Abstract
Fusarium wilt caused by Fusarium oxysporum f. sp. ciceris (Padwick) Matuo & K. Sato has become the main limiting factor for chickpea (Cicer arietinum L.) production around the world. Although the cultivation of this legume is recent in Brazil, there are reports on Fusarium spp. occurrence causing crop losses. Fourteen isolates obtained from roots of chickpea plants showing wilt and yellowing symptoms in Brazil were evaluated through phylogenetic analysis of the EF-1α region, morphological markers and pathogenicity tests. Three isolates were clustered within a distinct lineage from those already described for the FSSC. The remaining 11 isolates were clustered within the FOSC, in a different clade from F. oxysporum f. sp. ciceris. All isolates were pathogenic but showed differences in aggressiveness. Isolates of the different complexes elicited the same symptoms: yellowing, wilt and root rot of chickpea plants. Morphological markers allowed differentiating isolates from distinct complexes but not differentiating between lineages.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chickpea, c L., is a legume crop of great importance worldwide, especially within African and Asian countries due to the nutritional value of its seeds. This legume has been increasingly used as an alternative to animal protein, mostly within developing countries, and has become an important dietary component for people that are vegetarian by choice or due to economic reasons (Wood and Grusak 2007; Jukanti et al. 2012).
There are more than 170 reported plant pathogens associated with chickpea (Nene et al. 1996), among which Fusarium oxysporum (Padwick) f. sp. ciceris Matuo & Sato is considered as the most important pathogen. This fungus causes vascular wilt or Fusarium wilt, limiting chickpea production (Jiménez-Díaz et al. 2015). The disease has already being reported in production regions from Asia, Africa, Southern Europe and The Americas (Chand and Khirbat 2009). Black root rot caused by Fusarium solani, although considered of a secondary importance, also causes significant losses in production in areas where the fungus is present in the soil (Nene et al. 2012).
Fusarium oxysporum is considered as being a complex of different species indistinguishable by only morphological markers (Lievens et al. 2008). This complex comprehends nonpathogenic and pathogenic isolates designated as formae speciales, characterized based in their host specificity (Armstrong and Armstrong 1981). Members of various phylogenetic lineages of the Fusarium oxysporum species complex (FOSC) are known to cause important plant diseases such as vascular wilt in various plants of economic interest (Di Pietro et al. 2003).
Results of studies with multigenic analysis, AFLP and vegetative compatibility group analysis showed that formae speciales do not represent phylogenetic groups, evidencing that isolates of a determined formae specialis may have distinct evolutionary origin (Baayen et al. 2000). Studies performed in Brazil with de F. oxysporum f. sp. malvacearum and F. oxysporum f. sp. phaseolis isolates evidenced that isolates of both formae speciales are polyphyletic (Silva et al. 2014).
Fusarium solani is a widely used name for what is nowadays named Fusarium solani species complex - FSSC (O’Donnell 2000). Members of the FSSC include various names that correspond to approximately 50 phylogenetic lineages (O’Donnell et al. 2008). Within the Fusarium solani complex are included various species causing soybean sudden death syndrome (Aoki et al. 2005; Costa et al. 2016). Fusarium azukicola was described in azuki beans, causing root rot (Aoki et al. 2012). In large soybean producing centers as Unites States, Brazil and Argentina, outbreaks of red rot were reported as having species belonging to the FSSC as their etiological agents (Scandiani et al. 2004; O'Donnell et al. 2010; Aoki et al. 2005).
The pathogenic association of F. oxysporum f. sp. ciceris with chickpea plants was reported in 1985 in plantations from the cerrado region in the Distrito Federal, Brazil (Sharma and Cerkauskas 1985). Recently, reports of the occurrence of fusariosis were made in the states of Goiás and Minas Gerais, however, no research using modern molecular techniques was done in order to confirm the etiological agent (Unpublished data).
Therefore, due to the absence of studies concerning the characterization of Fusarium species associated with chickpea in Brazil, we aimed to identify species associated to the etiology of the chickpea wilt by using the concept of morphological and phylogenetic species, as well as pathogenicity tests.
