Abstract
Purpose of Review
This report broaches the topic of altered fidelity of DNA replication in herpesvirus mutants described over the past decades. Reduced genome replication fidelity of herpesvirus exonuclease mutants allows studying of virus population dynamics in the absence of exonucleolytic proofreading and can inform us on virus evolution in the face of error-prone genome replication.
Recent Findings
We recently found that mutations previously described to be lethal for herpes simplex type 1 (HSV-1) caused error-prone genome replication in Marek’s disease virus. This has allowed us to study the influence of augmented genetic diversity on viral population dynamics, replicative fitness, and virulence.
Summary
We conclude that the use of herpesvirus fidelity mutants allows unprecedented insights into virus evolution driven by low-fidelity replication. More than that, their use allows us to observe accelerated evolution, potentially enabling time-saving screens for the rise of drug- or vaccine-resistant mutants. In addition, we can infer that lethal or suicidal phenotypes observed in low-fidelity herpesvirus mutants are likely a consequence of error-prone genome replication, ultimately leading to lethal mutagenesis of small and isolated virus populations in cell culture.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Fidelity of DNA Replication
DNA replication is a process of remarkable accuracy. Nonetheless, accuracy is not perfect and the occasional misincorporation of bases cannot be avoided. Replication errors are important, however, as they constitute the molecular basis of biodiversity and are prerequisite for evolution. Replication errors can be single-nucleotide polymorphisms (SNPs) [1], single base deletions, or single base insertions (indels) [2]. The overall fidelity of DNA replication is primarily based on (1) nucleotide selectivity and (2) exonucleolytic proofreading [2].
Nucleotide selectivity is driven by the thermodynamic barrier presented by correct versus incorrect base pairing. Forming the correct hydrogen bond is thermodynamically favored but should still allow error rates of up to 1 in 100 [3]. A much higher accuracy of RNA polymerases strongly argues for other possibilities for base discrimination. It appears that the structural properties of the holoenzyme play an important role in enhancing nucleic acid replication fidelity, for example, by excluding water from the catalytic side [4] and by controlling for steric hindrances that would result from incorrect base pairing [5].
Intrinsic exonucleolytic proofreading is a unique enzymatic function of DNA polymerases [6•]. It is executed by a 3′– > 5′ exonuclease that is embedded in replicative DNA polymerases. The process of proofreading rests on the exonuclease removing erroneously incorporated bases, which allows the subsequent insertion of the correct nucleotide by the polymerase. The coordinated action of exonuclease and polymerase is essential for efficient and accurate DNA replication, and the proofreading process can lower the error rate of nucleic acid replication by 10–1000-fold [6•, 7, 8 ].
DNA Replication Fidelity in Herpesviruses
Herpesviruses are large dsDNA viruses that encode a family B replicative viral DNA polymerase with intrinsic 3′– > 5′ exonuclease activity, capable of exonucleolytic proofreading [9]. They are therefore believed to replicate their genomes with relatively high fidelity, comparable with other dsDNA microorganisms [10]. Most of the work on replication fidelity of herpesviruses was done for herpes simplex virus type 1 (HSV-1), and there is some ambiguous literature regarding the mutation rate of this particular alphaherpesvirus. In older reports, mutation rates of HSV-1 are estimated to be in the range of 0.003 mutations per genome and replication, equaling to a base substitution rate of around 2 × 10−8 or up to 5.9 × 10−8 based on slightly different calculations [1, 10]. These relatively low mutation rates are contrasted by a more recent study that suggests mutation rates of 3.6 × 10−4 per plaque transfer [11], the relatively rapid rise of drug-resistant mutants [12], and findings that indicate unexpected high diversity in herpesvirus infection scenarios [13,14,15,16, 17•, 18, 19]. The ambiguous data on herpesvirus mutation rates seems to be partly due to the use of marker genes (often the thymidine kinase gene or foreign genetic elements) for the estimation of mutation rates, and the overall lack of well-controlled and unbiased studies that examine mutation rates based on whole virus genomes. An informative review about procedures used to address HSV-1 mutation rates in the past can be found in the literature [20].
Structure and Function of Herpesvirus DNA Polymerase
Like all known nucleic acid polymerases, herpesvirus DNA polymerases can be described in analogy to the structure of a cupped right hand [21, 22]. Three main regions are recognized:
- 1.
The palm domain is home to the catalytic core harboring the highly conserved “DTDS” tetrapeptide found in all family B polymerases, where the aspartic acid residues are essential for the coordination of the divalent cations required for the “two-metal-ion-catalysis” and, thus, indispensable for the primary enzymatic function of the enzyme [23]. There is evidence for altered replication fidelity of palm domain mutants [24,25,26].
- 2.
The thumb domain, which contacts the template strand and guides it into the catalytic side. It is important for the transition between the polymerase and exonuclease active site. In family A and B polymerases, the thumb domain is thought to be important for the switch between polymerization and editing (exonuclease) mode. It has been speculated that the switch function could play a role for replication fidelity [27, 28].
