Abstract
Sustainable onion breeding relies on genetic values of the germplasm. Descriptive statistics for the breeding values for economical traits revealed the maximum variability among the 62 onion genotypes studied. The principle components among the genotypes contributed to 75.44% of the variance and their association was differed significantly (α = 0.05). Analysis of genetic diversity among the germplasm population aided for the classification of genotypes and identification of core collections with possible utility for specific breeding strategies. The sixty-two onion genotypes studied were grouped in to five clusters, cluster-I included nine genotypes, cluster-II had thirteen genotypes, cluster-III contained fourteen genotypes, cluster-IV consisted of twelve and cluster-V contained fourteen genotypes. The inter-cluster distance (56.54) was found maximum between cluster-I and -III. The cluster means reveals the cluster-I is the best for bulb weight, equatorial and polar diameter, stem girth and number of leaves, cluster-IV for plant height. The genotypes differed significantly for the bulb yield and their yield attributes. The ample variability and diversity among the genotype population could aid for genetic improvement of qualitative and quantitative traits of onion germplasm.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Onion (Allium cepa L.) is an antiquity spice; it is grown commercially as vegetable across the climates of temperate, sub-tropical and tropical regions of the world (Brewster, 2018; Corgan & Kedar, 2018; Currah & Proctor, 1990).The economic importance of the crop originated due to its nutritional properties (D C Manjunathagowda et al., 2019a) and health benefits (Benkeblia, 2005). Despite the significance of onion, the research on genetic improvement has significantly paused than the other major vegetable crops(McCallum et al., 2008), this could be due to the poor genetic resource representation among the germplasm (Cross, 1998; Kik, 2006; Manjunathagowda et al., 2021), hence there is a need to understand the population structure among the germplasm for better maintenance and exploitation of genotypes (McCallum et al., 2008).The crop improvement has related to the degree of its genetic variability existed among the genotypes (Mallor et al., 2014; Mallor Giménez et al., 2011; Manjunathagowda & Anjanappa, 2021a, 2021b), the efficiency of selection in the genotypes depends on the knowledge of genetic variability. The variety or hybrid breeding needs the knowledge about genetic variability and diversity, the genetic distinctionis the base for plant breeding and it makes ease to select the genotypes based on distinctiveness for varietal or hybrid development (Dangi et al., 2018; Manjunathagowda & Anjanappa, 2020; Manjunathagowda & Selvakumar, 2021).The management and evaluations of germplasm lead to the exploitation and transfer of useful genes into the desirable phenotypes.
Numerous methods are known for estimation of variance divergence among the genotypes through multivariate tools, among them the principal component (PC) analysis has the potential tool to transform possibly correlated variables into limited variables called principal components (Ziegel, 2002). It could reveal the dimensional data patterns in the identification of key components of variation in a data set. Each PC reveals the data properties and interpreted independently since PCs are orthogonal and independent of each other (Mohammadi & Prasanna, 2003). PCA could reveal the decisive traits among genotypes differentiation, it enables breeders to understand the impact and associations among different traits of the genotypes (Kovacic, 1994). The principal component analysis (PCA) was carried out in the Turkish onion of 96 genotypes for 31 quantitative traits revealed the maximum Eigen value and total variance in PC-I followed by PC-II and PC-III. The first nine principal components (PCs) with high Eigen values (> 1) contributed 71.84% of the variability. The bulb weight and pseudostem diameter traits were contributed more positively with PC1 (Hanci & Gokce, 2015).The principal components (PC1 to PC5) accounted of 78.5% total variation among the genotypes. The squared cosine value of bulb yield traits was with positive direction and plant growth traits were in the negative direction (Dangi et al., 2018).
During the process of breeding, the identification and selection of genetically varied psarents have an immense significance for genetic improvement through recombination breeding (Arunahalam, 1981). Among genotypes the diversity reveals the closeness; the better assortment of genotypes helps the plant breeder for strategic breeding of hybridization (Manjunathagowda et al., 2021).The economic traits in bulb yield in onion are polygenic character influenced by the environmental factors, hence understanding of genetic variability and magnitude of association of bulb yield with yielding traits are highly essential for genetic improvement of bulb yield through selection of better genotypes (Dod & Sharma, 2017).Therefore the study on strategic improvement of onions aided with variance, divergence and principal components analysis carried out for a better understanding of genetic variability and diversity among the genotypes.
