Abstract
In 2007, the new polyomaviruses WUPyV and KIPyV were identified in patients with acute respiratory infections. The aim of this study was to investigate these viruses in hospitalized patients with severe acute respiratory infection (SARI). A retrospective study was conducted with 251 patients, from April 2009 to November 2010, using nasopharyngeal aspirates, naso- and oropharyngeal swab samples from hospitalized patients (children < 12 years and adults) who had SARI within 7 days of the onset of symptoms, including fever (> 38.8 °C), dyspnea, and cough. Clinical and epidemiological information was obtained through standardized questionnaire. Enrolled patients were initially suspected to have influenza A(H1N1)pdm09 infections. WUPyV and KIPyV were detected by real-time PCR. Samples were also tested for influenza A and B viruses, human respiratory syncytial virus, rhinovirus, metapneumovirus, coronavirus, adenovirus, and parainfluenza viruses. WUPyV and KIPyV were detected in 6.77% (4.78% and 1.99%, respectively) of hospitalized patients with SARI. All samples from children showed coinfections (rhinovirus was the most commonly detected). Six adults had polyomavirus infection and four (1.6%) had monoinfection. Of them, 3 reported comorbidities including immunosuppression and 1 patient had worse outcome, requiring ICU admission. These preliminary data may suggest a possible role of polyomaviruses in SARI among immunocompromised adult patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In 2007, the two new polyomaviruses (PyV) KI and WU were described as probable causative agents of respiratory infections in humans [1, 2]. Currently, they are classified in the Polyomaviridae family, Betapolyomavirus genus, Human polyomavirus 3 (KIPyV), and Human polyomavirus 4 (WUPyV) species [3]. These two PyV have been detected globally in children and immunocompetent adults, with detection rates ranging from 1 to 16.4% for WUPyV and 0.45 to 12.14% for KIPyV [4,5,6,7,8].
There has been suggested a role for these viruses on the pathogenesis of respiratory infections in many studies so far demonstrating the presence of WUPyV and KIPyV in the respiratory tract of symptomatic and asymptomatic patients [4,5,6,7,8,9,10,11,12,13,14,15], although they are also found in other clinical samples such as stool and lymphoid tissue of immunocompromised and immunocompetent individuals [16,17,18,19,20].
Coinfection with other pathogens has been reported in 74% of patients with KIPyV, 68 to 79% of patients with WUPyV, and 10% of coinfection with KIPyV and WUPyV in the absence of other respiratory viruses [21]. In Brazil, KIPyV and WUPyV were found in the saliva of immunocompromised and immunocompetent patients [22, 23]. In this study, we investigated the PyV KI and WU in the occurence of severe respiratory diseases in hospitalized patients with acute respiratory infection (SARI).
Materials and methods
A retrospective study was conducted from April 2009 to November 2010 (19 months) using samples from hospitalized patients who had presented with SARI within 7 days of the onset of symptoms, including fever (> 38.8 °C), dyspnea, and cough. Children (< 12 years) and adults were included. The clinical and epidemiological information was obtained through a standardized questionnaire. Informed consent was previously obtained from all enrolled patients initially suspected to have influenza A(H1N1)pdm09 infections. This study was approved by the Ethics Committee of Sao Paulo Federal University (CEP 0090/2016). Nasopharyngeal aspirates were obtained from children under 2 years old. For the other patients, naso- and oropharyngeal swabs were collected, according to the Brazilian Ministry of Health Protocol for management of influenza A (H1N1) 2009 pandemics, and stored in a freezer at − 80 °C [24]. Total nucleic acids were extracted using the QIAamp Viral RNA Mini Kit (QIAGEN, Hilden, Germany), according to the manufacturer instructions. The specimens were then tested for the presence of respiratory viruses according to PCR protocols available at our laboratory for influenza A (FLU A) and B (FLU B) viruses [25, 26], human respiratory syncytial virus (HRSV) [27], human rhinovirus (HRV) [28], human metapneumovirus (hMPV) [29], human coronavirus (HCoV) [30], human adenovirus (HAdV) [31, 32], and parainfluenza virus (PIV 1,2,3,4) [33, 34].