- 3.
The finger domain contacts incoming nucleotides and undergoes considerable conformational changes during the catalytic process. The finger domain of herpesvirus DNA polymerases appears to be structurally different from other family B polymerases [29]. Mutations in this domain increase substrate specificity and are known to influence replication fidelity as well as conferring resistance against nucleoside analogs [30,31,32].
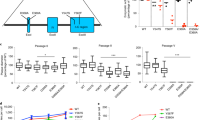
Beyond structural similarities to other polymerases, family B polymerases share seven conserved polymerase and three conserved exonuclease domains [33, 34]. These domains are named from I to VII (I to III in case of the exonuclease) according to the degree of similarity between different polymerases, I being the most conserved domain. Importantly, only the first conserved exonuclease domain, which harbors the charged amino acid residues coordinating the Mg2+ ion essential for catalytic activity, does not overlap with any conserved polymerase side. Exonuclease domains II and III overlap with the conserved Pol domains IV and the δ-C region, respectively (Fig. 1). Also known to be important for enzyme fidelity are the conserved polymerase domain I, located in the palm region and home to the catalytic core of the polymerase, and the conserved region III that harbors part of the finger domain of the enzyme. Another common feature of family B DNA polymerases that is shared by herpesvirus DNA polymerases is the association with an accessory subunit known as processivity factor [28]. Processivity factor is thought to influence replication fidelity by influencing DNA binding of the replication complex [20, 35,36,37,38].
Schematic representation of the HSV-1 DNA polymerase with the conserved Pol regions I–VII in yellow and the conserved Exo regions I–III in blue. Overlaps of both regions are indicated in green. Conserved residues that were mutated and studied with respect to their influence on enzymatic activity and replication fidelity are indicated
DNA Replication Fidelity Variants
Mutants that affect the fidelity of DNA replication are classified based on the differences in mutation rates compared with their respective parent. Usually, low-fidelity variants are described as mutator phenotypes and high-fidelity mutants as antimutator phenotypes.
Low-Fidelity Mutants
Mutations in the polymerase gene leading to mutator phenotypes have frequently been documented (Tables 1, 2, and 3). Low-fidelity variants in herpesviruses exclusively map to one of the exonuclease domains of the protein. The lower DNA replication fidelity of some exonuclease mutants can be explained by impaired proofreading of exonuclease-deficient mutants. It is important to note that, in the case of HSV-1, the majority of exonuclease mutants have only been generated experimentally and tested in vitro since the introduced mutations are lethal for virus replication (Table 1). There is some debate on why some of the exonuclease mutations are not well tolerated by HSV-1 [40, 50, 51•].
High-Fidelity Mutants
Mutations that result in higher DNA replication fidelity relative to parental wild-type viruses have been described far less frequently (Table 1, 2, and 3). Such mutations map to the finger and palm domains of the polymerase and cause resistance to antivirals [24, 32]. The relatively rare description of high-fidelity mutants argues for difficulties in further enhancing replication fidelity of herpesvirus DNA polymerases. For example, no exonuclease mutants with antimutator properties are described. It seems quite possible that exonucleolytic proofreading has reached an evolutionary maximum beyond which a further increase in exonuclease activity can interfere with efficient DNA replication.
Over the past decades, a number of herpesvirus fidelity mutants have been described. Most descriptions are based on HSV-1, but there is evidence in the literature for fidelity mutants in HSV-2, human cytomegalovirus (HCMV), and Marek’s disease virus (MDV). Since relatively little is known about replication fidelity in other herpesviruses, this report will focus on the viruses named above.
Herpes Simplex Viruses (HSV-1 and HSV-2)
In alphaherpesviruses, the DNA polymerase is encoded by UL30, the processivity factor by UL42 [52]. Before the structure and function of the viral polymerase were entirely understood, the discovery of fidelity mutants was based on mutations conveying resistance to antiviral drugs [24, 53]. This is still reflected in the nomenclature of some of the mutants described in the earlier literature (Table 1). With advancements in DNA sequencing and protein expression analyses, more systematic mutational analyses of the DNA polymerase became possible, which also included in vitro measurements of both enzymatic activities of the enzyme [39]. Most of the described mutants map to one of the exonuclease domains (Fig. 1, Table 1). Importantly, only relatively few exonuclease mutants can be reconstituted as viruses in cell culture, indicating a lethal effect of most of the mutations. Different reasons for the observed lethal phenotypes are discussed [20, 50, 51•]. The first and most obvious reason applies to Exo II and III mutants. Within these regions exists an interdependence of polymerase and exonuclease function, the exchange of conserved residues in these regions always affects both enzymatic activities [39, 41]. This interdependence seems to be more pronounced in Exo II mutants that always show strong reductions in polymerase processivity. Most Exo III mutants exhibit less of a reduction in polymerase activity (Table 1). The loss of polymerase function is an obvious explanation for detrimental or lethal effects observed in some of the described mutants. While large defects in polymerase processivity always have lethal outcomes, the more subtle reductions observed in most Exo III mutants do not seem to have a strong negative effect on virus growth. In fact, most published exonuclease mutants that gave rise to viable viruses in HSV-1 are Exo III mutants (Table 1). Although viruses with these mutations can replicate in cell culture, suppression of the mutator phenotype seems to be commonly observed [44]. In contrast to both Exo II and III mutations, mutations that reside within the first conserved exonuclease domain Exo I do not negatively affect polymerase processivity [39]. Instead, exonuclease function is completely abolished in Exo I mutants. These mutations affect the catalytic core of the exonuclease and replacement of the charged residues with other amino acids prevents coordination of Mg2+ and disables catalytic function. The reason for the lethality of these mutations is discussed extensively, but so far is not entirely understood [51•]. The lethal phenotype could be due to one or a combination of the following factors: (1) Important functions of the exonuclease other than proofreading are affected [41, 50]. (2) Certain mutations impact on the expression or folding of the protein [51•] (3) Mutagenesis is lethal due to a lack of proofreading in DNA replication [50]. We will discuss later why the latter explanation is favored by the authors of this report.
