Abstract
The aim of this study was to investigate the prevalence of thermophilic Campylobacter in the broiler production chain of southern Brazil, by evaluating broiler farms and slaughter line samples, and to determine the genetic diversity, antimicrobial resistance, and virulence genes of the isolates. Of the 140 samples investigated in this study, 75 (53.6%) were positive for thermophilic Campylobacter, and all isolates were identified by phenotypic and molecular tests as C. jejuni. The resistance to nalidixic acid was the most common (74%), followed by resistance to enrofloxacin (67.3%) and ciprofloxacin (37.1%). However, there was no resistance to the macrolides tested which are recommended for the treatment of human campylobacteriosis. The PFGE showed that the isolates were grouped in eight macrorestriction patterns (P1 to P8). A representative isolate of each macrorestriction pattern was investigated for the presence of virulence genes and all isolates carried the cadF, ciaB, cdtA, cdtB, cdtC, and flaA genes. The dnaJ gene was detected in 87.5% (7/8) of the isolates. The flhA and racR genes were detected in 75% (6/8), while the pldA gene was present in 62.5% (5/8) and the wlaN gene in 25% (2/8). The presence of C. jejuni in broiler farms and in the slaughterhouse is a hazard to consumer given that this pathogen can be maintained throughout the broiler production chain and contaminates the final product. Moreover, the presence of the major virulence genes in the isolates demonstrates that they have the ability to develop campylobacteriosis in humans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Campylobacter spp. is among the main pathogens that cause foodborne gastroenteritis in the world [1, 2]. Human campylobacteriosis is caused by the thermophilic Campylobacter species: C. jejuni, C. coli, C. lari, and C. upsaliensis [3]. It is estimated that Campylobacter spp. causes 1.5 million illnesses each year in the USA [4]. The notification rate in the European Union (EU) was 246,571 cases of human campylobacteriosis in 2018 [5]. Campylobacter jejuni species is currently attracting significant attention because it is involved in approximately 90% of human enteritis cases associated with campylobacteriosis [6].
The most common clinical symptoms of Campylobacter infections include diarrhea (frequently bloody), abdominal pain, fever, headache, nausea, and vomiting, which can progress to more severe cases, such as bacteremia, hepatitis, and pancreatitis, or to systemic infections that can lead to Guillain-Barré syndrome (GBS) and reactive arthritis [2]. The symptoms of campylobacteriosis are usually self-limiting, but in severe cases, it is necessary to introduce antimicrobial treatment [2]. In these cases, macrolides and fluoroquinolones are the frontline agents used in the treatment of the disease [7]. However, resistance to quinolones and macrolides has been reported and associated with the indiscriminate use of these antimicrobial agents in food-producing animals [8]. Therefore, antimicrobial susceptibility tests play a critical role, not only in determining the appropriate therapeutic intervention but also for the epidemiological surveillance of the resistance of this pathogen [9].
Domestic and wild animals are natural reservoirs of thermophilic Campylobacter. Birds, especially broiler chickens, are considered to be primary reservoirs of these microorganisms and may be their asymptomatic carriers [1, 2, 10]. Campylobacter jejuni colonizes the chicken gut, primarily the ceca at the distal end of the gastrointestinal tract to levels in excess of 109 CFU. g−1 [11]. This hampers control of this pathogen in broiler houses and, thus, introduces the potential risk of contamination of broiler meat as a result of contact with fecal content during the slaughtering stages [12]. The consumption of contaminated broiler meat is a major cause of human campylobacteriosis throughout the world [2].
Unlike other pathogens, such as Salmonella spp. and Escherichia coli, studies about thermophilic Campylobacter virulence mechanisms are still not very enlightening [6, 10]. Nevertheless, it is known that the main mechanisms involved with the virulence of these microorganisms are adhesion, invasion and toxin production, during pathogenesis in humans [6]. The cadF gene encodes a protein that interacts with the host fibronectin extracellular matrix, participating in the colonization of the cell surface [13]. The ciaB gene encodes a protein involved in cell invasion [14]. Campylobacter jejuni synthesizes the cytolethal toxin (CDT) that inside the host cell, this protein will potentiate the cell cycle block, resulting in DNA degradation [15].
In Brazil, there is still little data about thermophilic Campylobacter, especially due to difficulties with its isolation and identification, which is more complex than for other pathogens [16]. In this context, the aim of this study was to investigate the prevalence of thermophilic Campylobacter in the broiler production chain of southern Brazil, by evaluating broiler farms and slaughter line samples, and to determine the genetic diversity, antimicrobial resistance and virulence genes of the isolates.
Materials and methods
Sampling
Three sampling events (October, November, and December 2014) were carried out on commercial broiler farms located in the southern Brazil. A total of 45 floor feces samples and three drag swabs were collected from flocks with broilers aged between 40 and 45 days old from three different broiler farms (A, B, and C). The sampling comprised a drag swab sample and 15 floor feces sample pools (each pool containing three fresh fecal samples) on each farm. Fresh fecal samples were collected in sterile flasks and kept refrigerated. The drag swab was sampled using a hydrated sterile sponge (3M™, Brazil). The sponge was dragged along the henhouse. For the broiler slaughterhouse two sampling events (June and August 2015) were performed and four broiler flocks were sampled, totaling 92 samples. In the first sampling event, flocks I and II were sampled and in the second sampling event, flocks III and IV were sampled. Briefly, three broiler carcasses from each flock were sampled during the slaughtering process: after bleeding (n = 12), scalding (n = 12), defeathering (n = 12), evisceration (n = 12), and chilling (n = 12). Additionally, the cecum (n = 12) and the liver (n = 12) were extracted from each labeled carcass after the evisceration step. Furthermore, scalding water samples were collected before (n = 2) and after the slaughter (n = 2), and chiller water was collected before (n = 2) and after slaughter (n = 2). A surface smear technique was performed by rubbing the surfaces of the carcasses with a hydrated sterile sponge. All samples were packaged in sterile bags and refrigerated. Intestinal samples were placed in sterile bags, and cecal samples were collected with the aid of cloacal swabs. Water samples were collected in sterile flasks containing 1% sodium thiosulfate. The livers were also packed in sterile bags, and the sampling was carried out by weighing 10 g of each sample. The scalding and chiller water sampling process was performed with a 10 mL aliquot of the sample.