For the WUPyV and KIPyV specific detection, qualitative real-time PCR was performed with the primers PyVF (5’-TTGGATGAAAATGGCATTGG-3′) and PyVR (5’-TAACCCTTCTTTGTCTAAARTGTAGCC-3′), that amplify a fragment of the gene encoding the capsid protein VP1 only common to KIPyV and WUPyV, and two specific probes, WUPyVp (5’-FAM-CATAACTTGTGCTGACCTTTTGGGAGTTAAC-BHQ1–3′) and KIPyVp (3’-VIC-ACATTACTTGTGCAGATATGCTTGGAACAGC-BHQ1–3′), both within the VP1 gene, to differentiate the WUPyV and KIPyV viruses [35]. The concentration of primers and probes was 800 nM and 200 nM, respectively, in 25 μL of total reaction volume, with the use of AgPath-ID™ One-Step RT-PCR reagents (Applied Biosystems, Austin, USA) and addition of 5 μL of extracted nucleic acid. In parallel reactions, primers and specific probe were used to amplify a fragment of the gene glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as endogenous control and to assess the quality of the samples as described by Bergallo et al. [16] Thermocycling conditions were performed in an Applied Biosystems 7500 Real-Time PCR System and involved 10 min at 95 °C followed by 45 cycles of 10 s at 95 °C and 1 min at 55 °C (data collection). For both reactions, the cycle threshold value (Ct) of 39 was adopted as a cut-off. Chi-square analysis and Fisher’s exact test were used to analyze the occurrence of the investigated PyV. Student’s t test for independent samples and analysis of variance (ANOVA) were used to compare the mean ages. All statistical analyses were performed using OpenEpi version 2.3.1 (http://www.openepi.com), and a p value < 0.05 was considered statistically significant.
Results
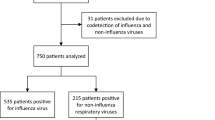
A total of 251 patients presenting with SARI suspected to be due to influenza A (H1N1)pdm09 treated at the Sao Paulo Hospital complex were enrolled; 159 patient samples were obtained in 2009, and 92 in 2010. Among those enrolled, 150 (59.8%) were children, 101 (40.2%) were adults, and (58.16%) were males. The mean age of the children was 2.4 years (median 1 year, range 0–12 years) and the mean age of the adults was 41.4 years (median 40, range 13–91 years). Viral detection was identified in 62.5% (157/251) of the patients, and the detection rate of WUPyV and KIPyV was 6.8% (17/251), with 12 cases of WUPyV and 5 of KIPyV.
The mean Ct value (± SD) for GAPDH endogenous control for all samples was 28.51 ± 3.16 (range 20.44–34.28). All positive samples have shown GAPDH Ct values ≤ 30.48, except for one sample positive for WUPyV (Ct = 32.46). The mean Ct values for WUPyV and KIPyV (range are shown in brackets) were 32.61 ± 4.86 (24.28–36.94) and 30.52 ± 6.95 (21.96–37.22), respectively.
The single viral detection rates were as follows: 4.78% (12/251) for WUPyV, 1.99 (5/251) for KIPyV, 7.2% (18/251) for FLU A, 4.0% (10/251) for FLU B, 1.6% (4/251) for HAdV, 0.8% (2/251) for HCoV, 8.4% (21/251) for hMPV, 3.6% (9/251) for PIV, 14.7% (37/251) for HRV, and 6.8% (17/251) for HRSV. The WUPyV and KIPyV coinfection rate was 5.2% (13/251). Table 1 compares the viral detection patterns within the 251 patients included in the study. Among the patients with viral infection, 18.5% (29/157) had a coinfection with 2 different viruses and 6.4% (10/157) with 3. Of the population with double viral coinfections, 44.8% (13/29) were also infected with WUPyV or KIPyV and 4 out of the 10 patients with triple coinfections were also positive for WUPyV or KIPyV. However, coinfection with WUPyV or KIPyV and other viruses was not correlated with severity of symptoms.
