Abstract
The interaction and binding characterization of a sulfa drug, sulfonamide derivative sulfathiazole (Stz) with calf thymus DNA (ctDNA), was investigated using ethidium bromide (EB) as a fluorescence probe by fluorescence spectroscopy, UV–Vis spectroscopy and molecular docking method. The fluorescence quenching mechanism of ctDNA/EB by Stz was static quenching. The UV–Vis absorption spectra, competitive assays with Hoechst 33,258 and ionic strength effect studies showed that the binding of Stz with ctDNA may be minor groove binding mode on the A-T-rich regions of DNA helix. The calculated thermodynamic parameters, ΔH and ΔS, suggested that the driven forces in binding mechanism are hydrogen bonds or van der Waals forces. The molecular docking results were found to be in agreement with the experimental results and confirmed the existence of hydrogen bonds between Stz and DNA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Sulfonamides, the active substance of the sulfa drugs, are synthetic antibiotics having para amino benzene sulfonylamide structure (Rajendiran and Thulasidhasan 2015; Braschi et al. 2010). They have been used as bacteriostatic agents in human and animal diseases. These drugs with a large number are used in the form of oral and injection and are harmless compounds at appropriate doses. Most of them are absorbed in the digestive tract, including the stomach and small intestine. Although they leave the body with body fluids such as urine, bile and tears, the most important way is the kidneys. In the case of acidic urine, the solubility of sulfonamides decreases, and they can cause nephrotoxic effects by obstructing the urinary tract. The interaction of sulfoamides with biomolecules is very important to explain the side effects, benefits and their biodegradable in body. Different sulfonamide derivatives are obtained by connecting different functional groups to the sulfonamide structure. Thus, their physical, chemical, pharmacological and anti-infectious properties also become different (Refat et al. 2016; Muthu Prabhu et al. 2010). In this study, sulfathiazole (Stz) is chosen as an antimicrobial short-acting sulfonamide drug for DNA binding. The biomolecule–drug interaction studies describe changes in the biological structure and conformation using many spectroscopic methods. Fluorescence spectroscopy is most preferred in this field. The interaction of a fluorescent structure and a molecule can be monitored on fluorescence quenching method. DNA is the nucleic acid that carries the genetic information necessary for biological developments in living organisms. The interaction of DNA with small molecules (drug, dye, fluorescence probe, etc.) and the examination of these interactions, their determination using different methods, are very important for the design and development of new drugs (Shahabadi et al. 2017; Ling et al. 2008; Wu et al. 2008; Zhao et al. 2013a). The vast majority of cancer drugs interact with the DNA molecule. The structure and functions of DNA, protein–nucleic acid interaction and DNA–drug interactions are explained by the studies of binding of a molecule to DNA. The interaction of drugs with DNA occurs through three non-covalent binding types: (1) Intercalation binding occurs when fluorescent compounds with planar chromophores are placed between nucleic acid base pairs, (2) groove binding, and (3) electrostatic external binding between anionic sugar–phosphate backbone of DNA and positively charge species (Williams et al. 2012; Suh et al. 2001; Das and Kumar 2008; Geng et al. 2013). The studies on DNA and sulfa group drug interactions are limited in the literature. In this work, the assays to examine DNA binding properties with Stz showed that the interaction could not be directly observed. Because fluorescence of DNA is very weak, also Stz has no fluorescence. So, ethidium bromide (EB), a well-known intercalator agent, was used for DNA as fluorescence probe (Zhang et al. 2012; Gao et al. 2015) which successfully carried out to monitoring the binding of DNA with Stz. The fluorescence change of DNA–EB system in the presence of Stz suggests that there is strong interaction between DNA and Stz. Here the binding properties of Stz with ctDNA was detailed presented by UV absorption, fluorescence quenching methods and computational results.
2 Materials and Method
2.1 Reagents
All reagents used in experimental studies were of analytical purity and were used without further purification. The working solutions were prepared daily by diluting the stock solutions in Tris–HCl buffer solution at pH 7.4. A sodium salt of calf thymus deoxyribonucleic acid (ctDNA, Type I, double-stranded, Sigma) was used to prepare the stock solution in Tris–HCl buffer solution. Absorption values of the solution at 260 nm (ε: 6600 M−1 cm−1) were used to get the concentration of the solution. The purity of the DNA was checked by monitoring the absorption ratio at 260/280 nm. The ratio was obtained as greater than 1.8 which showed that DNA was enough free from protein. Sulfathiazole (4-amino-N-(1,3-thiazol-2-yl) benzenesulfonamide, Sigma) was dissolved in DMSO and prepared as 5.0 mM stock solution. The stock solutions of 5.0 mM Hoechst 33,258 (Sigma) and 3.0 mM EB (Sigma) were prepared by dissolving their solid forms in double-distilled water. For effect of ionic strength assays, Na+ ion solutions were diluted from 1.0 M NaCl.
