Abstract
Success of crop improvement program depends on systematic exploitation of genetic diversity. Improved understanding on the genetic basis of traits contributing to yield and stress tolerance is necessary to accelerate development of resilient crop varieties. In this study, a subset of 102 diverse rice accessions was assembled after analysing population structure (K = 8) and removal of admixtures from a larger set of IRRI 3 K panel. The constructed subset showed adequate diversity in yield related traits. Genome wide association analysis using the genome wide SNP markers identified a total of 42 SNPs showing significant association with major yield traits. Out of the identified SNPs, 20 SNPs were found to be present in QTL or genes reported previously for yield traits. Remaining 22 loci were found to be novel and needs validation. Elite genetic stocks with increased yield potential will permit us to dissect out the physiological and molecular basis of spikelet number per panicle in rice and thereby accelerate yield enhancement in rice through haplotype based breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice, a staple food crop feeding majority of the world population, is cultivated in 162.05 million hectares with the production of 755.43 million tonnes (FAOSTAT, 2019). India contributes closer to 20% of the world’s rice production, and Indian rice production should increase by at least 50% by 2050 to meet the requirements of increasing population. But, any further increase in rice production is challenged by yield plateau, declining natural resources and increased occurrence in the frequency of biotic/abiotic stresses (Ray et al., 2013; IndiaStat, 2020). This warrants development of high yielding and climate resilient rice varieties through systematic exploitation of rice genetic diversity and effectively introgress the desirable alleles linked to grain yield and stress tolerance traits. Grain yield in rice is a complex traits contributed by several sub-components. There are several yield genes namely Gn1a, DEP1, DEP2, NAL1/SPIKE, etc., have been reported to control panicle related traits in rice and thereby controlling grain yield potential (Li et al., 2021). But, many of them have been discovered in japonica rice and their effects are very minimal in indica genotypes. Exploration of indica genetic diversity for grain yield traits will help us to generate improved knowledge on the physiological components of grain yield and thereby identify novel genes/alleles for rice improvement. In addition, rice genetic stocks differing in their grain number per panicle will allow us to understand the physiological relevance between sink size and dry matter partitioning. Hence, there is growing demand for discovering superior alleles of known candidate genes and also discovering novel genetic factors linked to grain yield traits in rice.
Mapping of genetic loci linked to any traits can be achieved through classical linkage mapping or through time saving Association mapping. Linkage disequilibrium based Association mapping strategies such as GWAS has several advantages over linkage mapping (Abdurakhmonov & Abdukarimov, 2008). Fundamental step in association mapping is to determine the genetic diversity and population structure of the study population (Kushwaha et al., 2017). GWAS has been successfully exploited in discovering genes specific markers associated with tiller number, yield and yield attributing traits and stress tolerance under different environmental conditions (Bhandari et al., 2020; Ren et al., 2021; Subedi et al., 2019).
Success and utility of GWAS depends on the availability of high density markers such as SNP markers. Rice being the model crop in monocots has been sequenced long back and recent efforts at IRRI, Philippines allowed generation of whole genome sequence information of 3024 diverse germplasm accessions from 89 countries (referred as 3 K Rice Panel) (RGP, 2014). Though such genomic dataset is readily available for public in Rice SNP-Seek Database (https://snp-seek.irri.org/), limited attempts have been made to exploit the genomic information for gene discovery (Abbai et al., 2019; Bhandari et al., 2020; Ren et al., 2021). Hence, it is assumed that exploitation of phenotypic diversity of 3 K panel and association mapping by utilizing the genomic information will identify novel genes/ alleles controlling grain yield traits in rice. In this study, efforts were made to analyze the genetic diversity and population structure of 217 diverse accessions of IRRI 3 K panel and a subset of 102 accessions was assembled for association mapping of panicle architecture and grain yield traits.
Materials and methods
Genetic materials and genomic data used
A set of 217 diverse rice genotypes of 3 K panel obtained from International Rice Research Institute, Philippines were used for genetic diversity and population structure analysis. CoreSNP Dataset (v0.4; 990 K) available at Rice SNP-Seek Database (http://snp-seek.irri.org) was used for analysis. Detailed information of this genotype file is described at (https://s3.amazonaws.com/3kricegenome/reduced/990k_3krg-snp-README.txt). Steps followed in performing genetic diversity, population structure and GWAS analysis are described in Fig. 1.