Fidelity measurements for the specific Exo III mutants Y577H and Y577H/D581A in HSV-1 vary dramatically and reflect the problems of mutation rate estimation in viruses. A good example is mutant Y577H, which proves to be highly mutagenic in a viral thymidine kinase (TK) assay [43], but shows only a slight increase in mutation frequencies in a supF mutagenesis assay [54] where mutations within a foreign genetic element inserted into the virus genome are measured. Another interesting case is the palm domain mutant R842S (PAAr5) that is described to be an antimutator based on a TK assay [55], but exhibits mutation frequencies slightly above wild-type (WT) in the supF assay [54]. This inconsistency in determining mutation frequencies has made the exploration of fidelity variants difficult and led to the conclusion that “the outcome of the fidelity of DNA replication is assay-dependent” [42]. With the advancements in sequencing technologies, a less-biased evaluation of mutation frequencies in HSV-1 has now become possible.
Two high-fidelity variants of the HSV-1 DNA polymerase are described to date. One, R842S that was originally described as phosphonoacetic acid-resistant clone 5 (PAAr5), maps to the palm domain of the polymerase and is thought to increase replication fidelity through enhanced nucleotide selectivity [20, 24, 56]. As discussed above, the antimutator properties of this mutant are not supported by a supF mutagenesis assay. The L774F high-fidelity variant has a mutation in the finger domain of the DNA polymerase, which is thought to increase substrate specificity through a slight structural change in the targeted domain [20, 32, 44]. Interestingly, this mutant was selected from a population established by hypermutator Y577H/D581A in cell culture [32] and was shown to at least partially restore DNA replication fidelity in the absence of exonucleolytic proofreading [44, 57].
In contrast to HSV-1, relatively little is known about fidelity variants in HSV-2. There are few descriptions of high-fidelity and low-fidelity mutants in literature [12, 58,54,55,61], each without detailed examination of the enzymatic properties of the mutants. Overall, spontaneous mutation rates in HSV-2 appear to be higher than those in HSV-1 [61].
In clinical isolates, a large number of polymerase mutations is found [62, 63]; some of them map to exonuclease domains and could potentially affect replication fidelity. However, a detailed examination of the phenotype of individual mutants from clinical samples is typically not available.
Marek’s Disease Virus
Marek’s disease virus (MDV), the gallid alphaherpesvirus type 2 (GaAHV-2), is the causative agent of a lymphoproliferative disease in chickens and known for its continuous evolution towards higher virulence [64]. Recently, DNA replication fidelity of the WT virus and a number of exonuclease mutants was examined by our laboratory [45]. To study the effects of error-prone genome replication, we constructed MDV polymerase mutants based on in vitro data for HSV-1 polymerase mutants with synonymous changes [39] (Fig. 2).
Alignment of conserved exonuclease domains of herpesviruses (Exo I–III). Highly conserved amino acids that were studied in more than one virus are displayed in color. Highlighted in red are conserved residues that were examined in all three viruses represented in this figure, the function of residues highlighted in yellow was examined in HSV-1 and MDV only. Shown in gray are conserved residues found in HCMV DNA polymerase that were not studied despite their identity to studied residues in HSV-1 and MDV
We found all of the tested mutants to result in viable viruses upon reconstitution in cell culture (Table 2). Among the tested mutants are the Exo I mutants that are reported to be lethal to HSV-1 virus replication. A suicidal phenotype of Exo I mutants D358A and E360A and the double mutant D358A/E360A as well as Exo III mutant Y567F was observed. The progressive growth deficit of these viruses correlated with drastically elevated mutation frequencies that were between 80 and 100 times higher when compared with WT virus. An Exo III mutant with only slightly impaired exonuclease activity was found to replicate stably and cause disease with WT-like kinetics while causing slightly elevated mutation frequencies of around 3× above WT. Interestingly, even strong hypermutators could overcome their suicidal phenotype and adopt WT-like growth as extremely diverse populations with only partially restored DNA replication fidelity. This result indicates the possibility of survival of herpesviruses in the presence of drastically elevated mutation rates and highlights the potential for using such mutants to study the implications and dynamics of low-fidelity nucleic acid replication in viruses.