2.2 Apparatus
The steady-state fluorescence measurements were taken on a spectrofluorimeter (Hitachi F-4500 with FL Solutions2.0 software, Japan) using 150 W xenon light source, 2.5 nm/2.5 nm of excitation/emission slits with PMT voltage 700 V. Fluorescent titration experiments were carried out in 1.0 cm quartz cells with a final volume of 3 mL of the working solution. UV absorption spectra were recorded using UV–visible spectrophotometer (Shimadzu UV-1700 PharmaSpec with UVProbe software, Japan) in the range at 200 and 350 nm. pH measurements were obtained from a Mettler Toledo (FiveEasy Plus) digital pH meter.
2.3 Fluorescence Studies
Ethidium bromide (EB) was used as fluorescent probe to investigate of interaction between Stz and ctDNA. Therefore first, the fluorescence data were obtained adding various concentration of DNA into 2.5 mL fixed concentration (3.0 μM) of EB solution at λex = 525 nm for ctDNA–EB binding assays. Then, to find the interaction of Stz with the ctDNA–EB system, various concentrations of Stz were added to the fixed ratio ctDNA–EB at λex = 286 nm. The effect quenching of Stz on the ctDNA–EB emission was examined using Stern–Volmer equation at different temperatures. To examine the effect of ionic strength on the binding mechanism, the fluorescence spectra of a fixed ctDNA–EB–Stz system were recorded in the presence of varying amount of NaCl. The competition experiments for DNA were carried out with Hoechst 33,258 as groove binder. The increasing concentrations of Stz solutions were added on the fixed amount of ctDNA–Hoechst 33,258 (10–5.0 μM) mixture at λex/λem = 343/470 nm at room temperature.
2.4 Absorption Studies
UV absorption spectra for ctDNA–Stz interactions were recorded by fixed amount of DNA (25 μM) while varying the drug concentration from 2.0 μM to 10 μM (ri = [complex/[DNA] = 0.08–0.4).
2.5 Computational Studies
The interaction mechanisms and possible orientations of Stz as ligand with in the target model DNA (ID: 1BNA, B form structure from Protein Data Bank) were evaluated by Discovery Studio (DS) (2018 software, Dassault Systemes Biovia) (Dassault Systemes Biovia 2016; Bernstein et al. 1977) The energy and geometry of ligand Stz were optimized by Gaussian 09 (G09) at DFT/B3LYP/6-31G* level (Frisch et al. 2009). Then it was used molecular electrostatic potential surface (MEPS) map for examining of electrophilic and nucleofilic sites of Stz compound. DNA as the target was minimized using the CHARMm force field based on the Newton–Raphson (ABNR) (Chattaraj et al. 2011) basis adopted to find the target for molecular docking target. Analyse Ligand Poses subprotocol in DS 2018 was applied to dock Stz into the double-stranded DNA binding sites. The best pose was described based on the lowest energy values for the Stz–DNA complex using the ten possible conformations of Stz at ligand position. Molecular docking results were obtained as root mean square deviation (RMSD) and energy value such as binding, CDOCKER and CDOCKER interaction.
3 Results and Discussion
3.1 Fluorescence Measurements
3.1.1 Determine of the Probe Concentration
EB is used as the most known DNA probe in the research of drug–DNA interaction. EB forms a strong complex with DNA by intercalation between two adjacent base pairs in double helix. It displays large enhancement in fluorescence upon binding to DNA (Qiao et al. 2008). In our experiments by titration method, to obtain optimal ratio of DNA/EB, various concentrations of ctDNA solution were added to constant concentration (3.0 μM) of EB at λex = 525 nm. According to the emission spectra shown in Fig. 1, both DNA and EB have low intrinsic fluorescence, but EB emission increases significantly in the presence of DNA. As shown in Fig. 2, for 3.0 μM EB at constant concentration, when 29.7 μM ctDNA was added, then the intensity of system was not enhanced any more. It shows the EB–DNA binding near upon saturation. Consequently, 3.0 μM EB and 29.7 μM ctDNA were selected as fixed system (ri = [DNA]/[EB] = 9.9) for interaction between Stz and ctDNA.
