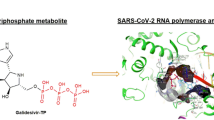
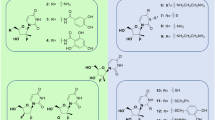
Abstract
The treatment of COVID-19 disease has been one of the most critical essential concerns of researchers in recent years. One of the most exciting and potential therapeutic targets for SARS-CoV-2 therapy progression is RNA-dependent RNA polymerase (RdRP), a viral enzyme for viral RNA replication throughout host cells. According to some research, Remdesivir suppresses RdRp. The nucleoside medication remdesivir has been authorized under an Emergency Use Authorization to treat COVID-19. Given the role of this enzyme in virus replication, our scientific question is whether Remdesivir is the most appropriate antiviral drug to inhibit this enzyme or not. Accordingly, this study aimed to repurpose antiviral drugs to inhibition of RdRp using virtual screening and Molecular Dynamics simulation methods. Five FDA-approved antiviral medications, including Elbasvir, Glecaprevir, Ledipasvir, Paritaprevir, and Simeprevir, had good interaction potential with RdRp. Also, the results show that the number of H-bonds and contacts and ∆G interactions between the protein and ligand in the Remdesivir complex is less than those of other complexes. According to the given data which shows the tendency of binding with RdRp for Paritaprevir, Simeprevir, Glecaprevir, and Ledipasvir and Elbasvir is more than Remdesivir and due to the fact that these five drugs have a high tendency to bind to other targets in the SARS-CoV-2, the use of Remdesivir as an antiviral drug in the treatment of COVID-19 should be considered more sensitively.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
COVID-19, as a respiratory illness induced by the severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), has quickly become a worldwide health concern. At the moment, the Remdesivir has been authorized to manage COVID-19 under an Emergency Use Authorization (Sharma et al. 2020). An attractive and potential therapeutic target for SARS-CoV-2 improvement is the RNA-dependent polymerase (RdRP), a viral enzyme that aids in viral RNA replication in host cells. Since RdRP is an enzyme with no host cell homologs, the inhibitors of this enzyme are safer and more effective therapeutics for the treatment and control of COVID-19 disease (V’kovski et al. 2021). RNA viruses are grouped with ssRNA (single-stranded RNA) and dsRNA (double-stranded RNA). ssRNA viruses can be either positive sense or negative sense. Coronaviruses are positive-sense RNA viruses containing an ssRNA genome that needs RdRP function during viral genome replication (Zhu et al. 2020). The viral genome encodes 29 proteins, including16 nonstructural proteins (nsps), 4 structural proteins, and 9 accessory proteins. The main structural proteins include envelope (E), spike (S), membrane (M), and nucleocapsid (N), occupying ∼33% of the virus genome (Zhu et al. 2020; Gorkhali et al. 2021). RdRp nsp12 is the main enzyme of virus replication. With the help of additional cofactors, such as nsp7 and nsp8, the nsp12 polymerase function is significantly increased, whereas the activity of nsp12 alone is restricted or non-existent.
To summarize, nsp12-nsp7-nsp8 are the essential components for viral RNA replication (Peng et al. 2020). The C-terminal catalytic domain of nsp12 (residues L366 to F920) connects to the nidovirus-specific extension domain (NiRAN, residues S115 to A250) through an interface subdomain (residues L251 to R365). The nsp7-nsp8 heterodimer binds above the subdomain. Nsp7 contributes to the binding of the heterodimer to nsp12. At the same time, nsp8 only contacts a few residues from nsp12 (Jiang et al. 2021). Given the role of this enzyme in virus replication, our scientific question is whether Remdesivir is the most appropriate antiviral drug to inhibit this enzyme. Accordingly, this study aimed to repurpose antiviral drugs to inhibition of RdRp using in silico methods.
2 Material and Methods
RdRp (PDB code: 7bv2) coordinates were obtained by using the Protein Data Bank (https://www.rcsb.org) (Yin et al. 2020). Water molecules were removed from proteins, and hydrogen atoms were introduced to better-functioning hydrogen bonds as a starting stage. The GROMACS 5.1.4 package (Spoel et al. 2005) was employed to reduce energy consumption during protein optimization. The Drug Bank database's small-molecule portions (https://www.drugbank.ca/) were utilized to acquire all antiviral medicines (91 compounds) (Wishart et al. 2018). The CB-Dock server was used to design the docking cell (Liu et al. 2020). Regarding ligand preparation, the Pyrx tool was used for energy minimization, and ligands converted into PDBQT format. Also, the Pyrx tool (AutoDock Vina) was used for virtual screening (Dallakyan and Olson 2015; Trott and Olson 2010).