All samples were transferred to the Food Microbiology Laboratory of the Department of Science and Agroindustrial Technology of FAEM/UFPel.
Isolation and identification of thermophilic Campylobacter
The isolation and phenotypic identification of thermophilic Campylobacter were performed in accordance with the International Organization for Standardization [17] with the addition of 2 mg. L−1 of potassium clavulanate (Sigma-Aldrich) to Bolton broth (Oxoid®). The samples were submitted for selective enrichment in Bolton broth (Oxoid®) at 42 °C for 24 h in microaerophilic conditions (5% O2, 10% CO2, and 85% N2) (White Martins®). The inoculation was then performed on mCCD agar (Oxoid®) and Preston agar (Oxoid®) supplemented with selective supplement SR0155E (Oxoid®, cefoperazone and amphotericin B) and SR0117E (Oxoid®, polymyxin B, rifampicin, trimethoprim, and cycloheximide), respectively, and with incubation at 42 °C under microaerophilic atmosphere for 48 h. The characteristics of the isolates were analyzed by morphological identification and by the phenotypic tests of oxidase, catalase, indoxyl acetate, and sodium hippurate. The isolates confirmed as thermophilic Campylobacter were preserved in a freezing medium (fetal bovine serum, nutrient broth with yeast extract, FBP supplement and glycerol) and stored at − 80 °C. The C. jejuni ATCC 33291, C. lari NCTC 11352, and C. coli CAMPY 1008 standard strains, which were provided by the Oswaldo Cruz Foundation (FIOCRUZ) in Rio de Janeiro, were used as positive controls for isolation and phenotypic identification of the isolates.
DNA extraction and species confirmation
The extraction of genomic DNA from the isolates was performed according to the protocol proposed by Sambrook and Russell [18]. The PCR conditions and primers used to confirm the genus and to differentiate between the species are listed in Table 1. The primers were previously described by Josefsen et al. [25], and the multiplex PCR protocol proposed by Maćkiw et al. [26]. The DNA of C. jejuni was used as a negative control for the C. coli primer, and the DNA of C. coli was used as a negative control for the C. jejuni primer.
Determination of antimicrobial resistance
Resistance to antimicrobials was tested using the Kirby-Bauer disk diffusion method [27]. The antimicrobials tested were ciprofloxacin (CIP 5 μg), enrofloxacin (ENR 5 μg), nalidixic acid (NAL 30 μg), azithromycin (AZI 15 μg), clarithromycin (CLA 15 μg), erythromycin (ERI 15 μg), doxycycline (DOX 30 μg), and tetracycline (TET 30 μg). The antimicrobial agents and criteria of interpretation were performed according to the European Committee on Antimicrobial Susceptibility Testing [28]. Enrofloxacin and nalidixic acid were evaluated according the recommendations of the Clinical and Laboratory Standards Institute [29].
PFGE
All isolates were subjected to pulsed-field gel electrophoresis (PFGE) as described by Ribot et al. [30] and recommended by PulseNet (http://www.cdc.gov/pulsenet/protocos.htm), using SmaI and KpnI as restriction endonucleases and Salmonella Braenderup H9812 as the reference marker (digested with XbaI). KpnI was used to verify the diversity of the isolates that had an indistinguishable pattern by SmaI digestion. The fragments generated were separated by electrophoresis using the CHEF system - DR II (Bio-Rad, USA), stained with ethidium bromide, and visualized with the photo-documentation L-Pix system (Loccus Biotecnologia®). The molecular patterns obtained were analyzed with BioNumerics software Version 7.1 (Applied Maths, Belgium) using the binary Dice similarity coefficient with 2% tolerance position, and the groups were determined by the unweighted pair-group method with arithmetic mean (UPGMA). The results were analyzed according to criteria established by Wieczorek et al. [31].
Detection of virulence genes by PCR
A representative isolate of each macrorestriction pattern identified in the PFGE was selected. PCR was performed to verify the genes involved with the virulence of thermophilic Campylobacter. The genes searched were docA, racR, dnaJ, cadF (responsible for colonization and adhesion), pldA, ciaB (responsible for invasion), cdtA, cdtB, cdtC (cytotoxin production), flhA, flaA (involved in motility), and wlaN (ganglioside mimics in Guillain-Barré syndrome (GBS)). PCR conditions and the primers used were previously described by the authors specified in Table 1. For separation of the amplified products, electrophoresis (80 V, 120 min) was performed on 1.5% agarose gel. The visualization and analysis of the amplification products were performed through the L-Pix photo-documentation system (Loccus Biotecnologia®). As a positive control of the reaction, the standard strain C. jejuni ATCC 33291 was used.
Statistical analysis
Tukey’s test at 5% significance level (Statistic Software 8.0) was used to determine if there are significant difference in the isolation rate of thermophilic Campylobacter between both the sampling events and the sampling points evaluated. The differences to compare the antimicrobial resistance rates were assessed by a chi-squared test. P values of less than 0.05 (p < 0.05) were deemed to be representative of significance.
Results
Thermophilic Campylobacter prevalence
The prevalence of thermophilic Campylobacter in the samples from broilers fecal pools, drag swabs, broiler carcasses, offal, scalding water, and chiller water was 53.6% (75/140). A total of 33.3% (16/48) of the farm samples was positive for thermophilic Campylobacter. On just one of three broiler farms (farm C), thermophilic Campylobacter was detected, and on this broiler farm, 100% (16/16) of the samples were positive for these microorganisms (Table 2). In the slaughterhouse, 64.1% (59/92) of the samples from broiler carcasses, offal and water assessed during the slaughtering steps were contaminated by thermophilic Campylobacter (Table 2). There was no significant difference in the isolation rate of the microorganism in the sampling points evaluated (p > 0.05). However, the average isolation rate of thermophilic Campylobacter differed significantly between the two collections performed at the slaughterhouse. The average isolation rate between the flocks also showed a significant difference (p > 0.05). The flock III differed from the flocks II and IV, which showed no difference between them. The flock I did not differ significantly from any evaluated lot.
