Abstract
Cellulose is considered to be one of the most underutilized biomass available on earth. These cellulosic resources, if utilized as the precursor of food, feed and biofuel can meet up the ever-increasing demands for food and energy. Since cellulase is an enzyme complex, fermentation of cellulose from agro-wastes and industrial effluents is a complex event. The physicochemical parameters are generally optimized in "one at a time" mode in fed-batch culture. But with the advent of time, statistical and mathematical modelling is used, like response surface methodology (RSM), artificial neural network (ANN), machine learning algorithm, and genetic algorithm to improve enzyme production. In bioreactor-based cellulase production, a LabVIEW-based intelligent system for monitoring bio-processing is used for the optimization of the target parameters. RSM and ANN are high-quality prediction mathematical models but ANN shows its superiority in context to the fitting of data as well as its estimation capabilities. The difference of ANN concerning RSM is its requirement of a large number of trained data. This review provides a comprehensive study of literature in context to various advanced mechanisms for optimization of cellulase production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Expansion of the human population with high growth rate results in an ever-increasing demand for food and energy (Lincoln 2013). Cellulose, being present in the cell wall structure of the green plants is almost ubiquitous and constitutes the major component of forest wastes and agricultural residues (Foyle et al. 2007). Since a large fraction of this polysaccharide remains unutilized, it is not fully exploited for the production of biofuels, food, and other value-added products (Mielenz 2001). On the other hand, the unutilized cellulosic raw material poses a real problem as uncared green garbage (Kumar et al. 2020). As the wastes containing celluloses are predominantly liberated by industrial, urban, sewage sludge and agricultural activities, the absence of a proper waste management system will give rise to enormous problems (Lohri et al. 2017). Hence judicious utilization of cellulosic raw materials is the need of the hour both for economic and ecological reasons. The major concern is the development of sustainable processes or technologies that can be both efficient and economical. The rise of environmental pollution is the biggest global concern as it has a direct effect on public health (Manisalidis et al. 2020).
Although cellulose can be treated with chemicals, bioprocessing with enzyme seems to be a much better way being non-toxic and environment-friendly approach (Rodhe et al. 2011). A large number of bacterial and fungal strains possess the ability to degrade cellulosic wastes with the help of a multi-enzyme system (Johnson et al. 1982). The cellulolytic microorganisms are mostly mesophilic or thermophilic, and aerobic or anaerobic (Kubicek 1993).
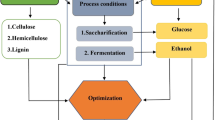
The enzyme, cellulase is responsible for the hydrolysis of β-1,4-glycosidic bond present in cellulose and complete enzymatic hydrolysis of the cellulose-containing three different types of the enzymes (Henrissat et al. 1998); namely endoglucanase, (1,4-d-glucan-4-glucanohydrolase; EC 3.2.1.4), exo-cellobiohydrolase (1,4-d-glucan glucohydrolase; EC 3.2.1.74) and β-glucosidase (d-glucoside glucohydrolase; EC 3.2.1.21). Optimization of media for the production of the enzyme from microbial sources is an important step in microbiological research and involves various types of challenges. Classical researches mainly implemented the "one variable-at-a-time" approach but it is very time-consuming as it requires the conduction of various experiments, lot of time, and is laborious (Panda et al. 2007). The classical mechanism of optimizing the production can be achieved when there are no interactions between the variables (Geiger et al. 1996). In recent times the advent of modern mechanisms of implying statistical or mathematical techniques for optimizing the media has been an effective, efficient, vibrant, and robust technology (Goukanapalle et al. 2020). The optimization of various parameters like temperature, pH, speed of agitation and various components of the media using the recent technological advancements has made productivity faster with better yield (Wang et al. 2005). Figure 1 showcases the schematic representation of the optimization techniques.
In this review, we focussed on the various optimization techniques for the production of the enzyme, their applicability, and feasibility to enhance yield by the involvement of fermentative processes. The discussion was also based on how techniques like response surface methodology (RSM), artificial neural network (ANN), machine learning, and other mathematical models have made the experimentation less laborious with reduced consumption of time. The present overview will also focus on the past, present, and future directions for optimization of process parameters, especially with the help of statistical modelling and computational biology for the production of microbial cellulases.
Need of optimizing media for the production of cellulase
Optimization of media is carried out in various industries to enhance the yield and desired product. Optimization of media essentially focuses on nutrient requirements for the growth of the microbial species that play an essential role in the formation of desired products. The major lookout of industries is to maximize the yield of the product with minimum cost or investment (Singh et al. 2017). Various strategies are involved in the process of optimization of media that are used frequently for better production of desired products. Figure 2 shows the schematic representations of various optimization techniques comprising both classical and modern.
Optimization of cellulase activity by one factor at a time
The mechanism of optimization of media by the technique of "one-factor-at-a-time" (OFAT) involves the use of one factor or variable in a particular experiment keeping other variables constant. This is followed by the change in the concentration of the components being present within the media up to the desired range. The dexterity and convenience of OFAT made it a choice for the researchers for designing suitable conditions for optimizing the production of enzymes (Gonzalez et al. 1995). Production of cellulase involves many steps and is followed by optimization of various factors. The most primitive method for optimizing a system conditioned by various factors is by OFAT approach. Numerous studies have been done on production optimization of microbial cellulase in fed-batch culture. Reviews on fermentation of agro-wastes or industrial effluent by microbes for cellulase production by Hussain et al. (2019) and Saranraj et al. (2012), clearly depicted various conditions optimized for the production of cellulase. Cultivation temperature, pH, substrate concentration, presence of an additive, and incubation time are known to be optimized for the production of cellulase (Karmakar and Ray 2010; Mukherjee et al. 2011) by the conventional method. A recent study by Goukanapalle et al. 2020 showed the optimization of cellulase from the endophytic fungi by OFAT (Goukanapalle et al. 2020). The negative aspects of this process are that estimation obtained from the factors are prone to experimental errors and time consuming.