Materials and methods
Fusarium isolates
Isolates of Fusarium were obtained from root and stem tissue fragments from plants of different chickpea cultivars exhibiting yellowing, wilt and vascular discoloration symptoms. Fragments were externally disinfected in alcohol 70% for 1 min, sodium hypochlorite 1% for 1 min and three consecutive washes in sterilized distilled water. Then, fragments were transferred to Petri dishes containing water-agar (AA) medium. After four days of incubation at 25 °C and photoperiod of 12 h, fragments from the periphery of typical Fusarium colonies were transferred to potato dextrose agar (PDA) medium in order to obtain pure cultures. Monosporic cultures were obtained from pure colonies before being cryopreserved according Smith and Onions (1994) and stored at Coleção Micológica de Lavras (CML) at Universidade Federal de Lavras (UFLA), Lavras, Minas Gerais, Brazil (http://www.dfp.ufla.br/cml). Fourteen isolates were studied, being 13 originated from plants from the county of Cristalina - GO, Brazil and one isolate from the county of Montes Claros – MG, Brazil (Table 1).
DNA extraction, PCR amplification and phylogenetic analysis
Isolates were grown in 2% malt extract broth medium for three days at 100 rpm and had their DNA extracted with the Wizard® Genomic DNA Purification (Promega) kit, following manufacturer’s instructions. PCR amplifications were performed using a GoTaq Colorless Master Mix (Promega) kit in a My Cycler (Bio-Rad) thermocycler. Partial sequences of the gene EF-1α were amplified using primers Ef1-F (5′-ATGGGTAAGGAGGACAAGAC-3′) and Ef2-R (5′-GGAAGTACCAGTGATCATGTT-3′), described by O’Donnell et al. (1998). PCR conditions were one cycle of 90 s at 94 °C; 35 cycles of 30 s at 94 °C, 45 s at 62 °C and 1 min at 72 °C; followed by one cycle of 5 min at 72 °C and a 4 °C soak (O’Donnell et al. 2008). PCR products were purified using a Wizard®SV Gel kit and PCR Clean-up System (Promega) and sequenced by Macrogen Inc. Consensus sequences were assembled from forward and reverse sequences using SeqAssem (Hepperle 2004). Edited sequences were compared with the GenBank database, from the National Center for Biotechnology Information – NCBI, using the Basic Local Alignment Search Tool – BLAST program (http://www.ncbi.nlm.nih.gov/cgi-bin/BLAST/). Phylogenetic analyses were performed for each species found, based in the results from BLAST. Sequences of the isolates of F. solani were compared with the sequences of phylogenetic lineages of the FSSC (O’Donnell et al. 2008). Isolates of F. oxysporum were compared with the sequences of the phylogenetic lineages of FOSC (O’Donnell et al. 2009) and sequences of F. oxysporum f. sp. ciceris - NRRL 32153, NRRL 32154, NRRL 32155, NRRL 32156 (Gurjar et al. 2009), Foc 11, Foc 19, Foc 122 and Foc 155 (Dubey et al. 2014 ). Multiple alignments of sequences were generated using the ClustalW as implemented by the software Mega 6 (Tamura et al. 2013). The Maximum Parsimony (MP) phylogenetic analysis was performed on MEGA 6 (Tamura et al. 2013), using the Close-Neighbor-Interchange algorithm. Clade support was inferred from 1000 bootstrap replications. A complementary Bayesian inference (BI) analysis employing a Markov Chain Monte Carlo method was performed. MrModeltest 2.3 (Nylander 2004) was used to determine the best nucleotide substitution model for each data set. According Akaike Information Criterion (AIC) were selected K80 + G for the FOSC dataset and GTR + G for FOSC dataset. Markov chains were run for 10.000.000 generations and sampled every 100th generation. From the resulting 100.000 trees 25% was discarded and the remaining 75.000 trees were used for the calculation of posterior probabilities of branches. The BI phylogenetic analyses were performed using the CIPRES webportal (Miller et al. 2010). The sequences generated in this study were deposited in the GenBank (http://www.ncbi.nlm.nih.gov) (Table 1) and the alignments were deposited in the TreeBASE under accession number 19949 (http://purl.org/phylo/treebase/phylows/study/TB2:S19949).
