieved from: https://file.qums.ac.ir/repository/mmrc/CLSI-2018-M100-S28.pdf. Accessed Mar 2022
INEI-ANLIS (2017) Administración Nacional de Laboratorios e Institutos de Salud. Dr. Carlos G. Malbrán. Servicio Antimicrobianos, Laboratorio Nacional de Referencia en Antimicrobianos, Método de Screening “COLISTIN AGAR-SPOT” Versión 2. Retrieved from: http://antimicrobianos.com.ar/ATB/wp-content/uploads/2017/09/Protocolo-Agar-spot-COL-2017-version2-Agosto2017.pdf. Accessed Mar 2022
Famiglietti A, Quinteros M, Vázquez M, Marin M, Nicola F, Radice M, Galas MF, Pasterán FG, Bantar C, Casellas JM, Kovensky Pupko J, Couto E, Goldberg M, Lopardo H, Gutkind G, Soloaga R (2005) Consenso sobre las pruebas de sensibilidad a los antimicrobianos en Enterobacteriaceae. Revista Argentina de Microbiología 37: 57–66. ISSN 0325–7541 ARTÍCULO ESPECIAL Revista Argentina de Microbiología 37: 57–66
Padola NL, Sanz ME, Blanco JE, Blanco M, Blanco J, Etcheverría AI, Arroyo GH, Usera MA, Parma AE (2004) Serotypes and virulence genes of bovine Shigatoxigenic Escherichia coli (STEC) isolated from a feedlot in Argentina. Vet Microbiol 100(1–2):3–9. https://doi.org/10.1016/S0378-1135(03)00127-5
Pizarro MA, Orozco JH, Degarbo SM, Calderón AE, Nardello AL, Laciar A, Rüttler ME (2014) Virulence profiles of Shiga toxin-producing Escherichia coli and other potentially diarrheagenic Ecoli of bovine origin, in Mendoza, Argentina. Braz J Microbiol 44(4):1173–1180. https://doi.org/10.1590/S1517-83822014005000010
Etcheverría AI, Lucchesi PMA, Krüger A, Bentancor AB, Padola N (2016) Escherichia coli in Animals. En Escherichia coli in the Americas, Torres AG (Ed.) Springer International Publishing, U.S.A., pp 149–72. https://doi.org/10.2174/97816080519221100101
Masana MO, Leotta GA, Del Castillo LL, D’Astek BA, Palladino PM, Galli L (2010) Prevalence, characterization and genotypic analysis of Escherichia coli O157:H7/NM from selected beef exporting abattoirs of Argentina. J Food Protect 73(4):649–56. https://doi.org/10.4315/0362-028x-73.4.649
Tanaro JD, Galli L, Lound LH, Leotta GA, Piaggio MC, Carbonari CC, Irino K, Rivas M (2012) Non-O157:H7 Shiga toxin–producing Escherichia coli in bovine rectums and surface water streams on a beef cattle farm in Argentina. Foodborne Pathog Dis 9(10):878–884. https://doi.org/10.1089/fpd.2012.1182
Masana M, D’Astek BA, Palladino PM, Galli L, del Castillo LL, Carbonari CC, Leotta GA, Vilacoba E, Irino K, Rivas M (2011) Genotypic characterization of non-O157 Shiga toxin–producing Escherichia coli in beef abattoirs of Argentina. J Food Protect 74(12):2008–17. https://doi.org/10.4315/0362-028X.JFP-11-189
Brusa V, Restovich V, Galli L, Teitelbaum D, Signorini M, Brasesco H, Londero A, García D, Padola NL, Superno V, Sanz M, Petroli S, Costa M, Bruzzone M, Sucari A, Ferreghini M, Linares L, Suberbie G, Rodriguez R, Leotta GA (2017) Isolation and characterization of non – O157 Shiga toxin-producing Escherichia coli from beef carcasses, cuts and trimmings of abattoirs in Argentina. PLoS One 12(8):e0183248. https://doi.org/10.1371/journal.pone.0183248
Cundon C, Carbonari CC, Zolezzi G, Rivas M, Bentancor A (2018) Putative virulence factors and clonal relationship of O174 Shiga toxin-producing Escherichia coli isolated from human, food and animal sources. Vet Microbiol 215:29–34. https://doi.org/10.1016/j.vetmic.2017.12.006
Rhades LC, Larzábal M, Bentancor A, Sabio y García J, Babineca FJ, Cataldi A, Amigo N, Baldone VN, Urquiza L, Delicia PJ, Forta MC (2019) A one-year longitudinal study of enterohemorrhagic Escherichia coli O157 fecal shedding in a beef cattle herd. Research in Veterinary Science 127:27–32. https://doi.org/10.1016/j.rvsc.2019.10.001
Beutin L, Geier D, Zimmermann S, Karch H (1995) Virulence markers of Shiga-like toxin-producing Escherichia coli strains originating from healthy domestic animals of different species. J Clin Microbiol 33(3):631–635. https://doi.org/10.1128/jcm.33.3.631-635.1995
