Summary
Background
Molecular technologies have paved the way to improved understanding of allergic diseases in many ways, ranging from molecular allergens to tailor-made tools for analytical, diagnostic, and therapeutic purposes. Engineering of such molecules has become a mainstay in most biotechnical and biomedical areas. A not so new kid on the block is the nanobody, a single-domain antibody obtained from primarily camelid species. Despite their large promise and potential, it took nanobodies a long time to also enter the stage in allergology.
Methods
This review summarizes the state of the art and the feasibility of engineering nanobody-based tools for applications in allergology.
Results
In recent years, nanobodies with specificity for allergens have been increasingly generated. In parallel, their molecular engineering has enabled the development of derivatives that offer many advantages compared to standard antibody approaches. Hence, different application forms of nanobody-based molecules have been developed and reported in proof-of-concept studies.
Discussion
Recent studies give a first glimpse of the future possibilities of nanobody technologies in a complex system such as allergic diseases. It has become clear that the simplicity of the approaches as compared to regular antibody technologies will both broaden and deepen the scope of applications in allergology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The central player in allergic diseases is the immunoglobulin E (IgE) antibody [1]. Allergen-specific IgE (sIgE) bound to the high-affinity IgE receptor FcεRI causes long-term sensitization of basophils and mast cells [2]. Crosslinking of the complexes by allergens triggers immediate allergic reactions and potential anaphylaxis [3]. CD23 is the low-affinity IgE receptor and crucial for facilitated antigen presentation, transport across the epithelium, regulation of IgE production, and epitope spread [3, 4]. IgE binding to both receptors and changes to these interactions are key in diagnosing disease and in therapeutic interventions.
Hence, sIgE is used as a biomarker of allergic sensitization and its measurement can provide information about the allergic state of a patient, the reactivity profile of the individual, potential cross-reactivity, and progression of the disease. Similarly, allergen-specific IgG levels are indicative of acquired protection induced i.a. by allergen immunotherapy (AIT). Hence, the different antibody isotypes involved in allergic disease represent a window for diagnostic as well as therapeutic options and therefore a focus area for basic and applied research.
In all conventional mammalian antibody isotypes, antigen recognition is driven by the variable regions of both the heavy and the light chains. By contrast, single-domain antibodies (sdabs), often called “nanobodies” or “single variable domain” (VHH), are the sole antigen-binding moiety of heavy-chain-only antibodies (HCAbs) occurring in camelid species and cartilaginous fish [5, 6]. Several benefits of the format render nanobodies highly versatile molecules in numerous biotechnological and biomedical applications [7]. So far, however, the use of nanobodies in the allergology field is scarce.
This review highlights the potential of using nanobodies and engineered nanobody formats in the context of allergology with a particular focus on diagnostic applications. Parts of this review have been presented at the International Symposium on Molecular Allergology meeting in 2022.
Allergy diagnostics: benefits and pitfalls of in vitro tests
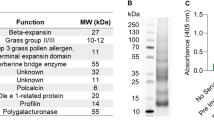
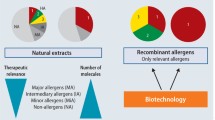
The diagnosis of allergy is currently based on several approaches including anamnesis, in vitro tests such as serological and cellular tests, and in vivo evaluation by skin tests and provocation tests. In the past few decades, knowledge on allergens has increased tremendously, providing clinicians and manufacturers of allergy tests with a variety of options to develop and use tailored tests for reliably determining the sensitization profile of the allergic patient. However, independently of the test, cross-reactivity of allergens based on sequence similarity or structural similarity complicates the read-out of serological tests significantly. The notorious IgE reactivity to cross-reactive carbohydrate determinants (CCD) complicates the picture when using extracts or natural allergens from a variety of sources [8]. Furthermore, extracts might suffer from quality problems arising from processing procedures. Aqueous extraction may result in loss of lipophilic components such as the oleosins in peanut [9], and follow-up purification and removal of small-molecular-weight components can lead to loss of smaller or sensitive components, e.g., as seen with the Api m 10 in honeybee venom (HBV; [10]). Hence, relevant components might be underrepresented or even missing in diagnostic and therapeutic extracts [11].
It is important to realize that serological in vitro tests represent sensitization, and that the results do not necessarily correlate with the clinical picture. In vitro tests typically only report concentrations and not other parameters, such as affinity, epitope usage, repertoire complexity, and the presence of blocking antibodies. More advanced, cellular tests such as the basophil activation test (BAT; [12, 13]) or the mast cell activation test (MAT; [14]) include a functional dimension in the form of degranulation in the test result and are therefore closer to the in vivo situation. Still, pronounced cellular activation does not necessarily imply allergic symptoms [15].
Today, more than 1000 allergenic molecules from a large number of animal and plant-derived species are listed in the IUIS/WHO database of allergenic molecules (www.allergen.org). The use of allergen components can provide technical and diagnostic benefits and often enables deeper insights into disease development and progression as well as on individual IgE reactivity profiles.
The clinical use of components, however, has been the subject of debate. An excellent example of the superiority of component testing is the yellow jacket major allergen Ves v 5, which shows a significantly higher sensitivity than the venom extract as such [16]. On the other hand, clinical use of components still raises doubts that components can provide a better handle for diagnosing than can the extract alone [17]. Often, components are recombinant proteins that might lack post-translational modifications (PTMs) or correct folding, or epitopes might be shielded by affinity tags, or the natural diversity of isoforms may not be represented.
Although this discussion might rely on either missing additional components or solvable technical aspects, it is undoubted that testing larger panels of allergen components can provide comprehensive insights into sensitization and cross-reactivity profiles. Hence, multiplexing technologies such as allergen arrays or similar products have potential beyond simple positive/negative assessment of single allergens, but the gain of in-depth information is somewhat questionable, as the costs are comparably high and information on potential other allergies might be obtained.