Evaluation of morphological markers
Morphological characterization was performed according to the characters described by Gerlach and Niremberg (1982) and Leslie and Summerell (2006). Isolates were cultivated in Petri dishes containing PDA medium, incubated at 25 °C in the dark for 4 days, with three replicates used to evaluate the mycelium growth rate. Colony colour and formation of aerial mycelium were observed after 14 days of incubation in PDA medium at 20 °C in a 12 h photoperiod under near UV and white fluorescent bulbs. In synthetic nutrient-poor agar medium (SNA), under the same previously mentioned conditions, were observed microconidial and macroconidial shape and septation, presence and color of sporodochia, arrangement of conidiogenous cells and presence or absence of chlamydospores. For each characterized structure, 30 measures were registered under a light microscope at 40 x magnification.
Pathogenicity test
The pathogenicity test was performed in a greenhouse from the Instituto de Ciências Agrárias from Universidade Federal de Minas Gerais Campus Montes Claros-MG. Isolates of Fusarium were cultivated in Petri dishes containing PDA medium and incubated for 15 days in the darkness at 25 °C temperature. Seven days old chickpea plants cv. Cícero, previously produced in polypropylene trays filled with commercial substrate Plantmax (Plantmax Sementes Ltda), had their roots wounded with a scalpel and then were immersed 10 min in 200 mL of a conidia suspension (1 × 106 conidia.mL−1) of each Fusarium isolate. Then, plants were transferred to 700 mL pots containing autoclaved substrate 1:1 (soil:sand). Twenty plants (replicates) were inoculated for each isolate. The control treatment consisted of wounded plants immersed in sterilized distilled water. The experiment was performed under a completely random design. Plants showing symptoms of wilt or yellowing were considered diseased. Disease incidence was evaluated during 42 days (6 weeks). Isolates aggressiveness was determined by the increase of weekly incidence, during the whole period of evaluation. The pathogen was recovered in order to complete Kock’s postulates and confirm pathogenicity. The pathogenicity test was repeated twice. Aggressiveness was determined based on the incidence observed during the first test.
Results
Phylogenetic analysis
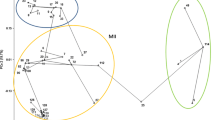
Phylogenetic trees were generated for each species complex, FOSC and FSSC. First the analysis of MP and BI were performed for all 22 partial sequences of the TEF-1α gene from F. oxysporum which contained 442 characters, being 378 conserved, 63 variable and within these, 24 were informative. In the resulting phylogram, the isolates of F. oxysporum (CML 2865, CML 2866, CML 2867, CML 2869, CML 2871, CML 2872, CML 2874, CML 2875, CML 2876, CML 2877 and CML 2878) made a distinct lineage within FOSC, strongly supported by both methods, clustering in a clade separated from F. oxysporum f. sp. ciceris isolates (Fig. 1).
Phylogram of maximum parsimony for EF-1α gene from lineages of the Fusarium oxysporum complex, including isolates from chickpea from Brazil. Length of the branches is shown by the scale in the tree base. Maximum parsimony bootstrap values ≥50% (1000 replicates) and posterior probabilities ≥0.90 are above the nodes. This tree has F. foetens NRRL38302 as root
Maximum parsimony and BI analysis was performed with 48 partial sequences of TEF-1α gene from F. solani contained 480 characters, being 304 conserved and 171 variable, with 103 being informative within these. The topology of the generated tree showed that isolates CML 2868, CML 2870 and CML 2873 did not cluster with any already described lineages for FSSC, constituting a clade strongly supported with a bootstrap of 97% and posterior probability equal to one, having the holotype of F. falciforme CBS 475–67 lineage FSSC 3 + 4 as a sister group (Fig. 2).