Blanco M, Blanco JE, Blanco J, Gonzalez EA, Mora A, Prado C, Fernández L, Rio M, Ramos J, Alonso MP (1996) Prevalence and characteristics of Escherichia coli serotype O157:H7 and other verotoxin-producing E. coli in healthy cattle. Epidemiology & Infection 117(2):251–7. https://doi.org/10.1017/s0950268800001424
Etcheverría AI, Padola NL (2013) Shiga toxin-producing Escherichia coli Factors involved in virulence and cattle colonization. Virulence 4(5):366–372. https://doi.org/10.4161/viru.24642
Fernández D, Rodriguez EM, Arroyo GH, Padola NL, Parma AE (2009) Seasonal variation of Shiga toxin-encoding genes (stx) and detection of E. coli O157 in dairy cattle from Argentina. J Appl Microbiol 106(4):1260–7. https://doi.org/10.1111/j.1365-2672.2008.04088.x
Fan R, Shao K, Yang X, Bai X, Fu S, Sun H, Xu Y, Wang H, Li Q, Hu B, Zhang J, Xiong Y (2019) High prevalence of non-O157 Shiga toxin producing Escherichia coli in beef cattle detected by combining four selective agars. BMC Microbiol 19:213. https://doi.org/10.1186/s12866-019-1582-8
Coombes BK, Wickham ME, Mascarenhas M, Gruenheid S, Finlay BB, Karmali MA (2008) Molecular analysis as an aid to assess the public health risk of non-O157 Shiga toxin-producing Escherichia coli strains. Appl Environ Microbiol 74(7):2153–2160. https://doi.org/10.1128/AEM.02566-07
Byrne L, Jenkins C, Launders N, Elson R, Adak GK (2015) The epidemiology, microbiology and clinical impact of Shiga toxin-producing Escherichia coli in England, 2009–2012. Epidemiology & Infection 143(16):3475–87. https://doi.org/10.1017/S0950268815000746
Valilis E, Ramsey A, Sidiq S, DuPont HL (2018) Non-O157 Shiga toxin-producing Escherichia coli - a poorly appreciated enteric pathogen: systematic review. Int J Infect Dis 76:82–87. https://doi.org/10.1016/j.ijid.2018.09.002
Rivero MA, Padola NL, Etcheverria AI, Parma AE (2004) Escherichia coli Enterohemorragica y Sindrome Urémico hemolítico en Argentina. MEDICINA 64: 352–356 ISSN 1669–9106
Fernández D, Krüger A, Polifroni R, Bustamante AV, Sanso AM, Etcheverría AI, Lucchesi PMA, Parma AE, Padola NL (2013) Characterization of Shiga toxin-producing Escherichia coli O130:H11 and O178:H19 isolated from dairy cows. Front Cell Infect Microbiol 2013(3):9. https://doi.org/10.3389/fcimb.2013.00009
Llorente P, Barnech L, Irino K, Rumi MV, Bentancor A (2014) Characterization of Shiga toxin-producing Escherichia coli isolated from ground beef collected in different socioeconomic strata markets in Buenos Aires, Argentina. Biomed Res Int 2014:795104. https://doi.org/10.1155/2014/795104
Bielaszewska M, Friedrich AW, Aldick T, Schurk-Bulgrin R, Karch H (2006) Shiga toxin activatable by intestinal mucus in Escherichia coli isolated from humans: predictor for a severe clinical outcome. Clin Infect Dis 43:1160–1167. https://doi.org/10.1086/508195
Giugno SM, Bibiloni N, Rahman R, Miliwebsky E, Chinen I, Rivas M (2007) Association between uremic hemolytic syndrome and infection by Shiga toxin-producing Escherichia coli. Acta Bioquímica Clínica Latinoamericana 41:27–33. ISSN 0325–2957; ISSN 1851–6114
Werber D, Beutin L, Pichner R, Stark K, Fruth A (2008) Shiga toxin-producing Escherichia coli serogroups in food and patients, Germany. Emerg Infect Dis 14:1803–1806. https://doi.org/10.3201/eid1411.080361
De Toni F, de Souza EM, Pedrosa FO, Klassen G, Irino K, Un Rigo L, Steffens MBR, Fialho OB, Farah SMSS, Fadel-Picheth CMT (2009) A prospective study on Shiga toxin-producing Escherichia coli in children with diarrhea in Parana State, Brazil. Lett Appl Microbiol 48:645–647. https://doi.org/10.1111/j.1472-765X.2009.02569.x