The fluorescence spectra were obtained with the addition at various amounts of Stz to containing constant concentrations of ctDNA–EB mixture. As shown in Fig. 3a, the intensity of ctDNA–EB fluorescence decreases with the increase in the Stz amount. Here, the quenching in this emission intensity of the system can be explained three possible reasons. Firstly, Stz competes to bind the regions where EB is bound to the DNA. The exclusion of the intercalated EB from duplex DNA decreases the fluorescence intensity of system. Secondly, the binding between Stz and EB may lead to the decrease in the DNA–EB emission. Thirdly, Stz could be bound to the DNA-EB system to create a formation of new complex structure that has no fluorescence and this may decrease of the emission of the system. When these reasons are examined, it is clear that there is no common intercalation binding between Stz and DNA. The binding constant of ctDNA with Stz was obtained as 3.24 × 105 M−1 while the binding constant (KEB) of EB–DNA was 1.25 × 106 M−1 (Zhao et al. 2013b). This points that Stz no enter the competition with EB because of low binding constant of ctDNA–Stz. Therefore, the first reason can be eliminated. Also, the second reason is not true because our experiments showed that Stz did not react with EB. The last reason is more suitable than the others, i.e., Stz is bound to ctDNA–EB to reduce the emission of system. The fluorescence quenching data were examined with Stern–Volmer equation as follows (Lakowicz 2006):
F0 and F are fluorescence intensities without and with quencher of ctDNA–EB system. [Q] is the quencher (Stz) concentration, and Ksv is quenching constant. The linear Stern–Volmer curves were obtained from titration data at three temperatures (Fig. 3b). Ksv constants were found from the slopes of graphs, in the range of 5.01–4.03 × 104 M−1, and they are given in Table 1. The quenching mechanisms are generally two categories: static or dynamic. They should be discriminated with their excited-state lifetimes and differing dependence on temperatures. The results suggest that only one type of quenching mechanism or interaction mode occurs. The type of quenching mechanism could be recognized in the following equation (Lakowicz 2006),
kq is the rate of quenching constant of biomolecule, and τ0 is the average lifetime of biomolecule in the absence quencher. Here τ0 value was counted as 10–8 s for DNA (Lakowicz and Weber 1973). kq values were calculated in the range of 5.01–4.03 × 1012 M−1 s−1, and the quenching constants are given in Table 1 at three temperature. The limit value of kq for interaction of biomolecule with small molecule in dynamic quenching process is 2 × 1010 M−1 s−1 (Lakowicz and Weber 1973). The kq data are much bigger than rate constant of maximum scatter collision—quenching indicates the possible mechanism is static quenching which indicates that a complex formation can be expressed. The decrease of Ksv results with increasing temperature is another evidence for static quenching. The binding site number (n) and binding constant (K) were obtained from log–log graphs plotted using Eq. (3) (Feng et al. 1998):
According to the results given in Table 1, it reveals that binding constant values decrease at high temperature, and it points to the partially fragmented the ctDNA–EB–Stz complex. n values are approximately 1, indicating that Stz has only one site binding with DNA–EB system.
3.1.2 Thermodynamic Studies and Binding Mode of Drug–DNA
The fluorescence quenching studies allow the explanation of binding mode in the biomolecule–small molecule interaction with the thermodynamic parameters (ΔH, ΔS and ΔG) obtained depending on the temperature (Ross and Subramanian 1981). The hydrogen bonding, electrostatic, hydrophobic and van der Waals forces are non-covalent interactions which are determined based on signs and magnitudes of ΔS and ΔH. Here, they were calculated using van’t Hoff equation given below:
K values are binding constants found in Eq. (3) for three temperatures. ΔG parameter is obtained using in the below equation:
van’t Hoff equation was obtained from graph of log K versus 1/T as log K = − 18.926 + 7276.8 1/T (R2 = 0.997) for ctDNA–EB–Stz system (Fig. 4). ΔS value was found from intercept and ΔH from slope of the graph. These parameters are given in Table 2. The negative sign of ΔH and ΔG indicates that the binding of ctDNA–EB and Stz is exothermic and spontaneous process, respectively. The negative sign of both ΔH (− 139.3 kJ/mol) and ΔS (− 362.4 J/molK) shows that van der Waals forces or hydrogen bonds play active roles in the bonding mechanism.
