Abstract
The microbial conversion of pentoses to ethanol is one of the major drawbacks that limits the complete use of lignocellulosic sugars. In this study, we compared the yeast species Spathaspora arborariae, Spathaspora passalidarum, and Sheffersomyces stipitis regarding their potential use for xylose fermentation. Herein, we evaluated the effects of xylose concentration, presence of glucose, and temperature on ethanol production. The inhibitory effects of furfural, hydroxymethylfurfural (HMF), acetic acid, and ethanol were also determined. The highest ethanol yield (0.44 g/g) and productivity (1.02 g/L.h) were obtained using Sp. passalidarum grown in 100 g/L xylose at 32 °C. The rate of xylose consumption was reduced in the presence of glucose for the species tested. Hydroxymethylfurfural did not inhibit the growth of yeasts, whereas furfural extended their lag phase. Acetic acid inhibited the growth and fermentation of all yeasts. Furthermore, we showed that these xylose-fermenting yeasts do not produce ethanol concentrations greater than 4% (v/v), probably due to the inhibitory effects of ethanol on yeast physiology. Our data confirm that among the studied yeasts, Sp. passalidarum is the most promising for xylose fermentation, and the low tolerance to ethanol is an important aspect to be improved to increase its performance for second-generation (2G) ethanol production. Our molecular data showed that this yeast failed to induce the expression of some classical genes involved in ethanol tolerance. These findings suggest that Sp. passalidarum may have not activated a proper response to the stress, impacting its ability to overcome the negative effects of ethanol on the cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Second-generation (2G) ethanol is produced from lignocellulosic biomass, a renewable energy source alternative to fossil fuels. This biomass is mainly composed of cellulose and hemicellulose, which can be hydrolyzed into hexoses and pentoses and then converted into ethanol by fermenting microorganisms [1,2,3,4]. One of the largest challenges for the efficient production of 2G ethanol is the microbial conversion of pentose into ethanol. These sugars comprise 25–40% of the lignocellulosic biomass, with xylose representing the main pentose present in lignocellulosic hydrolysates [5]. Ethanol production from xylose would largely contribute to the economic viability of 2G ethanol.
Strains of Saccharomyces cerevisiae, which are widely used in the industrial production of first-generation (1G) ethanol, do not naturally ferment pentoses due to the absence of specific transporters and the low expression of genes encoding the enzymes for xylose metabolism [6,7,8,9]. Additionally, the strains of S. cerevisiae genetically engineered for this purpose still present limitations, which results in xylose-fermenting rates not as high as those obtained with glucose [10]. Thus, to improve xylose use, it is important to enhance our knowledge about the complex regulation regarding the metabolism of this sugar. In this regard, the study and domestication of natural pentose-fermenting microorganisms have been considered as an interesting alternative in the field of 2G ethanol.
The yeasts Spathaspora arborariae, Scheffersomyces stipitis, and Spathaspora passalidarum can convert xylose into ethanol, making them potential candidates for lignocellulosic biomass fermentation [5, 11,12,13,14,15,16]. For example, Sp. arborariae was isolated from rotting wood samples collected at the National Park of Serra do Cipó and at the Rio Doce State Park [13]. The strain Sc. stipitis NRRL 7124 presents an endosymbiotic relationship with the beetle Odontotaenius disjunctus, which inhabits decaying wood [16, 17]. Spathaspora passalidarum NRRLY 27,907 has also been isolated from the intestine of wood-eating beetles [18]. The species Sc. stipitis and Sp. passalidarum are considered promising for industrial fermentation because they present good fermentative yields under anaerobic conditions [14, 19].
The species studied here were chosen according to the following criteria: Sc. stipitis for being the xylose-fermenting species with more scientific information accumulated so far, but with much yet to be investigated about its metabolism; the yeast Sp. passalidarum because it recently emerged as the most promising species for ethanol production from xylose; and Sp. arborariae, a yeast that belongs to the same genus as Sp. passalidarum but has few studies available in the literature. Thus, we suppose that a comparison with Sp. arborariae could provide important information about the physiological evolution of this genus of xylose-fermenting yeasts.
Besides the ability to generate high ethanol yields from xylose, other factors that influence ethanol production from hydrolysates should be considered when evaluating fermenting microorganisms. During fermentation, microorganisms are exposed to stressful industrial conditions that can compromise their growth and cellular metabolism, such as osmotic stress due to the high initial sugar concentrations, elevated temperatures, especially during the summer and in tropic countries, the generation of inhibitors from biomass pretreatment, including furfural, 5-hydroxymethylfurfural (HMF), and acetic acid, as well as ethanol accumulation in the culture medium [20].
In this context, our main research question was as follows: What are the physiological limitations of xylose-fermenting yeasts that impede their use for ethanol production? The hypothesis proposed and tested is that these yeasts can ferment xylose with high yields, but the low tolerance to fermentation conditions is the limiting factor in the process.
To date, respective studies have focused on the fermentative capacity of Sp. arborariae, Sp. passalidarum, and Sc. stipitis, while there are few comparative studies that show how those factors affect xylose fermentation in these yeasts. Furthermore, the relevant aspects of yeast physiology that will orientate strain selection or breeding programs are not widely addressed. Thus, the present work aimed to characterize the fermentative capacity of xylose-fermenting yeast strains (Sp. arborariae HM 19.1A, Sp. passalidarum NRRLY 27,907, and Sc. stipitis NRRL 7124) under different cultivation conditions. The results presented here contribute to the comprehension of the physiology of xylose fermentation and highlight the bottlenecks for future breeding programs.
Materials and methods
Strains and media
We used the yeasts Sp. arborariae HM 19.1A, Sp. passalidarum NRRLY 27,907, and Sc. stipitis NRRL 7124 in this study. For cell maintenance and activation, YPD medium [2% yeast extract, 1% peptone, and 2% glucose (w/v)] was used. To obtain solid medium, 2% agar (w/v) was added to YPD. All yeasts were stored and maintained in 20% glycerol (v/v) at -80 °C.