The proportion of children with WUPyV or KIPyV was 7.3% (11/150), and all samples had multiple detections. The mean age of the children coinfected with polyomaviruses was 1.6 years (median 1 year; range 0–4 years); of these 7 had WUPyV infection and 4 had KIPyV infection. The children coinfected with respiratory viruses other than PyV were younger than those coinfected with PyV, with rates among patients up to 6 months of age being 1/11 (9.1%) and 12/24 (50.0%), respectively (p < 0.027).
Two adults (aged 33 and 91 years) showed coinfections with WUPyV or KIPyV, respectively. Monoinfections were detected in 4 adults, and 3 of them had WUPyV infection (median age 30 years, range 21–51 years). These 4 patients were admitted with fever, cough, and dyspnea, and the symptoms prevailed for up to 4 days. Radiological chest examination, bacterial hemoculture and systemic antibiotic prophylaxis were conducted according to hospital proceedings for clinical bacterial pneumonia investigation and current guidelines. Three patients reported comorbidities and presented with different states of immunosuppression. The length of the period of hospitalization and ICU admission are shown in Table 2. The patient with systemic lupus erythematosus (SLE) was admitted to the ICU for mechanical ventilation due to nosocomial pneumonia with bacterial coinfection. The 28-year-old patient without comorbidities was hospitalized for 24 h and was discharged upon resolution of the dyspnea.
Discussion
The role of the PyV in the pathogenesis of respiratory infections is not well understood. Some studies have demonstrated the presence of both PyV WU and KI in the asymptomatic population [36, 37]. However, the rate of polyomavirus infection in adult patients hospitalized with SARI has not been established.
The proportion of respiratory samples from hospitalized patients with SARI that were positive for WUPyV or KIPyV was 6.8% with 4.0% (10/251) for WUPyV and 2.8% (7/251) for KIPyV, which is comparable to those obtained by other authors [4,5,6,7, 11].
The debate about the role of these viruses in respiratory infections is ongoing because of the high rates of coinfection with other respiratory viruses (approximately 80%) [9, 11, 12, 38, 39]. Recently, high viral loads of KIPyV were observed in children with severe or very severe pneumonia in the absence of other viral or bacterial respiratory infection, suggesting a potential pathogenic role for KIPyV in these patients [12].
A coinfection rate of 76.5% was observed in the present study, mainly among children, with the most common types being WUPyV/HRV in 43% of cases, KIPyV/HRV in 60% and KIPyV/AdV in 67%, which is in accordance with the results by Allander et al. [1] and Gaynor et al. [2] Nonetheless, the frequent coinfections with WUPyV and KIPyV with other respiratory viruses and variations in clinical presentation preclude any conclusion about the association between infection and disease in children. Of interest is the finding of lower rates among very young children that may posit the later acquisition of polyomavirus infection by older children as suggest by serological and molecular studies [19, 40,41,42].
The finding of monoinfection by polyomaviruses in 4 adult patients hospitalized with SARI may suggest a potential pathogenic role. Moreover, 3 out of these 4 patients were immunocompromised, and one patient with SLE had a severe outcome and required ICU admission and mechanical ventilation. We performed a retrospective study. Therefore, it was not possible to obtain samples from the lower respiratory tract from the immunocompromised patients for immunohistochemistry or in situ hybridization. Further studies are needed in order to understand the possible role of viral latency and reactivation of these DNA viruses in adult patients, especially among those who are immunocompromised. In this regard, prospective longitudinal studies with large sample sizes, including immunocompetent, immunocompromised, symptomatic, and asymptomatic patients could better answer some of these questions.