Similar to extracts, cross-reactivity occurs frequently in component testing. As mentioned above, an excellent example of the similarity of epitopes on different allergens is represented by CCDs, i.a. classic CCDs and alpha-Gal [8]. For the structurally highly defined CCD epitopes, equal IgE and IgG reactivity has been reported and highly specific antibodies have even been characterized in animals [18]. Hence, all proteins carrying such epitopes will show positive test results. Such a complete conservation of epitopes exemplifies the challenges both for extracts and for component testing.
Need for tools with defined specificity for allergens
While the value of in vitro tests is undoubted, the clinical reality of allergy testing is often different. Recent evaluations of sIgE proficiency testing have revealed a substantial variation of results between different test systems and between laboratories applying the same tests [19]. These studies assessed specifically important allergens such as birch pollen and Hymenoptera venom, resulting in a call by the authors for characterized standard material with known values of sIgE. A subsequent study with a focus on house dust mite and cat allergens corroborated the earlier findings [20].
As sIgE is not available for most allergens and in sufficient amounts, the determination of sIgE is based on heterologous calibration using a non-specific, quantified IgE preparation. Currently, the third international standard for serum IgE is the one in use [21]. The exact concentration of the non-specific IgE is measured by using an anti-IgE antibody instead of an allergen. This indirect method is reasonable, but far from ideal. In addition to a highly defined reference, national guidelines on diagnostic products might demand the availability of specific positive controls in the form of sera, indicating a proper function of the test. Similarly, the release of therapeutic products for AIT demands careful assessment with allergen-specific sera.
Overall, there is a clear demand for the availability of reagents that can provide a standardizable and reliable reactivity to all kinds of allergenic molecules. Considerations similar to the aforementioned aspects apply to other tests such as specific IgG4 or specific IgA tests.
Animal-derived reagents: antibodies and their coupling
For immunoassay establishment and validation, animal-derived reagents have been extensively used for more than a century. Hence, serum and polyclonal antibodies obtained by immunization of animals still represent efficient, simple, affordable, and easy-to-handle specific tools. A good example is the use of polyclonal antibody preparations in crossed immunoelectrophoresis for analysis of complex mixtures, e.g., therapeutic AIT preparations. In this technique, the main characteristic is the reactivity of a specific molecule or probe, independent of the origin and molecular context. Later, the advent of hybridoma technology nearly 50 years ago paved the way for specific, monoclonal antibodies for diagnostic and analytical purposes.
In a variety of settings, however, compatibility of tools with a human diagnostic assay set-up is crucial. Here, the presence and subsequent detection of a human IgE (or IgG, IgM, IgA) isotype is required. Such “humanized” tools require an allergen-specific binding moiety in conjunction with the human isotype of interest.
The most likely simplest approach relies on crosslinking an allergen-specific antibody with a human IgE molecule by common reagents, e.g., glutaraldehyde. Chemical derivatization, however, is often limited by problems with solubility, aggregation, and impaired reactivity with secondary reagents.
A real molecular linkage demands more sophisticated engineering of fusion molecules. Different approaches have been used for mimicking human serum with defined specificity, e.g., using single monoclonal IgE as described below [22,23,24] or IgE moieties forming complexes with animal-derived sera [23]. Here, fusion proteins of IgE with Fc receptors, i.a. FcγRI, were used for the formation of complexes with allergen-specific rabbit IgG. Binding of these complexes could then be detected using commonly used anti-human IgE antibodies [23]. Similar approaches using binders of animal-derived allergen-specific antibodies could in theory involve any other scaffold, which can be used in fusion approaches with human antibody isotypes.
Roads to allergen-specific IgE
Eventually, the best molecule for diagnostic standardization purposes would be allergen-specific human IgE. Generation of such defined IgE molecules has been pursued for many decades and only recently became more successful. The limited success is the result of several technical difficulties including the low frequency of IgE-producing memory B cells in peripheral blood and difficulties in identifying and growing IgE-producing B cells [25].
Due to these technical limitations, several types of allergen-specific surrogate human IgE have been generated using hybridoma technology and combinatorial antibody library technology. Most approaches rely on hybridoma technology and its use for generation of allergen-specific mouse IgG monoclonal antibodies. Recombinant chimeric mouse/human IgE monoclonal antibodies have been generated by combining the allergen-specific variable domains of mouse monoclonal IgG with human IgE domains [26, 27]. Chimeric mouse/human IgE have also been generated through the establishment of a knock-in mouse strain expressing human IgE (ε and κ constant regions). Here, allergen-specific chimeric mouse/human IgE is obtained by simple immunization [28].
Combinatorial antibody technologies enabled the generation of allergen-specific surrogate human IgE from synthetic combinatorial antibody libraries. Synthetic combinatorial single-chain antibody fragment (scFv) libraries have been used as the source for isolation of allergen-specific scFv, which were then fused to human IgE Fc for the generation of scFv-IgE [22].
In addition to synthetic libraries, combinatorial scFv and Fab IgE libraries have been generated from peripheral blood lymphocytes of allergic patients [29,30,31]. Again, selected IgE scFv or Fabs have subsequently been converted recombinantly to fully human IgE antibodies [31, 32].
Recent technical advances have enabled the direct generation of naturally occurring allergen-specific human IgE by using IgE hybridoma technology and single-cell transcriptomics [25, 33, 34]. These elegant approaches, however, demand high effort and are applicable for the most relevant allergens only.
Nanobodies: the promise of ease
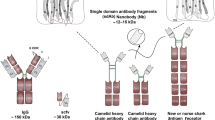
As a potential solution of the technical complexity of engineering monoclonal allergen-specific antibodies, we recently established the use of single-domain antibody technology for IgE engineering [35]. Sdabs are isolated from HCAbs found in animals of the family Camelidae. HCAbs lack light chains and therefore assemble as homodimers of heavy chains. Antigen recognition is mediated through a single VHH linked to the Fc domains by a hinge region ([5]; Fig. 1).