Population structure analysis and construction of a subset
Genotype data was filtered using TASSEL 5.0 (Bradbury et al., 2007) and used. SNPs with Minor Allele Frequency (MAF) ≥ 0.1 and heterozygous proportion ≤ 0.1 were retained, followed with pruning SNPs with minimum distance of 100,000 bp between the filtered SNPs. A set of 3449 SNPs covering all the 12 chromosomes were retained after filtering and pruning and used for inferring population structure. Population structure of 217 accessions was carried out using model-based maximum likelihood approach in ADMIXTURE 1.3 (Alexander et al., 2015) at different K values (K = 2 to 9). An optimum K with the least Cross-Validation (CV) error was fixed to retain pure accessions by eliminating admixtures.
Phenotyping of the subset for yield related traits
The subset of 102 rice accessions (Table 1) was evaluated for yield traits during October–January, 2020 at Paddy Breeding Station, Tamil Nadu Agricultural University, Coimbatore. Standard agronomic and nutrient management practices (150:50:50 N:P:K kg/ha) were followed. Observations on number of productive tillers/plant (NPT), number of primary branches/panicle (NPB), number of secondary branches/panicle (NSB), number of spikelets/panicle (NS) and single plant yield (SPY) were recorded. Observations were recorded in five randomly selected plants in a genotype. To assess the phenotypic variation and frequency distribution of evaluated traits in the subset, descriptive statistics and histogram were performed, respectively, using the Minitab 19.1(Allen, 2019).
GWAS analysis
Genotypic data pertaining to the subset of 102 accessions was prepared as described above. Missing data were imputed using Beagle 4.1 (Browning & Browning, 2016). After filtering, a total of 101,388 polymorphic SNPs were selected for GWAS analysis of panicle and yield traits using GAPIT version 3 of Fixed And Random Model Circulating Probability Unification (FarmCPU) pipeline (Wang & Zhang, 2021). Manhattan Plots were constructed using qqman package (Turner, 2014) with a significant threshold of −log10 p value ≥ 4 to detect the significant marker trait associations. Genes closer to the identified SNPs (± 10 kb) were identified using The Rice Annotation Project-Database (RAP-DB) (Sakai et al., 2013) and Information Commons for Rice (IC4R) (Sang et al., 2020).
Results and discussion
Population structure of 217 rice accessions
Genetic diversity and population structure analysis is one of the basic requirements in any crop improvement programs. Population structure has been demonstrated to interfere with discovery of marker-trait-associations (Zhao et al., 2011). The present study was initiated with the assembly of 217 diverse rice accessions containing different sub-populations including 183 indica accessions (ind1A-19, ind1B-21, ind2-52, ind3-31, indx-60), 17 aus accessions, 11 japonica accessions (japx-4, temp-2, trop-5), 4 aromatic accessions and 2 admix accessions. All the 217 accessions were subjected to population structure analysis to eliminate the admixtures and to construct a subset for GWAS.
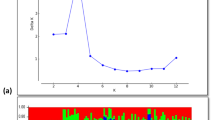
Model based maximum likelihood approach was used to infer population structure at different kinship (K) levels (K = 2 to 9) (Fig. 2). Though CV error gradually decreased with increasing K values, differences of CV error between each K were less and the least CV error was found at K = 8. Hence, K = 8 was considered as an optimum K to construct a subset of 102 accessions containing least admixtures providing a better statistical power for further GWAS analysis. At K = 2, the study population showed a clear distinctness between indica and other sub-populations. At K = 3, the study population showed distinctness within the indica sub-population where ind2 and other indica groups were distinctly separated into two sub-populations. On further increase of K, for example, at K = 5, the indica group of the study population showed distinct separation within their group. At K = 8, even within ind2 and ind3, there were distinct separation to different sub-populations. Lin et al. (2021) also inferred from population structure analysis of 2883 accessions that increased distinct separation of sub-populations can be observed when the K value increases. Though Lin et al. (2021) got an optimum K = 15 based on CV error, they fixed their own optimum at K = 5, due to prior knowledge of sub-populations in 3K-RGP and retained a total of 1378 accessions for GWAS. By following this approach, this study led to shortlisting of 102 accessions by retaining only pure types from 217 accessions with a minimum admixture of ≤ 0.2 ancestry proportions (0.0–1.0). Considering the need of pure types for perfect GWAS, this study has used K = 8 as optimum, admixtures were eliminated and pure types were retained (Table 1) and these 102 lines were used for GWAS.