From our work with MDV Pol mutants, we also conclude that lethal phenotypes of exonuclease mutants are most likely attributable to lethal mutagenesis caused by an accumulation of detrimental mutations generated through low-fidelity replication. Small and isolated populations of hypermutators are prone to extinction in the case of MDV. Considering that this virus replicates relatively slowly in cell culture, the phenotypic difference to the fast replicator HSV-1 could be explained by the speed of genome replication and thus speed of mutation accumulation. It seems possible that some HSV-1 exonuclease mutants cause an immediate collapse of virus populations through lethal mutagenesis immediately following reconstitution. We also acknowledge that there might be a considerable difference in the ability of different viruses to tolerate a substantially increased mutational load. An interesting example is the high viability of exonuclease mutants in the case of HCMV that is described in the following section.
Human Cytomegalovirus
In HCMV, the DNA polymerase is encoded by UL54, the processivity factor by UL44 [52]. Exonuclease mutations are more common in HCMV field isolates than in any other herpesvirus. One reason for this observation could be the frequent use of nucleoside analogues, mainly ganciclovir, in the treatment of HCMV infections, and the fact that certain exonuclease mutations confer resistance against nucleoside analogs. For ganciclovir, a novel mode of action was proposed that could explain resistance conferred by exonuclease-impaired mutants [46]. Upon incorporation of ganciclovir and in the presence of a functional exonuclease, the DNA polymerase is thought to switch to an idling mode where neither excision of ganciclovir by the exonuclease nor chain elongation by the polymerase is possible. In the absence of exonuclease activity, polymerization continues, allowing incorporation of ganciclovir in the nascent DNA double strand. This relatively novel mode of action for ganciclovir would explain the increased rate at which exonuclease mutations in HCMV are selected. The inevitable consequence of this mechanism would be stable incorporation of ganciclovir into the HCMV genome, and it remains unclear if and how this can be either tolerated or circumvented by the virus.
Most of the described HCMV mutants are mainly characterized with respect to their drug resistance [65,63,64,65,69]. Some of them are, however, known to increase mutation rates compared with WT [47], for others in vitro measurements showing an exonuclease defect are availabe [49] (Table 3). It is remarkable that HCMV seems to tolerate mutations in the exonuclease domain of the DNA polymerase much better than HSV-1. Notably, an Exo I mutation (D301N), which affects the aspartic acid within the highly conserved “FDIE” Exo I motif present in the catalytic core of the enzyme (Fig. 2), does not seem to negatively affect virus growth [68]. In contrast, the loss of the homologous D residue at position 368 is reported to be lethal to HSV-1 [51•] and the removal of the same amino acid in the MDV DNA polymerase causes a highly suicidal phenotype [45]. Mutation frequencies for Exo I mutants have not been systematically recorded and certainly require future analysis. Another interesting example that indicates that changes of conserved amino acids within the exonuclease are better tolerated by HCMV than by other herpesviruses, is Exo II D413A, originally identified in a clinical HCMV isolate [70], which results only in a mild growth deficit upon introduction into a HCMV laboratory strain [47]. The counterpart of this mutant in HSV-1 (D471A) cannot be reconstituted [50] and that of MDV (D461A) showed a severe and progressive growth deficit consistent with the expected consequences of hypermutation and loss of polymerase processivity (unpublished data). This polymerase mutant is so far the only mutant HCMV formally described to exhibit mutator properties [47], although exact mutation frequencies in this mutant were not recorded. Also, in the background of ganciclovir resistance, the only known herpesvirus mutant with enhanced exonuclease activity was described. The L501F mutation maps close to the proximal end of the conserved Exo III site and was described to show significantly enhanced exonuclease activity [48].
The viability of HCMV Pol mutants presents a very interesting difference to other herpesviruses. This difference could be a consequence of a much higher tolerance of HCMV to elevated mutation rates or could be explained by a less striking impact of these mutations on the replication fidelity accounted for by the HCMV DNA polymerase. A third possibility is that, in the case of HCMV, loss of replication fidelity can be readily compensated for by other domains of the polymerase, other viral proteins involved in DNA replication and repair, or even by other cellular gene products co-opted by the virus. The lack of whole genome sequencing studies involving virus populations established by exonuclease mutants so far prevents us from examining these possibilities.
The relative abundance of mutations in this region of the polymerase gene in clinical isolates highlights their clinical importance. It is also tempting to speculate that some of the high genetic variation in clinical HCMV isolates might be due to increased mutation rates acquired by exonuclease-deficient viruses selected under ganciclovir treatment. The formal analysis of mutation frequencies in exonuclease mutants seems to be an important area of study in the future.