Campylobacter jejuni identification
From 75 positive samples, two presumptive colonies of Campylobacter from each culture medium were evaluated and 116 isolates were obtained. They were submitted to phenotypic identification, molecular typing and evaluated for resistance to antimicrobial agents. In the phenotypic tests, all the isolates had typical morphology, gram negative staining, produced catalase and oxidase, were indoxyl acetate negative, and sodium hippurate positive. Preston agar was more effective (p < 0.05) than mCCD agar for the isolation of C. jejuni (Table 2). All 116 isolates were confirmed as C. jejuni by PCR assay.
Antimicrobial resistance
Among the 116 isolates, 74% (86/116) were resistant to at least one antimicrobial tested. The resistance to nalidixic acid was the most common (74%), followed by resistance to enrofloxacin (67.3%), ciprofloxacin (37.1%), tetracycline (25%), and doxycycline (25%). None of the isolates exhibited resistance to macrolides (azithromycin, clarithromycin, and erythromycin). The isolates were significantly more resistant to nalidixic acid and enrofloxacin. There was no statistically significant difference between the resistant and susceptible isolates to ciprofloxacin, tetracycline, and doxycycline. The antimicrobial resistance percentages are shown in Fig. 1. None of the C. jejuni isolates was characterized as multidrug-resistant.
PFGE typing
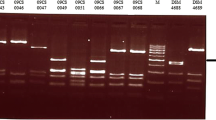
All the 116 isolates were submitted to macrorestriction analysis (PFGE). All the isolates from broiler farm C shared the same SmaI-macrorestriction pattern (P8). On the other hand, the isolates of the broiler slaughterhouse showed different clonal relationships between them and were grouped into seven distinct PFGE restriction patterns (P1 to P7). The isolates obtained in the first sampling of the slaughterhouse showed only two different restriction patterns (P2 and P7), while the isolates from the second sampling event exhibited a greater genetic variability, being grouped into five distinct genotypic patterns (P1, P3, P4, P5, and P6). Thus, eight different patterns were detected among the 116 isolates. Figure 2 shows the restriction patterns obtained by SmaI endonuclease. The restriction pattern of all isolates was confirmed by digestion with the restriction enzyme KpnI. Isolates from the same sample and different culture medium exhibited indistinguishable restriction patterns, with the exception of two isolates from the chiller water samples after slaughtering, two isolates from a carcass after evisceration and two isolates from a carcass after chilling, which also showed different PFGE patterns and antimicrobial resistance profiles (Table 2).
Dendgrogram and macroresctriction patterns of Campylobacter jejuni isolates from broiler farm (P8) and slaughterhouse line (P1–P7) obtained with the resctriction enzyme SmaI. The similarity analysis was performed using the Dice coefficient and UPGMA method (tolerance of 2%). The similarity of the Campylobacter jejuni isolates obained from from broiler farms and slaughterhouse line are given as percentages at the major junctions in the dendrogram
Virulence genes
All representative isolates of the eight macrorestriction patterns (P1 to P8) presented the genes docA, cadF, ciaB, cdtA, cdtB, cdtC, and flaA. The dnaJ gene was detected in 87.5% (7/8) of the isolates. The flhA and racR genes were detected in 75% (6/8), while the pldA gene was present in 62.5% (5/8) and the wlaN gene in 25% (2/8). Two isolates showed all the virulence genes studied. Three isolates showed all the genes except the wlaN (Fig. 3).
Discussion
The prevalence of C. jejuni in the samples from broilers fecal pools and drag swabs obtained in this study was 33% (16/48). This prevalence is quite variable in different studies. Høg et al. [32] isolated Campylobacter in 21% and 4% of the samples from broiler flocks in Denmark and Norway, respectively. Out of 101 cloacal swab samples, 44 (43.7%) were positive for Campylobacter from an abattoir in Denmark [33]. Thermophilic Campylobacter were isolated from 85%, 98%, and 80% of droppings, cecal content, and neck skin, respectively, from broilers on some farms and in slaughterhouses in Algeria [34]. These differences may be due to the type and number of samples, different sampling methods, transport conditions, isolation methodology and different sanitary conditions on broiler farms [16].
All isolates of Campylobacter were identified by phenotypic and molecular tests as C. jejuni, which is the species most commonly found along the productive chain of broilers. In addition, this is the main species associated with human campylobacteriosis [1, 2]. Campylobacter jejuni was isolated only on farm C among the three broiler farms evaluated. Farm C had two broiler houses, while the other two farms had only one. The number of broiler houses on the farm may have influenced the presence of Campylobacter. Høg and co-workers [32] performed a survey of risk factors for Campylobacter spp. colonization of broilers and demonstrated that an increased number of broiler houses on a farm was a risk factor. This can be explained by the transmission between the houses of broilers on the same property via footwear, clothes, unclean hands, or equipment [35].
It is noteworthy that on farm C, all samples were positive for C. jejuni. It is known that once a broiler is contaminated with Campylobacter, the pathogen quickly disseminates to all broilers in the flock and consequently to the farm environment [36]. Besides that, the high body temperature of the broilers, coprophagia and fecal excretion are also important factors for the dissemination of these microorganisms in broiler flocks [37].