Buvens G, Lauwers S, Pierard D (2010) Prevalence of subtilase cytotoxin in verocytotoxin-producing Escherichia coli isolated from humans and raw meats in Belgium. Eur J Clin Microbiol Infect Dis 29:1395–1399. https://doi.org/10.1007/s10096-010-1014-z
Miko A, Rivas M, Bentancor A, Delannoy S, Fach P, Beutin L (2014) Emerging types of Shiga toxin-producing E. coli (STEC) O178 present in cattle, deer, and humans from Argentina and Germany. Front Cell Infect Microbiol 4:78. https://doi.org/10.3389/fcimb.2014.00078
Qin X, Klein EJ, Galanakis E, Thomas AA, Stapp JR, Rich S, Buccat AM, Tarr PI (2005) Real-time PCR assay for detection and differentiation of Shiga toxin-producing Escherichia coli from clinical samples. J Clin Microbiol 53(7):2148–2153. https://doi.org/10.1128/JCM.00115-15
Wang L, Jokinen CC, Laing CR, Johnson RP, Ziebell K, Gannon V (2018) Multi-year persistence of verotoxigenic Escherichia coli (VTEC) in a closed Canadian beef herd: a cohort study. Front Microbiol 9:2040. https://doi.org/10.3389/fmicb.2018.02040
Delannoy S, Beutin L, Mariani-Kurkdjian P, Fleiss A, Bonacorsi S, Fach P (2017) The Escherichia coli serogroup O1 and O2 lipopolysaccharides are encoded by multiple O-antigen gene clusters. Front Cell Infect Microbiol 7:1–30. https://doi.org/10.3389/fcimb.2017.00030
Bürk C, Dietrich R, Açar G, Moravek M, Bülte M, Märtlbauer E (2003) Identification and characterization of a new variant of Shiga toxin 1 in Escherichia coli ONT:H19 of bovine origin. J Clin Microbiol 41(5):2106–2112. https://doi.org/10.1128/JCM.41.5.2106-2112.2003
Bai X, Zhang J, Ambikan A, Jernberg C, Ehricht R, Scheutz F, Xiong Y, Matussek A (2019) Molecular characterization and comparative genomics of clinical hybrid Shiga toxin-producing and enterotoxigenic Escherichia coli (STEC/ETEC) strains in Sweden. Sci Rep 9(1):5619. https://doi.org/10.1038/s41598-019-42122-z
Badouei MA, Morabito S, Najafifar A, Mazandarani E (2016) Molecular characterization of enterohemorrhagic Escherichia coli hemolysin gene (EHEC-hlyA)-harboring isolates from cattle reveals a diverse origin and hybrid diarrheagenic strains. Infect Genet Evol 39:342–348. https://doi.org/10.1016/j.meegid.2016.02.002
Johura F, Parveen R, Islam A, Sadique A, Rahim N, Monira S, Khan AR, Ahsan S, Ohnishi M, Watanabe H, Chakraborty S, George CM, Cravioto A, Navarro A, Hasan B, Alam M (2016) Occurrence of hybrid Escherichia coli strains carrying Shiga toxin and heat-stable toxin in livestock of Bangladesh. Front Public Health 4:287. https://doi.org/10.3389/fpubh.2016.00287
Akiyama Y, Futai H, Saito E, Ogita K, Sakae H, Fukunaga M, Tsuji H, Chikahira M, Iguchi A (2017) Shiga toxin subtypes and virulence genes in Escherichia coli isolated from cattle. J Infect Dis 70(2):181–185. https://doi.org/10.7883/yoken.JJID.2016.100
Cheasty T, Rowe B (1983) Antigenic relationships between the enteroinvasive Escherichia coli O antigens O28ac, O112ac, O124, O136, O143, O144, O152, and O164 and Shigella O antigens. J Clin Microbiol 17(4):681–684. https://doi.org/10.1128/jcm.17.4.681-684.1983
Beutin L, Gleier K, Kontny I, Echeverria P, Scheutz F (1997) Origin and characteristics of enteroinvasive strains of Escherichia coli (EIEC) isolated in Germany. Epidemiol Infect 118:199–205. https://doi.org/10.1017/s0950268897007413
da Silva LC, de Mello Santos AC, Silva RM (2017) Uropathogenic Escherichia coli pathogenicity islands and other ExPEC virulence genes may contribute to the genome variability of enteroinvasive E. coli. BMC Microbiology 17(1):68. https://doi.org/10.1186/s12866-017-0979-5
Beutin L, Montenegro MA, Orskov I, Orskov F, Prada J, Zimmermann S, Stephan R (1989) Close association of verotoxin (Shiga-like toxin) production with enterohemolysin production in strains of Escherichia coli. J Clin Microbiol 27(11):2559–2564. https://doi.org/10.1128/jcm.27.11.2559-2564.1989