Optimization of cellulase by RSM
As the conventional OFAT method becomes ineffective and time-consuming (Jeya et al. 2009), researchers are searching for a more advanced method to perform the optimization process within a short period. The use of statistical tools for designing the experiments is the latest trend in bioprocess development and its optimization. Techniques employing statistical tools can be used to check the correctness of the response and the importance of selected factors chosen for the experiment. The failure of the classical method of one-variable-at-a-time (OVAT) bioprocess design in analysis of the collective effect of all selected factors in the fermentation process shifts the focus towards statistical techniques (Selvendran 1985). One such statistical method is RSM which employs various mathematical and statistical techniques for planning experiments, developing models, exploring optimum conditions of independent variables for favorable responses, and assessing the comparative influences of various factors in the presence of complex interactions (Bezerra et al. 2008). It involves a collection of optimal production conditions and controllable variables for yielding a polynomial equation for the fermentation process that helps in the determination of interaction between observed results and governable variables (Zambare and Christopher 2011). It involves identifying the cumulative effect of the variables on the responses (Pandey et al. 1999; Mandal and Ghosh 2018). RSM helps in finding the optimal values for the time of fermentation, substrate amount, and range of temperature than a specific single value, enabling more flexibility during process advancement. A goodness of fit of the statistical models with the experimental data indicates that the empirical models derived from RSM can be used efficiently to describe the association between the dependent and independent variables.
RSM is an efficient and robust mathematical approach that comprises various statistical experiments and the utility of various regression analyses for assessing the optimal conditions for maximum yield (Franco-Lara et al. 2006). RSM requires an accurate model design on which depends that how correctly it will predict the relationship to the experimentally obtained data (Aanchal et al. 2016).
The RSM is an estimation of the response function (Y), taking various independent variables of cellulase like—pH temperature, concentration, etc., and taking the help of mathematics and statistical tools, which try to fit the empirical models for the experimentally obtained data (Aanchal et al. 2016). Various order designs are used for optimization using RSM. However, the optimization of the microorganism-produced cellulase is commonly done by first and second-order designs. The total ideal fit to the absolute function is important to be calculated in both domains, where two or more responses are present (Oehlert 2000). The various RSM designs are Plackett–Burman design (PBD), central composite design (CCD) (Somayajula et al. 2012; Ozer et al. 2009), Box-Behnken design (BBD) (Box and Hunter 1957), Doehlert design (Doehlert and Klee 1972), Taguchi design (Velazco 1991) that play a significant role in the mechanism of optimization for the maximum yield of the product. Some of these designs along with their applications have been discussed in Table 1.
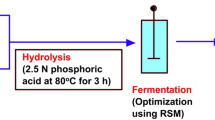
Generally, in experiments of optimization of cellulase production, some factors like enzyme concentration, pH, temperature, substrate concentration are selected based on the original center points (Jeya et al. 2009; Jabasingh and Valli 2012). Several trial runs include those for the factorial design, axial points, and replication of the central points. The experiment is repeated at least three times and the average value of the triplicate is the required response function. In an experiment by Jeya et al. (2009), statistical optimization of hydrolysis condition in the saccharification of rice straw using cellulase from T. hirsuta, by RSM, a CCRD (central composite rotary design) for four factors—enzyme concentration, pH, temperature, substrate concentration was selected. A total of thirty experimental trials were performed. The experiments were performed in triplicate and the average value of the data was the response value that gives the enhanced saccharification percentage (Jeya et al. 2009). An optimal substrate concentration, pH, and temperature were tested for optimizing the saccharification process by RSM, based on the initial results. The CCRD design responses were put in a second-order polynomial equation and the statistical significance of the equation was assessed by the F-test for variance analysis. This proved the regression to be significant statistically. The value that indicated the significance of the model and showed the confidence interval is the ‘Prob > F’. The coefficient determination indicated that the model was suitable for representing the real relationship between the selected reaction variables, quite adequately. Three-dimensional RSM plots were obtained by plotting any two variables, keeping the conversion percentage (i.e., the response) on the Z-axis while other variables were kept at their zero levels. The effects of substrate concentration and enzyme on the process of saccharification and its yield, along with other important results can be obtained from surface plots and iso-response contours.
RSM is used for the optimization of cellulase production from various wastes like water hyacinth (Karmakar and Ray 2011a, b, c), pineapple waste (Saravanan et al. 2013) rice bran, corn bran, sugarcane bagasse, rice straw and sawdust (Hareesh et al. 2016); rice bran and coconut water (Gozan et al. 2018) (Table 1). RSM has been the most commonly used experimental design to optimize the production of various products. As this process helps in the evaluation of multiple factors and their interactions on one or more response variables, the method shows its efficacy. The implementation of RSM has been shown to enhance the production of cellulase by diverse types of microbial species by deciphering the clues for selections that influence the factors affecting the activity of enzymes. Among various types of statistical packages, the most commonly used software is Design-Expert, which helps in easier optimization of variously used dependent and independent variables (Aanchal et al. 2016).