References
Allander T, Andreasson K, Gupta S et al (2007) Identification of a third human polyomavirus. J Virol 81(8):4130–4136. https://doi.org/10.1128/jvi.00028-07
Gaynor AM, Nissen MD, Whiley DM et al (2007) Identification of a novel polyomavirus from patients with acute respiratory tract infections. PLoS Pathog 3(5):e64. https://doi.org/10.1371/journal.ppat.0030064
Moens U, Calvignac-Spencer S, Lauber C et al (2017) ICTV virus taxonomy profile: polyomaviridae. J Gen Virol 98(6):1159–1160. https://doi.org/10.1099/jgv.0.000839
Teramoto S, Kaiho M, Takano Y et al (2011) Detection of KI polyomavirus and WU polyomavirus DNA by real-time polymerase chain reaction in nasopharyngeal swabs and in normal lung and lung adenocarcinoma tissues. Microbiol Immunol 55(7):525–530. https://doi.org/10.1111/j.1348-0421.2011.00346.x
Ren L, Gonzalez R, Xie Z et al (2008) WU and KI polyomavirus present in the respiratory tract of children, but not in immunocompetent adults. J Clin Virol 43(3):330–333. https://doi.org/10.1016/j.jcv.2008.08.003
Mourez T, Bergeron A, Ribaud P et al (2009) Polyomaviruses KI and WU in immunocompromised patients with respiratory disease. Emerg Infect Dis 15(1):107–109. https://doi.org/10.3201/eid1501.080758
Babakir-Mina M, Ciccozzi M, Dimonte S et al (2008) Identification of the novel KI polyomavirus in the respiratory tract of an Italian patient. J Med Virol 80(11):2012–2014. https://doi.org/10.1002/jmv.21303
Lin S-X, Wang W, Guo W et al (2017) A molecular epidemiological study of KI polyomavirus and WU polyomavirus in children with acute respiratory infection in Tianjin, China. Chinese J Contemp Pediatr 19(7):763–769. https://doi.org/10.7499/j.issn.1008-8830.2017.07.007
Han TH, Chung JY, Koo JW, Kim SW, Hwang ES (2007) WU polyomavirus in children with acute lower respiratory tract infections, South Korea. Emerg Infect Dis 13(11):1766–1768. https://doi.org/10.3201/eid1311.070872
Wattier RL, Vázquez M, Weibel C et al (2008) Role of human polyomaviruses in respiratory tract disease in young children. Emerg Infect Dis 14(11):1766–1768. https://doi.org/10.3201/eid1411.080394
Hansen-Estruch C, Coleman KK, Thoon KC, Low JG, Anderson BD, Gray GC (2018) Prevalence of respiratory polyomaviruses among pediatric patients with respiratory symptoms in Singapore. Front Pediatr ;6. https://doi.org/10.3389/fped.2018.00228
Rao S, Lucero MG, Nohynek H et al (2016) WU and KI polyomavirus infections in Filipino children with lower respiratory tract disease. J Clin Virol 82:112–118. https://doi.org/10.1016/j.jcv.2016.07.013
Gozalo-Margüello M, Agüero-Balbín J, Martínez-Martínez L (2015) WU and KI polyomavirus prevalence in invasive respiratory samples from transplant recipients in Cantabria, Spain. Transplant Proc 47(1):67–69. https://doi.org/10.1016/j.transproceed.2014.11.017
Essa S, Owayed A, Altawalah H, Khadadah M, Behbehani N, Al-Nakib W (2015) The prevalence of human bocavirus, human coronavirus-NL63, human metapneumovirus, human polyomavirus KI and WU in respiratory tract infections in Kuwait. Med Princ Pract 24(4):382–387. https://doi.org/10.1159/000381422
Iaria M, Caccuri F, Apostoli P et al (2015) Detection of KI WU and Merkel cell polyomavirus in respiratory tract of cystic fibrosis patients. Clin Microbiol Infect 21(6):603.e9–603.e15. https://doi.org/10.1016/j.cmi.2015.01.025