Conclusions
Mutants that affect the fidelity of viral DNA replication were initially regarded as an important tool to understand the mechanism of DNA replication and its fidelity. We believe, however, that herpesvirus fidelity mutants are important in many more aspects. Especially, low-fidelity variants allow us to interrogate the consequences of more error-prone genome replication and can be used as tools to better understand virus evolution. Stably replicating hypermutators like Y547S in MDV or D413A in HCMV can serve as models for a fast-forward evolution and could allow faster observation of adaptive changes, for example, in drug and vaccine resistance research. Finally, fidelity mutants in herpesviruses are frequently associated with altered susceptibilities to antiviral drugs. Especially in the case of HCMV, the understanding of the principles that lead to emergence of drug-resistant polymerase mutants seems of great clinical and therapeutic importance. More than that, higher mutation frequencies in these mutants may lead to accelerated development of multi-drug resistance and should be subject to close observation and systematic studies in the future.
Change history
27 January 2020
During the production process, the documents provided by the authors were inaccurately merged. As a result all 15 references provided in the tables of the paper are wrong.
27 January 2020
During the production process, the documents provided by the authors were inaccurately merged. As a result all 15 references provided in the tables of the paper are wrong.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance
Sanjuan R, Nebot MR, Chirico N, Mansky LM, Belshaw R. Viral mutation rates. J Virol. 2010;84(19):9733–48. https://doi.org/10.1128/jvi.00694-10.
Kunkel TA. DNA replication fidelity. J Biol Chem. 2004;279(17):16895–8.
Loeb LA, Kunkel TA. Fidelity of DNA synthesis. Annu Rev Biochem. 1982;51(1):429–57.
Petruska J, Goodman MF. Enthalpy-entropy compensation in DNA melting thermodynamics. J Biol Chem. 1995;270(2):746–50.
Kunkel TA, editor. Evolving views of DNA replication (in) fidelity. Cold Spring Harbor symposia on quantitative biology; 2009: Cold Spring Harbor Laboratory Press.
• Bebenek A, Ziuzia-Graczyk I. Fidelity of DNA replication-a matter of proofreading. Curr Genet. 64(2018, 5):985–96. https://doi.org/10.1007/s00294-018-0820-1Very informative review on DNA replication fidelity with special emphasis on exonucleolytic proofreading.
McCulloch SD, Kunkel TA. The fidelity of DNA synthesis by eukaryotic replicative and translesion synthesis polymerases. Cell Res. 2008;18(1):148–61.
Bebenek K, Joyce C, Fitzgerald MP, Kunkel T. The fidelity of DNA synthesis catalyzed by derivatives of Escherichia coli DNA polymerase I. J Biol Chem. 1990;265(23):13878–87.
Knopf CW. Evolution of viral DNA-dependent DNA polymerases. Virus Genes. 1998;16(1):47–58.
Drake JW, Hwang CB. On the mutation rate of herpes simplex virus type 1. Genetics. 2005;170(2):969–70.
Jaramillo N, Domingo E, Muñoz-Egea MC, Tabarés E, Gadea I. Evidence of Muller’s ratchet in herpes simplex virus type 1. J Gen Virol. 2013;94(2):366–75. https://doi.org/10.1099/vir.0.044685-0.
Sarisky RT, Nguyen TT, Duffy KE, Wittrock RJ, Leary JJ. Difference in incidence of spontaneous mutations between herpes simplex virus types 1 and 2. Antimicrob Agents Chemother. 2000;44(6):1524–9. https://doi.org/10.1128/aac.44.6.1524-1529.2000.
Renzette N, Bhattacharjee B, Jensen JD, Gibson L, Kowalik TF. Extensive genome-wide variability of human cytomegalovirus in congenitally infected infants. PLoS Pathog. 2011;7(5):e1001344. https://doi.org/10.1371/journal.ppat.1001344.
Szpara ML, Gatherer D, Ochoa A, Greenbaum B, Dolan A, Bowden RJ, et al. Evolution and diversity in human herpes simplex virus genomes. J Virol. 2014;88(2):1209–27. https://doi.org/10.1128/JVI.01987-13.
Akhtar LN, Bowen CD, Renner DW, Pandey U, Della Fera AN, Kimberlin DW et al. Genotypic and phenotypic diversity of herpes simplex virus 2 within the infected neonatal population. mSphere. 2019;4(1). doi:https://doi.org/10.1128/mSphere.00590-18.
Shipley MM, Renner DW, Ott M, Bloom DC, Koelle DM, Johnston C, et al. Genome-wide surveillance of genital herpes simplex virus type 1 from multiple anatomic sites over time. J Infect Dis. 2018;218(4):595–605. https://doi.org/10.1093/infdis/jiy216.
• Renner DW, Szpara ML. Impacts of genome-wide analyses on our understanding of human herpesvirus diversity and evolution. J Virol. 2017;92(1):e00908–17. https://doi.org/10.1128/JVI.00908-17This review article provides an up-to-date discussion on whole genome sequencing, bioinformatic analysis, and genome diversity in human herpesviruses.