Phylogram of maximum parsimony for EF-1α gene from lineages of the Fusarium solani complex, including isolates from chickpea from Brazil. Length of the branches is shown by the scale in the tree base. Maximum parsimony bootstrap values ≥50% (1000 replicates) and posterior probabilities ≥0.90 are above the nodes. This tree has F. plagianthi NRRL22632 and F. iludens NRRL22090 as root
Evaluation of morphological markers
Within the 14 characterized Fusarium isolates, 11 isolates showed typical morphological markers for F. oxysporum and three for F. solani (Table 2).
Isolates of F. oxysporum on PDA showed sparse and cotton-like aerial mycelium, white or cream in color, with pigmentation at the colony’s bottom ranging from violet to cream. Growth rate ranged from 9.7 to 14.4 mm/day after 4 days at 25 °C in the dark. On SNA were observed microconidia aggregated in false heads on short phialides in the aerial mycelium (Fig. 3a). Presence of simple and short monophialides, 4–32 μm length (Fig. 3c). Microconidia cylindrical to kidney shaped, with 0–2 septs, 3–20 × 1–3 μm (Fig. 3b). Macroconidia falcate to straight in shape, with curved apical cell and evident basal foot cell, 17–55 × 2–5 μm, 3–7 septs, (Fig. 3d). All isolates showed globose chlamydospores, single or in chains, intercalary or terminal, with smooth or rough walls, 3–13.5 μm wide (Fig. 3e).
Morphological characteristics of Fusarium oxysporum isolates: a False heads on short phialides, bar = 50 μm; b Microconidia, bar = 10 μm; c Simple and short monophialides, bar = 10 μm; d Macroconidia, bar = 10 μm; e Chlamidospores, bar = 20 μm. Symptoms triggered by F. oxysporum isolates in chickpea plants: f Yellowing; g-h Root rot; and darkening of the vascular tissues
Isolates of F. solani showed white colored cotton-like aerial mycelium, with brown pigmentation at the colony’s bottom. Colonies on PDA showed mycelial growth rate varying from 9.9 to 10.7 mm/day. On SNA were observed microconidia produced in false heads on long monophialides, 22.5–172.5 μm length (Fig. 4a and b). Microconidia globose, fusiform and reniform in shape, 0–2 septate, 3–27 × 1.5–5 μm (Fig. 4c). All F. solani isolates showed abundant sporodochia with cream to pale orange coloration. Macroconidia showed cylindrical shape, with slight curved and rounded apical cell and slightly protruding basal foot cell, 21–51 × 3.5–5 μm, 3–5 septate (Fig. 4d). Chlamydospores were observed in pairs or in chains, globose shape, rough-walled, 4–10.5 μm wide (Fig. 3f and g).
Morphological characteristics of Fusarium solani isolates: a False heads on long phialides, bar = 50 μm; b Long monophialide, bar = 20 μm; c Microconidia, bar =10 μm; d Macroconidia, bar = 10 μm; e Macro and Microconidia, bar = 20 μm; f Chlamidospores, bar = 20 μm; g Chain of chlamidospores, bar = 10 μm. Symptoms triggered by F. solani isolates in chickpea plants: h Yellowing; i Root rot
Pathogenicity test
All Fusarium isolates studied were pathogenic to chickpea. All Infected plants showed rapid yellowing starting from the leaf blade with later wilt and plant death. F. solani and F. oxysporum isolates triggered similar aerial symptoms (Fig. 3f and 4h). Regarding to the radicular system, roots of all infected plants were rotted or showed blackened tissue (Fig. 3g- h and 4i). Some plants infected with F. oxysporum showed darkening of the vascular tissues, but always associated to root rot (Fig. 3o and p). Plants infected with F. solani don’t showed darkening of vascular tissues. Isolates varied in aggressiveness according to the velocity they caused disease symptoms (Fig. 5). In general, F. oxysporum isolates were more aggressive when compared to F. solani isolates. Isolate CML 2878 was more aggressive and caused high disease incidence since the second week after inoculation, while isolates CML 2865, CML 2866, CML 2869, CML 2872 and CML 2873 were less aggressive. Isolates CML 2867, CML 2868, CML CML 2870, CML 2871, CML 2874, CML 2875, CML 2876 and CML 2877 showed an intermediary aggressiveness (Fig. 5). The highest disease incidence was observed in plants inoculated with the isolate CML 2877.