Nanobodies are single-domain antibodies derived from heavy-chain-only antibodies. a Schematic representation of a heavy-chain-only antibody compared to a conventional IgG antibody. b,c Cartoon representation of a crystal structure (PDB:1ZVY) and sequence of a lysozyme-specific nanobody [38]. Complementarity-determining regions (CDRs) are colored according to IMGT numbering (CDR1: green, CDR2: yellow, CDR3: orange)
The VHH, often referred to as a nanobody, has a size of only ~ 15 kDa. Despite their small size, nanobodies retain high affinities in the nanomolar range, and they are therefore considered the smallest natural antigen-binding fragment [36]. Nanobodies are structurally similar to the heavy-chain variable (VH) domain of human antibodies with a typical immunoglobulin fold, comprising four conserved framework regions and three complementarity-determining regions (CDR1–3) [36]. Nanobodies involve predominantly CDR3 for antigen interactions [37, 38], which is on average longer than the CDR3 of conventional antibodies [37, 39]. The long CDR3 loop and the prolate shape of nanobodies result in a convex paratope that can reach into clefts (e.g., active sites and binding pockets) and bind epitopes less accessible for conventional antibodies [38].
A major difference between nanobodies and conventional antibodies is four conserved hydrophobic residues that guide the contact of the VH and the VL. These are substituted in nanobodies by smaller and/or more hydrophilic amino acids increasing solubility in polar solutions [36].
Nanobodies exhibit pronounced stability and an excellent shelf life and can resist chemical and thermal denaturation due to efficient refolding. Furthermore, they can be produced with high yields in microbial hosts, e.g., Escherichia coli, Saccharomyces cerevisiae, and Pichia pastoris. Signal sequences often enable secretion into the culture medium even in bacteria and straightforward purification, making production and downstream characterization highly convenient [36, 40].
Generation and engineering of nanobodies
The straightforward approach for the generation of nanobodies is based on the ease of handling and manipulation on a molecular level. Nanobody repertoires can be generated with relatively limited effort. The bottleneck of repertoire generation in scFv and Fab formats of regular antibodies is the combination of heavy and light chains, which results in predominantly improper assembly. As the single domain of the eponymous sdabs does not need another domain for proper function, all molecules in the library are readily represented. Isolation of individual nanobodies from these cloned repertoires is then performed by using combinatorial selection approaches (Fig. 2).
Generation/selection of nanobodies by combinatorial technologies. Schematic overview of the generation and selection of allergen-specific nanobodies. Repertoire nanobody libraries are typically made either by immunizing a camelid (e.g., a llama) with the allergen of interest (immune library) or built synthetically using a pre-designed nanobody scaffold (synthetic library) in which diversity is introduced. Either way, to retrieve nanobodies specific to the allergen of interest, the nanobody repertoire is cloned into a phage display library or a yeast surface display library. In phage display, the screening is carried out on surface-immobilized allergens, whereas in yeast surface display the screening is carried out against soluble, natively folded allergens and binders are isolated by use of magnetic-activated cell sorting (MACS) or fluorescence-activated cell sorting (FACS). In both approaches, after clonal selection, enriched nanobodies undergo initial characterization by sequencing, recombinant expression, and functional testing
Repertoire libraries are typically generated either by immunization of a camelid (immune library) or built synthetically. For an immune library, the animal is immunized with the target antigen four to eight times. As antigen, purified or recombinant protein (soluble, properly folded) is preferred [41]. Peripheral blood lymphocytes are isolated from a blood sample and the mRNA serves as a template for PCR amplification of the VHH region, which is cloned into a suitable vector [36, 41].
If the target antigen is non-immunogenic, unstable, toxic, or a conformation-specific binder is aimed for, synthetic libraries are suitable. Such libraries are constructed using special pre-designed nanobody scaffolds [42,43,44]. However, synthetic libraries typically need to be larger than immune libraries (108–1010 clones for synthetic libraries compared to 106–108 clones for immune libraries) and require a larger number of selection rounds in order to retrieve high-affinity binders [41, 44].
Various combinatorial biology methods exist to retrieve antigen-specific nanobodies from a library, and all approaches that have been established for other antibody and ligand types can in principle be used for the selection of nanobodies (Fig. 2). Nevertheless, phage display (biopanning) on surface-immobilized antigens is the best established and most common approach [41]. The method enables affinity-based enrichment of binders in an iterative manner using the target of interest [45]. This platform has been used for a plethora of applications in allergology, ranging from cloned allergen repertoires [46] to allergen-specific IgE repertoires [29, 31].
Increasingly more popular is the use of cellular display technologies, which are enhanced by the power of cellular analysis and sorting techniques adding a functional layer during selection against soluble antigens. A particularly versatile technology is the yeast surface display (YSD) technology [47, 48]. Here, the binder is displayed on the yeast surface as fusion with, e.g., the Aga‑2 protein. Combined detection of the binder and antigen binding enables the gating and sorting of yeast cells of desired characteristics, such as high affinity, precise specificity, expression yield, etc. [49]. Furthermore, YSD has the advantage of a eukaryotic protein expression machinery, which allows for complex PTMs and quality control of protein folding, qualifying the binder further [50]. By contrast, also fully cell-free technologies like the ribosome display enable selection of in particular large synthetic nanobody repertoires, since cellular transformation can be circumvented [51].
After enrichment, individual nanobodies are expressed in small scale and evaluated for basic characteristics such as expression level, specificity, and affinity. Especially nanobodies originating from immune libraries are mostly of good quality and suitable for downstream applications [36, 41]. However, selected nanobodies may be further improved by various in vitro approaches [36, 52, 53].
Nanobodies are inherently limited by their small size, half-life, monovalency, and lack of functions beyond antigen binding. Thus, the availability of approaches for overcoming these limitations is needed and is crucial for more sophisticated applications (Fig. 3).