Fermentation assays
Yeasts were grown overnight in YPX (20 g/L yeast extract, 10 g/L peptone, and 20 g/L xylose) under agitation of 120 rpm at 28 °C. The cells were centrifuged and inoculated into the fermentation medium (5.0 g/L yeast extract, 5.0 g/L peptone, 2.0 g/L NH4Cl, 1.0 g/L KH2PO4, 0.3 g/L MgSO4.7H2O) [16] containing xylose (40, 80, or 100 g/L). The initial OD600 nm was adjusted to 2 [21], corresponding to 3.8 g/L of cell dry weight (CDW) for Sp. arborariae, 1.5 g/L for Sp. passalidarum, and 3.9 g/L for Sc. stipitis. The flasks were incubated at 28ºC for 70 h at 120 rpm. The best concentration of xylose for ethanol production was determined and used to evaluate the effects of different temperatures (28, 32, and 35 °C) on the fermentation efficiency of the yeasts. Co-fermentation was carried out by adding glucose (20 or 40 g/L) to the medium containing a predetermined xylose concentration and at a certain temperature.
Tolerance assays
Tolerance tests to furfural, HMF, and acetic acid were carried out at the temperature at which the optimum fermentative parameters were determined (see above). The assays were performed in 125-mL Erlenmeyer flasks containing fermentation medium supplemented with xylose (20 g/L) and one of the inhibitors (2.5 g/L furfural, 0.5 g/L HMF, or 3 g/L acetic acid) at 120 rpm for 60 h. The inhibitor concentrations were chosen based on the literature [5, 22]. Ethanol tolerance assays were performed under the same conditions, except for the addition of 2, 4, or 6% (v/v) ethanol instead of the pretreatment compounds.
Fermentation parameters
Ethanol yield (YP/S, g/g) was estimated using Eq. 1, in which Pf is ethanol final mass (g), Pi is ethanol initial mass (g), Si is sugar initial mass (g), and Sf is sugar final mass (g).
Ethanol volumetric productivity (QP, g/L.h) was estimated using Eq. 2, in which Pf is ethanol final concentration (g/L), Pi is ethanol initial concentration (g/L), and t is fermentation period (h).
Fermentation efficiency (Y%) was estimated using Eq. 3, in which the experimental YP/S (g/g) was divided by the theoretical YP/S (0.51 g/g), and the result multiplied by 100.
Total sugar consumption rate (g/L.h) was estimated using Eq. 4, in which Sf is sugar final concentration (g/L), Si is sugar initial concentration (g/L), and t is consumption period (h).
A calibration curve correlating OD600 nm with CDW (g/L) was constructed for each carbon source and for each yeast to evaluate their growth. The specific growth rate (h−1) was determined by the angular coefficient from a linear regression of the plot ln OD (600 nm) versus time (h) during the yeasts’ exponential (log) growth phase. The consumption yield (g/g) was determined by the angular coefficient from a linear regression of the plot CDW (g/L) versus the sugars’ concentrations (glucose or xylose) (g/L). The specific sugar consumption rate (gsugar/gCDW.h) was determined by multiplying the consumption yields (g/g) for each sugar (xylose and glucose) by the specific growth rate (h−1) of each yeast. Unpaired two-tailed t test (GraphPad Prism 5) was used for the statistical analysis.
Analytical methods
Quantitative determination of substrate consumption and metabolite production was carried out by high-performance liquid chromatography (HPLC) using the Bio-Rad Aminex HPX-87 column at 45 °C. Sulfuric acid (5 mM), applied at a flow rate of 0.7 mL/min, was used as the eluent for separation. The column was coupled to an RID-10A refractive index detector.
Total RNA extraction and quantitative real-time PCR (qPCR) analysis
Sp. passalidarum cells pre-cultured in YPX medium overnight were transferred to 125-mL Erlenmeyer flasks containing 20 mL of fresh medium (initial OD600 nm ~ 0.20). The flasks were incubated at 32 °C at 150 rpm until the cells reached the mid-exponential growth phase (OD600 nm ~ 1.0). The cultures’ volumes were divided into two new flasks, with one containing the cells growing without ethanol and the other having the cells treated with ethanol to a final concentration of 4% (v/v). The flasks were incubated for 2 h in the conditions described above. Three independent biological replicates were performed.
Total RNA extraction was performed with RNeasy Mini kit (Qiagen). The samples were quantified by Qubit 3.0 Fluorometer (Invitrogen) and treated with RNase-free DNase I (Promega). cDNA synthesis was performed with the High-Capacity cDNA Reverse Transcription kit (Applied Biosystems) according to the following cycle: 10 min at 25 °C, 120 min at 37 °C, and 5 min at 85 °C. The qPCR analysis was performed using Power SYBR Green PCR Master Mix (Applied Biosystems) with StepOne™ Real-Time PCR System (Applied Biosystems). The assays were carried out in technical duplicate using the conditions as follows: 10 min at 95ºC and 40 cycles of 15 s at 95ºC, 1 min at 60ºC. Eight Sp. passalidarum genes orthologs of S. cerevisiae genes that are involved in ethanol response (TDH1, ADH1, ADH7, TPS1, HSP30, HSP104, SOD1, and MSN2/4) were selected for qPCR analysis. Primers were designed with Primer3Plus [23], and nucleotide specificity was evaluated by Primer-BLAST (https://www.ncbi.nlm.nih.gov/tools/primer-blast/index.cgi) [22].
Relative mRNA quantification was performed by the standard curve method. A standard curve was obtained for each gene by plotting the average Ct versus the log10 of different cDNA concentrations from control sample (0.39 – 50.0 ng/µL). The results were normalized using ACT1 and RDN18 [24] as internal controls. Unpaired two-tailed t test (GraphPad Prism 5) was used for the statistical analysis. Primer sequences and genes accession numbers are available in Supplementary File S1.