Renzette N, Pokalyuk C, Gibson L, Bhattacharjee B, Schleiss MR, Hamprecht K, et al. Limits and patterns of cytomegalovirus genomic diversity in humans. Proc Natl Acad Sci U S A. 2015;112(30):E4120–8. https://doi.org/10.1073/pnas.1501880112.
Sijmons S, Thys K, Mbong Ngwese M, Van Damme E, Dvorak J, Van Loock M, et al. High-throughput analysis of human cytomegalovirus genome diversity highlights the widespread occurrence of gene-disrupting mutations and pervasive recombination. J Virol. 2015;89(15):7673–95. https://doi.org/10.1128/JVI.00578-15.
Hwang CB-C. DNA Replication Fidelity of Herpes Simplex Virus. In: Kusic-Tisma J, editor. DNA Replication and Related Cellular Processes. IntechOpen; 2011. https://doi.org/10.5772/23548. Available from: https://www.intechopen.com/books/dna-replication-and-relatedcellular-processes/dna-replication-fidelity-of-herpes-simplex-virus.
Ollis DL, Brick P, Hamlin R, Xuong NG, Steitz TA. Structure of large fragment of Escherichia coli DNA polymerase I complexed with dTMP. Nature. 1985;313(6005):762–6.
Steitz TA. DNA polymerases: structural diversity and common mechanisms. J Biol Chem. 1999;274(25):17395–8. https://doi.org/10.1074/jbc.274.25.17395.
Steitz TA. A mechanism for all polymerases. Nature. 1998;391(6664):231–2. https://doi.org/10.1038/34542.
Hall JD, Furman PA, St Clair MH, Knopf CW. Reduced in vivo mutagenesis by mutant herpes simplex DNA polymerase involves improved nucleotide selection. Proc Natl Acad Sci U S A. 1985;82(11):3889–93.
Foury F, Szczepanowska K. Antimutator alleles of yeast DNA polymerase gamma modulate the balance between DNA synthesis and excision. PLoS One. 2011;6(11):e27847-e. https://doi.org/10.1371/journal.pone.0027847.
Jacewicz A, Makiela K, Kierzek A, Drake JW, Bebenek A. The roles of Tyr391 and Tyr619 in RB69 DNA polymerase replication fidelity. J Mol Biol. 2007;368(1):18–29. https://doi.org/10.1016/j.jmb.2007.01.067.
Wu P, Nossal N, Benkovic SJ. Kinetic characterization of a bacteriophage T4 antimutator DNA polymerase. Biochemistry. 1998;37(42):14748–55.
Johansson E, Dixon N. Replicative DNA polymerases. Cold Spring Harb Perspect Biol. 2013;5(6):a012799. https://doi.org/10.1101/cshperspect.a012799.
Luczkowiak J, Álvarez M, Sebastián-Martín A, Menéndez-Arias L. Chapter 4 - DNA-dependent DNA polymerases as drug targets in herpesviruses and poxviruses. In: Gupta SP, editor. Viral Polymerases. Academic Press; 2019. p. 95–134. ISBN 9780128154229, https://doi.org/10.1016/B978-0-12-815422-9.00004-8.
Hwang C, Ruffner KL, Coen DM. A point mutation within a distinct conserved region of the herpes simplex virus DNA polymerase gene confers drug resistance. J Virol. 1992;66(3):1774–6.
Hwang YT, Liu BY, Hong CY, Shillitoe EJ, Hwang CB. Effects of exonuclease activity and nucleotide selectivity of the herpes simplex virus DNA polymerase on the fidelity of DNA replication in vivo. J Virol. 1999;73(7):5326–32.
Hwang YT, Zuccola HJ, Lu Q, Hwang CB. A point mutation within conserved region VI of herpes simplex virus type 1 DNA polymerase confers altered drug sensitivity and enhances replication fidelity. J Virol. 2004;78(2):650–7.
Larder BA, Kemp SD, Darby G. Related functional domains in virus DNA polymerases. EMBO J. 1987;6(1):169–75.
Wong SW, Wahl AF, Yuan PM, Arai N, Pearson BE, Arai K, et al. Human DNA polymerase alpha gene expression is cell proliferation dependent and its primary structure is similar to both prokaryotic and eukaryotic replicative DNA polymerases. EMBO J. 1988;7(1):37–47.
Song L, Chaudhuri M, Knopf CW, Parris DS. Contribution of the 3'- to 5'-exonuclease activity of herpes simplex virus type 1 DNA polymerase to the fidelity of DNA synthesis. J Biol Chem. 2004;279(18):18535–43. https://doi.org/10.1074/jbc.M309848200.
Chaudhuri M, Song L, Parris DS. The herpes simplex virus type 1 DNA polymerase processivity factor increases fidelity without altering pre-steady-state rate constants for polymerization or excision. J Biol Chem. 2003;278(11):8996–9004. https://doi.org/10.1074/jbc.M210023200.