Optimization by molecular docking
The extent of interaction between the enzymes and the target substrate regulates the degradation (by the enzymes) and can be presumed through in-silico analysis by molecular docking methods using various tools (Table 2). During molecular docking analysis, the molecules interact with one another in a favorable configuration resulting in the development of a perfect-fit interactive complex (Kim et al. 2006). Docking interaction between cellulose and related enzymes, with respective substrates can be studied using various molecular docking softwares like Schrodinger, AutoDock Vina, Haddock, etc. Computational tools like PyMol, RasMol, Avogadro can be used as molecular visualization tool to understand the chemistry and structures of the enzyme macromolecules. The ligand and substrate binding active sites can be determined by the analysis of the region in proximity to the bound ligands or macromolecules. The value of energy minimization provides the value of the molecular stability and flexibilities of both the enzyme and substrate that play an important role in the docking interaction. The variation in the interactions provides different values for Gibb's energy and thus helps in the optimization of the molecular interaction. A large number of docking studies have been performed between β-glucosidase and cellobiose, cellotetriose, and cellotetraose to reveal the key residues interacting with the substrates through the formation of hydrogen bonds (Mazlan and Khairudin 2010; Kar et al. 2017; Wickramasinghe et al. 2017). In a study conducted by Selvam et al. (2017), the substrate specificity of cellulase from Acinetobacter sp. was determined by 'flexible ligand, rigid receptor' docking studies, where the binding affinities of the enzyme were tested with the substrates laminaribiose cellotetraose, cellobiose, and cellotetriose. In the study, cellulase showed highest affinity to cellotetraose with the binding energy of − 7.87 kJ/mol. Similar docking analyses were performed by Paul et al. (2020) involving five microbial cellulases with β-d glucose to understand binding affinity and the enzyme–substrate interaction that is associated with cellulose hydrolysis. Their study revealed that the endoglucanase-1 enzyme from Streptomyces sp. had the highest affinity of − 5.61 kcal/mol for cellulose.
These in-silico dynamics and molecular docking approaches can be very useful in real-time screening of cellulases having high catalytic activity and stability, as compared to the laboratory methods, as they save time and effort and are comparatively economical. Molecular docking analysis is often a convenient and handy method of assessing whether mutant enzyme variants serve better catalytic efficiencies as compared to wild-type enzymes without the need for an experimental design involving conventional colorimetric sugar assays, and has been extensively used in research in recent times (Table 2).
Optimization of cellulase production by machine learning
The activity of an enzyme can be governed by several factors such as growth and stress associated environmental conditions that control the ultimate yield of the enzyme.
Even though RSM has proven significant for the optimization of cellulase production, limited research has been done to date on the usage of artificial intelligence (AI), an inner subset of machine learning (ML) tools. In the ML approach, scientists choose specific environmental factors based on which an orthogonal experimental design is formulated. Process variables can be analyzed and shortlisted based on their response to enzyme production by proper parameter design and subsequent logical steps (Cheng et al. 2011). There are various recursive cycles of logical step algorithms and statistical models designed for such purpose of screening process variables and optimizing the parameters; one such widely used is the backpropagation algorithm by ANN. ML uses training datasets to make predictions and decisions by which a foreign dataset can be screened and parameters can be optimized. Directed evolution of enzymes and wet laboratory experimental analyses produce huge data with the potential to be used as training datasets (Li et al. 2019). Therefore, ML techniques are often favored in protein engineering and statistical optimization of enzymes such as cellulases. ML techniques were applied to study the effect of different process conditions on the fermentation of sugarcane bagasse by Saccharomyces cerevisiae (Fischer et al. 2017). The ML-algorithm (MLA) was used to analyze the nature of the cellulase enzyme (Sambasivarao et al. 2014). Recent studies have shown that the use of the ML-based approach for quantitative structural property and activity relation has shown its efficiency and accuracy. The dual-use of ML-based prediction and experimental library generation has markedly enhanced the production of enzymes. To develop the sequence-function relationships of enzyme activity, ML needs a substantial amount of information both in the terms of quality and quantity and thus this results in the uncertainty of the model thereby brings about its impact on the success of the function. Moreover, the data obtained on the enzyme descriptors should possess a correlation with the observed fitness of the mathematical function that helps in the efficient prediction at sufficient levels. Thus, a good amount of information is needed for ML-based predictive tools for optimizing enzyme production and activity (Siedhoff et al. 2020). ML was applied for the identification of unknown cellulolytic microbes and the prediction of their properties by metagenomic analysis (Foroozandeh Shahraki et al. 2020)
In biological studies, ANN methods were often used in the mid-1980s, with high applicability in mathematical methods (Bhat et al. 2000) as MLA. With the use of a computing system for the improvement of the research, ANN has shown a potential application in the procedure of standardization of regulating parameters for enzyme production (Himmel et al. 1999). Despite high demand in industries, the production of cellulase on a large scale is hampered due to inadequate optimization of the enzyme production (Wu et al. 2005). To enhance the yield of cellulase and to make it available at a nominal cost, attention was given to the production parameter optimization process (Filos et al. 2006). But sometimes it faces severe setbacks due to various production constraints. However, this problem can be solved by the integration of an ANN in it. This technique is more accurate and gives the best possible results (Li et al. 2007). Many studies also confirm that this technique is more fruitful than RSM because the representation of non-linearity is done in a much better way in the case of ANN technology (Desai et al. 2008). As a very popular and widely used tool, ANN is applied in the construction of many new models of some bioprocesses like in the functional analyzing of different proteomic or genomic sequences (Liao et al. 2008). This method is generally taken as fitness function and is used for the process of construction of the genetic algorithms (GA) (Wu et al. 2005). It uses the natural selection method which follows the evolution and by this method, the results are well obtained. Recently, ANN was jointly used with GA for the process of optimization. This has become a very friendly way to resolve the problems of optimizations in different bioprocess progressions. These can be mechanistic or theoretical or both. This is eventually used as a new coupling condition method, as a technology based on the ANN, which is more useful compared to RSM processes, and is used for the improvement of the yield of the cellulase by the process of immobilization (Himmel et al. 1999). The advantage of ANN-based prediction is that it allows the mechanism of rational optimization of various types of variables and also possesses the ability to predict the optimization conditions when a sufficient amount of information or data is not available (Bhotmange and Shastri 2011). The process of ANN can be used to stimulate the process of hydrolysis of enzymes for a range of various input variables and help in determining the optimization conditions and complex interactions of variables within the same set of time (Bhotmange and Shastri 2011). Thus, ANN provides a potential complex system that helps in better prediction of optimization parameters (Gama et al. 2017a, b). Improved cellulase activity was found to be achieved using ANN (Fig. 3). In various studies conducted over the years, the ANN architecture is comprised of three neurons namely, the pH, carbodimide concentration, and coupling time which are there in the input layer, the neurons which are hidden in the interior layer and one neuron, which is the activity yield (Roy et al. 2003). Now, the new scaled data are taken for training in an ANN model by using a back-propagation algorithm. The tangent sigmoid and pure linear functions are used to transfer functions in the hidden and output layers of the ANN (Rajoka et al. 2007). So, the mean square error which is taking place between the observations for the output neurons and the actual outputs is usually calculated and propagated backward by the network itself. This algorithm is then adjusted through the weight of each of the other neurons (Taniguchi et al. 1989). After getting the mean square error, we can calculate the training time, and eventually, then we can end it and build the corresponding ANN (Gaur and Lata Khare 2005) (Figs. 3, 4).