Fungi were recovered from symptomatic plants and showed the same morphological characteristics from the originally inoculated isolates, thus confirming their pathogenicity. Control plants showed no symptoms at all. Inoculations were performed twice, showing similar results.
Discussion
The presence of Fusarium oxysporum f.sp. ciceris infecting chickpea plants in the cerrado’s areas from the Distrito Federal was reported in Brazil (Sharma and Cerkauskas 1985). However, the identification was based only in morphological markers, which are not enough to make distinctions between species within the FOSC. In the present study, isolates identified as belonging to the FOSC formed a clade with highly supported, clearly separated from F. oxysporum f. sp. ciceris, which is a monophyletic origin group (Jiménez-Gasco et al. 2002; Demers et al. 2014). This result corroborates the information that different FOSC phylogenetic lineages are able to parasite the same host. Phylogenetic analysis of Fusarium oxysporum f. sp. phaseoli and Fusarium oxysporum f. sp. malvacearum isolates confirmed that both populations are constituted by distinct phylogenetic lineages (Silva et al. 2014). Recently, phylogenetic studies evidenced that Fusarium oxysporum f. sp. lycopersi, agent of tomato wilt, actually consists of four distinct phylogenetic lineages (Nirmaladevi et al. 2016). These information shows that formae specialis is an obsolete taxonomical terminology and may exist many species of the FOSC to be described.
After phylogenetic analysis, F. solani isolates (CML 2868, CML 2870 and CML 2873) clustered together forming a strongly supported clade, having as a brother group the F. falciforme CBS 475–67 holotype, lineage FSSC 3 + 4 (Fig. 2). F. falciforme consists of a diverse group not yet resolved phylogenetically, associated to soils and human infections (Zhang et al. 2006; O’Donnell et al. 2008; Nalim et al. 2011). The isolates of Fusarium solani studied had no phylogenetic relation with the main species causing root rot in various legumes. Species triggering root rot in legumes crops are included in clade 2 which gathers pathogens causing red rot in soy bean and beans (Aoki et al. 2005; Covert et al. 2007; Aoki et al. 2012). Recently, Fusarium paranaense Costa & Pfenning was described as being pathogenic to soy bean, however, this species belongs to clade three (Costa et al. 2016).
The morphological characteristics evaluated allowed to differentiate between FOSC and FSSC but were not sufficient to differentiate lineages within each complex. The limited number of existent morphological characters is often not enough to express the great diversity within the genus Fusarium (Leslie and Summerell 2006), once individuals sharing the same morphology may belong to distinct phylogenetic species (O’Donnell 2000). Fusarium tucumaniae Aoki, member of the FSSC, shows distinct markers from the others species of the FSSC, such as macroconidia morphology and production of blue colored sporodochia (Aoki et al. 2005). Recently, Fusarium secorum Secor, a new member from the Fusarium fujikuroi complex was described. Based on morphological markers, this fungus was previously named Fusarium oxysporum f. sp. betae (Secor et al. 2014). Fusarium foetens Schroers and Fusarium oxysporum are distinct species, however they share similar morphological markers (Schroers et al. 2004). These observations evidence the importance for using the phylogenetic concept to identify fungi from the genus Fusarium.