The prevalence of C. jejuni in the broiler slaughterhouse obtained was 64.1% (59/92). This is of concern because of the risk of human campylobacteriosis associated with the consumption of contaminated broiler meat and its by-products. It is estimated that 20 to 30% of campylobacteriosis cases in humans in the EU can be attributed to the consumption and handling of broiler meat [38]. Similar results to those found in this research were reported in other studies that assessed the presence of thermophilic Campylobacter in a broiler slaughterhouse [34]. In contrast, García-Sánchez et al. [39] evaluated a total of 494 samples (defeathering machine, evisceration machine, floor, sink, conveyor belt, shackles, and broiler meat) in order to isolate C. jejuni and the prevalence was 94.5% in a poultry slaughterhouse in the North of Spain. In other study, the prevalence of Campylobacter spp. in broiler ceca and broiler skin samples were 11.2% (11/98) and 51% (50/98), respectively, in a slaughterhouse in Thailand [40].
Among the thermophilic Campylobacter species, C. jejuni is the species most isolated in the productive chain of broilers [41, 42] and is also responsible for 90% of cases of human campylobacteriosis [6]. Our results showed that 75% (9/12) of the cecum samples were positive for C. jejuni. The presence of this pathogen in the cecum contents postmortem suggests that the high isolation rate of this microorganism in the carcasses along the slaughtering line may be related to their presence in the broiler intestine before slaughter. Seliwiorstow et al. [43] revealed that the higher the level of colonization of Campylobacter in the broiler’s cecum, the greater the number of carcasses contaminated after plucking, evisceration, washing, and refrigeration. According to Bull et al. [44], after Campylobacter spp. colonizes the intestine of broiler, it is excreted, leading to the rapid spread of microorganisms throughout the entire flock in the period before slaughter. Moreover, the presence of the microorganism in the carcasses after bleeding indicates that the feathers may be contaminated, which may have occurred during transportation to the slaughterhouse [45].
Our results are similar to the findings of Gruntar et al. [46], in a study which also isolated the bacteria in the broiler carcasses after the scalding and defeathering steps. Besides this, the microorganism was found in the scalding water sampled after slaughter in this study. The presence of C. jejuni in the carcasses after evisceration may be related to the leakage of intestinal contents, which is reinforced by the fact that the pathogen was isolated in the liver of broilers; as such, this contamination can be associated with disruption of the intestine during evisceration. According to Rosenquist et al. [47], the rupture of the intestines of broilers during evisceration is not rare in slaughterhouses because the equipment used does not usually fit natural variations in the size of the carcasses. Moreover, the vertical position of the carcass during slaughter steps can facilitate the spread of fecal contamination from its surface.
There was a high prevalence (83.3%) of C. jejuni in the carcasses after cooling by immersion, although the water temperature of the chiller (2.8 °C) was in accordance with the standards stipulated by the current Brazilian legislation which requires that the water temperature does not exceed 4 °C and allows the addition of up to 5 ppm of chlorine in the water. Similar results were found by Franchin et al. [48], who reported a contamination rate of 91% after this step and affirmed that the low temperature and the presence of chlorine were insufficient to prevent contamination by this microorganism. It is important to note that C. jejuni was isolated in the chiller water sampled after slaughter; however, there was no presence of the pathogen in the water before the start of the process. This indicates that cross-contamination occurs in this step and that the chiller water may contribute to the redistribution of C. jejuni contamination among the carcasses.
Resistance to quinolones (ciprofloxacin, nalidixic acid, and enrofloxacin) and tetracyclines (tetracycline and doxycycline) was observed among the C. jejuni isolates. Despite this, the isolates were susceptible to macrolides. Similar results were also observed in other countries [34, 49]. Resistance to quinolone and tetracyclines has increased among Campylobacter spp. isolates worldwide [26, 38]. This has become a public health concern because fluoroquinolones, such as ciprofloxacin, are recommended for treatment of campylobacteriosis in humans [50]. The main cause of the increase in quinolone resistance is the abusive use in veterinary medicine, especially in the broiler production chain, for purposes of growth promotion, disease prevention, treatment, or control [34, 51].
The antimicrobial resistance observed in this study highlights the importance of adopting restrictive measures to control the use of these antimicrobials. It is noteworthy that enrofloxacin was administered to the broilers belonging to the flocks evaluated in this study; as such, it is likely that the quinolone resistance observed is related to the medication administered, as reported in other studies [52, 53]. Several authors have described how the use of antimicrobials as therapeutic, prophylactic, or growth promoter agents in farmed animals can contribute to the development of resistant strains. These can reduce the effectiveness of veterinary and human pharmaceutical products and can promote selective pressure, favoring the multiplication of isolates resistant to antimicrobials [34, 51].
In Brazil, antimicrobial agents, such as enrofloxacin, are freely marketed to producers without veterinary prescription. This may contribute to the selection of isolates resistant to the active principle. In Australia, for example, where fluoroquinolones are not licensed for use in broilers, Miflin et al. [54] found no ciprofloxacin resistance in isolates of C. jejuni and C. coli. The lowest percentages of resistance (25%) were observed for tetracycline class drugs (tetracycline and doxycycline), which is an interesting result from a clinical point of view, since they are alternative antimicrobial agents that may be administered in the treatment of campylobacteriosis. Furthermore, the isolates did not show resistance to the three macrolides (azithromycin, clarithromycin, and erythromycin) tested, which are recommended for the treatment of human campylobacteriosis, and none had multidrug resistance.
The molecular typing of the isolates from farm C by PFGE showed that all of them (28) shared the same SmaI-macrorestriction pattern. Even when the digestion with KpnI endonuclease was performed, different patterns were not obtained among the isolates, a finding which was also reported in other studies [55, 56]. A clonal profile among C. jejuni isolates from the same flock was also previously described [55]. According to Workman et al. [57], this clonal relationship among the C. jejuni isolates may indicate a common contamination origin in positive broiler farms.