Chang et al. (2011) and Gautam et al. (2011) optimized the cellulase production by solid-state fermentation of Trichoderma viride using wheat straw and wheat bran, using AI-based techniques, where a backpropagation network was designed with Levenberg-Marquardt training algorithm. The pure linear functions and tangent sigmoid were used as the transfer functions in the output layers and hidden layers of ANN respectively. The GA coupled with ANN was used to optimize the process parameters, to detect the concentration of various suitable factors. Saravanan et al. (2012a, b) used BBD statistical tool with GA to optimize the parameters for cellulase production by Trichoderma reesei using pineapple waste as a substrate, where they found the BBD model better than GA. For minimizing the number of experiments but with the accurate result, an ANN was used to model the enzymatic hydrolysis process involved during enzymatic degradation of apple pomace by various fungal cellulases (Gama et al. 2017a, b) Such result effectively could curtail the time and cost of experimentation. Caramihai and Severin (2018) optimized cellulase production in fed-batch culture of Aspergillus niger using an agricultural residue as the substrate with the application of ANN (Caramihai and Severin 2018). ANN was also applied to optimize cellulose and xylanase production by T. ressei utilizing bagasse as a sole carbon source by Singh et al. (2008). They found ANN as a good prediction tool for complex fermentation process involving two enzymes. Figure 4 shows optimization by ANN for the production of cellulase by solid-state fermentation (SSF).
RSM was applied for optimization of various parameters in a submerged fermentation medium for cellulase production by Trichoderma harzanium ATCC 20846 followed by a computational fluid dynamic (CFD) simulation for additional factor (Kumar et al. 2020)
Application of genetic algorithm for optimization of cellulase production
GA is a type of evolutionary algorithm (EA) that works based on Darwin's theory of natural selection. This algorithm helps in expressing the genes as a combination of digits '1' and '0'. Chromosomes comprising a set of genes are referred to as strings. GA helps in matching and mixing independent variables to develop superior offspring resulting in the enhancement of flexibility, efficacy, and efficiency of the optimized model.
The process parameters of cellulase production by Ochrobactrum haematophilum, a cellulolytic bacterium was developed by simulated annealing (SA) algorithm to increase the production, where after comparison it was found that the SA-based RSM model can give better predictions for cellulase production than the GA-based RSM models or sole RSM models (Parkhey et al. 2017).
Exoglucanase activity from Penicillium roqueforti by SSF was achieved by the use of simplex-centroid design along with the coupling of ANN and GA. It was found that such optimization predicted an increase of production by up to 1263% when a mixture of agro-wastes namely, corn cob, green coconut shell, and sugarcane bagasse was used in comparison to the usage of the individual substrate alone (da Silva et al. 2020).
Sirohi et al. (2018) opined that GA may be used as a tool to determine the optimized conditions for the production of enzymes from wastes after its successful application using mathematical and genetic optimizers for standardizing the parameters for cellulase production by T. reesei using pea hull as carbon source.
Application of LabVIEW-based intelligent system for monitoring of bioprocesses for optimization of cellulase production
LabVIEW is a program that helps to automate the implementation of various software-associated sensors resulting in the development of a graphical programming environment. It possesses a library of functions and subroutines for various types of programming tasks. It also helps in providing library-specific data acquisition, data processing, control of instruments, analysis, and storage (Alford 2006).
Cellulase production by SSF of Aspergillus niger was evaluated by statistical design methodology. The measurements were stored in real-time by using the graphical interface programming package LabVIEW 8.2 (Farinas et al. 2011). On the other hand, in another study Myceliophthora sp. I-1D3b was used by SSF, at pilot-scale, taking sugar cane bagasse (SCB) and wheat bran (WB) as substrates and the bioreactor was forcefully aerated packed bed bioreactor (PBB) connected to LabVIEW 8.5 routine (Casciatori et al. 2013).
