Abstract
Erwinia pyrifoliae, the Gram-negative bacterial pathogen responsible for black shoot blight, exhibits symptoms similar to E. amylovora (fire blight), though with distinct molecular characteristics. Given its prevalence primarily in South Korea and the availability of only nine assembled genomes, there is a lack of high-quality genome sequences and annotated genetic information for E. pyrifoliae. We present the sequencing and assembly of a Korean E. pyrifoliae strain, YKB12327, isolated from a diseased apple tree branch, using a combination of long Oxford Nanopore Technologies and short Illumina sequence reads. This genome comprises a circular chromosome and three plasmid sequences, totaling 4,061,634 bp. Annotation of YKB12327 identifies 3123 coding sequence protein-coding genes, 22 rRNA genes (5S, 16S, and 23S), and 76 tRNA genes. Our sequence data will enrich the current E. pyrifoliae genome resources and facilitate in understanding its evolution, diversity and structural variations, as well as the molecular basis of pathogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Erwinia pyrifoliae, commonly known as ‘black shoot blight’, is a Gram-negative bacterium belonging to the Erwiniaceae family (Thompson et al. 2019; Lee et al. 2020). This bacterium predominantly infects plants, particularly pears, leading to conditions known as “Bacterial Shoot Blight” or “Asian Pear Blight” (Kim et al. 2001; Lee et al. 2020). Initially reported in pear trees in South Korea in 1995, the disease exhibits symptoms similar to those of Erwinia amylovora (fire blight). However, molecular analysis distinguished E. pyrifoliae as a distinct species (Kim et al. 1999). Subsequent investigations revealed that some bacterial samples from Japan, previously thought to be E. amylovora (Beer et al. 1996), were in fact E. pyrifoliae (Geider et al. 2009; Thapa et al. 2013). In 2013, its unexpected discovery in strawberries in the Netherlands underscored the pathogen’s host non-specificity and geographic versatility (Wenneker and Bergsma-Vlami 2015).
Unlike E. amylovora, little is known about the genetic basis of virulence and environmental adaptation in E. pyrifoliae. The genome sequence of E. pyrifoliae consists of a circular chromosome and plasmids, containing typical bacterial genomic elements such as genes for metabolism, replication, and cell division, alongside pathogenicity-related genes (Smits et al. 2010; Llop et al. 2012; Lee et al. 2020). Previous research has yielded only nine assembled sequences of the E. pyrifoliae genome available/released in National Center for Biotechnology Information (NCBI) GenBank (https://www.ncbi.nlm.nih.gov/datasets/genome/?taxon=79967). There is a pressing need for additional genomic information and gene annotation to enrich the genomic data of E. pyrifoliae. Meanwhile, breakthroughs have been made in previous study of the genetic diversity of this pathogen, which will enable us to better understand the genetic characteristics and pathogenic genes of this pathogen (Ham and Park 2024).
Branch of an apple (Malus domestica cv. Fuji) tree showing black shoot blight were collected from Idong-myeon, Pocheon-si, Gyeonggi-do, Republic of Korea (38°05′24″N, 127°38′52″E). The apple branches were surface sterilized using 70% ethanol and were dissected into 3 ~ 5 mm samples. The dissected samples were immersed in 10 mM phosphate-buffered saline (PBS) (pH 7.2) (Biosesang, Yongin, Korea) in a sterile mortar. Single colonies observed after 48 h of incubation on Nutrient Broth (NB) medium at 27 °C were isolated and subsequently purified. The purified strain was presumptively identified as E. pyrifoliae by internal transcribed spacer (ITS) region sequencing. The primer pair ITS-F (5′-AGAGTTTGATCMTGGCTCAG-3′) and ITS-R (5′-TACGGYTACCTTGTTACGACTT-3′), was used to identify the strain as E. pyrifoliae. Total genomic DNA was extracted using a PureLink™ Genomic DNA Mini Kit (Cat. No. K182002, Thermo Fisher Scientific Inc., Waltham, MA, USA) following the manufacturer’s instructions. Genomic DNA integrity was assessed via 1% agarose gel electrophoresis, while DNA purity was evaluated using a NanoDrop UV–Vis Spectrophotometer (Cat. No. ND-2000, Thermo Fisher Scientific). DNA concentrations were quantified using a Qubit dsDNA HS Quantification Assay Kit (Cat. No. Q32854, Thermo Fisher Scientific) and measured using a Qubit 4 Fluorometer (Cat. No. Q33238, Thermo Fisher Scientific). Libraries for long-read sequencing were prepared through end-repair and dA-tailing, barcode and adapter ligation, and purification of ligated DNA using the NEBNext® Ultra™ II End Repair/dA-Tailing Module [Cat. No. E7546, New England Biolabs Co. (NEB), Ipswich, MA, USA], FFPE Repair Mix NEBNext® Quick Ligation Module (Cat. No. E6056, NEB), and Native Barcoding Kit [Cat. No. SQK-NBD114.24, Oxford Nanopore Technologies Co. (ONT), Oxfordshire, UK], respectively, following recommendations by MinION. Genomic DNA long-read sequencing was conducted using the MinION Mk1C device R10.4.1 (Cat. No. MIN-101 C, ONT) with a SpotON Flow Cell (Cat. No. FLO-MIN114, ONT) according to the manufacturer’s instructions and managed using MinKNOW software v4.1.23 (ONT). Short-read sequencing was performed to refine genomic sequences. Genomic DNA paired-end libraries with 350-bp inserts were generated using the TruSeq Nano DNA High Throughput Preparation Kit (Cat. No. 20015965, Illumina Inc., San Diego, CA, USA). These paired-end libraries were sequenced at Macrogen Co. (Seoul, Korea) using Illumina Sequencing by Synthesis (SBS) Technology (Illumina).