Material and methods
The trial was conducted in late-kharif 2019 at ICAR-Directorate of Onion and Garlic Research, Rajgurunagar, Pune, Maharashtra, India. The experimental layout as followed as Randomized Complete Block Design (RCBD) with three replications. The plots of size of 1.5 m length and 1.2 m width with the spacing of 15 cm × 10 cm apart between plants was followed. Crop production and management practices were performed with the guidelines set by the ICAR-Directorate of Onion and Garlic Research, Rajgurunar, Pune, Maharashtra, India (DOGR, 2020). The 45 days old seedling was transplanted in the experimental field, the observations on plant height (cm), number of leaves, stem girth (cm), equatorial diameter (cm), polar diameter (cm) and bulb weight (g) recorded from randomly selected five plants per treatment during full crop growth at 75 days after transplanting, which were subjected for statistical analysis. The performance of 62 genotypes for plant growth and bulb yield traits was assayed and resulted presented in Supplementary Table 1, majority of the genotypes were evaluated for male fertility maintainer and restorer lines (Dalasanuru Chandregowda Manjunathagowda & Selvakumar, 2021). The data were subjected to One-Way-ANOVA per the procedure of the Turkeys test by using MiniTab software (MiniTab, 2020). Genetic variability assays was carried out by OPSTAT open access software (Sheoran et al. 1998). The k-mean analysis of divergence and principle component analysis was performed with the help of XLSTAT Software (XLSTAT, 2020).
Result and discussion
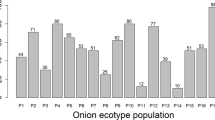
The performance of genotypes for plant growth, bulb yield and its attributing traits was differed significantly (Supplementary Table 1). Genotypes namely DOGR-006 and DOGR-132 differed significantly with the maximum bulb weight, equatorial bulb diameter, stem girth and number of leaves per plant. Similarly, the performance onion genotypes was differed significantly for their plant growth and bulbing traits in kharif season(D C Manjunathagowda et al., 2019b). The significant difference could be due to the response of genotypes to the photosynthetic activity and mobility of availed nutrient within the plant, as this helped in better accumulation of photosynthesis in the bulbs (Sharma, 2009). The significant difference for bulb yield and their attributing traits could be influenced by the genotypic characters, which cause for increased bulb morphological traits, as these traits influenced the bulb weight, and ultimately contributed to bulb yield (Hosamani et al., 2010; Umamaheswarappa et al., 2015).
The descriptive statistics of growth and bulbing traits were revealed in Table 1.The results revealed the considerable variability existed among the genotypes. The highest variability was noted for plant height followed by the number of leaves, stem girth, equatorial and polar diameter, and bulb weight respectively. The principal components of individual values contributed to the cent percentage to cumulative variation among the genotypes (Table 1). Low and high estimates of coefficient reveal the low and high genetic variability among the genotypes, assortment helps to the effective selection of suitable genotypes for breeding (Gurjar & Singhania, 2006; D. Manjunathagowda et al., 2021), the higher variation indicates maximum variation for different growth, yield and quality parameters. These traits afford sufficient opportunity for proficient selection and genetic improvement of genotypes (Ananthan & Balakrishnamoorthy, 2007; Haydar et al., 2007; Hosamani et al., 2010; Ram et al., 2011).
Varied significant variance aid for the exploitation of breeding potential among genotypes. The variance of traits were varied from 7.94 to 28.90 per cent for GCV, 10.41 to 31.19 per cent for PCV, heritability 52.44 to 85.89 per cent and 13.73 to 55.19 per cent for genetic advance mean, respectively (Table 2), the variance ranged from medium to high. The bulb weight acknowledge for high variance for the studied genetic variables (Table 2).These traits were for genotypic and phenotypic variation leads to subsistence variation of traits, and medium to high co-efficient heritability and GAM could contribute for additive-gene-action indicates selection either by mass selection or simple selection could be suitable for genetic improvement of onion genotypes (Dhotre et al., 2010; Manjunathagowda & Anjanappa, 2021b).
The Pearson’s coefficients of correlation among the traits results revealed that, the plant growth and bulb yielding traits were highly significantly and positively correlated to bulb weight (Table 3). The studied traits are significantly influenced to increase bulb weight. The correlation studies among the twelve cultivars of onion revealed a significant positive association of bulb yield (Rashid, 2006). Plant height, number of leaves per plant and stem girth were primary yield determining traits in onion, thus these traits can be improved through direct selection (Raghuwanshi et al., 2016). Bulb yield has a positive significant correlation with equatorial and polar bulb diameter (Patil et al., 1990), equatorial bulb diameter (Dhotre et al., 2010), and the number of leaves (Mahanthesh et al., 2008; Mohanty, 2001; Pal & Singh, 1988).The results divulged the presence of significant genetic variability among the onion genotypes, which can be exploited for genetic improvement of onion.