Results
Effects of xylose concentration and temperature on ethanol production
Xylose fermentation by Sp. arborariae, Sp. passalidarum, and Sc. stipitis was evaluated at different concentrations of xylose at 28ºC (Fig. 1). The species Sp. arborariae produced the largest amount of biomass and the lowest ethanol concentration and was the only yeast that produced xylitol under the tested conditions (Fig. 1a-c). The effects of different xylose concentrations on the fermentation parameters were analyzed (Table 1). Ethanol volumetric productivity (QP) was higher in Sp. passalidarum, which is consistent with the lowest biomass achieved. Fermentation efficiencies (Y%) did not vary as a function of xylose concentration.
Fermentation kinetics in 40, 80, or 100 g/L of xylose. Spathaspora arborariae (a, b, and c), Spathaspora passalidarum (d, e, and f), and Scheffersomyces stipitis (g, h, and i). Fermentation lasted 50 h in (a) and 70 h in the other experiments (b-i). Xylose (●); xylitol (
 ); glycerol (
); glycerol (
 ); ethanol (
); ethanol (
 ); cell dry weight (
); cell dry weight (
 )
)
The highest ethanol concentration (g/L) was achieved when 100 g/L of xylose was used. The highest ethanol yields were obtained at 40 h of fermentation for Sp. passalidarum (YP/S 0.40 g/g), 70 h for Sp. arborariae (YP/S 0.32 g/g), and 50 h for Sc. stipitis (YP/S 0.35 g/g). Overall, xylose was completely consumed in less than 70 h, except for Sp. arborariae grown in 100 g/L, which consumed about 92% of xylose (Fig. 1). As 100 g/L sugar favored the production of high ethanol concentrations, it was chosen to evaluate the effects of different temperatures in further fermentation assays.
For Sp. arborariae, biomass and ethanol production were lower at 35 °C (Fig. 2a-c, Table 2). At 32 and 35 °C, the yeast presented higher yields of xylitol (0.28 and 0.34 g/g, respectively) compared to ethanol (0.19 and 0.18 g/g, respectively). The highest ethanol yield (0.32 g/g) was achieved at 28 °C after 70 h of fermentation (Table 2). Biomass formation and ethanol production by Sp. passalidarum were similar regardless of the temperature (Fig. 2d-f). In contrast to Sp. arborariae, Sp. passalidarum produced more ethanol than xylitol (Fig. 2a-f). Importantly, ethanol volumetric productivity (QP) and fermentation efficiency (Y%) were higher at 32ºC when compared to the other two species (Table 2). The species Sc. stipitis also produced similar levels of ethanol and biomass at the evaluated temperatures (Fig. 2g-i, Table 2). The highest ethanol concentration (36.8 g/L) was obtained after 60 h of fermentation at 32ºC, whereas xylitol production was temperature-dependent, as observed in Sp. arborariae. However, xylitol concentration did not exceed ethanol concentration (Fig. 2g-i).
Fermentation kinetics in fermentation medium containing 100 g/L of xylose. Fermentation was carried out at different temperatures by Spathaspora arborariae (a, b, and c), Spathaspora passalidarum (d, e, and f), and Scheffersomyces stipitis (g, h, and i). Xylose (●); xylitol (
 ); glycerol (
); glycerol (
 ); ethanol (
); ethanol (
 ); cell dry weight (
); cell dry weight (
 )
)
The yeasts showed faster xylose consumption at 28 °C compared to the other temperatures (Fig. 2a, d and g), and the sugar was completely consumed by Sp. passalidarum and Sc. stipitis within 40 h of fermentation (Fig. 2d and g). On the other hand, Sp. arborariae did not completely assimilate xylose at any temperature (Fig. 2a-c). The species Sp. passalidarum consumed 100% of xylose within 60 h at 32 °C and 92% within 70 h at 35 °C (Fig. 2e and f). Similarly, Sc. stipitis assimilated 97 and 94% of xylose at 32 and 35 °C, respectively (Fig. 2h and i). Based on the presented results, further experiments were carried out at 28ºC for Sp. arborariae and at 32ºC for Sp. passalidarum and Sc. stipitis.
Glucose and xylose co-fermentation
In the co-fermentation assays, the yeasts consumed all glucose between 20 and 30 h of fermentation (Fig. 3). The rates of total sugar consumption were higher in Sp. passalidarum and Sc. stipitis than in Sp. arborariae (Table 3).
In media containing 20 g/L glucose, ethanol production by Sp. passalidarum and Sc. stipitis was 29.28 and 29.27 g/L, respectively, whereas 25.25 and 32.62 g/L were achieved when 40 g/L glucose was used. The amount of ethanol produced by Sp. arborariae was similar under both conditions. Glucose decreased the fermentation efficiency (Y%) of Sp. arborariae and Sp. passalidarum in the co-fermentation assays, although this parameter was unaffected in Sc. stipitis. Ethanol productivity by yeasts was statistically higher in the absence of glucose (Table 4).
Xylitol production in 20 or 40 g/L glucose was higher in Sp. arborariae (13.3 and 15.8 g/L), followed by Sp. passalidarum (6.5 and 8.1 g/L) and Sc. stipitis, which produced xylitol concentrations below 6 g/L (Fig. 3). Again, Sp. arborariae produced more xylitol when compared to the other yeasts.
Effects of compounds generated from pre-treatment on the fermentative capacity of yeasts
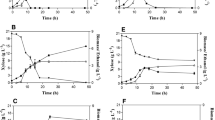
The effects of furfural, HMF, and acetic acid on the fermentative capacity of Sp. arborariae, Sp. passalidarum, and Sc. stipitis were analyzed (Fig. 4). In the absence of inhibitors, Sp. arborariae consumed all xylose and reached the maximum ethanol production (7.7 g/L) within 30 h (Fig. 4a). In the presence of furfural, the yeast had a long lag phase, and biomass was effectively produced only after 30 h of cultivation (Fig. 4d). Growth coincided with xylose consumption and depletion of furfural. In the presence of HMF, biomass production was similar to that of the control, xylose consumption occurred in the initial hours of cell cultivation, and ethanol production was not impaired (Fig. 4g).