The macrorestriction patterns of the C. jejuni isolates identified in the first sampling event from the slaughterhouse (P2 and P7) were genetically distinct from those observed in the second sampling event (P1, P3, P4, P5, and P6), as shown in Fig. 2. The results obtained through the analysis of the genetic diversity of C. jejuni indicate that the contamination of the broiler carcasses may occur during all slaughter operations, and that it is dependent on the previous colonization of the intestines of broiler. A similar observation was made by Gruntar et al. [55]. This shows that the control of this microorganism will require interventions both in slaughterhouses and on broiler farms, for example, through the implementation of biosecurity programs. The search for the main virulence genes in C. jejuni demonstrated the virulence potential of these isolates, since the P4 and P5 patterns included isolates that presented all 11 virulence genes investigated in the present study, and in the P1, P2, and P7 restriction patterns the isolates only did not carry the wlaN gene.
The isolates from the same sample, but from different culture media, exhibited indistinguishable genotypic profiles, with the exception of the chiller water samples after slaughter, carcasses after evisceration, and carcasses after chilling. This result is particularly interesting, reinforcing the importance of using more than one culture medium for the isolation of C. jejuni and of evaluating all isolates in order to obtain more accurate answers. The isolates obtained from the chiller water sample after slaughter, for example, besides presenting distinct genotypic restriction patterns after analysis by the PFGE technique, had different antimicrobial resistance profiles: the isolate obtained from the MCCD agar was susceptible to all antibiotics tested, while the isolate from the Preston agar showed resistance to nalidixic acid, enrofloxacin, and ciprofloxacin.
The flaA, ciaB and cadF genes are widely present in C. jejuni [6, 58], but despite this, there are divergences regarding the constitutive presence of these genes in this microorganism. Although the flaA gene is considered a conservative gene in Campylobacter spp., it is reported that not all the isolates present this gene [58]. In any case, studies using mutant strains show that the flaA gene is essential for the pathogen colonization process [59], demonstrating the virulence potential of the isolates obtained in this study, since all carry this gene. The presence of the cadF gene is of extreme importance for the pathogenesis of the microorganism, since adhesion is probably a prerequisite for the invasion of epithelial cells by any bacterial pathogen. The ciaB gene is considered to be constitutive in C. jejuni; however, Samad et al. [60] found that only 76.25% (61/80) of C. jejuni isolates from broiler meat carried this gene. It is known that mutants with deletions in the cadF and ciaB genes had a reduction in their ability for adhesion and internalization in cellular models [61, 62].
The docA, dnaJ, and racR genes are associated with the pathogen adhesion process to the host. The pldA gene is related to the mechanism of cell invasion and synthesis of outer membrane phospholipases. Konkel et al. [63] evaluated the adhesion capacity of wild and mutant isolates (dnaJ−) in broilers and verified that adhesion capacity was compromised when the infection occurred by the mutant isolates. There are also some studies that demonstrate the deficient colonization of C. jejuni when there is deletion of the genes racR and docA [64].
The presence of the cdtABC operon in all isolates of the present study demonstrates its high virulence potential, since the CDT toxin promotes DNA damage; therefore, its presence is supposed to be associated with the severity of the disease caused by C. jejuni. Besides that, Gagnaire et al. [65] report that chronic exposure of a tissue to bacteria that secrete CDT increases the likelihood of tumor development. In addition, He et al. [66] recently demonstrated that a C. jejuni isolate was able to promote colorectal cancer in mice through the genotoxic action of cdtB.
It is known that the molecular mimicry of host structures by the saccharide portion of C. jejuni lipopolysaccharides is associated with the development of autoimmune sequelae seen in GBS. The wlaN gene is responsible for expressing these lipopolysaccharides. In this study, the wlaN gene was detected with a low frequency, in agreement with other reports [21].
In broiler slaughterhouses, the contamination by C. jejuni was dependent on the previous colonization of the intestine of the broilers, typically determined by the slaughter flock. The isolates were resistant to nalidixic acid, ciprofloxacin and enrofloxacin. This resistance is likely to result from the medication administered to animals on broiler farms. However, the isolates did not show resistance to the macrolides tested (azithromycin, clarithromycin, and erythromycin), which are recommended for the treatment of campylobacteriosis in humans, and did not present multidrug-resistance.
The molecular restriction patterns obtained by PFGE showed that the contamination came from a single source on the broiler farm. Besides that, the presence of C. jejuni in broilers during the period of production on farms represents a potential risk for the introduction of this pathogen to the slaughterhouse environment. This could be observed, given that C. jejuni contamination was identified in broiler carcasses at all stages of the slaughter process, as well as in offal, scalding water and chilled water in the slaughterhouse that received broilers from the evaluated farms. Moreover, the presence of the major virulence genes demonstrates that these isolates have the ability to develop campylobacteriosis in humans.