Engineered formats of nanobodies. Schematic representation of exemplified formats of engineered nanobodies, including linking of nanobodies into dimers or trimers with either mono-specific, bi-specific, or tri-specific functions, and Fc-fusions to human IgG or IgE, and fusion directly to larger proteins such as the human serum albumin (HSA)
Nanobodies can be engineered into dimers, trimers, oligomers, or multimers, either with mono-specific capabilities, bi-specific, or multi-specific properties. Genetic fusion of nanobodies is typically spaced by a linker, where the threefold glycine-serine linker (G4S)3 is the most commonly used enabling sufficient flexibility [54]. However, it might be necessary to test different linker lengths and order of nanobodies. Fusion of two (or more) identical nanobodies generates a monospecific construct that most likely will have enhanced affinity due to avidity effects, whereas the fusion of nanobodies with distinct epitopes will form bi-specific or multi-specific constructs of broader specificity [55].
Many strategies have been developed to extend the half-life of nanobodies, either by coupling to a second nanobody directed against HSA, or coupled to larger proteins such as HSA or fused to the human Fc domain either to the N‑ or C‑terminal end [35, 56, 57]. As for dimeric and multimeric nanobodies alone, constructs fused to HSA or Fc domains of IgG can be mono-specific or bi-specific.
Constructing bi-specific Fc fusions relies on the Fc heterodimerization, which can be realized via different approaches. An example is creating knobs-into-holes mutations in the Fc domains. A knob is created by replacing an amino acid with a small side chain with an amino acid with a larger side chain, and vice versa to create a hole in the complementary chain. Traditionally, a T366Y mutation in one CH3 domain has been used to create a knob while an a Y407T mutation gives rise to a hole [58]. Another example for construction of bi-specific Fc fusions is the use of DEKK mutations, where double mutations are incorporated into the two heavy chains with pairs consisting of substitutions L351D and L368E in one chain and L351K and T366K exchanges in the other chain [59].
Expression of Fc-fused nanobodies typically requires a more demanding expression system than the bacterial expression system, which could include the human embryonic kidney cells (HEK293) or adapted variants of this—FreeStyle293F or Expi293F, or Chinese hamster ovary (CHO) cells [60]. However, nanobody-based Fc fusions have also been expressed successfully in a variety of microbial hosts for the purpose of in vitro applications such as fusions to tags or as a secondary antibody [56].
Nanobodies with specificity for allergenic molecules
Currently, only a few nanobodies with specificity for allergens have been reported. Nanobodies against hen egg lysozyme, Gal d 4, have been developed, primarily for gaining insight into the structural understanding of the recognition mode of nanobodies [61]. More recently, nanobodies against the peanut allergen Ara h 3, the macadamia nut allergen Mac i 1, and the lupin seed allergen Lup an 1 were generated with the purpose of establishing immunoassays for allergen detection in food [62,63,64]. Furthermore, three nanobodies directed against the birch pollen allergen Bet v 1 were developed and found to recognize an important IgE epitope and thereby were able to reduce the binding of patient IgE to the allergen [65].
Recently, we have added to the list panels of nanobodies against individual Hymenoptera venom [35] and grass pollen allergens [66]. These targets as either single allergens or natural extracts were chosen deliberately, since they represent important areas within allergology. Venom allergens are often molecularly unique. By contrast, timothy grass is the best characterized allergenic grass since most of the commercial components in grass pollen allergy originate from timothy grass [67].
Application and gain in diagnostics and internal standardization
As described before, samples of well-characterized recombinant allergen-specific human IgE surrogates would be ideal tools for the determination of sIgE concentration, as their sIgE concentration by definition is equivalent to the total IgE (tIgE). Their potential has been demonstrated in a plethora of studies, i.a. a study by Wood et al. [24]. Here, chimeric mouse/human IgE directed against the allergens Bet v 1 and Der p 2 revealed a large deviation of sIgE from tIgE in two out of three systems tested [24].
Taking into consideration the large number of allergens, the selection and production of recombinant allergen-specific human IgE for standardization purposes should be as simple and efficient as possible. Therefore, we recently established the generation of nanobody-based human IgE (nb-hIgE; [35]). One major advantage of the nanobody-based IgE format is the simple structure. In contrast to the complex heterotetrameric structure of human or chimeric IgE, nb-hIgE is homodimeric, rendering secretion, assembly, and expression processes highly efficient, resulting in significantly increased yields (Fig. 3).
Recently, we generated nb-hIgE directed against the four allergens Api m 1, Api m 2, Phl p 4, and Phl p 6 [35, 66]. All the generated nb-hIgE are compatible with commercial test systems such as the ImmunoCap system. When the nb-hIgEs were applied for the determination of tIgE and sIgE using both single components and allergen extracts, we found a good consistency between tIgE and sIgE when testing with the single allergens (Fig. 4).
Diagnostic application of nb-IgE. Allergen-specific IgE (sIgE) and total IgE were measured on the ImmunoCap test system using four different allergen-specific nanobody-based human IgE formats. The test was performed using individual allergen components and allergen extracts. The data are reprinted from previous publications [35, 66]. Dashed lines indicate the optimal correlation between sIgE and total IgE
This emphasizes the quality of the single component in sIgE determinations. We also found a good consistency between tIgE and Api m 1 sIgE when testing with HBV extract. The recombinant Api m 1 ImmunoCap test sensitivity has previously been discussed [68, 69]. However, in our study we observed that the Api m 1 sIgE values were comparable to the values measured with HBV extract as well as to the measured tIgE, corroborating that the recombinant Api m 1 ImmunoCap test is reliable for Api m 1 sIgE determination.