3.1.3 Effect of Ionic Strength
NaCl, a strong electrolyte, is used to investigate the effect of ionic strength on binding in drug–DNA interaction studies. The anionic phosphate groups on the DNA helix interact electrostatically with Na+ ions. If drug molecule is bound with groove binding to DNA, for binding mechanism, the narrower and deeper grove will be favorable; then, fluorescence intensity of system will decrease as the concentration of Na+ ions increases. If drug molecule is an intercalator between DNA base pairs, it is not affected by environmental changes because small molecule protects of DNA base pairs above and below (Bi et al. 2019). The various concentrations of (0–0.5 M) NaCl were added on the system with a constant amount of DNA–EB–Stz and the emission of the system decreased with the increase in the NaCl amount (Fig. 5). This situation points that there is a more efficient interaction of drug with ctDNA–EB system and also possible binding mode should be groove binding.
3.1.4 Competitive Binding Between Drug and Hoechst 33,258
The groove binding is related to the two types of grooves in DNA helix: major and minor binding. Hoechst 33,258 is well known as minor groove binder for DNA using fluorescence probe for competitive displacement experiments to determine the binding type of small molecule to DNA. It binds strongly to DNA helix with specifically for A-T-rich sequences (Rehman et al. 2015; Banerjee and Pal 2006). Our experiments showed that Stz had no effect on Hoechst 33,258 emission and thus no chemical interaction between the two species. Hoechst 33,258 fluorescence intensity was enhanced with DNA addition at 470 nm when excited at 343 nm. The fluorescence intensity of the ctDNA/Hoechst 33,258 system was found to decrease remarkably with increasing amounts of Stz (Fig. 6). These experiments showed that Stz competes with Hoechst 33,258 for the binding sites of ctDNA, further supporting that Stz may act as a minor groove binder.
3.2 Absorption Spectra of Drug–DNA
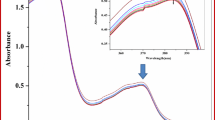
UV absorption spectroscopy is a widely used method to investigate the studies about the interaction of DNA with small molecules. The absorption spectra of DNA show some shifts due to the interaction of DNA with drugs, resulting from hypochromic and hyperchromic effects including hydrogen bond, electrostatic, stacking and hydrophobic interactions. Therefore, the change in the conformation and structure of the DNA causes the spectral behavior of the DNA to change (Ramana et al. 2016; Bi et al. 2008). Absorption spectra were obtained by adding various amounts of Stz solutions on ctDNA of 25 μM ctDNA at constant concentration. As shown in the spectra in Fig. 7, the absorption peak of DNA increased at 260 nm with the addition of drug amount and no change or shift in the peak position was observed. It reveals that a change of secondary structure with hyperchromic effect. The data indicated that Stz does not bind to DNA intercalation. Therefore, groove binding might be more acceptable.
3.3 Computational Studies
3.3.1 MEPS Map of Stz Compound
The electrophilic and nucleophilic sites are figured out with MEPS plot is useful in chemical property relationship in the molecular interaction studies. After Stz compound optimized, its MEPS map was examined by using G09 software program. As shown in Fig. 8, the neutral, negative and positive charged regions of the compound are depicted in some colors on the map (Mulla et al. 2023; Gayathri and Arivazhagan 2011). According to the color analysis, the red color on Stz shows negative regions for electrophilic attacks such as sulfonyl oxygen atoms and nitrogen atom of amide group. The blue color shows positive regions for nucleophilic attacks such as nitrogen atom of amine moiety. And also, the green color indicates neutral regions of compound surface.