The path matrics of bulb weight revealed that, the trait was influenced by directly with positive effects viaplant height, number of leaves, neck girth, polar diameter and equatorial diameter (Table 4). Thus, it could aid for onion genetic improvement by direct mass selection of studied traits could be more effective. The path analysis results revealed desirable positive direct effect on bulb yield via plant height, number of leaves, neck girth, polar diameter and equatorial diameters (Dhotre et al., 2010; Manjunathagowda & Anjanappa, 2021a; Raghuwanshi et al., 2016).
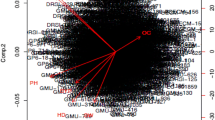
The high positive component values in the principal component (Fig. 1) were noted for bulb weight, equatorial diameter, polar diameter, stem girth, number of leaves and plant height in PC1. The plant height, stem girth and the number of leaves in PC2, and the number of leaves in PC3, stem girth in PC4, polar diameter in PC5 and equatorial diameter in PC6 (Table 5). The principal component coefficients with equal or more than 0.3 were taken to finalize the limiting factor for the coefficients of the vector (Raji, 2002), the traits with higher positive, or negative value revealed the existence of more genetic diversity (Hanci & Gokce, 2015).The PCAof 96 Turkish onion genotypes for 31 quantitative traits revealed the maximum Eigen value with total variance in PC-I, which was followed by PC-II and PC-III. The nine PCs contributed 71.84% of the variability(high Eigen values > 1), the bulb weight and pseudostem diameter traits were contributed more positively with PC1 (Hanci & Gokce, 2015). However, in our study high Eigen values less than one. The principal components (PC1 to PC5) accounted of 78.5% total variation among the genotypes. The squared cosine value of bulb yield traits was with positive direction and plant growth traits were in the negative direction (Dangi et al., 2018).
The PCA simplified the composite data into principal components with a smaller number of variables through the transformation of correlated variables, the first principal component (PC1) reported the highest variability in the data as regarded by subsequent principal components (Leilah & Al-Khateeb, 2005). The results of the study revealed the identification of genetic variability among the onion genotypes based on these traits, which could be accurate for genetic improvement of onion.
The sixty-two onion genotypes were grouped in to five clusters presented in Table 6, cluster-I included nine genotypes, cluster-II had thirteen genotypes, cluster-III contained fourteen genotypes, cluster-IV consisted of twelve and cluster-V contained fourteen genotypes. The inter-cluster distance (56.54) was found maximum between cluster-I and –III (Table 7). The cluster means revealed the cluster-I have the best for bulb weight, equatorial and polar diameter, stem girth and number of leaves, cluster-IV for plant height, and the mean contribution of all traits studied are presented in the Table 8.Genetic diversity has necessarily assembled the genotypes based on the mean similarity, it could diversify goals of plant breeding, and hence the selection of genotypes from the cluster-I and III was most appropriate for hybridization breeding program for better exploitation of heterosis. The assortment of genetically assorted parents in breeding is of immense importance for successful recombination and is vital for breeding (Mohanty, 2001; Patil et al., 1990),the highest diversity showed its greatest potential for improving qualitative and quantitative traits (Singh et al., 2013).
Conclusion
Among the genotypes, the DOGR-006 and DOGR-132 differed significantly for growth and yield, agronomic performance was found to be superior to the other genotypes. The genotypes divulged considerable variability, the high positive component values in the principal components were attributed to the precise selection of genotypes. The significant correlation for growth and bulbing traits in the onion genotypesis attributed to breeding and genetic improvement of onion. The inter-cluster distance was found maximum between cluster-I and –III, hence selecting parents from these groups is appropriate for heterosis breeding of onion for the development of hybrids.
References
Ananthan, M., & Balakrishnamoorthy, G. (2007). Genetic variability and correlation studies in onion (Allium cepa var. cepa L.) for economic dry matter yield. Agricultural Science Digest, 27(3), 190–193.
Arunahalam, V. (1981). Genetic distances in plant breeding. Indian Journal of Genetics, 4, 226–236.
Benkeblia, N. (2005). Free-radical scavenging capacity and antioxidant properties of some selected onions (Allium cepa L.) and garlic (Allium sativum L.) extracts. Brazilian Archives of Biology and Technology, 48, 753–759.
Brewster, J. L. (2018). Cultural systems and agronomic practices in temperate climates. In Onions and allied crops (pp. 1–30). CRC Press.