Conclusion and future prospects
The solid wastes, generated from agricultural activities or forests pose serious environmental problems. Major part os these wastes are biodegradable cellulosic and lignocellulose substances. Bioconversion of cellulosic wastes is usually left unattended as garbage and needs intensive research, as production of value-added materials from these wastes can be the most important way of waste management. Researches on optimization procedure before enzyme production from agricultural wastes reveal that application of a combined procedure of RSM, ANN or GA can lead to the improvement in product yield. Increased utilization of cellulosic waste not only makes it available at a nominal cost but also will show a new vista for a sustainable way of waste utilization. All the aforementioned optimization methods are found useful and more effective than OFAT method due to many reasons as discussed above. But no particular method can be said to be most preferable, as it is case-specific. The RSM is found to be advantageous over OFAT where several process parameters are involved, which are difficult to handle at a time by OFAT. On the other hand, ANN is proved to be useful, when there is insufficient detailed knowledge of the underlying process, and formulation of a reaction mechanism is not possible as it can simulate an arbitrary bioprocess. ML methods have been increasingly applied to find patterns in data that help predict enzyme protein structures, improve enzyme stability, solubility, and function, predict substrate specificity and guide rational protein design of efficient biocatalysts. But ML requires good information content in terms of the quality and quantity of the enzyme protein. Although molecular docking is not directly applied for optimization of process parameters, it can indicate the activation energy and substrate affinity of the enzyme. Similarly, the involvement of optimization for enzyme production based on genetics lacks a mathematical foundation, but it greatly improves the prediction capability of computer models. Prediction bioprocess thus can remove major bottlenecks in improving the scale-up technology for the production of microbial cellulase from left-overs. Advanced and modern technologies and high-throughput techniques of optimization are of immense importance for the production of novel microbial cellulases with greater potentiality.
References
Aanchal, Akhtar N, Kainka, Goyal D, Goyal A (2016) Response surface methodology for optimization of microbial cellulase production. Rom Biotechnol Lett 21(5):11832–11841
Akhtar N, Anchal A, Goyal D, Goyal A (2015) Simplification and optimization of media ingredients for enhanced production of CMCase by newly isolated Bacillus subtilis NA15. Environ Prog Sustain Energy 34(2):533–541
Alford JS (2006) Bioprocess control: advances and challenges. Comput Chem Eng 30:1464–1475
Ali SBR, Muthuvelayudham R, Viruthagiri T (2013) Enhanced production of cellulase from tapioca stem using response surface methodology. Innova Roman Food Biotechnol 12:41–51
Badhan A, Chadha B, Kaur J, Saini H, Bhat M (2007) Production of multiple xylanolytic and cellulolytic enzymes by thermophilic fungus Myceliophthora sp. IMI 387099. Bioresour Technol 98(3):504–510
Baraa R, Anwar R (2017) Avogadro program for chemistry education: to what extent can molecular visualization and three-dimensional simulations enhance meaningful chemistry learning? World J Chem Educ 5:136–141
Bezerra MA, Santelli RE, Oliveira EP, VillarEscaleira LSLA (2008) Response surface methodology (RSM) as a tool for optimization in analytical chemistry. Talanta 76:965–977
Bhachoo J, Beuming T (2017) Investigating protein–peptide interactions using the Schrödinger computational suite. In: Schueler-Furman O, London N (eds) Modeling peptide–protein interactions. Methods in molecular biology, vol 1561. Humana Press, New York
Bhat MK (2000) Cellulases and related enzymes in biotechnology. Biotechnol Adv 18:355–383
Bhotmange M, Shastri P (2011) Application of artificial neural networks to food and fermentation technology. In: Suzuki K (ed) Artificial neural networks—industrial and control engineering applications. InTech, Rijeka, pp 201–222 (ISBN 978-953-307-220-3)
Box G, Hunter JS (1957) Multifactor experimental designs for exploring response surfaces. Ann Math Stat 28:195–241
Caramihai M, Severin I (2018) Enzyme production modeling simulation using neural techniques. Int J Biol Biomed 3:26–29
Casciatori FP, Casciatori PA, Thoméo JC (2013) Cellulase production in packed bed bioreactor by solid-state fermentation. In: 21st European biomass conference and exhibition, 3–7 June 2013, Copenhagen, Denmark
Chang C, Guizhuan X, Junfang Y, Wang D (2011) Optimization of cellulase production using agricultural wastes by artificial neural network and genetic algorithm. Chem Prod Process Model 6(1)
Cheng SW, Wang YF, Hong B (2012) Statistical optimization of medium compositions for chitosanase production by a newly isolated streptomyces albus. Braz J Chem Eng 29(04):691–698
da Silva Nunes N, Carneiro LL, de Menezes LHS, de Carvalho MS, Pimentel AB, Silva TP, Pacheco CSV, Tavares IMC, Santos PH, das Chagas TP, da Silva EGP, de Oliveira JR, Bilal M, Franco M (2020) Simplex-centroid design and artificial neural network-genetic algorithm for the optimization of exoglucanase production by Penicillium roqueforti ATCC 10110 through solid-state fermentation using a blend of agroindustrial wastes. BioEnergy Res 13:1130–1143. https://doi.org/10.1007/s12155-020-10157-0