Vieira MA, Andrade JR, Trabulsi LR, Rosa AC, Dias AM, Ramos SR, Frankel G, Gomes TAT (2001) Phenotypic and genotypic characteristics of Escherichia coli strains of non-enteropathogenic E. coli (EPEC) serogroups that carry eae and lack the EPEC adherence factor and Shiga toxin DNA probe sequences. J Infect Dis 183(5):762–72. https://doi.org/10.1086/318821
Cookson AL, Bennett J, Thomson-Carter F, Attwood GT (2007) Molecular subtyping and genetic analysis of the Enterohemolysin Gene (ehxA) from Shiga toxin-producing Escherichia coli and atypical enteropathogenic E. coli. Appl Environ Microbiol 73(20):6360–9. https://doi.org/10.1128/AEM.00316-07
Zhang S, Yang G, Huang Y, Zhang J, Cui L, Wu Q (2018) Prevalence and characterization of atypical enteropathogenic Escherichia coli isolated from retail foods in China. J Food Prot 81(11):1761–1767. https://doi.org/10.4315/0362-028X.JFP-18-188
Colello R, Velez MV, Gonzáleza J, Montero DA, Bustamante AV, Del Canto F, Etcheverría AI, Vidal R, Padola NL (2018) First report of the distribution of locus of adhesion and autoaggregation (LAA) pathogenicity island in LEE-negative Shiga toxin-producing Escherichia coli isolates from Argentina. Microb Pathog 123:259–263. https://doi.org/10.1016/j.micpath.2018.07.011
Montero DA, Del Canto F, Velasco J, Colello R, Padola NL, Salazar JC, San Martin C, Oñate A, Blanco J, Rasko DA, Contreras C, Puente JL, Scheutz F, Franz E, Vidal RM (2019) Cumulative acquisition of pathogenicity islands has shaped virulence potential and contributed to the emergence of LEE-negative Shiga toxin-producing Escherichia coli strains. Emerg Microbes Infect 8(1):486–502. https://doi.org/10.1080/22221751.2019.1595985
Mosquito S, Ruiz J, Pons MJ, Durand D, Barletta F, Ochoa TJ (2012) Molecular mechanisms of antibiotic resistance in diarrhoeagenic Escherichia coli isolated from children. Int J Antimicrob Agents 40:544–554. https://doi.org/10.1016/j.ijantimicag.2012.07.021
Gonzales L, Joffre E, Rivera R, Sjöling Å, Svennerholm AM, Iñiguez V (2013) Prevalence, seasonality and severity of disease caused by pathogenic Escherichia coli in children with diarrhoea in Bolivia. J Med Microbiol 62:1697–1706. https://doi.org/10.1099/jmm.0.060798-0
Pantozzi FL, Moredo FA, Vigo GB, Giacoboni GI (2010) Antimicrobial resistance in indicator and zoonotic bacteria isolated from domestic animals in Argentina. Revista Argentina de Microbiología 42(1):49–52. https://doi.org/10.1590/S0325-75412010000100011
Boletín Informativo Resistencia Antimicrobiana (2019) Retrieved from: folletoRAM2019Final.pdf (conicet.gov.ar)
Ribeiro LF, Barbosa MMC, Pinto FR, Lavezzo LF, Rossi GAM, Almeida HMS, Amaral LA (2019) Diarrheagenic Escherichia coli in raw milk, wáter and cattle feces in non-technified dairy farms. Ciência Animal Brasileira 20:e-47449. https://doi.org/10.1590/1089-6891v20e-47449
Calderon Toledo C, Arvidsson I, Karpman D (2011) Cross-reactive protection against enterohemorrhagic Escherichia coli infection by enteropathogenic E. coli in a mouse model. Infect Immun 79(6):2224–33. https://doi.org/10.1128/IAI.01024-10
Acknowledgements
This study was performed at the Universidad de Buenos Aires, Facultad de Ciencias Veterinarias. The authors would like to thank the University and the Microbiology department for the assistance during the whole period of work.
Funding
This research was funded by a magister fellowship by Universidad de Buenos Aires, UBACyT 20020190100320, and PICT 2017–3360.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Responsible Editor: Waldir P. Elias
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bonino, M.P., Crivelli, X.B., Petrina, J.F. et al. Detection and analysis of Shiga toxin producing and enteropathogenic Escherichia coli in cattle from Tierra del Fuego, Argentina. Braz J Microbiol 54, 1257–1266 (2023). https://doi.org/10.1007/s42770-023-00958-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-00958-8