References
EFSA (2018) The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2017. doi: https://doi.org/10.2903/j.efsa.2018.5500
WHO (2018) Campylobacter. https://www.who.int/news-room/fact-sheets/detail/campylobacterjejuni
Romero MR, D’agostino M, Arias AP, Robles S, Casado CF, Iturbe LO, Lerma OG, Andreou M, Cook N (2016) An immunomagnetic separation/loop-mediated isothermal amplification method for rapid direct detection of thermotolerant Campylobacter spp. during poultry production. J Appl Microbiol 120:469–477. https://doi.org/10.1111/jam.13008
CDC (2019) Campylobacter (Campylobacteriosis) https://www.cdc.gov/campylobacter/index.html
EFSA (2019) The European Union One Health 2018 Zoonoses Report. EFSA Journal 17(12):5926
Bolton DJ (2015) Campylobacter virulence and survival factors. Food Microbiol 48:99–108. https://doi.org/10.1016/j.fm.2014.11.017
Devi A, Mahony TJ, Wilkinson JM, Vanniasinkam T (2019) Antimicrobial susceptibility of clinical isolates of Campylobacter jejuni from New South Wales, Australia. J Glob Antimicrob Re 16:76–80. https://doi.org/10.1016/j.jgar.2018.09.011
Whitehouse CA, Zhao S, Tate H. (2018) Antimicrobial resistance in Campylobacter species: mechanisms and genomic epidemiology
Ge B, Wang F, Sjeolund-Karlsson M, Mcdermott PF (2013) Antimicrobial resistance in Campylobacter: susceptibility testing methods and resistance trends. J Microbiol Meth 95:57–67. https://doi.org/10.1016/j.mimet.2013.06.021
Abu-Madi M, Behnke JM, Sharma A, Bearden R, Al-Banna N (2016) Prevalence of virulence/stress genes in Campylobacter jejuni from chicken meat sold in Qatari retail outlets. PLoS One 11:e0156938. https://doi.org/10.1371/journal.pone.0156938
Humphrey S, Chaloner G, Kemmett K, Davidson N, Williams N, Kipar A, Humphrey T, Wigleya P (2014) Campylobacter jejuni is not merely a commensal in commercial broiler chickens and affects bird welfare. Amer Soc Microbiol 5(4):e01364–e01314. https://doi.org/10.1128/mBio.01364-14
Newell DG, Shreeve JE, Toszeghy M, Domingue G, Bull S, Humphrey T, Mead G (2001) Changes in the carriage of Campylobacter strains by poultry carcasses during processing in abattoirs. Appl Environ Microbiol 67:2636–2640. https://doi.org/10.1128/AEM.67.6.2636-2640.2001
Monteville MR, Yoon JE, Konkel ME (2002) Maximal adherence and invasion of INT 407 cells by Campylobacter jejuni requires the CadF outer membrane protein and microfilament reorganisation. Microbiol 149:153–165
Rivera-Amill V, Kim BJ, Seshu J, Konkel ME (2001) Secretion of the virulence associated Campylobacter invasion antigens from Campylobacter jejuni requires a stimulatory signal. J Infec Dis 183:1607–1616
Smith JL, Bayles DO (2006) The contribution of cytolethal distending toxin to bacterial pathogenesis. Crit Rev Microbiol 32:227–248
Panzenhagen PHN, Aguiar WS, Frasão BS, Pereira VLA, Abreu DLC, Rodrigues DP, do Nascimento ER, de Aquino MHC (2016) Prevalence and fluoroquinolones resistance of Campylobacter and Salmonella isolates from poultry carcasses in Rio de Janeiro, Brazil. Food Control 61:243–247. https://doi.org/10.1016/j.foodcont.2015.10.002
ISO (International Organization for Standard) (2006) Microbiology of FOOD and animal feeding stuffs – Horizontal method for detection and enumeration of Campylobacter spp. – Part 1: Detection method. (ISO 10272-1:2006 [E]). 16 p
Sambrook J, Russell D (2001) Molecular cloning: a laboratory manual. V. 1, Chapter 6, Protocol 7, Third edn. Cold Spring Harbor Laboratory Press, New York
Müller J, Schulze F, Müller W, Hänel I (2006) PCR detection of virulence-associated genes in Campylobacter jejuni strains with differential ability to invade Caco-2 cells and to colonize the chick gut. Vet Microbiol 113:123–129. https://doi.org/10.1016/j.vetmic.2005.10.029
Linton D, Gilbert M, Hitchen PG, Dell A, Morris HR, Wakarchuk WW, Gregson NA, Wren BW (2000) Phase variation of a beta-1,3 galactosyltransferase involved in generation of the ganglioside GM1-like lipo-oligosaccharide of Campylobacter jejuni. Mol Microbiol 37(3):501–514
Datta S, Niwa H, Itoh K (2003) Prevalence of 11 pathogenic genes of Campylobacter jejuni by PCR in strains isolated from humans, poultry meat and broiler and bovine faeces. J Med Microbiol 52:345–348. https://doi.org/10.1099/jmm.0.05056-0
Wieczorek K, Kania I, Osek J (2009) Identification of main virulence markers of food-borne pathogens recovered from bovine carcasses as an aid in assessing public health risk. B Vet I Pulawy 53:425–432
Zheng J, Meng JH, Zhao SH, Singh R, Song WX (2006) Adherence to and invasion of human intestinal epithelial cells by Campylobacter jejuni and Campylobacter coli isolates from retail meat products. J Food Protect 69:768–774
Konkel ME, Kim BJ, Rivera-Amill V, Garvis SG (1999) Bacterial secreted proteins are required for the internalization of Campylobacter jejuni into cultured mammalian cells. Mol Microbiol 32:691–701. https://doi.org/10.1046/j.1365-2958.1999.01376.x
Josefsen MH, Lübeck PS, Hansen F, Hoorfar J (2004) Towards an international standard for PCR-based detection of foodborne thermotolerant campylobacters: interaction of enrichment media and pre-PCR treatment on carcass rinse samples. J Microbiol Meth 58:39–48. https://doi.org/10.1016/j.mimet.2004.03.001
Maćkiw E, Korsak D, Rzewuska K, Tomczuk K, Rozynek E (2012) Antibiotic resistance in Campylobacter jejuni and Campylobacter coli isolated from food in Poland. Food Control 23:297–301. https://doi.org/10.1016/j.foodcont.2011.08.022
Bauer AW, Kirby WMM, Sherris JC, Turck M (1966) Antimicrobial susceptibility testing by a standardized single disk method. Am J Clin Pathol 45:493–496