Desai KM, Survase SA, Saudagar PS, Lele SS, Singhal RS (2008) Comparison of artificial neural network (ANN) and response surface methodology (RSM) in fermentation media optimization: case study of fermentative production of scleroglucan. Biochem Eng J 41:266–273
Doehlert DH, Klee VL (1972) Experimental designs through level reduction of the d-dimensional cuboctahedron. Discrete Math 2:309–334
Dominguez C, Boelens R, Bonvin AM (2003) HADDOCK: a protein–protein docking approach based on biochemical or biophysical information. J Am Chem Soc 125(7):1731–1737
Farinas CS, Vitcosque GL, Fonseca RF, Neto VB, Couri S (2011) Modeling the effects of solid state fermentation operating conditions on endoglucanase production using an instrumented bioreactor. Ind Crops Prod 34(2011):1186–1192
Filos G, Tziala T, Lagios G, Vynios DH (2006) Preparation of cross-linked cellulases and their application for the enzymatic production of glucose from municipal paper wastes. Prep Biochem Biotechnol 36:111–125
Fischer J, Lopes VS, Cardoso SL, Coutinho FU, Cardoso VL (2017) Machine learning techniques applied to lignocellulosic ethanol in simultaneous hydrolysis and fermentation. Braz J Chem Eng 34(1):53–63
Forli S, Huey R, Pique ME, Sanner MF, Goodsell DS, Olson AJ (2016) Computational protein-ligand docking and virtual drug screening with the AutoDock suite. Nat Protoc 11(5):905–919
Foroozandeh Shahraki M, Ariaeenejad S, Fallah Atanaki F, Zolfaghari B, Koshiba T, Kavousi K, Salekdeh GH (2020) MCIC: automated identification of cellulases from metagenomic data and characterization based on temperature and pH dependence. Front. Microbiol. 11: https://doi.org/10.3389/fmicb.2020.567863
Foyle T, Jennings L, Mulcahy P (2007) Compositional analysis of lignocellulosic materials: evaluation of methods used for sugar analysis of waste paper and straw. Biores Technol 98(16):3026–3036
Franco-Lara E, Link H, Weuster-Botz D (2006) Evaluation of artificial neural networks for modelling and optimization of medium composition with a genetic algorithm. Process Biochem 41:2200–2206. https://doi.org/10.1016/j.procbio.2006.06.024)
Gama R, Van Dyk JS, Burton MH (2017a) Using an artificial neural network to predict the optimal conditions for enzymatic hydrolysis of apple pomace. 3 Biotech 7:138
Gama R, Van Dyk JS, Burton MH, Pletschke BI (2017b) Using an artificial neural network to predict the optimal conditions for enzymatic hydrolysis of apple pomace. 3. Biotech 7(2):138. https://doi.org/10.1007/s13205-017-0754-1
Gaur R, Lata Khare SK (2005) Immobilization of xylan-degrading enzymes from Scytalidium thermophilum on Eudragit L-100. World J Microbiol Biotechnol 21:1123–1128
Gautam SP, Bundela PS, Pandey AK, Khan J, Awasthi MK, Sarsaiya S (2011) Optimization for the production of cellulase enzyme from municipal solid waste residue by two novel cellulolytic fungi. Biotechnol Res Int 2011:1–8. https://doi.org/10.4061/2011/810425
Geiger EO, Vogel HC, Todaro CL (1996) Fermentation and biochemical engineering handbook; ed., vol 2. Noyes Publications, pp 161–180
Gonzalez R, Islas L, Obregon A-M, Escalante L, Sanchez S (1995) Gentamicin formation in Micromonospora purpurea: stimulatory effect of ammonium. J Antibiot 48:479–483
Goukanapalle PKR, Kanderi DK, Rajoji G (2020) Optimization of cellulase production by a novel endophytic fungus Pestalotiopsis microspora TKBRR isolated from Thalakona forest. Cellulose 27:6299–6316. https://doi.org/10.1007/s10570-020-03220-8
Gozan M, Harahap AF, Bakti CP, Setyahadi S (2018) Optimization of cellulase production by Bacillus sp. BPPT CC RK2 with pH and temperature variation using response surface methodology. E3S Web Conf 67:02051
Grosdidier A, Zoete V, Michielin O (2011) SwissDock, a protein-small molecule docking web service based on EADock DSS. Nucleic Acids Res 39:W270–W277
Hareesh ES, Faisal PA, Benjamin S (2016) Optimization of parameters for the production of cellulase from Achromobacter xylosoxidans BSS4 by solid-state fermentation. Electron J Biol 12:4
Henrissat B, Teeri TT, Warren RAJ (1998) A scheme for designating enzymes that hydrolyse the polysaccharides in the cell walls of plants. FEBS Lett 425(2):352–354
Himmel ME, Ruth MF, Wyman CE (1999) Cellulase for commodity products from cellulosic biomass. Curr Opin Biotechnol 10(4):358–364
Hoa BT, Hung PV (2013) Optimization of nutritional composition and fermentation conditions for cellulase and pectinase production by Aspergillus oryzae using response surface methodology. Int Food Res J 20(6):3269–3274
Hussain Z, Sajjad W, Khan T (2019) Production of bacterial cellulose from industrial wastes: a review. Cellulose 26:2895–2911
Imran M, Anwar Z, Irshad M, Javid A, Hussain A, Ali S (2017) Optimization of cellulase production from a novel strain of Aspergillus Tubingensis IMMIS2 through response surface methodology. Biocatal Agric Biotechnol 12:191–198. https://doi.org/10.1016/j.bcab.2017.10.005
Jabasingh SA (2011) Response surface methodology for the evaluation and comparison of cellulase production by Aspergillus nidulans SU04 and Aspergillus nidulans MTCC344 cultivated on pretreated sugarcane bagasse. Chem Biochem Eng 25(4):501–511
Jabasingh S, Valli N (2012) Optimization of cellulase synthesis by RSM and evaluation of ethanol production from enzymatically hydrolyzed sugarcane bagasse using Saccharomyces cerevisiae. J Sci Ind Res 71(5)