Corgan, J. N., & Kedar, N. (2018). Onion Cultivation in Subtropical Climates. Onions and Allied Crops. https://doi.org/10.1201/9781351075152-2
Cross, R. J. (1998). Global genetic resources of vegetables. Plant Varieties and Seeds (United Kingdom).
Currah, I., & Proctor, F. J. (1990). Onion in tropical region. National Research Institute Kent. UK Bulletin, 20–35.
Dangi, R., Kumar, A., & Khar, A. (2018). Genetic variability, heritability, and diversity analysis studies in short day tropical onion (Allium cepa L.). Indian Journal of Agricultural Sciences, 88, 948–957.
Dhotre, M., Allolli, T. B., Athani, S. I., & Halemani, L. C. (2010). Genetic variability, character association and path analysis studies in Kharif onion (Allium cepa var. cepa L.). Asian Journal of Horticulture, 5(1), 143–146.
Dod, V. N., & Sharma, M. (2017). Variability studies in Rabi onion (Allium cepa var. cepa L) for yield and yield contributing traits. International Journal of Farm Sciences, 7(1), 123–126.
DOGR. (2020). Climate & Soil. https://dogr.icar.gov.in/index.php?option=com_content&view=article&id=110&Itemid=156&lang=en. Accessed 23 March 2020
Gurjar, R. S. S., & Singhania, D. L. (2006). Genetic variability, correlation and path analysis of yield and yield components in onion. Indian Journal of Horticulture, 63(1), 53–58.
Hanci, F., & Gokce, A. F. (2015). Genetic diversity evaluations in Turkish onion (Allium cepa L.) genotypes: principal component analyses (PCA) for breeding strategies. In VII International Symposium on Edible Alliaceae 1143 (pp. 227–234).
Haydar, A., Sharker, N., Ahmed, M. B., Hannan, M. M., Razvy, M. A., Hossain, M., et al. (2007). Genetic variability and interrelationship in onion (Allium cepa L.). Middle-East Journal of Scientific Research, 2(3–4), 132–134.
Hosamani, R. M., Patil, B. C., & Ajjappalavara, P. S. (2010). Genetic variability and character association studies in onion (Allium cepa L.). Karnataka Journal of Agricultural Sciences, 23(2).
Kik, C. (2006). Allium genetic resources with particular reference to onion. In XXVII International Horticultural Congress-IHC2006: International Symposium on Cultivation and Utilization of Asian, 770 (pp. 135–138).
Kovacic, Z. (1994). Multivariate analysis. Faculty of Economics. University of Belgrade. Serbian. p, 293.
Leilah, A. A., & Al-Khateeb, S. A. (2005). Statistical analysis of wheat yield under drought conditions. Journal of Arid Environments, 61(3), 483–496.
Mahanthesh, B., Sajjan, M. R. P., Harshavardhan, M., & Janardhan, G. (2008). Evaluation of different onion (Allium cepa L.) genotypes for yield and processing quality parameters in kharif season under irrigated condition. Asian Journal of Horticulture, 3(1), 5–9.
Mallor, C., Arnedo-Andrés, M. S., & Garcés-Claver, A. (2014). Assessing the genetic diversity of Spanish Allium cepa landraces for onion breeding using microsatellite markers. Scientia Horticulturae, 170, 24–31.
Mallor Giménez, C., Carravedo Fantova, M., Estopañán Muñoz, G., & Mallor Giménez, F. (2011). Characterization of genetic resources of onion (Allium cepa L.) from the Spanish secondary centre of diversity. Spanish Journal of Agricultural Research, 9(1), 144–155.
Manjunathagowda, D., Jayaswall, K., & Kumar, A. (2021). Variability and genetic diversity among selfed lines (S1) of onion (Allium cepa L.). Indian Journal of Traditional Knowledge (IJTK), 20(2), 563–568.
Manjunathagowda, D. C., Anjannappa, M., Lingaiah, H. B., Rao, E. S., Shankarappa, K. S., & Jayappa, J. (2019a). Performance of short-day onion genotypes for nutritional quality traits of bulb. Journal of Pharmacognosy and Phytochemistry, 8(5), 185–187.
Manjunathagowda, D. C., Anjannappa, M., Lingaiah, H. B., Rao, E. S., Shankarappa, K. S., & Jayappa, J. (2019b). Performance of open pollinated onion (Allium cepa L.) genotypes under southern dry zone of Karnataka. Journal of Pharmacognosy and Phytochemistry, 8(6), 2493–2497.