Isolates of both complexes were pathogenic and triggered similar aerial symptoms of yellowing and root rot in chickpea plants. But only F. oxysporum isolates triggered darkening of the vascular tissues. Studies performed in California-USA, showed that although chickpea plants inoculated with F. oxysporum f. sp. ciceris and F. solani had similar symptomatology in the aerial portion of the plant, F. oxysporum f. sp. ciceris caused vascular discoloration and F. solani black root rot without vascular discoloration (Westerlund et al. 1974). In a different study, Trapero-Casas and Jiménez-Díaz (1985) reported that F. oxysporum isolates caused leaf yellowing and wilt associated to vascular discoloration, as well as rotting in roots and in the lap of the chickpea plants. Isolates of F. oxysporum from various formae speciales when infecting plant tissues shows systemic colonization, observed by the darkening of conducting vessels tissues (Leslie and Summerell 2006). While species from FSSC show local colonization, mainly causing rot roots, specifically in leguminous. Due to tissues colonization, the plants exhibit reflexes symptoms of yellowing (Aoki et al. 2005; Aoki et al. 2012).
Differences observed in aggressiveness from isolates may be a result of genetic variability within the population, once isolates were obtained from different chickpea cultivars. For F. oxysporum f. sp. ciceris, differences in aggressiveness between isolates may be explained by the existence of different pathotypes and races of the pathogen (Jiménez-Gasco et al. 2004). Studies must be performed to evaluating the pathogenic capacity of the isolates to different cultivars of chickpea. Future research is needed to exploit the diversity of Fusarium obtained from different geographical regions from Brazil, aiming to confirm the existence of different phylogenetic species associated to fusariosis in this culture.
References
Aoki T, O’Donnell K, Scandiani MM (2005) Sudden death syndrome of soybean in South America is caused by four species of Fusarium: Fusarium brasiliense sp. nov., F. cuneirostrum sp. nov., F. tucumaniae, and F. virguliforme. Mycoscience 46:162–183
Aoki T, Tanaka F, Suga H, Hyakumachi M, Scandiani MM, O’ Donnel K (2012) Fusarium azukicola sp. nov., an exotic azuki bean root-rot pathogen in Hikkaido, Japan. Mycologia 104:1068–1084
Armstrong GM, Armstrong JK (1981) Formae speciales and races of Fusarium oxysporum causing wilt diseases. In: Nelson PE, Toussoun TA, Cook RJ (eds) Fusarium: diseases, biology, and taxonomy. Pennsylvania State University Press, University Park, PA, pp 391–399
Baayen RP, O’Donnell K, Bonants PJM, Cigelnik E, Kroon LPNM, Roebroeck EJA, Waalwijk C (2000) Gene genealogies and AFLP analyses in the Fusarium oxysporum complex identify monophyletic and nommonophyletic Formae Speciales causing wilt and rot disease. Phytophatology 90:891–900
Chand H, Khirbat SK (2009) Chickpea wilt and its management-a review. Agric Rev 30:1–12
Costa SS, Matos KS, Tessmann DJ, Seixas CDS, Pfenning LH (2016) Fusarium paranaense sp. nov., a member of the Fusarium solani species complex causes root rot on soybean in Brazil. Fungal Biol 120:51–60
Covert SF, Aoki T, O’Donnell K, Starkey D, Holliday A, Geiser DM, Cheung F, Town C, Strom A, Juba J, Scandiani M, Yang XB (2007) Sexual reproduction in the soybean sudden death syndrome pathogen Fusarium tucumaniae. Fungal Genet Biol 44:799–807
Demers JE, Garzón CD, Jiménez-Gasco MDM (2014) Striking genetic similarity races of Fusarium oxysporum f. Sp. ciceris confirms a monophyletic origin and clonal evolution of the chickpea vascular wilt pathogen. Eur J Plant Pathol 139:303–318