2.1 Molecular Dynamics Simulation
Once all the FDA-approved drugs were docked with RdRp, medications with the desirable delta G (∆G < − 9 kcal/mol) with RdRp were chosen for additional analysis. At this point, we wanted to find drugs that could better attach to the protein. The free energy of binding, the number of hydrogen bonds, and the number of contacts between drugs and RdRp were calculated using molecular dynamics (MD) simulations.
2.1.1 MD Systems Setup
The software GROMACS 5.1.4 was used for MD simulation throughout this study (Berendsen and Spoel 1995). We also used the gromos 54a7 force field (Schmid et al. 2011). All modeling packages were handled with the necessary quantities of chloride ions and sodium to neutralize the platform. The Periodic Boundary Condition (PBC) was used along each box's axial direction in each modeling framework, and the SP3 water simulation was used for systemic solubilization (Hess 2008). All covalent bonds were restricted using the LINCS algorithms. The modelings were brought on by a short-range electrostatic contact and a van der Waals distance cutoff of 1.2 nm (Darden et al. 1993). The long-range electrostatic connection was computed using the Particle Mesh Ewald (PME) method. Energy reduction for all systems was achieved using the sharpest descending method, followed by equilibration across all systems using the NVT ensemble. Following that, the NPT ensemble gradually guided each system's balance, preserving the Nose–Hoover algorithm temperature (Hoover 1985; Nose 1984) and maintaining the temperature at 310 K. The Parrinello-Rahman barostat (Parrinello and Rahman 1981) kept the pressures constant at 1 bar throughout the NPT equilibration. The MD simulation of the complexes was finished after 100 ns.
2.1.2 Calculation
Gromacs utilities analyzed and assessed each trajectory's outcomes after running MD calculations. Binding free energy was evaluated with a calculation of nonpolar and polar interactions between RdRp and drugs. The g_mmpbsa instrument computed the binding free energy by using the MM-PBSA technique (Kumari et al. 2014).
The addition of the nonpolar association free-energy (∆G nonpolar) and the polar association free-energy (∆G polar) resulted in the overall quantity of bonding free-energy (∆G), as seen below:
where \(\Delta G_{{{\text{elec}}}}\), \(\Delta G_{{{\text{ps}}}}\), \(\Delta G_{{{\text{vdW}}}}\), \(\Delta G_{{{\text{nps}}}}\) are the electrostatic energy, polar solubilization energy, van der Waals energy, and nonpolar solubilization energy.
3 Results
In the current research, we used the drug repurposing approach based on virtual screening and MD simulation to inhibit RdRp.
3.1 Results of Virtual Screening
Five FDA-approved antiviral medications, including Elbasvir, Glecaprevir, Ledipasvir, Paritaprevir, and Simeprevir, had good interaction potential with RdRp (∆G < − 9 kcal/mol) as a consequence of virtual screening, according to the findings. As a result, these five medications were chosen for additional study and comparison with remdesivir utilizing MD simulation. The BIOVIA Discovery Studio Visualizer was used to display 2D and 3D images of protein/ligand interactions (Figs. 1, 2, 3,4, 5 and 6).
3.2 MD Simulation
3.2.1 Root-Mean-Square Deviation (RMSD) and Root Mean Square Fluctuation (RMSF) Evaluation
The complexes' adaptability and durability were assessed using RMSD and RMSF. Six complexes were examined to see how stable they were in terms of conformation using C-alpha RMSD analyses. Figure 7 depicts the complexes' RMSD. It was possible to see how flexible the protein arrangement was by looking at the C-alpha RMSF of each residue throughout the complex. The high RMSF value indicates more flexibility, whereas the low RMSF value indicates limited motion. The RMSF graphs of protein for six complexes are shown in Fig. 8. The RMSF analysis revealed that the protein in the Elbasvir complex is more flexible than other complexes during the trajectory.