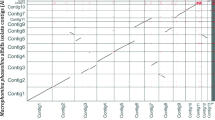
Nanopore long-read sequencing yielded 120,000 raw reads with an N50 length of 14,677 bp and a total sequence length of 754.3 Mb (188.6× coverage), while Illumina short-read sequencing generated 1204 Mb (150.5×coverage) of paired-end sequences (Table 1). We utilized the Trimmomatic v0.38 tool to assess the quality of Nanopore long-read sequences, specifically targeting the removal of adapters, low-quality reads (defined as those containing “N” in > 10% of nucleotides), and duplicated reads. The resultant clean reads underwent assembly using the Long Read Support (beta) plugin 23.0 within the Qiagen CLC Genomics Workbench v.23.0.4 software (Qiagen, Hilden, Germany), employing default parameters. The raw assembly was subjected to two rounds of polishing with short reads from Illumina sequencing. The genomic sequence of YKB12327 was assembled into one chromosome with a size of 4,018,953 bp (53.4% G + C content) and three circular plasmids with sizes of 34,650 bp (49.7% G + C content), 4942 bp (53.4% G + C content), and 3089 bp (49.0% G + C content) (Fig. 1). Validation of the genome assemblies was performed using the BUSCO v4.1.4 software (Simão et al. 2015), leveraging 1614 Nb of BUSCO markers in Embryophyta (odb10). The assembly quality was evidenced by the detection of 124 (100%) complete and single-copy Benchmarking Universal Single-Copy Orthologs (BUSCOs) in the assembled genome, indicating high-quality assembly. Compared with the assembled genomes, the genomic sequence of YKB12327 exhibited high homology to four strains: EpK1/15, YKB12328, Ep1/96, and DSM 12163 (Supplementary Fig. S1). Whole-genome alignment analysis revealed that the chromosome sequence of YKB12327 closely resembled those of the four neighboring genomes, while the plasmids varied significantly in both length and number.
The genome sequences were annotated for CDSs, ribosomal RNA (rRNA) genes, and transfer RNA (tRNA) genes using using the NCBI Prokaryotic Genome Annotation Pipeline (PGAP) (https://www.ncbi.nlm.nih.gov/genome/annotation_prok/) (Haft et al. 2018). A total of 3123 protein-coding genes, 22 rRNA genes (5S, 16S, and 23S), and 76 tRNA genes were predicted. Moreover, referring to information reported by Kube et al. (2010), we identified 26 disease-causing genes for subsequent investigation (Supplementary Table 1).
In summary, our study sequenced, assembled, and annotated the genome of E. pyrifoliae YKB12327, offering insights into its genetic makeup. We believe that the availability of the complete genome sequence of strain YKB12327 will further support studies to understand evolution, diversity and structural variations of E. pyrifoliae strains, as well as the molecular basis of pathogenesis.
Circular representation of E. pyrifoliae YKB12327 genome. The innermost circle is the ideogram of chromosome and plasmids in Mb scale, surrounding concentric circles of Prokka on the forward and reverse strand (blue); mobile genetic elements Alien Hunter (orange), MobileOG (purple), and Phigaro (sky blue) on the forward (inner) and reverse strand (outside); positive (green) and negative (red) GC skew; and GC content (black)
Data availability
All relevant data are within the article and its supplementary information files.