By contrast, we found that the concentrations of Api m 2 sIgE and Phl p 4 sIgE were underestimated compared to the tIgE measurements when testing with HBV extract and timothy grass extract, respectively. Similar underestimation of sIgE observed when testing with allergen extract has previously been reported for Ves v 5 [16]. The sensitivity of the test could in this case be increased by spiking the venom with recombinant Ves v 5 and the currently available YJV test does contain added Ves v 5. Underestimation of IgE values might be caused by limited amounts of the allergen in the extract, insufficient coupling of the allergens to the solid surface, or inaccessibility of important epitopes.
The aforementioned examples underline the need for better standardization and demonstrate the quality of the nb-hIgE formats for sIgE determination on the ImmunoCap test system. As patient serum contains a complex mix of IgE antibodies recognizing different epitopes with different affinities, it would be ideal to mix nb-IgEs with different specificities and affinities for the generation of an artificial serum pool that more closely resembles the complexity of patient serum. Such well-characterized artificial serum pools would not only be valuable for the calibration of diagnostic tests but could also be applied for quality control of allergen extracts for diagnostics as well as immunotherapy.
Application areas of nanobodies beyond diagnostics
Targeting allergens or other relevant molecules with nanobodies offers a variety of benefits in other areas in allergology besides diagnostics. Nanobodies as ready-to-use building blocks allow for a plethora of advanced derivatives. One area of major interest is using nanobodies in a therapeutic setting, in which soluble targets such as allergens and cytokines or cellular targets such as receptors can be addressed.
There is a broad range of soluble targets of nanobodies. As an example, it has been shown that a trimeric nanobody targeting the fusion protein (F protein) in respiratory syncytial virus (RSV) could efficiently neutralize the virus in vitro and effectively reduce both nasal and lung RSV titers when delivered directly to the tissue in the nose or in the lungs of rats [70].
Targeting the key mediator of allergic reactions, IgE, with nanobodies has been demonstrated. The potential of anti-IgE nanobodies in the context of HCAbs has been shown by immunization of camels and analysis of inhibition by the natural antibodies [71]. For isolated and monoclonal nanobodies, it has been shown that the anti-IgE nanobody 026 exhibited not only blocking but also pronounced disruption of the interaction of IgE to FcεR1 [72]. The structural peculiarities of the nanobody/antigen interaction as observed here emphasize the large potential of nanobodies for inducing conformational changes.
Recently, data have suggested that allergen-specific blocking by IgG antibodies may be a highly protective mechanism in AIT. For example, a cocktail of human IgG antibodies directed against the cat allergen Fel d 1 significantly reduced the allergic response in patients with cat allergy [73, 74]. Similarly, a cocktail consisting of three human IgG4 antibodies developed against the major birch pollen allergen, Bet v 1, has been shown to significantly reduce symptoms in birch pollen allergic patients [75]. These proof-of-concept studies with human antibodies can easily be considered as trailblazers for the utilization of blocking nanobodies in the IgG format in AIT. Enabled by the versatility of nanobody technologies, it is of course tempting to speculate and foresee a combination of blocking function and cellular receptor targeting, and other options.
Recently, a nanobody targeting the intercellular adhesion molecule 1 (ICAM-1) on the surface of respiratory epithelial cells in allergic patients exhibited potential for blocking allergen entry into the epithelial cells [76]. Combining the different targeting approaches in a complementary manner might hold potential for the development of highly efficient therapeutic molecules.
Perspectives
The molecular engineering of nanobodies has advanced significantly in the past few years and continues evolving into a platform applicable in a variety of areas. The main difference to other technologies is the ease of handling, which enables also non-specialized laboratories to develop, engineer, and apply nanobodies in all kinds of applications in the field, including analytical, research-based, diagnostic, and even therapeutic use.
This unique versatility to generate a large set of downstream molecules will expand in the years to come, increasingly replacing standard antibodies in many areas. The use of synthetic libraries for the development of binders, which had been hampered by the complexity of multi-domain molecules, will further expand the toolbox and thereby contribute to boosting efficient nanobody generation. All aspects of conventional antibody technologies, from affinity maturation to enhanced effector function, are applicable to nanobodies, but can be performed in an easier manner.
Hence, it is easily foreseeable that complex immunological settings such as allergology and clinical immunology will particularly benefit and be boosted by nanobody technologies. Availability of large numbers of allergen-specific tools, exploration of novel targeting and delivery concepts, and many other fascinating approaches come into reach, which ultimately will benefit the patient.
Abbreviations
- AIT:
-
allergen immunotherapy
- BAT:
-
basophil activation test
- CCD:
-
cross-reactive carbohydrate determinants
- CDR:
-
complementarity-determining region
- HBV:
-
honeybee venom
- HCAbs:
-
heavy chain-only antibodies
- HEK:
-
human embryonic kidney
- HSA:
-
human serum albumin
- IgE:
-
immunoglobulin E
- MAT:
-
mast cell activation test
- nb-hIgE:
-
nanobody-based human IgE
- PTM:
-
post-translational modification
- RSV:
-
respiratory syncytial virus
- scFv:
-
single-chain antibody fragment
- sdabs:
-
single-domain antibodies
- sIgE:
-
specific IgE
- tIgE:
-
total IgE
- VH:
-
Heavy-chain variable domain
- VHH:
-
single variable domain
- YSD:
-
yeast surface display
References
Finkelman FD, Boyce JA, Vercelli D, Rothenberg ME. Key advances in mechanisms of asthma, allergy, and immunology in 2009. J Allergy Clin Immunol. 2010;125:312–8.
Chang TW. The pharmacological basis of anti-IgE therapy. Nat Biotechnol. 2000;18:157–62.
Gould HJ, Sutton BJ. IgE in allergy and asthma today. Nat Rev Immunol. 2008;8:205–17.
Clement MJ, Fortune A, Phalipon A, Marcel-Peyre V, Simenel C, Imberty A, et al. Toward a better understanding of the basis of the molecular mimicry of polysaccharide antigens by peptides: the example of Shigella flexneri 5a. J Biol Chem. 2006;281:2317–32.