3.3.2 Molecular Docking Results
In the first part of the study, the interaction mechanism with DNA was tried to be explained by using EB as a reference ligand. The docking data indicate that electrostatic attractions are most influential than other interaction types in ctDNA/EB system. EB compound has six electrostatic attractions with, B:DT20, B:DT19 and A:DG10 nucleotides, two hydrogen bonds with A:DC9 and two hydrophobic interactions with A:DC9, B:DT19 nucleotides of ctDNA (seen in Fig. 9). The real aim of this study is: Molecular docking was implemented to probe the relative orientation and affinity of Stz against to ctDNA. Figure 9 shows the best pose of the ligand, Stz and superimposed form with EB in active site of ctDNA. It is known from the literature that EB exhibits intercalation modes with ctDNA by using non-bonding interactions. This was supported by implemented with the help of molecular docking. On the other side, it displayed that Stz did groove binding mode with DNA in this study. The generated ctDNA–Stz complex has four hydrogen bonds with active nucleotides such as B:DT19, A:DT7, A:DC9 and A:DG10 in DNA. Table 3 gives the non-binding interaction kinds and distances of the compound Stz with DNA. The data revealed that the hydrogen bonds were important parameters in the binding process, which was coherent with the experiment results. Table 4 presents docking energy values of Stz–DNA which agree with the experimental section having thermodynamic results.
4 Conclusion
The binding properties of sulfa drug sulfathiazole (Stz) with calf thymus DNA (ctDNA) were investigated using ethidium bromide (EB) as a fluorescence probe by fluorescence spectroscopy and UV–Vis spectroscopy. Also molecular docking calculations are verified the results of spectral methods which are indicated that the binding form between Stz and ctDNA may be groove binding. The negative values of thermodynamic parameters (ΔH < 0 and ΔS < 0) suggest that ctDNA could bind to Stz mainly through hydrogen bonds and van der Waals forces. Furthermore, the calculation of binding energy (− 6.650 kcal/mol) and docking interactions with ctDNA CDOCKER in DS 2018 exhibited that the hydrogen bond was effective parameters in the binding process, which was correlative with the other results in this study. The study may provide a good model to explain binding properties of sulfonamide like compounds to DNA. In addition, studying the binding mechanism of target sulfonamides with DNA is one of the key steps in their biomolecular interaction and designing new sulfa drugs.
Data Availability
Not applicable.
References
Banerjee D, Pal SK (2006) Ultrafast charge transfer and solvation of DNA minor groove binder: Hoechst 33258 in restricted environments. Chem Phys Lett 432:257–262
Bernstein FC, Koetzle TF, Williams GJ, Meyer EE, Brice MD, Rodgers JR, Kennard O, Shimanouchi T, Tasumi M (1977) The protein data bank: a computer-based archival file for macromolecular structures. J Mol Biol 112:535–542
Bi S, Zhang H, Qiao C, Sun Y, Liu C (2008) Studies of interaction of emodin and DNA in the presence of ethidium bromide by spectroscopic method. Spectrochim Acta Part A 69:123–129
Bi S, Zhou H, Wu J, Sun X (2019) Micronomicin/tobramycin binding with DNA: fluorescence studies using of ethidium bromide as a probe and molecular docking analysis. J Biomol Struct Dyn 37:1464–1476
Braschi I, Blasioli S, Gigli L, Gessa CE, Alberti A, Martucci A (2010) Removal of sulfonamide antibiotics from water: evidence of adsorption into an organophilic zeolite Y by its structural modifications. J Hazard Mater 178:218–225
Chattaraj PK, Giri S, Duley S (2011) Update 2 of: electrophilicity Index. Chem Rev 111:43–75
Das S, Kumar GS (2008) Molecular aspects on the interaction of phenosafranine to deoxyribonucleic acid: model for intercalative drug-DNA binding. J Mol Struct 872:56–63
Dassault Systemes Biovia (2016) Discovery studio modeling environment release 2017. Dassault Systemes, San Diego
Feng X-Z, Lin Z, Yang L-J, Whang C, Bai C-L (1998) Investigation of the interaction between acridine orange and bovine serum albümin. Talanta 47:1223–1229
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA et al (2009) Gaussian09 RevisionE.01. Gaussian, Inc., Wallingford