Jeya M, Zhang YW, Kim IW, Lee JK (2009) Enhanced saccharification of alkali-treated rice straw by cellulase from Trametes hirsuta and statistical optimization of hydrolysis conditions by RSM. Biores Technol 100(21):5155–5161
Johnson EA, Sakajoh M, Halliwell G, Madia A, Demain AL (1982) Saccharification of complex cellulosic substrates by the cellulase system from Clostridium thermocellum. Appl Environ Microbiol 43:1125–1132
Kar B, Verma P, Patel GK, Sharma AK (2017) Molecular cloning, characterization and in silico analysis of a thermostable β-glucosidase enzyme from Putranjiva roxburghii with a significant activity for cellobiose. Phytochemistry 140:151–165
Karmakar M, Ray RR (2010) Extra cellular endoglucanase production by Rhizopus oryzae in solid and liquid state fermentation of agro wastes. Asian J Biotechnol 2(1):27–36
Karmakar M, Ray RR (2011a) Optimization of endoglucanase production in liquid state fermentation from waterhyacinth by Rhizopus oryzae using response surface methodology. Aust J Basic Appl Sci 5(3):713–720
Karmakar M, Ray RR (2011b) A statistical approach for optimization of simultaneous production of β-glucosidase and endoglucanase by Rhizopus oryzae from solid-state fermentation of water hyacinth using central composite design. Biotechnol Res Int 2011:574983–574990
Karmakar M, Ray RR (2011c) Optimization of production conditions of extra cellular β-glucosidase in submerged fermentation of waterhyacinth by Rhizopus oryzae using response surface methodology. Res J Pharm Biol Chem Sci 2:299–308
Kim D, Park BH, Jung BW, Kim MK, HongLee SI (2006) Identification and molecular modeling of a family 5 endocellulase from Thermus caldophilus GK24, a cellulolytic strain of Thermus thermophiles. Int J Mol Sci 7:571–589
Kubicek C (1993) From cellulose to cellulase inducers: facts and fiction. In: Proceedings of the second Tricel symposium on Trichoderma reesei cellulases and other hydrolases, Espoo, Finland. Foundation for Biotechnical and Industrial Fermentation Research, Helsinki, pp 181–188
Kumar NR, Sambavi TR, Baskar G, Renganathan S (2020) International experimental validation of optimization by statistical and CFD simulation methods for cellulase production from waste lignocellulosic mixture. J Mod Sci Technol 5(2):45–58
Laskowski RA, Jabłońska J, Pravda L, Vařeková RS, Thornton JM (2018) PDBsum: structural summaries of PDB entries. Protein Sci 27(1):129–134
Li Y, Irwin DC, Wilson DB (2007) Processivity, substrate binding, and mechanism of cellulose hydrolysis by Thermobifida fusca Cel9A. Appl Environ Microbiol 73(10):3165–3172
Li G, Dong Y, Reetz MT (2019) Can machine learning revolutionize directed evolution of selective enzymes? Adv Synth Catal 361:2377–2386
Liao L, McClatchy DB, Yates JR (2009) Shotgun proteomics in neuroscience. Neuron. 63(1):12–26. https://doi.org/10.1016/j.neuron.2009.06.011
Lincoln T (2013) Agriculture, plant physiology, and human population growth: past, present, and future. Theor Exp Plant Physiol 25(3):167–181
Lindahl E, Azuara C, Koehl P, Delarue M (2006) NOMAD-Ref: visualization, deformation and refinement of macromolecular structures based on all-atom normal mode analysis. NAR 34:W52–W56
Lohri CR, Diener S, Zabaleta I (2017) Treatment technologies for urban solid biowaste to create value products: a review with focus on low- and middle-income settings. Rev Environ Sci Biotechnol 16:81–130
Lua RC, Lichtarge O (2010) PyETV: a PyMOL evolutionary trace viewer to analyze functional site predictions in protein complexes. Bioinformatics (oxford, England) 26(23):2981–2982
Mandal M, Ghosh U (2018) Application of statistical tool for optimization of physical parameters for cellulase production under solid state fermentation. Indian J Biotechnol 17(3):441–447
Manisalidis I, Stavropoulou E, Stavropoulos A, Bezirtzoglou E (2020) Environmental and health impacts of air pollution: a review. Front Public Health 8:14
Mazlan NSF, Khairudin NBA (2010) Docking Study of β-glucosidase B (BglB) from P. polymyxca with cellobiose and cellotetrose. J Med Bioeng 3:78–83
Mielenz JR (2001) Ethanol production from biomass: technology and commercialization status. Curr Opin Microbiol 4(3):324–329
Mukherjee S, Karmakar M, Ray RR (2011) Production of extra cellular exo glucanase by Rhizopus oryzae from submerged fermentation of agro wastes. Recent Res Sci Technol 3(3):69–75
Oehlert GW (2000) Design and analysis of experiments: response surface design. W. H. Freeman Company, New York, pp 509–542
Ozer A, Gurbuz G, Calmila A, Korbaht BK (2009) Biosorption of copper (II) ions on Enteromorpha prolifera: application of response surface methodology (RSM). Chem Eng J 146(3):377–387
Panda BP, Ali M, Javed S (2007) Fermentation process optimization. Res J Microbiol 2:201–208
Pandey A, Selvakumar P, Soccol CR, Nigam P (1999) Solid state fermentation for the production of industrial enzymes. Curr Sci 77:149–162
Parkhey P, Gupta P, Eswari JS (2017) Optimization of cellulase production from isolated cellulolytic bacterium: comparison between genetic algorithms, simulated annealing, and response surface methodology. Chem Eng Commun 204(1):28–38
Paul M, Panda G, Mohapatra PKD, Thatoi H (2020) Study of structural and molecular interaction for the catalytic activity of cellulases: an insight in cellulose hydrolysis for higher bioethanol yield. J Mol Struct 1204(15):127547
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612