References
Beer SV, Kim JH, Zumoff CH, Bogdanove AJ, Laby RJ, Gustafson HL, Momol T, Aldwinckle HS, Tanii A, Tamura O (1996) Characterization of bacteria that cause bacterial shoot blight of pear in Japan. Acta Hortic 411:179–182. https://doi.org/10.17660/ActaHortic.1996.411.36
Geider K, Auling G, Jakovljevic V, Völksch B (2009) A polyphasic approach assigns the pathogenic Erwinia strains from diseased pear trees in Japan to Erwinia pyrifoliae. Lett Appl Microbiol 48:324–330. https://doi.org/10.1111/j.1472-765X.2008.02535.x
Haft DH, DiCuccio M, Badretdin A et al (2018) RefSeq: an update on prokaryotic genome annotation and curation. Nucleic Acids Res 46:D851–D860. https://doi.org/10.1093/nar/gkx1068
Ham H, Park DS (2024) New insights and approach toward the genetic diversity and strain typing of Erwinia pyrifoliae based on rsxC, an Electron Transport Gene. Plant Dis 108:296–301. https://doi.org/10.1094/PDIS-03-23-0475-SC
Kim W, Gardan L, Rhim S, Geiderl K (1999) Erwinia pyrifoliae sp. nov., a novel pathogen that affects Asian pear trees (Pyrus pyrifolia Nakai). Int J Syst Bacteriol 49:899–906. https://doi.org/10.1099/00207713-49-2-899
Kim W-S, Hildebrand M, Jock S, Geider K (2001) Molecular comparison of pathogenic bacteria from pear trees in Japan and the fire blight pathogen Erwinia amylovora. Microbiology 147:2951–2959. https://doi.org/10.1099/00221287-147-11-2951
Kube M, Migdoll AM, Gehring I et al (2010) Genome comparison of the epiphytic bacteria Erwinia billingiae and E. tasmaniensis with the pear pathogen E. pyrifoliae. BMC Genomics 11:1–15. https://doi.org/10.1186/1471-2164-11-393
Lee GM, Ko S, Oh EJ et al (2020) Comparative genome analysis reveals natural variations in the genomes of Erwinia pyrifoliae, a black shoot blight pathogen in apple and pear. Plant Pathol J 36:428–439. https://doi.org/10.5423/PPJ.OA.06.2020.0097
Llop P, Barbé S, López MM (2012) Functions and origin of plasmids in Erwinia species that are pathogenic to or epiphytically associated with pome fruit trees. Trees (Berl West) 26:31–46. https://doi.org/10.1007/s00468-011-0630-2
Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31:3210–3212. https://doi.org/10.1093/bioinformatics/btv351
Smits THM, Rezzonico F, Kamber T et al (2010) Complete genome sequence of the fire blight pathogen Erwinia amylovora CFBP 1430 and comparison to other Erwinia spp. Mol Plant-Microbe Interact 23:384–393. https://doi.org/10.1094/MPMI-23-4-0384
Thapa SP, Park DH, Kim WS et al (2013) Comparative genomics of Japanese Erwinia pyrifoliae strain Ejp617 with closely related erwinias. Genome 56:83–90. https://doi.org/10.1139/gen-2012-0094
Thompson DW, Casjens SR, Sharma R, Grose JH (2019) Genomic comparison of 60 completely sequenced bacteriophages that infect Erwinia and/or Pantoea bacteria. Virology 535:59–73. https://doi.org/10.1016/j.virol.2019.06.005
Wenneker M, Bergsma-Vlami M (2015) Erwinia pyrifoliae, a new pathogen on strawberry in the Netherlands. J Berry Res 5:17–22. https://doi.org/10.3233/JBR-140086
Acknowledgements
This work was supported by grants from the Agenda program (PJ015594) of the Rural Development Administration and the Korean Research Institute of Bioscience and Biotechnology Initiative Program. The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Nucleotide sequence accession number
This genome sequence has been deposited at DNA Data Bank of National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/) repository, under the BioProject PRJNA1044457 and BioSample SAMN38393563.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nie, H., Lee, S., Ko, SR. et al. Complete genome sequence of Erwinia pyrifoliae strain YKB12327, isolated from an apple tree in Korea. J Plant Pathol (2024). https://doi.org/10.1007/s42161-024-01708-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42161-024-01708-x