EUCAST (European Committee on Antimicrobial Susceptibility Testing) (2015) Breakpoint tables for interpretation of MICs and zone diameters. (http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_5.0_Breakpoint_Table_01.pdf)
CLSI (2015) M100–S25. Performance standards for antimicrobial susceptibility testing; twenty-fifth informational supplement. Wayne, PA, USA: clinical and laboratory standards institute
Ribot EM, Fitzgerald C, Kubota K, Swaminathan B, Barrett TJ (2001) Rapid pulsed-field gel electrophoresis protocol for subtyping of Campylobacter jejuni. J Clin Microbiol 39:1889–1894. https://doi.org/10.1128/JCM.39.5.1889-1894.2001
Wieczorek K, Denis E, Osek J (2015) Comparative analysis of antimicrobial resistance and genetic diversity of Campylobacter from broilers slaughtered in Poland. Int J Food Microbiol 210:24–32. https://doi.org/10.1016/j.ijfoodmicro.2015.06.006
Høg B, Sommer HM, Larsen LS, Sorensen AIV, David B, Hofshagen M, Rosenquist H (2016) Farm specific risk factors for Campylobacter colonisation in Danish and Norwegian broilers. Prev Vet Med 130:137–145. https://doi.org/10.1016/j.prevetmed.2016.04.002
Krause M, Josefsen MH, Lund M, Jacobsen NR, Brorsen L, Moos M, Stockmarr A, Hoorfar J (2006) Comparative, collaborative, and on-site validation of a TaqMan PCR method as a tool for certified production of fresh, Campylobacter-free chickens. Appl Environ Microb 72(8):5463–5468. https://doi.org/10.1128/AEM.00291-06
Messad S, Hamdi TM, Bouhamed R, Ramdani-Bouguessa N, Tazir M (2014) Frequency of contamination and antimicrobial resistance of thermotolerant Campylobacter isolated from some broiler farms and slaughterhouses in the region of Algiers. Food Control 40:324–328. https://doi.org/10.1016/j.foodcont.2013.12.016
Sommer HM, Heuer OE, Sørensena AIV, Madsenc M (2013) Analysis of factors important for the occurrence of Campylobacter in Danish broiler flocks. Prev Vet Med 111:100–111. https://doi.org/10.1016/j.prevetmed.2013.04.004
Battersby T, Whyte P, Bolton D (2016) Protecting broilers against Campylobacter infection by preventing direct contact between farm staff and broilers. Food Control 69:346–351. https://doi.org/10.1016/j.foodcont.2016.04.053
Humphrey T, O’Brien S, Madsen M (2007) Campylobacters as zoonotic pathogens: a food production perspective. Int J Food Microbiol 117:237–257. https://doi.org/10.1016/j.ijfoodmicro.2007.01.006
EFSA (European Food Safety Authority) (2012) The European Union summary report on antimicrobial resistance in zoonotic and indicator bacteria from humans, animals and food in 2010. EFSA J 10:2598. https://doi.org/10.2903/j.efsa.2012.2598
García-Sánchez L, Melero B, Jaime I, Hänninen ML, Rossi M, Rovira J (2017) Campylobacter jejuni survival in a poultry processing plant environment. Food Microbiol 65:185–192. https://doi.org/10.1016/j.fm.2017.02.009
Chokboonmongkol C, Patchanee P, Golz G, Zessin KH, Alter T (2013) Prevalence, quantitative load, and antimicrobial resistance of Campylobacter spp. from broiler ceca and broiler skin samples in Thailand. Poultry Sci 92:462–467. https://doi.org/10.3382/ps.2012-02599
Gharbi M, Béjaoui A, Hmada CB, Jouini A, Ghedira K, Zrelli C, Hamrouni S, Aouadhi C, Bessoussa G, Ghram A, Maaroufi A (2018) Prevalence and antibiotic resistance patterns of Campylobacter spp. isolated from broiler chickens in the north of Tunisia. BioMed Res Int 2018:7943786–7943787. https://doi.org/10.1155/2018/7943786
Oliveira MG, Rizzi C, Galli V, Lopes GV, Haubert L, Dellagostin OA, Silva WP (2019) Presence of genes associated with adhesion, invasion, and toxin production in Campylobacter jejuni isolates and effect of temperature on their expression. Can J Microbiol 65:4. https://doi.org/10.1139/cjm-2018-0539
Seliwiorstow T, Baré J, Berkvens D, Van Damme I, Uyttendaele M, de Zutter L (2016) Identification of risk factors for Campylobacter contamination levels on broiler carcasses during the slaughter process. Int J Food Microbiol 226:26–32. https://doi.org/10.1016/j.ijfoodmicro.2016.03.010
Bull SA, Allen VM, Domingue G, Jorgensen F, Frost JA, Ure R, Whyte R, Tinker D, Corry JEL, Gillard-King J, Humphrey TJ (2006) Sources of Campylobacter spp. colonizing housed broiler flocks during rearing. Appl Environ Microbiol 72:645–652. https://doi.org/10.1128/AEM.72.1.645-652.2006
Berrang ME, Northcutt JK, Fletcher DL, Cox NA (2003) Role of dump cage fecal contamination in the transfer of Campylobacter to carcasses of previously negative broilers. J Appl Poult Res 12:190–195. https://doi.org/10.1093/japr/12.2.190
Gruntar I, Biasizzo M, Kušar D, Pate M, Ocepek M (2015) Campylobacter jejuni contamination of broiler carcasses: population dynamics and genetic profiles at slaughterhouse level. Food Microbiol 50:97–101. https://doi.org/10.1016/j.fm.2015.03.007
Rosenquist H, Sommer HM, Nielsen NL, Christensen BB (2006) The effect of slaughter operations on the contamination of chicken carcasses with thermotolerant Campylobacter. Int J Food Microbiol 108:226–232. https://doi.org/10.1016/j.ijfoodmicro.2005.12.007
Franchin PR, Ogliari PJ, Batista CRV (2007) Frequency of thermophilic Campylobacter in broiler chickens during industrial processing in a southern Brazil slaughterhouse. Brit Poultry Sci 48:127–132. https://doi.org/10.1080/00071660701261286
Nobile CGA, Costantino R, Bianco A, Pileggi C, Pavia M (2013) Prevalence and pattern of antibiotic resistance of Campylobacter spp. in poultry meat in Southern Italy. Food Control 32(2):715–718. https://doi.org/10.1016/j.foodcont.2013.02.011