Yeast fermentation profiles in the absence and presence of inhibitors. Furfural or hydroxymethylfurfural (HMF) was added into the YPX media at different concentrations. Spathaspora arborariae (a, d, and g), Spathaspora passalidarum (b, e, and h), and Scheffersomyces stipitis (c, f, and i). Xylose (●); ethanol (
 ); furfural (
); furfural (
 ); hydroxymethylfurfural (
); hydroxymethylfurfural (
 ); cell dry weight (
); cell dry weight (
 )
)
In the control condition, almost all xylose was consumed by Sp. passalidarum within 9 h of fermentation, and ethanol production reached 10.2 g/L (Fig. 4b). Furfural also extended the yeast’s lag phase and slowed xylose consumption. Effective biomass production occurred only after the inhibitor was completely depleted (20 h). The ethanol concentration reached its maximum (9.1 g/L) at 30 h (Fig. 4e). Hydroxymethylfurfural concentration was reduced by 42.4% within 3 h. After 12 h, a complete depletion of xylose was seen and 11.8 g/L of ethanol had been produced (Fig. 4h). No significant difference in ethanol yield (YP/S) was found when comparing the control and HMF conditions.
The species Sc. stipitis produced 8.1 g/L of ethanol after 30 h when grown in the presence of only xylose (Fig. 4c). Unlike the other species, Sc. stipitis reduced 60% of furfural within 3 h and 99% within 6 h; ethanol production was 8.23 g/L at 30 h (Fig. 4f). In the presence of HMF, xylose was completely consumed within 20 h, when the maximum ethanol production of 10.2 g/L was achieved. The HMF was depleted after 3 h (Fig. 4i).
The yeasts were not able to grow and produce ethanol in the presence of (3 g/L) acetic acid, and therefore, its concentration did not change over time (data not shown). During the experiment, only small amounts of xylose were consumed, especially in the early hours. For these reasons, the fermentative parameters were not determined.
Effect of ethanol on fermentation
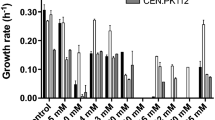
The species Sp. arborariae, Sp. passalidarum, and Sc. stipitis consumed all xylose within 30, 12, and 20 h, respectively, when grown in media containing 2% (v/v) ethanol (Fig. 5a-c). The presence of 4% (v/v) of ethanol reduced the yeasts growth (Fig. 5d-f). At this condition, Sp. arborariae did not consume all sugar within the 60 h of fermentation (Fig. 5a). Xylose consumption was also delayed in Sp. passalidarum and Sc. stipitis in this ethanol concentration. However, Sp. passalidarum consumed all sugar within 20 h, whereas Sc. stipitis consumed about 98% of xylose within 60 h (Fig. 5b and c). The species Sp. arborariae, Sp. passalidarum, and Sc. stipitis did not grow at 6% (v/v) of ethanol (Fig. 5d-f), and under this condition, xylose concentration was reduced by 26, 36.7, and 22.5%, respectively (Fig. 5a-c).
Effect of ethanol on the expression of classical stress response genes in Sp. passalidarum
The yeast Sp. passalidarum was selected for further molecular studies in the presence of ethanol stress. The expression of eight canonical genes with well-documented roles in different mechanisms of stress response in S. cerevisiae was analyzed in Sp. passalidarum under ethanol stress [25, 26]. These genes are involved in several processes such as carbon metabolism and fermentation (ADH1, ADH7, TDH1, and TPS1), protein disruption (HSP104), ATPase regulation (HSP30), superoxide detoxification (SOD1), and regulation of transcription in response to different stresses (MSN2/4). Under stress, the expression of the genes TDH1, ADH1, TPS1, HSP30, and MSN2/4 was down-regulated, while the expression of SOD1 was up-regulated. No significant difference was detected in the expression of ADH7 and HSP104 (Fig. 6).
Expression of genes related to ethanol response in Spathaspora passalidarum. The expression profile of genes involved in S. cerevisiae response to ethanol (TDH1, ADH1, ADH7, TPS1, HSP30, HSP104, SOD1, and MSN2/4) was analyzed in Sp. passalidarum. Total RNA was extracted from cells grown in the absence (white column) or presence (black column) of 4% (v/v) ethanol after 2 h of stress exposure. The graphs represent the estimated mean and standard deviation values. Asterisks denote statistical significance by the T test (*, p-value < 0.05; **, p-value < 0.01; ***, p-value < 0.001)
Discussion
Ethanol production of Sp. arborariae HM19.1A, Sp. passalidarum NRRLY 27,907, and Sc. stipitis NRRL 7124 was evaluated at three different xylose concentrations. In agreement with other reports [14, 27], we observed that the yeasts consumed all substrate present in the medium and ethanol was the main product formed. Overall, Sp. passalidarum presented the best yields and ethanol production values, followed by Sc. stipitis. Veras and collaborators [14] also reported good performances of Sp. passalidarum and Sc. stipitis upon the comparison of the fermentative capacity of different xylose-consuming yeasts. Similarly, Cadete and collaborators [24] showed that Sp. passalidarum stands out as the best xylose fermenter when compared to other yeasts of the same genus.
Spathaspora arborariae had the lowest ethanol yield and presented the highest xylitol production. Xylitol is a by-product of the xylose assimilation pathway. First, xylose is reduced to xylitol by the enzyme xylose reductase (XR), which is then oxidized to xylulose by the enzyme xylitol dehydrogenase (XDH). Xylulose is phosphorylated by the enzyme xylulokinase, forming xylulose-5-phosphate, which is metabolized by the pentose-phosphate pathway and then directed to glycolysis, where ethanol is produced [28]. Other studies have also demonstrated xylitol production in synthetic media and lignocellulosic hydrolysates by Sp. arborariae [5, 29]. In the present study, higher xylitol concentrations were obtained with increasing temperatures. Overall, other yeast species that produce xylitol have also shown better yields at cultivation temperatures ranging from 30 to 37 °C [30], which can be due to changes in the enzymatic activities of xylose reductase and xylitol dehydrogenase. Several optimal temperature values for the XR enzyme are reported in the literature, varying according to the yeast species [31]. Future studies with Sp arborariae, the yeast that produced the highest xylitol amounts, should investigate this interesting feature aiming at xylitol production.