Bergallo M, Terlizzi ME, Astegiano S et al (2009) Real time PCR TaqMan assays for detection of polyomaviruses KIV and WUV in clinical samples. J Virol Methods 162(1–2):69–74. https://doi.org/10.1016/j.jviromet.2009.07.016
Sadeghi M, Wang Y, Ramqvist T et al (2017) Multiplex detection in tonsillar tissue of all known human polyomaviruses. BMC Infect Dis 17(1):409. https://doi.org/10.1186/s12879-017-2479-5
Comar M, Zanotta N, Rossi T, Pelos G, D’Agaro P (2011) Secondary lymphoid tissue as an important site for WU polyomavirus infection in immunocompetent children. J Med Virol 83(8):1446–1450. https://doi.org/10.1002/jmv.22124
Herberhold S, Hellmich M, Panning M et al (2017) Human polyomavirus and human papillomavirus prevalence and viral load in non-malignant tonsillar tissue and tonsillar carcinoma. Med Microbiol Immunol 206(2):93–103. https://doi.org/10.1007/s00430-016-0486-6
Bialasiewicz S, Whiley DM, Lambert SB, Nissen MD, Sloots TP (2009) Detection of BK, JC, WU, or KI polyomaviruses in faecal, urine, blood, cerebrospinal fluid and respiratory samples. J Clin Virol 45(3):249–254. https://doi.org/10.1016/j.jcv.2009.05.002
Sloots TP, Whiley DM, Lambert SB, Nissen MD (2008) Emerging respiratory agents: new viruses for old diseases? J Clin Virol 42(3):233–243. https://doi.org/10.1016/j.jcv.2008.03.002
Robaina TF, Mendes GS, Benati FJ et al (2013) Shedding of polyomavirus in the saliva of immunocompetent individuals. J Med Virol 85(1):144–148. https://doi.org/10.1002/jmv.23453
Robaina TF, Mendes GS, Benati FJ et al (2013) Polyomavirus in saliva of HIV-infected children, Brazil. Emerg Infect Dis 19(1):155–157. https://doi.org/10.3201/eid1901.120563
MS. Protocolo para o Enfrentamento à Pandemia de Influenza Pandêmica (H1N1) 2009: Ações da Atenção Primária à Saúde. Epidemiológica D de V, ed. 2010:38. http://bvsms.saude.gov.br/bvs/publicacoes/protocolo_enfrentamento_influenza_2009.pdf
CDC. CDC Protocol of realtime RT-PCR for influenza A (H1N1). World Health Organization. http://www.who.int/csr/resources/publications/swineflu/CDCRealtimeRTPCR_SwineH1Assay-2009_20090430.pdf. Published 2009
Perosa AH, Watanabe AS, Guatura SB, Silva ER, Granato C, Bellei N (2013) Comparison of the direct fluorescence assay and real-time polymerase chain reaction for the detection of influenza virus A and B in immunocompromised patients. Clin (Sao Paulo) 68(9):1206–1209. https://doi.org/10.6061/clinics/2013(09)05
Fry AM, Chittaganpitch M, Baggett HC et al (2010) The burden of hospitalized lower respiratory tract infection due to respiratory syncytial virus in rural Thailand. PLoS One 5(11):e15098. https://doi.org/10.1371/journal.pone.0015098
Savolainen C, Mulders MN, Hovi T (2002) Phylogenetic analysis of rhinovirus isolates collected during successive epidemic seasons. Virus Res 85(1):41–46 https://www.ncbi.nlm.nih.gov/pubmed/11955637
Falsey AR, Erdman D, Anderson LJ, Walsh EE (2003) Human metapneumovirus infections in young and elderly adults. J Infect Dis 187(5):785–790. https://doi.org/10.1086/367901
Dare RK, Fry AM, Chittaganpitch M, Sawanpanyalert P, Olsen SJ, Erdman DD (2007) Human coronavirus infections in rural Thailand: a comprehensive study using real-time reverse-transcription polymerase chain reaction assays. J Infect Dis 196(9):1321–1328. https://doi.org/10.1086/521308
Heim A, Ebnet C, Harste G, Pring-Akerblom P (2003) Rapid and quantitative detection of human adenovirus DNA by real-time PCR. J Med Virol 70(2):228–239. https://doi.org/10.1002/jmv.10382