3.2.2 The Number of H-Bonds, the Number of Contacts, and ∆G Interactions
The number of H-bonds, contacts and free energy of interaction between RdRp and ligands have essential roles in stabilizing the complexes. The total number of H-bonds and connections in six ligands and RdRp versus time at 310 K are shown in Figs. 9 and 10. The number of H-bonds and contacts between RdRp and ligand in the Elbasvir complex is higher than the other complexes. Calculation of ∆G for the polar and nonpolar interactions between RdRp and six ligands was shown in Table 1. The binding free energy of Paritaprevir and Elbasvir complexes are more favorable than the other complexes.
4 Discussion
The viral RdRp is required for the replication of SARS-CoV-2. Remdesivir is the FDA-approved drug for the treatment of COVID-19 patients. Some studies have shown that Remdesivir inhibits RdRp of coronaviruses, including SARS-CoV-2 (Kokic et al. 2021). Seven conserved catalytic motifs A-G make up the active site of the SARS-CoV-2 RdRp. The palm subdomain contains five of these motifs (A-E), whereas the finger subdomain has the remaining two (F and G) (Jiang et al. 2021). There is a conserved aspartic acid (D618) in the catalytic motif A (residues T611 to M626) of the majority of viral polymerases, such as the hepatitis C virus and poliovirus (Jiang et al. 2021). Glecaprevir (Fig. 2), Ledipasvir (Fig. 3), Paritaprevir (Fig. 4), Simeprevir (Fig. 6) and Remdesivir (Fig. 5) interact with LYS621 in Motif A. Also, Remdesivir interacts with TYR 619 in Motif A (Fig. 5).
A hinge for the conformational organization associated with template RNA and substrate attachment is found in Motif B's flexible loop (G678–T711 residues). Ledipasvir interacts with N 691 and T680 in Motif B. Also, Simeprevir interacts with N 691 in Motif B (Fig. 6). For the metal ion to be bound, motif C (residues F753 to N767) has the catalytic motif in residues S759 to D761. In the Ledipasvir /RdRp complex, the ligand establishes hydrogen bonds with S 759 in Motif C (Fig. 3). The recent structures of RdRp also confirm that the D760 and D761 coordinate two magnesium ions at the catalytic center. Glecaprevir and Simeprevir interact with D760 of RdRp (Figs. 2 and 6). The phosphate group of NTP interacts with Motif F (residues L544 to V557). In the Elbasvir/RdRp complex, the ligand establishes hydrogen bonds with K545 and S549 in the F motif (Fig. 1).
The RNA template may be directed to the active catalytic site by motif G (residues D499 to L514), which interacts with the template strand. Glecaprevir (Fig. 2), Simeprevir (Fig. 6), and Ledipasvir (Fig. 3) establish a hydrogen bond with the enzyme at position R555. Also. Ledipasvir and Paritaprevir interact with the enzyme at position K551 (Figs. 3 and 4).
The results of our study show that among the five drugs Paritaprevir, Simeprevir, Glecaprevir, Ledipasvir and Elbasvir, Elbasvir with the highest number of contacts, the highest hydrogen bond and optimal binding energy can establish a more stable complex with RDRP. Also, the results show that the number of H-bonds and the number of contacts between the protein and ligand in the Remdesivir complex are less than in other complexes. Therefore, it seems that the Remdesivir binding tendency to the RdRp is less than the other complexes.