The simultaneous consumption of glucose and xylose is an important feature for microorganisms that ferment lignocellulosic biomass as these two sugars are the most abundant carbohydrates in this substrate [32]. Our results demonstrate that glucose reduced the rate of xylose consumption in the co-fermentation assays. Increasing concentrations of glucose resulted in a greater reduction in the xylose consumption rate of all yeasts tested. This might be due to sugar competition by transporters [16] and inhibition of the synthesis and/or activity of xylose metabolism enzymes [16, 32]. However, even after glucose depletion, the estimated xylose consumption rates were lower when compared to the fermentation assay containing only xylose. Thus, the presence of glucose possibly affects the cellular metabolism in a global way, as suggested by Conrad and collaborators [33]. Collectively, our results show that the presence of glucose decreased ethanol productivity of the yeasts and reduced ethanol yields. In fact, it was recently shown that glucose inhibits xylose metabolization in Sp. passalidarum by decreasing the expression of genes and the activity of key enzymes of xylose metabolism [34].
The final ethanol concentration produced in the presence of furfural and HMF was similar to the amount observed in the control condition. The presence of 2.5 g/L of furfural delayed yeast cell growth and ethanol production until the inhibitor could no longer be detected in the medium. Some S. cerevisiae strains can reduce furfural to furfuryl alcohol, which is less toxic to the cell, a process known as in situ detoxification [35]. Probably, the xylose-fermenting yeasts evaluated in the present study have a similar reducing mechanism. Some methods of lignocellulosic biomass pretreatment can also minimize furfural concentration in the raw material. Tanaka and collaborators [36] demonstrated that a pretreatment with two-step acid hydrolysis at a moderate temperature reduced furfural content in the hydrolysate. Another alternative to minimize inhibitor effect is to increase the initial cell concentration for fermentation with non-detoxified hydrolysates. A higher initial cell concentration of Blastobotry adeninivorans degraded furfural in non-detoxified acid hydrolysate more quickly, leading to complete consumption of sugars and increased ethanol production [37].
Fermentation was not inhibited by 0.5 g/L HMF. Our study shows that the yeasts were able to metabolize HMF and consume xylose simultaneously with no influence on the lag phase, indicating that a detoxifying mechanism is present in the yeasts. According to the literature, some strains of S. cerevisiae can convert hydroxymethylfurfural to 2,5-bis-hydroxymethylfuran, a compound that is less toxic to the cells [38]. Despite our results showing that HMF did not affect the yeasts fermentation in synthetic medium, the synergistic effect of this compound along with the other pretreatment inhibitors in lignocellulosic hydrolysates can interfere with yeasts growth and fermentation abilities. Thus, the generation of microorganism with evolved higher tolerance to pretreatment inhibitors is an important strategy to overcome this issue [5, 39, 40].
Acetic acid can be highly toxic to yeasts due to diffusion through the plasma membrane and dissociation in the cytoplasm, which reduces the intracellular pH and affects cell metabolism [41]. Our results show that acetic acid affected cell growth, xylose consumption, and ethanol production of the xylose-fermenting yeasts. Other authors have also reported a high toxicity of acetic acid for yeasts such as S. cerevisiae and Sc. (Pichia) stipitis [41,42,43]. The detoxification of hydrolysates can occur before fermentation to remove or decrease the concentration of the inhibitory compounds, reducing their harmful effects on the process [44].
One of the most common stresses that yeasts face during fermentation is the accumulation of ethanol. At high concentrations, ethanol promotes a complex stress that affects different cellular processes and structures, which can end up limiting yeast cell growth, viability, and fermentative activity [45]. The yeasts studied in this work have a low ethanol tolerance profile as they were unable to grow at 6% (v/v) ethanol. The species Sp. passalidarum was less affected by 4% (v/v) ethanol and presented a higher ethanol tolerance compared to the other yeasts. Still, the improvement of Sp. passalidarum ethanol tolerance would largely contribute to accelerating its biotechnological use for 2G bioethanol production since the ethanol titer obtained in these fermentations varies between 5 and 7% [20]. Such ethanol concentrations are higher than the ones this yeast can support and remain fermenting.
Based on the screening we have performed in this work with the three xylose-fermenting yeasts, we noticed that their low ethanol tolerance was an interesting subject to further our studies. Ethanol is the final product of fermentative metabolism. Thus, the production and tolerance to high ethanol concentrations are desirable traits for fermentative processes. Much effort has been employed so far to understand the stress response of S. cerevisiae [45,46,47,48,49,50,51,52,53,54,55,56]. The phenotype of low ethanol tolerance presented by non-conventional yeasts has not received much attention. Our results pointed out that Sp. passalidarum was the best candidate for the context of xylose fermentation among the yeasts studies. Thus, we performed gene expression analysis with Sp. passalidarum growing in a medium containing 4% (v/v) ethanol for 2 h to investigate the expression of classical genes involved in yeast ethanol response. The qPCR results showed that the genes encoding the glycolytic enzyme glyceraldehyde-3-phosphate dehydrogenase 1 (TDH1) and the fermentation enzyme alcohol dehydrogenase 1 (ADH1) were down-regulated by ethanol, while the expression of the gene encoding ADH7, another alcohol dehydrogenase, was not significantly altered. This profile suggests that even in mild ethanol concentrations, Sp. passalidarum reduces glycolytic flux and ethanol production to minimize the stress effect on the cells that would enhance with ethanol accumulation.
TPS1 and HSP30 genes were also down-regulated in Sp. passalidarum under stress, which is the opposite to the demonstrated in the literature for S. cerevisiae [46]. Trehalose accumulation in S. cerevisiae increases resistance to several adverse environmental conditions, and the overexpression of TPS1 (trehalose-6-phosphate synthase) improves ethanol tolerance [47]. Hsp30p is a membrane heat-shock protein that regulates H + -ATPase Pma1p activity, having roles in preventing ATP depletion during cytosolic deacidification in the presence of ethanol [48, 49]. The expression of HSP104—a chaperone with disaggregase function that is responsive to a wide range of environmental stimuli [50]—was not significantly altered by ethanol stress in our assay. In S. cerevisiae, the expression of HSP104 increases with increasing ethanol concentrations [51], and its expression is regulated by several transcription factors, such as Msn2/4p and Hsf1p [52]. In Sp. passalidarum, the transcription of this gene was less effective and the expected up-regulation was not observed.