Jiang C, Hwang YT, Randell JC, Coen DM, Hwang CB. Mutations that decrease DNA binding of the processivity factor of the herpes simplex virus DNA polymerase reduce viral yield, alter the kinetics of viral DNA replication, and decrease the fidelity of DNA replication. J Virol. 2007;81(7):3495–502. https://doi.org/10.1128/jvi.02359-06.
Jiang C, Komazin-Meredith G, Tian W, Coen DM, Hwang CBC. Mutations that increase DNA binding by the processivity factor of herpes simplex virus affect virus production and DNA replication fidelity. J Virol. 2009;83(15):7573–80. https://doi.org/10.1128/JVI.00193-09.
Kühn FJ, Knopf CW. Herpes simplex virus type 1 DNA polymerase. Mutational analysis of the 3'-5'-exonuclease domain. J Biol Chem. 1996;271(46):29245–54.
Baker RO, Hall JD. Impaired mismatch extension by a herpes simplex DNA polymerase mutant with an editing nuclease defect. J Biol Chem. 1998;273(37):24075–82.
Gibbs JS, Weisshart K, Digard P, Knipe D, Coen D. Polymerization activity of an alpha-like DNA polymerase requires a conserved 3'-5'exonuclease active site. Mol Cell Biol. 1991;11(9):4786–95.
Hwang YT, Hwang CB. Exonuclease-deficient polymerase mutant of herpes simplex virus type 1 induces altered spectra of mutations. J Virol. 2003;77(5):2946–55.
Hwang YT, Liu B-Y, Coen DM, Hwang C. Effects of mutations in the Exo III motif of the herpes simplex virus DNA polymerase gene on enzyme activities, viral replication, and replication fidelity. J Virol. 1997;71(10):7791–8.
Tian W, Hwang YT, Lu Q, Hwang CB. Finger domain mutation affects enzyme activity, DNA replication efficiency, and fidelity of an exonuclease-deficient DNA polymerase of herpes simplex virus type 1. J Virol. 2009;83(14):7194–201.
Trimpert J, Groenke N, Kunec D, Eschke K, McMahon DP, Osterrieder N. A proofreading-impaired herpesvirus generates populations with quasispecies-like structure. Nat Microbiol. 2019. https://doi.org/10.1038/s41564-019-0547-x.
Chen H, Beardsley GP, Coen DM. Mechanism of ganciclovir-induced chain termination revealed by resistant viral polymerase mutants with reduced exonuclease activity. Proc Natl Acad Sci U S A. 2014;111(49):17462–7. https://doi.org/10.1073/pnas.1405981111.
Chou S, Marousek GI. Accelerated evolution of maribavir resistance in a cytomegalovirus exonuclease domain II mutant. J Virol. 2008;82(1):246–53. https://doi.org/10.1128/jvi.01787-07.
Kariya M, Mori S, Eizuru Y. Comparison of human cytomegalovirus DNA polymerase activity for ganciclovir-resistant and -sensitive clinical strains. Antivir Res. 2000;45(2):115–22.
Cihlar T, Fuller MD, Mulato AS, Cherrington JM. A point mutation in the human cytomegalovirus DNA polymerase gene selected in vitro by cidofovir confers a slow replication phenotype in cell culture. Virology. 1998;248(2):382–93. https://doi.org/10.1006/viro.1998.9299.
Hall JD, Orth KL, Sander KL, Swihart BM, Senese RA. Mutations within conserved motifs in the 3'-5' exonuclease domain of herpes simplex virus DNA polymerase. J Gen Virol. 1995;76(Pt 12):2999–3008. https://doi.org/10.1099/0022-1317-76-12-2999.
• Lawler JL, Coen DM. HSV-1 DNA polymerase 3'-5' exonuclease-deficient mutant D368A exhibits severely reduced viral DNA synthesis and polymerase expression. J Gen Virol. 2018;99(10):1432–7. https://doi.org/10.1099/jgv.0.001138This work features interesting observations regarding the phenotype of exonuclease (ExoI) mutants in HSV-1 and includes an up-to-date discussion of the lethal phenotypes of exonuclease mutants.
Zarrouk K, Piret J, Boivin G. Herpesvirus DNA polymerases: structures, functions and inhibitors. Virus Res. 2017. https://doi.org/10.1016/j.virusres.2017.01.019.
Hall JD, Coen DM, Fisher BL, Weisslitz M, Randall S, Almy RE, et al. Generation of genetic diversity in herpes simplex virus: an antimutator phenotype maps to the DNA polymerase locus. Virology. 1984;132(1):26–37. https://doi.org/10.1016/0042-6822(84)90088-6.
Hwang YT, Liu BY, Hwang CB. Replication fidelity of the supF gene integrated in the thymidine kinase locus of herpes simplex virus type 1. J Virol. 2002;76(8):3605–14.
Hwang CB, Chen HJ. An altered spectrum of herpes simplex virus mutations mediated by an antimutator DNA polymerase. Gene. 1995;152(2):191–3. https://doi.org/10.1016/0378-1119(94)00712-2.