Some of the antiviral drugs studied in this manuscript bind to other targets in addition to RdRp. For example, based on target-based computational drug screening, it was shown that elbasvir to have a very high affinity for essential SARS-CoV-2 proteins, including RNA-dependent RNA polymerase, helicase, papain-like proteinase, and the viral S protein (Balasubramaniam and Reis 2020). Another study showed that elbasvir had promising activity in the low micromolar range against RdRp (Milani et al. 2021). Also, using computational methods, it was demonstrated that Glecaprevir is the best inhibitor of SARS-CoV-2 main protease. Glecaprevir binds to the substrate-binding pocket of SARS-CoV-2 main protease and forms a significant number of interactions (Shamsi et al. 2020). We have shown in previous studies that ledipasvir has a high affinity for the ACE2 (ACE2 acts as a cellular doorway—a receptor—for the virus that causes COVID-19.) (Mahdian et al. 2021). The 2020 study found that Paritaprevir, simeprevir, Glecaprevir, and Ledipasvir had good binding energy with 3-chymotrypsin-like protease (3CLpro), Papain-Like protease (PLpro), cleavage site, HR1, and RBD in Spike protein. Also, Paritaprevir, simeprevir, Glecaprevir, and Ledipasvir had good binding energy with ACE2 and TMPRSS2. ACE2 and TMPRSS2 activate the viral S protein, which facilitates virus-cell membrane fusion (Mahdian et al. 2020). A new study aimed at finding hub genes in the development of coronavirus disease shows that colony-stimulating factor (CSF3) is a potential drug Target for the treatment of COVID-19. They using computational methods showed that Elbasvir, Ritonavir have the ability to bind favorably with CSF3 and these two antiviral drugs significantly inhibited CSF3 protein expression (Fang et al. 2021). Another study was conducted with the aim of repurposing the FDA drugs and targeting the SARS-CoV-2 nonstructural protein 15 (Nsp15). In this study, Paritaprevir and Elbasvir, both currently approved for the treatment of hepatitis C, were shown to have favorable binding energies with NSP15 (Sixto-López and Martínez-Archundia 2021).
5 Conclusion
According to the above data in this study, which shows the tendency of binding with RdRp for Paritaprevir, Simeprevir, Glecaprevir, and Ledipasvir and Elbasvir is more than Remdesivir and due to the fact that these five drugs have a high tendency to bind to other targets in the SARS-CoV-2, the use of Remdesivir as an antiviral drug in the treatment of COVID-19 should be considered more sensitively.
References
Balasubramaniam M, Reis RJS (2020) Computational target-based drug repurposing of elbasvir, an antiviral drug predicted to bind multiple SARS-CoV-2 proteins. ChemRxiv. https://doi.org/10.26434/chemrxiv.12084822
Berendsen HJC, van der Spoel D (1995) Gromacs: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 1995(91):43–56. https://doi.org/10.1016/0010-4655(95)00042-E
Dallakyan S, Olson AJ (2015) Small-molecule library screening by docking with PyRx. In: Chemical biology. Humana Press, New York, NY, pp 243–250
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N⋅ log (N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092. https://doi.org/10.1063/1.464397
Fang C, Mei J, Tian H, Liou YL, Rong D, Zhang W, Liao Q, Wu N (2021) CSF3 is a potential drug target for the treatment of COVID-19. Front Physiol 11:605792. https://doi.org/10.3389/fphys.2020.605792
Gorkhali R, Koirala P, Rijal S, Mainali A, Baral A, Bhattarai HK (2021) Structure and function of major SARS-CoV-2 and SARS-CoV proteins. Bioinform Biol Insights 15:11779322211025876
Hess B (2008) P-Lincs: a parallel linear constraint solver for molecular simulation. J Chem Theory Comput 4:116–122. https://doi.org/10.1021/ct700200b
Hoover WG (1985) Canonical dynamics: equilibrium phase-space distributions. Phys Rev A 31(3):1695–1697. https://doi.org/10.1103/physreva.31.1695
Jiang Y, Yin W, Xu HE (2021) RNA-dependent RNA polymerase: structure, mechanism, and drug discovery for COVID-19. Biochem Biophys Res Commun 538:47–53. https://doi.org/10.1016/j.bbrc.2020.08.116 (Epub 2020 Sep 4)
Kokic G, Hillen HS, Tegunov D, Dienemann C, Seitz F, Schmitzova J, Cramer P (2021) Mechanism of SARS-CoV-2 polymerase stalling by remdesivir. Nat Commun 12(1):1–7. https://doi.org/10.1038/s41467-020-20542-0