Hamers-Casterman C, Atarhouch T, Muyldermans S, Robinson G, Hamers C, Songa EB, et al. Naturally occurring antibodies devoid of light chains. Nature. 1993;363:446–8.
Ward ES, Gussow D, Griffiths AD, Jones PT, Winter G. Binding activities of a repertoire of single immunoglobulin variable domains secreted from Escherichia coli. Nature. 1989;341:544–6.
Konning D, Zielonka S, Grzeschik J, Empting M, Valldorf B, Krah S, et al. Camelid and shark single domain antibodies: structural features and therapeutic potential. Curr Opin Struct Biol. 2016;45:10–6.
Platts-Mills TA, Hilger C, Jappe U, van Hage M, Gadermaier G, Spillner E, et al. Carbohydrate epitopes currently recognized as targets for IgE antibodies. Allergy. 2021;76:2383–94.
Pons L, Chery C, Romano A, Namour F, Artesani MC, Gueant JL. The 18 kDa peanut oleosin is a candidate allergen for IgE-mediated reactions to peanuts. Allergy. 2002;57(72):88–93.
Blank S, Seismann H, Michel Y, McIntyre M, Cifuentes L, Braren I, et al. Api m 10, a genuine A. mellifera venom allergen, is clinically relevant but underrepresented in therapeutic extracts. Allergy. 2011;66:1322–9.
Frick M, Fischer J, Helbling A, Rueff F, Wieczorek D, Ollert M, et al. Predominant Api m 10 sensitization as risk factor for treatment failure in honey bee venom immunotherapy. J Allergy Clin Immunol. 2016;138:1663–1671.e9.
Santos AF, Alpan O, Hoffmann HJ. Basophil activation test: mechanisms and considerations for use in clinical trials and clinical practice. Allergy. 2021;76:2420–32.
Erdmann SM, Sachs B, Schmidt A, Merk HF, Scheiner O, Moll-Slodowy S, et al. In vitro analysis of birch-pollen-associated food allergy by use of recombinant allergens in the basophil activation test. Int Arch Allergy Immunol. 2005;136:230–8.
Bahri R, Custovic A, Korosec P, Tsoumani M, Barron M, Wu J, et al. Mast cell activation test in the diagnosis of allergic disease and anaphylaxis. J Allergy Clin Immunol. 2018;142:485–96.e16.
Eberlein B, Krischan L, Darsow U, Ollert M, Ring J. Double positivity to bee and wasp venom: improved diagnostic procedure by recombinant allergen-based IgE testing and basophil activation test including data about cross-reactive carbohydrate determinants. J Allergy Clin Immunol. 2012;130:155–61.
Vos B, Kohler J, Muller S, Stretz E, Rueff F, Jakob T. Spiking venom with rVes v 5 improves sensitivity of IgE detection in patients with allergy to Vespula venom. J Allergy Clin Immunol. 2013;131:1225–7.e1.
Vachova M, Panzner P, Kopac P, Bidovec Stojkovic U, Korosec P. Routine clinical utility of honeybee venom allergen components. J Allergy Clin Immunol Pract. 2018;6:2121–2123.e1.
Prenner C, Mach L, Glossl J, Marz L. The antigenicity of the carbohydrate moiety of an insect glycoprotein, honey-bee (Apis mellifera) venom phospholipase A2. The role of alpha 1,3-fucosylation of the asparagine-bound N‑acetylglucosamine. Biochem J. 1992;284(2):377–80.
Wojtalewicz N, Goseberg S, Kabrodt K, Schellenberg I. Six years of INSTAND e. V. sIgE proficiency testing: an evaluation of in vitro allergy diagnostics. Allergo J Int. 2017;26:43–52.
Wojtalewicz N, Kabrodt K, Goseberg S, Schellenberg I. Evaluation of the manufacturer-dependent differences in specific immunoglobulin E results for indoor allergens. Ann Allergy Asthma Immunol. 2018;121:490–5.
Thorpe SJ, Heath A, Fox B, Patel D, Egner W. The 3rd international standard for serum IgE: international collaborative study to evaluate a candidate preparation. Clin Chem Lab Med. 2014;52:1283–9.
Braren I, Blank S, Seismann H, Deckers S, Ollert M, Grunwald T, et al. Generation of human monoclonal allergen-specific IgE and IgG antibodies from synthetic antibody libraries. Clin Chem. 2007;53:837–44.
Offermann N, Plum M, Hubner U, Rathloff K, Braren I, Fooke M, et al. Human serum substitution by artificial sera of scalable allergen reactivity based on polyclonal antibodies and chimeras of human FcgammaRI and IgE domains. Allergy. 2016;71:1794–9.
Wood RA, Segall N, Ahlstedt S, Williams PB. Accuracy of IgE antibody laboratory results. Ann Allergy Asthma Immunol. 2007;99:34–41.
Smith SA, Chruszcz M, Chapman MD, Pomes A. Human monoclonal IgE antibodies‑a major milestone in allergy. Curr Allergy Asthma Rep. 2023;23:53–65.
Schuurman J, Perdok GJ, Lourens TE, Parren PW, Chapman MD, Aalberse RC. Production of a mouse/human chimeric IgE monoclonal antibody to the house dust mite allergen Der p 2 and its use for the absolute quantification of allergen-specific IgE. J Allergy Clin Immunol. 1997;99:545–50.
Furtado PB, McElveen JE, Gough L, Armour KL, Clark MR, Sewell HF, et al. The production and characterisation of a chimaeric human IgE antibody, recognising the major mite allergen Der p 1, and its chimaeric human IgG1 anti-idiotype. Mol Pathol. 2002;55:315–24.