Manjunathagowda, D. C., & Anjanappa, M. (2020). Identification and development of male sterile and their maintainer lines in short-day onion (Allium cepa L.) genotypes. Genetic Resources and Crop Evolution, 67(2), 357–365. https://doi.org/10.1007/S10722-019-00879-2
Manjunathagowda, D. C., & Anjanappa, M. (2021a). Genetic variability studies for yield and yield contributing traits in onion (Allium cepa L.). Vegetos, 34(1), 174–182. https://doi.org/10.1007/S42535-020-00170-1
Manjunathagowda, DC, & Selvakumar, R. (2021). Marker-assisted selection of Ms locus responsible for male fertility restoration in onion (Allium cepa L.). Genetic Resources and Crop Evolution, 1–5.
McCallum, J., Thomson, S., Pither-Joyce, M., Kenel, F., Clarke, A., & Havey, M. J. (2008). Genetic diversity analysis and single-nucleotide polymorphism marker development in cultivated bulb onion based on expressed sequence tag–simple sequence repeat markers. Journal of the American Society for Horticultural Science, 133(6), 810–818.
MiniTab. (2020). Data Analysis, Statistical & Process Improvement Tools | Minitab. https://www.minitab.com/en-us/. Accessed 23 September 2020
Mohammadi, S. A., & Prasanna, B. M. (2003). Analysis of genetic diversity in crop plants—salient statistical tools and considerations. Crop Science, 43(4), 1235–1248.
Mohanty, B. K. (2001). Analysis of genetic divergence in kharif onion. Indian Journal of Horticulture, 58(3), 260–263.
Pal, N., & Singh, N. B. (1988). Correlation and path coefficient studies in onion. Indian Journal of Horticulture, 45(3and4), 295–299.
Patil, A. H., Dod, V. N., Kale, P. B., & Kulwal, L. V. (1990). Performance of onion varieties with reference to seed production. PKV Research Journal, 14(2), 123–126.
Raghuwanshi, O. S., Jain, P. K., Sengupta, S. K., Dangi, A. S., Verma, N. R., & Sunil, P. (2016). Correlation and path analysis study in diverse onion (Allium cepa L.) genotypes. Asian Journal of Horticulture, 11(1), 19–24.
Raji, A. A. (2002). Assessment of genetic diversity and heterotic relationships in African improved and local cassava (Manihot esculenta Crantz) germplasm. Unpublished doctoral dissertation). University of Ibadan, Ibadan, Nigeria.
Ram, R. B., Bharti, N., Meena, M. L., Lata, R., & Babu, M. (2011). Genetic variability and correlation studies in onion (Allium cepa L.). Vegetos, 24(1), 152–156.
Rashid, M. (2006). Genetic variability and induction of male sterility in onion (Allium cepa L.). MSc thesis (Salna, Gazipur, Bangladesh: Bangabandhu Sheikh Mujibur Rahman ….
Sharma, A. K. (2009). Evaluation of onion varieties in kharif season under submontane low hill conditions of Himachal Pradesh. Annals of Horticulture, 2(2), 191–193.
Sheoran, O. P., Tonk, D. S., Kaushik, L. S., Hasija, R. C., & Pannu, R. S. (1998). Statistical software package for agricultural research workers. In Recent advances in information theory, statistics & computer applications by DS Hooda & RC Hasija Department of Mathematics Statistics (pp. 139–143), CCS HAU, Hisar.
Singh, S. R., Ahmed, N., Lal, S., Ganie, S. A., Amin, M., Jan, N., & Amin, A. (2013). Determination of genetic diversity in onion (Allium cepa L.) by multivariate analysis under long day conditions. African Journal of Agricultural Research, 8(45), 5599–5606.
Umamaheswarappa, P., Naik, A. H. K., & Nataraja, M. (2015). Evaluation of onion (Allium cepa L.) genotypes for growth, yield and quality parameters under central dry zone of Karnataka. Environment and Ecology, 33(2A), 992–995.
XLSTAT. (2020). XLSTAT | Statistical Software for Excel. https://www.xlstat.com/en/. Accessed 23 September 2020
Ziegel, E. (2002). Editor’s report on Encyclopedia of Environmetrics. Technometrics, 1–4.
Acknowledgements
Author acknowledge to Major Singh, Janhavi V, Kuldip for extending the research support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Manjunathagowda, D.C. Genetic enhancement of onion germplasm through population improvement. Plant Physiol. Rep. 27, 73–80 (2022). https://doi.org/10.1007/s40502-022-00646-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40502-022-00646-z