Interestingly, SOD1 (Cu/Zn superoxide dismutase), which is involved in the detoxification of superoxides, was the only up-regulated gene among the ones analyzed in our study. The expression of this gene is induced by oxidative stress, which is also caused by ethanol [53]. The formation of superoxides can promote cellular apoptosis and affect the structures of lipids, proteins, and nucleic acids [54]. The overexpression of SOD1 improves ethanol tolerance in S. cerevisiae [55]. Thus, Sp. passalidarum probably reduced part of the damage caused by reactive oxygen species (ROS) and maintained cell viability by a similar mechanism.
The MSN-like gene of Sp. passalidarum was down-regulated under ethanol stress. Msn2p and Msn4p are partially redundant stress-responsive transcription factors of S. cerevisiae that regulate the expression of more than 200 genes in response to various environmental stresses, including ethanol [46, 56]. Sp. passalidarum presents only one copy of these transcription factors. The down-regulation of this gene in Sp. passalidarum may have affected the gene expression profile of this yeast and its final response to ethanol stress.
Conclusion
Among the three xylose-fermenting yeasts evaluated in the present study, Sp. passalidarum stood out as the best candidate to in-depth the studies regarding ethanol production from xylose based on its fermentative parameters in the conditions assessed. When analyzing the physiological responses of Sp. passalidarum in different sugar concentrations and temperatures, presence of glucose, furfural, HMF, acetic acid, and ethanol, our data showed that the negative effect of glucose on xylose metabolism, and the low tolerance to acetic acid and ethanol appear as the most important aspects to be improved to increase its performance for the production of 2G ethanol.
Regarding the ethanol stress response of Sp. passalidarum, our expression analysis showed that some genes important for cells facing the effects of ethanol challenge were down-regulated in this yeast subjected to 4% (v/v) ethanol. It suggests that Sp. passalidarum may not have activated a proper response to cope with ethanol stress, which could have contributed to the low tolerance profile exhibited by the yeast. Future studies investigating how Sp. passalidarum regulates gene expression in response to ethanol stress can contribute to obtaining more robust strains of this yeast with increased ethanol tolerance.
Availability of data and material
Not applicable.
Code availability
Not applicable.
References
Sun Y, Cheng J (2002) Hydrolysis of lignocellulosic materials for ethanol production: a review. Bioresour Technol 83:1–11. https://doi.org/10.1016/S0960-8524(01)00212-7
Soccol CR, Vandenberghe LPDS, Medeiros ABP et al (2010) Bioethanol from lignocelluloses: Status and perspectives in Brazil. Bioresour Technol 101(13):4820–4825. https://doi.org/10.1016/j.biortech.2009.11.067
Limayem A, Ricke SC (2012) Lignocellulosic biomass for bioethanol production: Current perspectives, potential issues and future prospects. Prog Energy Combust Sci 38(4):449–467. https://doi.org/10.1016/j.pecs.2012.03.002
Canilha L, Chandel AK, Suzane Dos Santos Milessi T et al (2012) Bioconversion of sugarcane biomass into ethanol: An overview about composition, pretreatment methods, detoxification of hydrolysates, enzymatic saccharification, and ethanol fermentation. J Biomed Biotechnol 1-15.https://doi.org/10.1155/2012/989572
da Cunha-Pereira F, Hickert LR, Sehnem NT et al (2011) Conversion of sugars present in rice hull hydrolysates into ethanol by Spathaspora arborariae, Saccharomyces cerevisiae, and their co-fermentations. Bioresour Technol 102(5):4218–4225. https://doi.org/10.1016/j.biortech.2010.12.060
Cadete RM, Rosa CA (2017) The yeasts of the genus Spathaspora: potential candidates for second-generation biofuel production. Yeast Primer 35(2):191–199. https://doi.org/10.1002/yea.3279
Kuyper M, Hartog MMP, Toirkens MJ et al (2005) Metabolic engineering of a xylose-isomerase-expressing Saccharomyces cerevisiae strain for rapid anaerobic xylose fermentation. FEMS Yeast Res 5:399–409. https://doi.org/10.1016/j.femsyr.2004.09.010
Nevoigt E (2008) Progress in metabolic engineering of Saccharomyces cerevisiae. Microbiol Mol Biol Rev 72(3):379–412. https://doi.org/10.1128/MMBR.00025-07
Salusjärvi L, Kankainen M, Soliymani R et al (2008) Regulation of xylose metabolism in recombinant Saccharomyces cerevisiae. Microb Cell Fact 16:1–16. https://doi.org/10.1186/1475-2859-7-18
Sharma S, Arora A (2020) Tracking strategic developments for conferring xylose utilization/fermentation by Saccharomyces cerevisiae. Ann Microbiol 70(50):1–17. https://doi.org/10.1186/s13213-020-01590-9
Agbogbo FK, Coward-Kelly G, Torry-Smith M, Wenger KS (2006) Fermentation of glucose/xylose mixtures using Pichia stipitis. Process Biochem 41(11):2333–2336. https://doi.org/10.1016/j.procbio.2006.05.004
Hickert LR, De S-C, Rosa CA, Ayub MAS (2013) Simultaneous saccharification and co-fermentation of un-detoxified rice hull hydrolysate by Saccharomyces cerevisiae ICV D254 and Spathaspora arborariae NRRL Y-48658 for the production of ethanol and xylitol. Bioresour Technol 143:112–116. https://doi.org/10.1016/j.biortech.2013.05.123