Huang L, Ishii KK, Zuccola H, Gehring AM, Hwang CB, Hogle J, et al. The enzymological basis for resistance of herpesvirus DNA polymerase mutants to acyclovir: relationship to the structure of alpha-like DNA polymerases. Proc Natl Acad Sci U S A. 1999;96(2):447–52. https://doi.org/10.1073/pnas.96.2.447.
Tian W, Hwang YT, Hwang CB. The enhanced DNA replication fidelity of a mutant herpes simplex virus type 1 DNA polymerase is mediated by an improved nucleotide selectivity and reduced mismatch extension ability. J Virol. 2008;82(17):8937–41. https://doi.org/10.1128/jvi.00911-08.
Abbotts J, Nishiyama Y, Yoshida S, Loeb LA. On the fidelity of DNA replication: herpes DNA polymerase and its associated exonuclease. Nucleic Acids Res. 1987;15(3):1185–98. https://doi.org/10.1093/nar/15.3.1185.
Nishiyama Y, Suzuki S, Yamauchi M, Maeno K, Yoshida S. Characterization of an aphidicolin-resistant mutant of herpes simplex virus type 2 which induces an altered viral DNA polymerase. Virology. 1984;135(1):87–96. https://doi.org/10.1016/0042-6822(84)90119-3.
Nishiyama Y, Yoshida S, Tsurumi T, Yamamoto N, Maeno K. Drug-resistant mutants of herpes simplex virus type 2 with a mutator or antimutator phenotype. Microbiol Immunol. 1985;29(4):377–81. https://doi.org/10.1111/j.1348-0421.1985.tb00837.x.
Duffy KE, Quail MR, Nguyen TT, Wittrock RJ, Bartus JO, Halsey WM, et al. Assessing the contribution of the herpes simplex virus DNA polymerase to spontaneous mutations. BMC Infect Dis. 2002;2:7.
Bohn K, Zell R, Schacke M, Wutzler P, Sauerbrei A. Gene polymorphism of thymidine kinase and DNA polymerase in clinical strains of herpes simplex virus. Antivir Ther. 2011;16(7):989–97. https://doi.org/10.3851/imp1852.
Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes of herpes simplex virus types 1 and 2. J Antimicrob Chemother. 2016;71(1):6–16. https://doi.org/10.1093/jac/dkv285.
Osterrieder N, Kamil JP, Schumacher D, Tischer BK, Trapp S. Marek’s disease virus: from miasma to model. Nat Rev Microbiol. 2006;4(4):283–94. https://doi.org/10.1038/nrmicro1382.
Komatsu TE, Pikis A, Naeger LK, Harrington PR. Resistance of human cytomegalovirus to ganciclovir/valganciclovir: a comprehensive review of putative resistance pathways. Antivir Res. 2014;101:12–25. https://doi.org/10.1016/j.antiviral.2013.10.011.
Topalis D, Gillemot S, Snoeck R, Andrei G. Distribution and effects of amino acid changes in drug-resistant alpha and beta herpesviruses DNA polymerase. Nucleic Acids Res. 2016;44(20):9530–54. https://doi.org/10.1093/nar/gkw875.
Chou S, Ercolani RJ, Lanier ER. Novel cytomegalovirus UL54 DNA polymerase gene mutations selected in vitro that confer brincidofovir resistance. Antimicrob Agents Chemother. 2016;60(6):3845–8. https://doi.org/10.1128/AAC.00214-16.
Chou S, Lurain NS, Thompson KD, Miner RC, Drew WL. Viral DNA polymerase mutations associated with drug resistance in human cytomegalovirus. J Infect Dis. 2003;188(1):32–9. https://doi.org/10.1086/375743.
Lurain NS, Chou S. Antiviral drug resistance of human cytomegalovirus. Clin Microbiol Rev. 2010;23(4):689–712. https://doi.org/10.1128/cmr.00009-10.
Marfori JE, Exner MM, Marousek GI, Chou S, Drew WL. Development of new cytomegalovirus UL97 and DNA polymerase mutations conferring drug resistance after valganciclovir therapy in allogeneic stem cell recipients. J Clin Virol. 2007;38(2):120–5. https://doi.org/10.1016/j.jcv.2006.11.005.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised: During the production process, the documents provided by the authors were inaccurately merged. As a result all 15 references provided in the tables of the paper are wrong. The authors identified the error during the proof stage; however, the requested changes were not incorporated in the final published version. Likewise, some other corrections requested by the authors in the proof process remained unchanged in the published version of the manuscript. The publisher apologizes and takes full responsibility for the problems that result in the changes to this article.
This article is part of the Topical Collection on Virology
Rights and permissions
About this article
Cite this article
Trimpert, J., Osterrieder, N. Herpesvirus DNA Polymerase Mutants—How Important Is Faithful Genome Replication?. Curr Clin Micro Rpt 6, 240–248 (2019). https://doi.org/10.1007/s40588-019-00135-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40588-019-00135-2