Lu CS, Hung AF, Lin CJ, Chen JB, Chen C, Shiung YY, et al. Generating allergen-specific human IgEs for immunoassays by employing human epsilon gene knockin mice. Allergy. 2015;70:384–90.
Steinberger P, Kraft D, Valenta R. Construction of a combinatorial IgE library from an allergic patient. Isolation and characterization of human IgE Fabs with specificity for the major timothy grass pollen allergen, Phl p 5. J Biol Chem. 1996;271:10967–72.
Jakobsen CG, Bodtger U, Kristensen P, Poulsen LK, Roggen EL. Isolation of high-affinity human IgE and IgG antibodies recognising bet v 1 and humicola lanuginosa lipase from combinatorial phage libraries. Mol Immunol. 2004;41:941–53.
Hecker J, Diethers A, Schulz D, Sabri A, Plum M, Michel Y, et al. An IgE epitope of Bet v 1 and fagales PR10 proteins as defined by a human monoclonal IgE. Allergy. 2012;67:1530–7.
Hecker J, Diethers A, Etzold S, Seismann H, Michel Y, Plum M, et al. Generation and epitope analysis of human monoclonal antibody isotypes with specificity for the Timothy grass major allergen Phl p 5a. Mol Immunol. 2011;48:1236–44.
Croote D, Darmanis S, Nadeau KC, Quake SR. High-affinity allergen-specific human antibodies cloned from single IgE B cell transcriptomes. Science. 2018;362:1306–9.
Wurth MA, Hadadianpour A, Horvath DJ, Daniel J, Bogdan O, Goleniewska K, et al. Human IgE mAbs define variability in commercial Aspergillus extract allergen composition. JCI Insight. 2018;3(20):e123387. https://doi.org/10.1172/jci.insight.123387.
Aagaard JB, Sivelle C, Fischer M, Byskov K, Laursen NS, Pfutzner W, et al. Nanobody-based human antibody formats act as IgE surrogate in hymenoptera venom allergy. Allergy. 2022;77:2859–62.
Muyldermans S. Nanobodies: natural single-domain antibodies. Annu Rev Biochem. 2013;82:775–97.
Zavrtanik U, Lukan J, Loris R, Lah J, Hadzi S. Structural basis of epitope recognition by heavy-chain camelid antibodies. J Mol Biol. 2018;430:4369–86.
De Genst E, Silence K, Decanniere K, Conrath K, Loris R, Kinne J, et al. Molecular basis for the preferential cleft recognition by dromedary heavy-chain antibodies. Proc Natl Acad Sci U S A. 2006;103:4586–91.
Mitchell LS, Colwell LJ. Comparative analysis of nanobody sequence and structure data. Proteins. 2018;86:697–706.
Harmsen MM, De Haard HJ. Properties, production, and applications of camelid single-domain antibody fragments. Appl Microbiol Biotechnol. 2007;77:13–22.
Muyldermans S. A guide to: generation and design of nanobodies. FEBS J. 2021;288:2084–102.
McMahon C, Baier AS, Pascolutti R, Wegrecki M, Zheng S, Ong JX, et al. Yeast surface display platform for rapid discovery of conformationally selective nanobodies. Nat Struct Mol Biol. 2018;25:289–96.
Moutel S, Bery N, Bernard V, Keller L, Lemesre E, de Marco A, et al. NaLi-H1: A universal synthetic library of humanized nanobodies providing highly functional antibodies and intrabodies. Elife. 2016;5:e16228.
Zimmermann I, Egloff P, Hutter CAJ, Kuhn BT, Brauer P, Newstead S, et al. Generation of synthetic nanobodies against delicate proteins. Nat Protoc. 2020;15:1707–41.
Smith GP. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science. 1985;228:1315–7.
Crameri R, Walter G. Selective enrichment and high-throughput screening of phage surface-displayed cDNA libraries from complex allergenic systems. Comb Chem High Throughput Screen. 1999;2:63–72.
Boder ET, Wittrup KD. Yeast surface display for screening combinatorial polypeptide libraries. Nat Biotechnol. 1997;15:553–7.
Ryckaert S, Pardon E, Steyaert J, Callewaert N. Isolation of antigen-binding camelid heavy chain antibody fragments (nanobodies) from an immune library displayed on the surface of Pichia pastoris. J Biotechnol. 2010;145:93–8.
Sivelle C, Sierocki R, Ferreira-Pinto K, Simon S, Maillere B, Nozach H. Fab is the most efficient format to express functional antibodies by yeast surface display. mAbs. 2018;10:720–9.
Kang BH, Lax BM, Wittrup KD. Yeast surface display for protein engineering: library generation, screening, and affinity maturation. Methods Mol Biol. 2022;2491:29–62.
Chen X, Gentili M, Hacohen N, Regev A. A cell-free nanobody engineering platform rapidly generates SARS-CoV‑2 neutralizing nanobodies. Nat Commun. 2021;12:5506.
Koide A, Tereshko V, Uysal S, Margalef K, Kossiakoff AA, Koide S. Exploring the capacity of minimalist protein interfaces: interface energetics and affinity maturation to picomolar KD of a single-domain antibody with a flat paratope. J Mol Biol. 2007;373:941–53.
Yau KY, Dubuc G, Li S, Hirama T, Mackenzie CR, Jermutus L, et al. Affinity maturation of a V(H)H by mutational hotspot randomization. J Immunol Methods. 2005;297:213–24.
Chen X, Zaro JL, Shen WC. Fusion protein linkers: property, design and functionality. Adv Drug Deliv Rev. 2013;65:1357–69.
Conrath KE, Lauwereys M, Wyns L, Muyldermans S. Camel single-domain antibodies as modular building units in bispecific and bivalent antibody constructs. J Biol Chem. 2001;276:7346–50.
Djender S, Schneider A, Beugnet A, Crepin R, Desrumeaux KE, Romani C, et al. Bacterial cytoplasm as an effective cell compartment for producing functional VHH-based affinity reagents and camelidae IgG-like recombinant antibodies. Microb Cell Fact. 2014;13:140.