Piovesan D, Minervini G, Tosatto SC (2016) The RING 2.0 web server for high quality residue interaction networks. Nucleic Acids Res 44(W1):W367–W374
Rajoka MI, ZiaKhalil-ur-Rehman Y (2007) A surface immobilization method of endoglucanase from Cellulomonas biazotea mutant improved catalytic properties of biocatalyst during processing. Protein Pept Lett 14:734–741
Rodhe AV, Sateesh L, Sridevi J, Venkateswarlu B, Rao LV (2011) Enzymatic hydrolysis of sorghum straw using native cellulase produced by T. reesei NCIM 992 under solid state fermentation using rice straw. 3 Biotech 1:207–215
Roger S, Bissell A (1992) RasMol: a program for fast realistic rendering of molecular structures with shadows. In: Proceedings of the 10th eurographics UK '92 conference, University of Edinburgh, Scotland
Roy I, Gupta A, Khare SK, Bisaria VS, Gupta MN (2003) Immobilization of xylan-degrading enzymes from Melanocarpus albomyces IIS 68 on the smart polymer Eudragit L-100. Appl Microbiol Biotechnol 61:309–313
Sambasivarao SV, Granum DM, Wang H, Mark Maupin C (2014) Identifying the enzymatic mode of action for cellulase enzymes by means of docking calculations and a machine learning algorithm. AIMS Mol Sci 1(1):59–80. https://doi.org/10.3934/molsci.2014.1.59
Sandhu SK, Oberoi HS, Babbar N, Miglani K, Chadha BS, Nanda DK (2013) Two-stage statistical medium optimization for augmented cellulase production via solid-state fermentation by newly isolated Aspergillus niger HN-1 and application of crude cellulase consortium in hydrolysis of rice straw. J Agric Food Chem 61:12653–12661
Saranraj P, Stella D, Reetha D (2012) Microbial cellulases and its applications: a review. Int J Biochem Biotechnol Sci 1:1–12
Saravanan P, Muthuvelayudham R, Rajesh K, Kannan R, Viruthagiri T (2012a) Optimization of cellulase production using Trichoderma reesei by RSM and comparison with genetic algorithm. Front Chem Sci Eng 6(4):443–452
Saravanan P, Muthuvelayudham R, Rajesh Kannan R (2012b) Optimization of cellulase production using Trichoderma reesei by RSM and comparison with genetic algorithm. Front Chem Sci Eng 6:443–452
Saravanan P, Muthuvelayudham RT, Viruthagiri T (2013) Enhanced production of cellulase from pineapple waste by response surface methodology. J Eng 2013:1–8
Selvendran RR (1985) Developments in the chemistry and biochemistry of pectin and hemicellulosic polymers. J Cell Sci 2:51–88
Selvam K, Senbagam D, Selvankumar T, Sudhakar C, Kamala-Kannan S, Senthilkumar B, Govarthanan M (2017) Cellulase enzyme: homology modeling, binding site identification and molecular docking. J Mol Struct 1150:61–67
Shankar T, Isaiarasu L (2012) Statistical optimization for cellulase production by Bacillus pumilus EWBCM1 using response surface methodology. Glob J Biotechnol Biochem 7(1):1–6
Siedhoff NE, Schwaneberg U, Davari MD (2020) Machine learning-assisted enzyme engineering. Methods Enzymol. https://doi.org/10.1016/bs.mie.2020.05.005
Singh J, Sharma A (2012) Application of response surface methodology to the modelling of cellulase purification by solvent extraction. Adv Biosci Biotechnol 3(4):408–416
Singh A, Tatewar D, Shastri PN, Pandharipande SL (2008) Application of ANN for prediction of cellulase and xylanase production by Trichoderma ressei under SSF condition. Indian J Chem Technol 15:53–58
Singh V, Haque S, Niwas R, Srivastava A, Pasupuleti M, Tripathi CK (2017) Strategies for fermentation medium optimization: an in-depth review. Front Microbiol 7:2087
Sirohi R, Singh A, Tarafdar A, Shahi NC (2018) Application of genetic algorithm in modelling and optimization of cellulase production. Bioresour Technol 270:751–754
Somayajula A, Asaithambi P, Susree M, Matheswaran M (2012) Sonoelectrochemical oxidation for decolorization of reactive red 195. Ultrasonics Sonochem 19(4):803–811
Taniguchi M, Kobayashi M, Fujii M (1989) Properties of a reversible soluble insoluble cellulase and its application to repeated hydrolysis of crystalline cellulose. Biotechnol Bioeng 34(8):1092–1097
Velazco EE, Bendell A, Disney J, Pridmore WA (1991) Taguchi methods: applications in world industry. Interfaces (Providence) 21:99–101
Wang J-C, Hu S-H, Liang Z-C, Yeh C-J (2005) Optimization for the production of water-soluble polysaccharide from Pleurotus citrinopileatus in submerged culture and its antitumor effect. Appl Microbiol Biotechnol 67:759–766
Wickramasinghe GHIM, Rathnayake PPAMSI, Chandrasekharan NV (2017) Trichoderma virens β-glucosidase I (BGLI) gene; expression in Saccharomyces cerevisiae including docking and molecular dynamics studies. BMC Microbiol 17:137
Wu LL, Yuan XY, Sheng J (2005) Immobilization of cellulase in nanofibrous PVA membranes by electrospinning. J Membr Sci 250(1):167–173
Zambare V, Christopher L (2011) Statistical analysis of cellulase production in Bacillus amyloliquefaciens. ELBA Bioflux 3:38–45
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors do not have any conflict of statement.
Ethical statement
The work was conducted under guidelines of Institutional Ethics Committee MAKAUT: IEC-(18-19)/02.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lahiri, D., Nag, M., Mukherjee, D. et al. Recent trends in approaches for optimization of process parameters for the production of microbial cellulase from wastes. Environmental Sustainability 4, 273–284 (2021). https://doi.org/10.1007/s42398-021-00189-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42398-021-00189-3