Gao Y, Li J, Huang G, Yan L, Dong Z (2015) Spectroscopic studies on the interaction between anthragallol and DNA using of ethidium bromide as a fluorescence probe. Spectrochim Acta Part A 141:239–243
Gayathri R, Arivazhagan M (2011) Vibrational spectroscopy investigation and HOMO, LUMO analysis using DFT (B3LYP) on the structure of 1,3-dichloro 5-nitrobenzene. Spectrochim Acta A 81:242–250
Geng S, Liu G, Li W, Cui F (2013) Molecular interaction of ctDNA and HSA with sulfadiazine sodium by multispectroscopic methods and molecular modeling. Luminescence 28:785–792
Lakowicz JR (2006) Principles of fluorescence spectroscopy. Plenum, New York
Lakowicz JR, Weber G (1973) Quenching of fluorescence by oxygen. A probe for structural fluctuations in macromolecules. Biochemistry 12:4161–4170
Ling X, Zhong W, Huang Q, Ni K (2008) Spectroscopic studies on the interaction of pazufloxacin with calf thymus DNA. J Photochem Photobiol B Biol 93:172–176
Mulla BBA, Nesagari AR, Pasha KM, Pusar MS, Kamble RR, Sidarai AH (2023) Experimental and theoretical spectroscopic investigation on coumarin based derivatives for non-linear optoelectronics application. J Fluoresc 33:161–175
Muthu Prabhu AA, Venkatesh G, Rajendiran N (2010) Spectral characteristics of sulfa drugs: effect of solvents, pH and β-cyclodextrin. J Solut Chem 39:1061–1086
Qiao C, Bi S, Sun Y, Song D, Zhang H, Zhou W (2008) Study of interactions of anthraquinones with DNA using ethidium bromide as a fluorescence probe. Spectrochim Acta Part A 70:136–143
Rajendiran N, Thulasidhasan J (2015) Interaction of sulfanilamide and sulfamethoxazole with bovine serum albumin and adenine: spectroscopic and molecular docking investigations. Spectrochim Acta Part A 144:183–191
Ramana MMV, Betkar R, Nimkar A, Ranade P, Mundhe B, Pardeshi S (2016) Synthesis of a novel 4H-pyran analog as minor groove binder to DNA using ethidium bromide as fluorescence probe. Spectrochim Acta Part A 152:165–171
Refat MS, Sharshar T, Elsabawy KM, El-Sayed MY, Adam AMA (2016) Synthesis, physicochemical characterization and anticancer screening of sulfa drug ruthenium complexes as anticancer agent. J Mol Liq 222:334–349
Rehman SU, Sarwar T, Ishgi HM, Husain MA, Hasan Z, Tabish M (2015) Deciphering the interactions between chlorambucil and calf thymus DNA: a multi-spectroscopic and molecular docking study. Arch Biochem Biophys 566:7–14
Ross PD, Subramanian S (1981) Thermodynamics of protein association reactions: forces contributing to stability. Biochemistry 20:3096–3102
Shahabadi N, Falsafi M, Maghsudi M (2017) DNA-binding study of anticancer drug cytarabine by spectroscopic and molecular docking techniques. Nucleosides Nucleotides Nucleic Acids 36:49–65
Suh D, Oh Y-K, Chaires JB (2001) Determining the binding mode of DNA sequence specific compounds. Process Biochem 37:521–525
Williams AK, Dasilva SC, Bhatta A, Rawal B, Liu M, Korobkova EA (2012) Determination of the drug-DNA binding modes using fluorescence-based assays. Anal Biochem 422:66–77
Wu M, Wu W, Gao X, Lin X, Xie Z (2008) Synthesis of a novel fluorescent probe based on acridine skeleton used for sensitive determination of DNA. Talanta 75:995–1001
Zhang G, Hu X, Fu P (2012) Spectroscopic studies on the interaction between carbaryl and calf thymus DNA with the use of ethidium bromide as a fluorescence probe. J Photochem Photobiol B Biol 108:53–61
Zhao J, Li W, Ma R, Chen S, Ren S, Jiang T (2013a) Design, synthesis and DNA interaction study of new potential DNA bis-intercalators based on glucuronic acid. Int J Mol Sci 14:16851–16865
Zhao X-L, Li Z-S, Zhang Z-B, Zhang A-G, Whang K-Z (2013b) pH luminescence switch, DNA binding and photocleavage, and cytotoxicity of a dinuclear ruthenium complex. Dalton Trans 42:5764–5777
Funding
The study was supported by the Hacettepe University Scientific Research Fund (Project no: FBA2017-12854).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gökoğlu, E., Kıpçak, F. & Taskin Tok, T. Deciphering of Biophysical Interaction of Calf Thymus DNA with a Sulfa Drug: Spectroscopic and Molecular Docking Study. Iran J Sci 47, 1451–1459 (2023). https://doi.org/10.1007/s40995-023-01520-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40995-023-01520-9