Ruiz-Palacios GM (2007) The health burden of Campylobacter infection and the impact of antimicrobial resistance: playing chicken. Clin Infect Dis 44:701–703. https://doi.org/10.1086/509936
Avrain L, Humbert F, L’Hospitalier R, Sanders P, Vernozy-Rozand C, Kempf I (2003) Antimicrobial resistance in Campylobacter from broilers: association with production type and antimicrobial use. Vet Microbiol 96(3):267–276. https://doi.org/10.1016/j.vetmic.2003.07.001
Chen X, Naren GW, Wu CM, Wang Y, Dai L, Xia LN (2010) Prevalence and antimicrobial resistance of Campylobacter isolates in broilers from China. Vet Microbiol 144:133–139. https://doi.org/10.1016/j.vetmic.2009.12.035
Farnell MB, Donoghue AM, Cole K, Reyes-Herrera I, Blore PJ, Donoghue DJ (2005) Campylobacter susceptibility to ciprofloxacin and corresponding fluoroquinolone concentrations within the gastrointestinal tracts of chickens. J Appl Microbiol 99:1043–1050. https://doi.org/10.1111/j.1365-2672.2005.02712.x
Miflin JK, Templeton JM, Blackall PJ (2007) Antibiotic resistance in Campylobacter jejuni and Campylobacter coli isolated from poultry in the South-East Queensland region. J Antimicrob Chemoth 59:775–778. https://doi.org/10.1093/jac/dkm024
Gruntar I, Ocepek M, Avbersek J, Mićunović J, Pate M (2010) A pulsed-field gel electrophoresis study of the genetic diversity of Campylobacter jejuni and Campylobacter coli in poultry flocks in Slovenia. Acta Vet Hung 58(1):19–28. https://doi.org/10.1556/AVet.58.2010.1.2
Lindmark H, Harbom B, Thebo L, Andersson L, Hedin G, Osterman B, Lindberg T, Andersson Y, Westöö A, Engvall EO (2004) Genetic characterization and antibiotic resistance of Campylobacter jejuni isolated from meats, water, and humans in Sweden. J Clin Microbiol 42(2):700–706. https://doi.org/10.1128/JCM.42.2.700-706.2004
Workman SN, Mathison GE, Lavoie MC (2008) An investigation of sources of Campylobacter in a poultry production and packing operation in Barbados. Int J Food Microbiol 121:106–111. https://doi.org/10.1016/j.ijfoodmicro.2007.10.014
Ghorbanalizadgan M, Bakhshi B, Najar-Peerayeh S (2018) Heterogeneity of cytolethal distending toxin sequence types of Campylobacter jejuni and correlation to invasion/ cytotoxicity potential: The first molecular survey from Iran. Microb Pathogenesis 114:213–218. https://doi.org/10.1016/j.micpath.2017.11.035
Jones MA, Marston KL, Woodall CA, Maskell DJ, Linton D, Karlyshev AV, Dorrell N, Wren BW, Barrow PA (2004) Adaptation of Campylobacter jejuni NCTC11168 to high level colonization of the avian gastrointestinal tract. Infect Immun 72:3769–3776. https://doi.org/10.1128/IAI.72.7.3769-3776.2004
Samad A, Abbas F, Ahmed Z, Akbar A, Naeem M, Sadiq MB, Ali I, Roomeela S, Bugti FS, Achakzai SK (2018) Prevalence, antimicrobial susceptibility, and virulence of Campylobacter jejuni isolated from chicken meat. J Food Safety 36:e12600. https://doi.org/10.1111/jfs.12600
Konkel ME, Gray SA, Kim BJ, Garvis SG, Yoon J (1999) Identification of the enteropathogens Campylobacter jejuni and Campylobacter coli based on the cadF virulence gene and its product. J Clin Microbiol 37(3):510–517 http://www.ncbi.nlm.nih.gov/pmc/articles/PMC84446
Monteville MR, Yoon JE, Konkel ME (2003) Maximal adherence and invasion of INT 407 cells by Campylobacter jejuni requires the CadF outer membrane protein and microfilament reorganization. Microbiol 149:153–165. https://doi.org/10.1099/mic.0.25820-0
Konkel ME, Kim BJ, Klena JD, Young CR, Ziprin R (1998) Characterization of the thermal stress response of Campylobacter jejuni. Infect Immun 66(8):3666–3672
Brás AM, Chatterjee S, Wren BW, Newell DG, Ketley JM (1999) A novel Campylobacter jejuni two-component regulatory system important for temperature-dependent growth and colonization. J Bacteriol 181:3298–3302 https://www.ncbi.nlm.nih.gov/pubmed/10322038
Gagnaire A, Nadel B, Raoult D, Neefjes J, Gorvel JP (2017) Collateral damage: insights into bacterial mechanisms that predispose host cells to cancer. Nat Rev Microbiol 15:109–128. https://doi.org/10.1038/nrmicro.2016.171
He Z, Gharaibeh RZ, Newsome RC, Pope JL, Dougherty MW, Tomkovich S, Pons B, Mirey G, Vignard J, Hendrixson DR, Jobin C (2019) Campylobacter jejuni promotes colorectal tumorigenesis through the action of cytolethal distending toxin. Gut 68:289–300
Acknowledgments
This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior-Brasil (CAPES)-Finance Code 001. The authors extend their thanks to the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) (Process n. 309101/2016-6) and to the Fundação de Amparo à Pesquisa do Estado do Rio Grande do Sul (FAPERGS) (Process n. 17/2551-0000956-8).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by Tassiana Ramires, Mauricéia Greici de Oliveira, Natalie Rauber Kleinubing, Simone de Fátima Rauber Würfel, Marcia Magalhães Mata, Mariana Almeida Iglesias, and Graciela Volz Lopes. The first draft of the manuscript was written by Tassiana Ramires and Mauricéia Greici de Oliveira, and all authors commented on previous versions of the manuscript. The manuscript was reviewed by Odir Antônio Dellagostin and Wladimir Padilha da Silva. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict interest.
Additional information
Responsible Editor: Mariza Landgraf.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ramires, T., de Oliveira, M.G., Kleinubing, N.R. et al. Genetic diversity, antimicrobial resistance, and virulence genes of thermophilic Campylobacter isolated from broiler production chain. Braz J Microbiol 51, 2021–2032 (2020). https://doi.org/10.1007/s42770-020-00314-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-020-00314-0