Puerari D, Camargo C, Gratura S, Watanabe AS, Granato C, Bellei NC (2015) Application of molecular assay for adenovirus detection among different pediatric patients. Rev Paul Pediatr 33(2):136–141. https://doi.org/10.1016/j.rpped.2014.09.004
Kim C, Ahmed JA, Eidex RB et al (2011) Comparison of nasopharyngeal and oropharyngeal swabs for the diagnosis of eight respiratory viruses by real-time reverse transcription-PCR assays. PLoS One 6(6):e21610. https://doi.org/10.1371/journal.pone.0021610
van de Pol AC, van Loon AM, Wolfs TF et al (2007) Increased detection of respiratory syncytial virus, influenza viruses, parainfluenza viruses, and adenoviruses with real-time PCR in samples from patients with respiratory symptoms. J Clin Microbiol 45(7):2260–2262. https://doi.org/10.1128/JCM.00848-07
Lindau C, Tiveljung-Lindell A, Goh S, Ramqvist T, Allander T (2009) A single-tube, real-time PCR assay for detection of the two newly characterized human KI and WU polyomaviruses. J Clin Virol 44(1):24–26. https://doi.org/10.1016/j.jcv.2008.09.006
Abed Y, Wang D, Boivin G (2007) WU polyomavirus in children, Canada. Emerg Infect Dis 13(12):1939–1941. https://doi.org/10.3201/eid1312.070909
Norja P, Ubillos I, Templeton K, Simmonds P (2007) No evidence for an association between infections with WU and KI polyomaviruses and respiratory disease. J Clin Virol 40(4):307–311. https://doi.org/10.1016/j.jcv.2007.09.008
Bialasiewicz S, Whiley DM, Lambert SB et al (2008) Presence of the newly discovered human polyomaviruses KI and WU in Australian patients with acute respiratory tract infection. J Clin Virol 41(2):63–68. https://doi.org/10.1016/j.jcv.2007.11.001
Le BM, Demertzis LM, Wu G et al (2007) Clinical and epidemiologic characterization of WU polyomavirus infection, St. Louis, Missouri. Emerg Infect Dis 13(12):1936–1938. https://doi.org/10.3201/eid1312.070977
Gossai A, Waterboer T, Nelson HH et al (2016) Seroepidemiology of human polyomaviruses in a US population. Am J Epidemiol 183(1):61–69. https://doi.org/10.1093/aje/kwv155
Abedi Kiasari B, Vallely PJ, Corless CE, Al-Hammadi M, Klapper PE (2008) Age-related pattern of KI and WU polyomavirus infection. J Clin Virol 43(1):123–125. https://doi.org/10.1016/j.jcv.2008.05.003
White MK, Gordon J, Khalili K (2013) The rapidly expanding family of human polyomaviruses: recent developments in understanding their life cycle and role in human pathology. Hobman TC, ed. PLoS Pathog ;9(3):e1003206. https://doi.org/10.1371/journal.ppat.1003206
Funding
This work was supported by the Fundação de Amparo à Pesquisa do Estado de São Paulo – FAPESP [grant number 2016/09279-6.]. DBC e LKSL são bolsistas da Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Brasil.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This study was approved by the Ethics Committee of Sao Paulo Federal University (CEP 0090/2016).
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible Editor: Giliane Trindade
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Caldeira, D.B., de Souza Luna, L.K., Watanabe, A. et al. The occurrence of polyomaviruses WUPyV and KIPyV among patients with severe respiratory infections. Braz J Microbiol 50, 133–137 (2019). https://doi.org/10.1007/s42770-018-0038-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-018-0038-x