Di Pietro AMP, Caracuel ZM, Delgado-Jarana J, Roncero MIG (2003) Fusarium oxysporum: exploring the molecular arsenal of a vascular wilt pathogen. Mol Plant Pathol 4:315–325
Dubey SC, Priyanka K, Singh V (2014) Phylogenetic relationship between different race representative populations of Fusarium oxysporum f. Sp. ciceris in respect of translation elongation factor-1α, β-tubulin, and internal transcribed spacer region genes. Arch Microbiol 196:445–452
Gerlach W, Niremberg H (1982) The genus Fusarium: a pictorial atlas. Biologischen Bundesanstalt fur Land-und Forstwirtschaft, Berlin
Gurjar GS, Barve MP, Giri AP, Gupta VS (2009) Identification of Indian pathogenic races of Fusarium oxysporum f. Sp. ciceris with gene specific, ITS and random markers. Mycologia 101:484–495
Hepperle D (2004) SeqAssem©: a sequence analysis tool conting assembler and trace data visualization tool for molecular sequences: win32-version. Sequentx, Klein Raden
Jiménez-Díaz RM, Castillo P, Jiménez-Gasco MM, Landa BB, Navas-Cortés JA (2015) Fusarium wilt of chickpeas: biology, ecology and management. Crop Prot 73:16–27
Jiménez-Gasco MM, Milgroom MG, Jiménez-Díaz RM (2002) Gene genealogies support Fusarium oxysporum f. Sp. ciceris as a monophyletic group. Plant Pathol 51:72–77
Jiménez-Gasco MM, Navas-Cortes J, Jiménez-Díaz RM (2004) The Fusarium oxysporum f. Sp. ciceris/Cicer arietinum: a case study of the evolution of plant-pathogenic fungi into races and pathotypes. Int Microbiol 7:95–104
Jukanti AK, Gaur PM, Gowda CLL, Chibbar RN (2012) Nutritional quality and health benefits of chickpea (Cicer arietinum L.): a review. Br J Nutr 108:11–26
Leslie JF, Summerell BA (2006) The Fusarium laboratory manual. Blackwell Publishing Asia, Australia
Lievens B, Rep M, Thomma BPHJ (2008) Recent developments in the molecular discrimination of formae speciales of Fusarium oxysporum. Pest Manag Sci 64:781–788
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: Proceedings of the Gateway Computing Environments Workshop (GCE). New Orleans, pp. 1–8
Nalim FA, Samuels GJ, Wijesundera RL, Geiser DM (2011) New species from the Fusarium solani species complex derived from perithecia and soil in the old world tropics. Mycologia 103:1302–1330
Nene YL, Sheila YK, and Sharma SB (1996) A world list of chickpea and pigeonpea pathogens. 5th edn. Patancheru 502 324, Andhra Pradesh, India: International Crops Research Institute for the Semi-Arid Tropics. (Semi-formal publication.), 27p
Nene YL, Reddy MV, Haware MP, Ghanekar AM, Amin KS, Pande S, Sharma M (2012) Field Diagnosis of Chickpea Diseases and their Control, Information Bulletin No. 28. (revised). International Crops Research Institute for the Semi-Arid Tropics
Nirmaladevi D, Venkataramana M, Srivastava RK, Uppalapati SR, Gupta VK, Yli-Mattila T, Clement Tsui KM, Srinivas C, Niranjana SR, Chandra NS (2016) Molecular phylogeny, pathogenicity and toxigenicity of Fusarium oxysporum f.sp. lycopersici. Sci Rep 6:21367
Nylander JAA (2004) MrModeltest ver. 2.3. Program distributed by the author. Evolutionary Biology Centre, Uppsala University
O’Donnell K (2000) Molecular phylogeny of the Nectria haematococca–Fusarium solani species complex. Mycologia 92:919–938
O’Donnell K, Kistler HC, Cigelnik E, Ploetz RC (1998) Multiple evolutionary origins of the fungus causing Panama disease of banana: concordant evidence from nuclear and mitochondrial gene genealogies. Proc Natl Acad Sci 95:2044–2049
O’Donnell K, Sutton DA, Fothergill A, Mccarthy D, Rinaldi MG, Brandt ME, Zhang N, Geiser DM (2008) Molecular phylogenetic diversity, multilocus haplotype nomenclature, and in vitro antifungal resistance within the Fusarium solani species complex. J Clin Microbiol 46:2477–2490