Kumari R, Kumar R, Lynn A et al (2014) g_mmpbsa A GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54:1951–1962. https://doi.org/10.1021/ci500020m
Liu Y, Grimm M, Dai WT, Hou MC, Xiao ZX, Cao Y (2020) CB-Dock: a web server for cavity detection-guided protein–ligand blind docking. Acta Pharmacol Sin 41(1):138–144. https://doi.org/10.1038/s41401-019-0228-6 (Epub 2019 Jul 1)
Mahdian S, Ebrahim-Habibi A, Zarrabi M (2020) Drug repurposing using computational methods to identify therapeutic options for COVID-19. J Diabetes Metab Disord 19:691–699. https://doi.org/10.1007/s40200-020-00546-9
Mahdian S, Zarrabi M, Panahi Y, Dabbagh S (2021) Repurposing FDA-approved drugs to fight COVID-19 using in silico methods: Targeting SARS-CoV-2 RdRp enzyme and host cell receptors (ACE2, CD147) through virtual screening and molecular dynamic simulations. Inform Med Unlocked 23:100541. https://doi.org/10.1016/j.imu.2021.100541 (Epub 2021 Feb 25)
Milani M, Donalisio M, Bonotto RM, Schneider E, Arduino I, Boni F, Mastrangelo E (2021) Combined in silico and in vitro approaches identified the antipsychotic drug lurasidone and the antiviral drug elbasvir as SARS-CoV2 and HCoV-OC43 inhibitors. Antivir Res 189:105055. https://doi.org/10.1016/j.antiviral.2021.105055
Nose S (1984) A unified formulation of the constant temperature molecular dynamics methods. J Chem Phys 81:511–519. https://doi.org/10.1063/1.447334
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182. https://doi.org/10.1063/1.328693
Peng Q, Peng R, Yuan B, Zhao J, Wang M, Wang X, Shi Y (2020) Structural and biochemical characterization of the nsp12-nsp7-nsp8 core polymerase complex from SARS-CoV-2. Cell Rep 31(11):107774. https://doi.org/10.1016/j.celrep.2020.107774 (Epub 2020 May 30)
Schmid N, Eichenberger AP, Choutko A, Riniker S, Winger M, Mark AE, van Gunsteren WF (2011) Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur Biophys J 40(7):843–856
Shamsi A, Mohammad T, Anwar S, AlAjmi MF, Hussain A, Rehman MT, Hassan MI (2020) Glecaprevir and Maraviroc are high-affinity inhibitors of SARS-CoV-2 main protease: possible implication in COVID-19 therapy. Biosci Rep 40(6):BSR20201256. https://doi.org/10.1042/BSR20201256
Sharma A et al (2020) Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2): a global pandemic and treatment strategies. Int J Antimicrob Agents 56(2):106054. https://doi.org/10.1016/j.ijantimicag.2020.106054
Sixto-López Y, Martínez-Archundia M (2021) Drug repositioning to target NSP15 protein on SARS-CoV-2 as possible COVID-19 treatment. J Comput Chem 42(13):897–907. https://doi.org/10.1002/jcc.26512
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem 31:455–461. https://doi.org/10.1002/jcc.21334
V’kovski P, Kratzel A, Steiner S, Stalder H, Thiel V (2021) Coronavirus biology and replication: implications for SARS-CoV-2. Nat Rev Microbiol 19(3):155–170. https://doi.org/10.1038/s41579-020-00468-6
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26(16):1701–1718
Wishart DS, Feunang YD, Guo AC, Lo EJ, Marcu A, Grant JR, Assempour N (2018) DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res 46(D1):D1074–D1082. https://doi.org/10.1093/nar/gkx1037
Yin W, Mao C, Luan X, Shen DD, Shen Q, Su H, Wang X, Zhou F, Zhao W, Gao M, Chang S, Xie YC, Tian G, Jiang HW, Tao SC, Shen J, Jiang Y, Jiang H, Xu Y, Zhang S, Zhang Y, Xu HE (2020) Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 368:1499–1504. https://doi.org/10.1126/science.abc1560 (Epub 2020 May 1)
Zhu W, Chen CZ, Gorshkov K, Xu M, Lo DC, Zheng W (2020) RNA-dependent RNA polymerase as a target for COVID-19 drug discovery. SLAS Discov Adv Sci Drug Discov 25(10):1141–1151. https://doi.org/10.1177/2472555220942123
Acknowledgements
We are grateful to everyone who helped us by giving valuable comments.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
Soodeh Mahdian conceived the original idea. Seyed Shahriar Arab developed the theory. Soodeh Mahdian wrote the manuscript. All authors discussed the results and contributed to the final manuscript. Seyed Shahriar Arab supervised the project.
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mahdian, S., Arab, S.S. Effectiveness of Remdesivir in Comparison with Five Approved Antiviral Drugs for Inhibition of RdRp in Combat with SARS-CoV-2. Iran J Sci Technol Trans Sci 46, 1359–1367 (2022). https://doi.org/10.1007/s40995-022-01364-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40995-022-01364-9