Cadete RM, Santos RO, Melo MA et al (2009) Spathaspora arborariae sp. nov., a D-xylose-fermenting yeast species isolated from rotting wood in Brazil. FEMS Yeast Res 9(8):1338–1342. https://doi.org/10.1111/j.1567-1364.2009.00582.x
Veras HCT, Parachin NS, Almeida JRM (2017) Comparative assessment of fermentative capacity of different xylose-consuming yeasts. Microb Cell Fact 16(153):1–8. https://doi.org/10.1186/s12934-017-0766-x
Lachke A (2002) Biofuel from D-xylose – The second most abundant sugar. Reson 7(5):50–58. https://doi.org/10.1007/BF02836736
Hou X (2012) Anaerobic xylose fermentation by Spathaspora passalidarum. Appl Microbiol Biotechnol 94(1):205–214. https://doi.org/10.1007/s00253-011-3694-4
Pignal M-C (1967) Une nouvelle espece de levure isole e de larves d’insectes: Pichia stipitis. Bull Mens Soc Linn Lyon 36:163–168. https://doi.org/10.3406/linly.1967.5915
Nguyen NH, Suh SO, Marshall CJ, Blackwell M (2006) Morphological and ecological similarities: wood-boring beetles associated with novel xylose-fermenting yeasts, Spathaspora passalidarum gen. sp. nov. and Candida jeffriesii sp. nov. Mycol Res 110(10):1232–1241. https://doi.org/10.1016/j.mycres.2006.07.002
Hou X, Yao S (2012) Improved inhibitor tolerance in xylose-fermenting yeast Spathaspora passalidarum by mutagenesis and protoplast fusion. Appl Microbiol Biotechnol 93(6):2591–2601. https://doi.org/10.1007/s00253-011-3693-5
Deparis Q, Claes A, Foulquié-Moreno MR, Thevelein JM (2017) Engineering tolerance to industrially relevant stress factors in yeast cell factories. FEMS Yeast Res 17(4):1–17. https://doi.org/10.1093/femsyr/fox036
de Souza CJA, Costa DA, Rodrigues MQRB et al (2012) The influence of pre saccharification, fermentation temperature and yeast strain on ethanol production from sugarcane bagasse. Bioresour Technol 109:63–69. https://doi.org/10.1016/j.biortech.2012.01.024
Ye J, Coulouris G, Zaretskaya I, Cutcutache I et al (2012) Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatic 13(134):1–11. https://doi.org/10.1186/1471-2105-13-134
Untergasser A, Nijveen H, Rao X, Bisseling T (2007) Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Res 35:71–74. https://doi.org/10.1093/nar/gkm306
Cadete RM, de las Heras AM, Sandström AG et al (2016) Exploring xylose metabolism in Spathaspora species: XYL1.2 from Spathaspora passalidarum as the key for efficient anaerobic xylose fermentation in metabolic engineered Saccharomyces cerevisiae. Biotechnol Biofuels 9(167):1–14. https://doi.org/10.1186/s13068-016-0570-6
Caspeta L, Castillo T, Nielsen J (2015) Modifying yeast tolerance to inhibitory conditions of ethanol production processes. Front Bioeng Biotechnol 3:1–15. https://doi.org/10.3389/fbioe.2015.00184
Ma M, Liu LZ (2010) Quantitative transcription dynamic analysis reveals candidate genes and key regulators for ethanol tolerance in Saccharomyces cerevisiae. BMC Microbiol 10(169):1–20. https://doi.org/10.1186/1471-2180-10-169
Nakanishi SC, Soares LB, Biazi LE et al (2017) Fermentation strategy for second generation ethanol production from sugarcane bagasse hydrolyzate by Spathaspora passalidarum and Scheffersomyces stipitis. Biotechnol Bioeng 114(10):2211–2221. https://doi.org/10.1002/bit.26357
Kim SR, Park YC, Jin YS, Seo JH (2013) Strain engineering of Saccharomyces cerevisiae for enhanced xylose metabolism. Biotechnol Adv 31:851–861. https://doi.org/10.1016/j.biotechadv.2013.03.004
Cadete RM, Melo MA, Dussán KJ et al (2012) Diversity and Physiological Characterization of D-Xylose- Fermenting Yeasts Isolated from the Brazilian Amazonian Forest. PLoS One 7(8):e43135. https://doi.org/10.1371/journal.pone.0043135
de Albuquerque TL, da Silva IJ, de Macedo GR, Rocha MPV (2014) Biotechnological production of xylitol from lignocellulosic wastes: A review. Process Biochem 49(11):1779–1789. https://doi.org/10.1016/j.procbio.2014.07.010
Lugani Y, Puri M, Singh B (2021) Recent insights, applications and prospects of xylose reductase: a futuristic enzyme for xylitol production. Eur Food Res Technol 247(4):921–946. https://doi.org/10.1007/s00217-020-03674-x
Harner NK, Wen X, Bajwa PK et al (2015) Genetic improvement of native xylose-fermenting yeasts for ethanol production. J Ind Microbiol Biotechnol 42(1):1–20. https://doi.org/10.1007/s10295-014-1535-z
Conrad M, Schothorst J, Kankipati HN et al (2014) Nutrient sensing and signaling in the yeast Saccharomyces cerevisiae. FEMS Microbiol Rev 38:254–299. https://doi.org/10.1111/1574-6976.12065
Ribeiro LE, Albuini FM, Castro AG et al (2021) Influence of glucose on xylose metabolization by Spathaspora passalidarum. Fungal Genet Biol 157:103624. https://doi.org/10.1016/j.fgb.2021.103624
Liu ZL, Slininger PJ, Gorsich SW (2005) Enhanced biotransformation of furfural and hydroxymethylfurfural by newly developed ethanologenic yeast strains. Appl Biochem Biotechnol 121:451–460. https://doi.org/10.1385/ABAB:121:1-3:0451
Tanaka K, Koyama M, Thi P et al (2019) Production of high-concentration bioethanol from cassava stem by repeated hydrolysis and intermittent yeast inoculation. Int Biodeterior Biodegrad 138:1–7. https://doi.org/10.1016/j.ibiod.2018.12.007
Pham P, Tuyet-Le D, ThuHuong L et al (2019) Recycling cassava stem to bioethanol by inoculating a novel xylose – glucose fermenting yeast at high initial concentration. Environ Prog Sustain Energy 39(1):e13286. https://doi.org/10.1002/ep.13286