O’Donnell K, Gueidan C, Sink S, Johnston PR, Crous PW, Glenn A, Riley R, Zitomer NC, Colyer P, Waalwijk C, Van Der Lee T, Moretti A, Kang S, Kim HS, Geiser DM, Juba JH, Baayen RP, Cromey MG, Bithell S, Sutton DA, Skovgaard KR, Kistler P, Elliott HC, Davis M, Sarver MBAJ (2009) A two- locus DNA sequence database for typing plant and human pathogens within the Fusarium oxysporum species complex. Fungal Genet Biol 46:936–948
O'Donnell K, Sink S, Scandiani MM, Luque A, Colletto A, Biasoli M, Lenzi L, Salas G, González V, Ploper LD, Formento N, Pioli RN, Aoki T, Yang XB, Sarver BAJ (2010) Soybean sudden death syndrome species diversity within north and South America revealed by multilocus genotyping. Phytopathology 100:58–71
Scandiani M, Ruberti D, O’Donnell, Aoki T, Pioli R, Giorda L, Luque A, Biasoli M (2004) Recent outbreak of soybean sudden death syndrome caused by Fusarium virguliforme and F. tucumaniae in Argentina. Plant Dis 88:1044
Schroers HJ, Baayen RP, Meffert JP, Gruyter J, Hooftman M, O’Donnell K (2004) Fusarium foetens, a new species pathogenic to begonia elatior hybrids (Begonia x hiemalis) and the sister taxon of the Fusarium oxysporum species complex. Mycologia 96:393–406
Secor GA, Rivera- Varas V, Christ DS, Mathew FM, Khan MF, Varrelmann M, Bolton MD (2014) Characterization of Fusarium secorum, a new species causing Fusarium yellowing decline of sugar beet in north Central USA. Fungal Biol 118:764–775
Sharma RD, Cerkauskas RF (1985) Interação entre Meloidogyne javanica e Fusarium oxysporum f. sp. ciceris sobre o grão-de-bico. Nematol Bras 9:113–121
Silva FP, Vechiato MH, Harakava R (2014) EF-1 gene and IGS sequencing of Fusarium oxysporum f.Sp. vasinfectum and F. oxysporum f.Sp. phaseoli reveals polyphyletic origin of strains. Trop Plant Pathol 39:64–73
Smith D, Onions AHS (1994) The preservation and maintenance of living fungi. Kew: Commonwealth Mycological Institute
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Trapero-Casas A, Jiménez-Díaz RM (1985) Fungal wilt and root rot diseases of chickpea in southern Spain. Phytopathology 75:1146–1151
Westerlund FV, Campbell RN, Kimble KA (1974) Fungal root rots wilt of chickpea in California. Phytopathology 64:432–436
Wood JA, Grusak MA (2007) Nutritional value of chickpea. In: Redden RR, Chen W, Sharma B (eds) Yadav SS. CAB International, Chickpea Breeding and Management, pp 101–142
Zhang N, O’Donnell K, Sutton DA, Nalim FA, Summerbell RC, Padhye AA, Geiser DM (2006) Members of the Fusarium solani species complex that cause infections in both humans and plants are common in environment. J Clin Microbiol 44:2186–2190
Acknowledgements
The first author acknowledges the financial support provided by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior CAPES/PNPD. Authors are grateful to Osmar Artiaga for kindly allowing collect samples in his chickpea production field. We would like to express our sincere thanks to all team members from the Laboratório de Fitopatologia-UFMG and Labroratório de Ecologia e Sistemática de Fungos-UFLA for the voluntary help. Special thanks to Edson Luis Rezende and Everson for their support and technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Olinto L. Pereira
Rights and permissions
About this article
Cite this article
Azevedo, D.M.Q., Rocha, F.S., Costa, C.A. et al. Etiology of root rot and wilt disease of chickpea in Brazil. Trop. plant pathol. 42, 273–283 (2017). https://doi.org/10.1007/s40858-017-0145-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-017-0145-5