Shen ZL, Xiang YF, Vergara S, Chen AP, Xiao ZY, Santiago U, et al. A resource of high-quality and versatile nanobodies for drug delivery. Iscience. 2021;24(9):103014. https://doi.org/10.1016/j.isci.2021.103014.
Ridgway JBB, Presta LG, Carter P. ‘Knobs-into-holes’ engineering of antibody C(H)3 domains for heavy chain heterodimerization. Protein Eng. 1996;9:617–21.
De Nardis C, Hendriks LJA, Poirier E, Arvinte T, Gros P, Bakker ABH, et al. A new approach for generating bispecific antibodies based on a common light chain format and the stable architecture of human immunoglobulin G(1). J Biol Chem. 2017;292:14706–17.
de Marco A. Recombinant expression of nanobodies and nanobody-derived immunoreagents. Protein Expr Purif. 2020;172:105645.
Akiba H, Tamura H, Kiyoshi M, Yanaka S, Sugase K, Caaveiro JMM, et al. Structural and thermodynamic basis for the recognition of the substrate-binding cleft on hen egg lysozyme by a single-domain antibody. Sci Rep. 2019;9:15481.
Chen F, Ma H, Li Y, Wang H, Samad A, Zhou J, et al. Screening of nanobody specific for peanut major allergen Ara h 3 by phage display. J Agric Food Chem. 2019;67:11219–29.
Hu Y, Wu S, Wang Y, Lin J, Sun Y, Zhang C, et al. Unbiased immunization strategy yielding specific nanobodies against macadamia allergen of Vicilin-like protein for immunoassay development. J Agric Food Chem. 2021;69:5178–88.
Hu Y, Zhang C, Yang F, Lin J, Wang Y, Wu S, et al. Selection of specific nanobodies against lupine allergen Lup an 1 for immunoassay development. Foods. 2021;10(10):2428.
Zettl I, Ivanova T, Strobl MR, Weichwald C, Goryainova O, Khan E, et al. Isolation of nanobodies with potential to reduce patients IgE binding to Bet v 1 (68/100 characters). Allergy. 2022;77(6):1751–60. https://doi.org/10.1111/all.15191.
Aagaard JB, Fischer M, Lober J, Neumann FB, Allahverdi D, Sivelle C, et al. Extract-shaped immune repertoires as source for nanobody-based human IgE in grass pollen allergy. Mol Biotechnol. 2023; https://doi.org/10.1007/s12033-023-00664-8.
Matricardi PM, Kleine-Tebbe J, Hoffmann HJ, Valenta R, Hilger C, Hofmaier S, et al. EAACI molecular allergology user’s guide. Pediatr Allergy Immunol. 2016;27(23):1–250.
Korosec P, Valenta R, Mittermann I, Celesnik N, Erzen R, Zidarn M, et al. Low sensitivity of commercially available rApi m 1 for diagnosis of honeybee venom allergy. J Allergy Clin Immunol. 2011;128:671–3.
Schrautzer C, Bokanovic D, Hemmer W, Lang R, Hawranek T, Schwarz I, et al. Sensitivity and specificity of hymenoptera allergen components depend on the diagnostic assay employed. J Allergy Clin Immunol. 2016;137:1603–5.
Detalle L, Stohr T, Palomo C, Piedra PA, Gilbert BE, Mas V, et al. Generation and characterization of ALX-0171, a potent novel therapeutic nanobody for the treatment of respiratory syncytial virus infection. Antimicrob Agents Chemother. 2016;60:6–13.
Khaled AQ, Sana Y, Abdulrahman R, Raida K, Sami AH. Blocking of histamine release and IgE binding to FcepsilonRI on human basophils by antibodies produced in camels. Allergy Asthma Immunol Res. 2015;7:583–9.
Jabs F, Plum M, Laursen NS, Jensen RK, Molgaard B, Miehe M, et al. Trapping IgE in a closed conformation by mimicking CD23 binding prevents and disrupts Fc epsilon RI interaction. Nat Commun. 2018;9(1):7. https://doi.org/10.1038/s41467-017-02312-7.
Orengo JM, Radin AR, Kamat V, Badithe A, Ben LH, Bennett BL, et al. Treating cat allergy with monoclonal IgG antibodies that bind allergen and prevent IgE engagement. Nat Commun. 2018;9:1421.
Shamji MH, Singh I, Layhadi JA, Ito C, Karamani A, Kouser L, et al. Passive prophylactic administration with a single dose of anti-Fel d 1 monoclonal antibodies REGN1908-1909 in cat allergen-induced allergic rhinitis - a randomized, double-blind, placebo-controlled clinical trial. Am J Respir Crit Care Med. 2021;204:23–33.
Gevaert P, De Craemer J, De Ruyck N, Rottey S, de Hoon J, Hellings PW, et al. Novel antibody cocktail targeting Bet v 1 rapidly and sustainably treats birch allergy symptoms in a phase 1 study. J Allergy Clin Immunol. 2022;149:189–99.
Zettl I, Ivanova T, Zghaebi M, Rutovskaya MV, Ellinger I, Goryainova O, et al. Generation of high affinity ICAM-1-specific nanobodies and evaluation of their suitability for allergy treatment. Front Immunol. 2022;13:1022418. https://doi.org/10.3389/fimmu.2022.1022418.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
J. Baunvig Aagaard, A.-S. Ravn Ballegaard, P. Ommen Andersen and E. Spillner declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Baunvig Aagaard, J., Ravn Ballegaard, AS., Ommen Andersen, P. et al. Molecular engineering of nanobodies as tools in allergology: diagnostics and beyond. Allergo J Int 32, 240–250 (2023). https://doi.org/10.1007/s40629-023-00261-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40629-023-00261-w