Liu ZL (2011) Molecular mechanisms of yeast tolerance and in situ detoxification of lignocellulose hydrolysates. Appl Microbiol Biotechnol 90(3):809–825. https://doi.org/10.1007/s00253-011-3167-9
Soares LB, Bonan CIDG, Biazi LE et al (2020) Biomass and bioenergy investigation of hemicellulosic hydrolysate inhibitor resistance and fermentation strategies to overcome inhibition in non-Saccharomyces species. Biomass Bioenergy 137:105549. https://doi.org/10.1016/j.biombioe.2020.105549
Pacheco TF, Machado BRC, Júnior WGDM (2021) Enhanced tolerance of Spathaspora passalidarum to sugarcane bagasse hydrolysate for ethanol production from xylose. Appl Biochem Biotechnol. https://doi.org/10.1007/s12010-021-03544-6
van Zyl C, Prior BA, du Preez JC (1991) Acetic acid inhibition of d-xylose fermentation by Pichia stipitis. Enzyme Microb Technol 13(1):82–86. https://doi.org/10.1016/0141-0229(91)90193-E
Chen Y, Sheng J, Jiang T et al (2016) Transcriptional profiling reveals molecular basis and novel genetic targets for improved resistance to multiple fermentation inhibitors in Saccharomyces cerevisiae. Biotechnol Biofuels 9(9):1–18. https://doi.org/10.1186/s13068-015-0418-5
Wang X, Jin M, Balan V et al (2014) Comparative metabolic profiling revealed limitations in xylose-fermenting yeast during co-fermentation of glucose and xylose in the presence of inhibitors. Biotechnol Bioeng 111(1):152–164. https://doi.org/10.1002/bit.24992
Jönsson LJ, Alriksson B, Nilvebrant N-O (2013) Bioconversion of lignocellulose: inhibitors and detoxification. Biotechnol Biofuels 6(16):1–10. https://doi.org/10.1186/1754-6834-6-16
Stanley D, Bandara A, Fraser S et al (2010) The ethanol stress response and ethanol tolerance of Saccharomyces cerevisiae. J Appl Microbiol 109(1):13–24. https://doi.org/10.1111/j.1365-2672.2009.04657.x
Ma M, Liu ZL (2010) Mechanisms of ethanol tolerance in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 87(3):829–845. https://doi.org/10.1007/s00253-010-2594-3
Divate NR, Chen G, Divate RD et al (2017) Metabolic engineering of Saccharomyces cerevisiae for improvement in stresses tolerance. Bioengineered 8(5):524–535. https://doi.org/10.1080/21655979.2016.1257449
Aguilera F, Peinado RA, Millán C et al (2006) Relationship between ethanol tolerance, H + -ATPase activity and the lipid composition of the plasma membrane in different wine yeast strains. Int J Food Microbiol 110:34–42. https://doi.org/10.1016/j.ijfoodmicro.2006.02.002
Piper PW, Ortiz-Calderon C, Holyoak C et al (1997) Hsp30, the integral plasma membrane heat shock protein of Saccharomyces cerevisiae, is a stress-inducible regulator of plasma membrane H(+)-ATPase. Cell Stress Chaperones 2(1):12–24
Bond U (2006) Stressed out ! Effects of environmental stress on mRNA metabolism. FEMS Yeast Res 6:160–170. https://doi.org/10.1111/j.1567-1364.2006.00032.x
Kempf C, Lengeler K, Wendland J (2017) Differential stress response of Saccharomyces hybrids revealed by monitoring Hsp104 aggregation and disaggregation. Microbiol Res 200:53–63. https://doi.org/10.1016/j.micres.2017.03.009
Grably MR, Stanhill A, Tell O, Engelberg D (2002) HSF and Msn2/4p can exclusively or cooperatively activate the yeast HSP104 gene. Mol Microbiol 44(1):21–35. https://doi.org/10.1046/j.1365-2958.2002.02860.x
Bleoanca I, Rita A, Silva C et al (2013) Relationship between ethanol and oxidative stress in laboratory and brewing yeast strains. J Biosci Bioeng 116(6):697–705. https://doi.org/10.1016/j.jbiosc.2013.05.037
Bailey SM, Pietsch EC, Cunningham CC (1999) Ethanol stimulates the production of reactive oxygen species at mitochondrial complexes I and II. Free Radic Biol Med 27:891–900. https://doi.org/10.1016/s0891-5849(99)00138-0
Wang H, Ji B, Ren H (2014) The relationship between lysine 4 on histone H3 methylation levels of alcohol tolerance genes and changes of ethanol tolerance in Saccharomyces cerevisiae. Microb Biotechnol 7(4):307–314. https://doi.org/10.1111/1751-7915.12121
Gasch AP, Spellman PT, Kao CM et al (2000) Genomic Expression Programs in the Response of Yeast Cells to Environmental Changes. Mol Biol Cell 11:4241–4257. https://doi.org/10.1091/mbc.11.12.4241
Acknowledgements
This work was supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG, grant numbers APQ-01525-14 and APQ-01229-15), and the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES, Finance Code 001).
Author information
Authors and Affiliations
Contributions
LF, CA, and WS conceived and designed the research. VC, LR, FA, and PF conducted the experiments. AG performed the statistical analysis. LF, VC, LR, FA, and AG analyzed the data. All authors contributed to the writing of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Campos, V.J., Ribeiro, L.E., Albuini, F.M. et al. Physiological comparisons among Spathaspora passalidarum, Spathaspora arborariae, and Scheffersomyces stipitis reveal the bottlenecks for their use in the production of second-generation ethanol. Braz J Microbiol 53, 977–990 (2022). https://doi.org/10.1007/s42770-022-00693-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-022-00693-6







 ); xylose (●); xylitol (
); xylose (●); xylitol (
 ); glycerol (
); glycerol (
 ); ethanol (
); ethanol (
 ); cell dry weight (
); cell dry weight (
 )
)

 ); ethanol 2% (
); ethanol 2% (
 ); ethanol 4% (
); ethanol 4% (
 ); ethanol 6% (
); ethanol 6% (
 )
)