Phenotypic diversity for yield traits among the subset
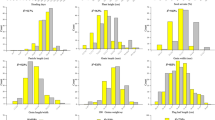
The subset of 102 rice accessions constructed as above was evaluated for yield related traits. Descriptive statistics and frequency distribution of all traits were worked out to evaluate the trait diversity (Table 2 and Fig. 3). The panel showed diversification for all the evaluated traits. It was observed that single plant yield (SPY) exhibited maximum diversity in the study population followed by number of secondary branches per panicle (NSB), number of productive tillers per plant (NPT), number of spikelets per panicle (NS) and number of primary branches per panicle (NPB). Frequency distribution of all the five traits was found to be normal which indicated the suitability of the subset for GWAS.
GWAS identified novel genetic factors linked to yield related traits in rice
GWAS analysis was performed using the genotypic and phenotypic data of the subset of 102 genetically pure accessions so as to minimize false discovery rate (Korte & Farlow, 2013; Tian et al., 2008). GWAS using FarmCPU resulted in the identification of 42 SNPs significantly (threshold of –log10 p ≥ 4) linked to five different target traits. Among the 42 identified loci, 22 SNPs were linked to number of primary branches in the panicle (NPB) followed by 8 SNPs showing significant association with number of secondary branches per panicle (NSB), 6 SNPs with number of productive tillers per plant (NPT), 4 SNPs with number of spikelets per panicle (NS) and 2 SNPs with single plant yield (SPY) (Fig. 4 and Table 3). It was found that SPY and NS showed less SNPs association owing to their complex architecture as reported earlier (Korte & Farlow, 2013). Number of primary branches per panicle showed more number of associations due to the balanced distribution of NPB when compared to all other traits involved (Fig. 3) (Asimit & Zeggini, 2010). Amongst the 42 SNPs identified in this study, 20 SNPs were found to be located closer or co-localized with previously reported QTLs/genes related to panicle traits.
SNP # 23943442 showed significant association with Number of secondary branches/ panicle. This SNP was found to be present within the candidate gene OsWD40-17 (LOC_Os01g42260) that encodes WD40/YVTN repeat-like-containing domain with varied functions including adaptor/regulatory modules in signal transduction, pre-mRNA processing and cytoskeleton assembly and acting as a transcriptional co-repressor modulating flower development (Gonzalez et al., 2007). Similarly SNP # 157578471 was identified as a significant SNP associated with number of productive tillers/plant. This encodes for OsMADS58 (LOC_Os05g11414), a MADS-box domain containing transcriptional factor and important for homeotic regulation in plants. This gene determines the floral meristem formation and involves in carpel development. SNP # 186260109 was found to be significantly associated with number of productive tillers/plant and found nearer to OsSTA173 (LOC_Os06g10140), which encodes Osfbl-29—F-Box Domain and LRR Containing Protein involved in the anther dehiscence, male fertility and pollen germination and determines the spikelet fertility (Ling et al., 2015).
Apart from this, the current study has identified 22 different novel genetic loci linked to various yield related traits. Phenotypic evaluation of the subset provided enough phenotypic variance for GWAS analysis. Among the 42 different genetic loci identified, 20 are co-localized with already reported QTLs/genes. These novel genetic factors may require validation through classical linkage mapping involving contrasting genotypes or validated in a large set of independent population. The subset of 102 rice accessions can be used for mapping agronomic and physiological traits such as nitrogen use efficiency, photosynthetic efficiency and so on. Superior genetic stocks and genes/alleles identified in this study will permit us to generate improved knowledge on the physiological and molecular basis of spikelet number per panicle in rice.
References
Abbai, R., Singh, V. K., Nachimuthu, V. V., Pallavi, S., Ramchander, S., Abhilash, K. V., Singh, A.K., Singh, U.M., Varshney, R.K., & Kumar, A. (2019). Haplotype analysis of key genes governing grain yield and quality traits across 3K RG panel reveals scope for the development of tailor-made rice with enhanced genetic gains. Plant Biotechnology Journal. https://doi.org/10.1111/pbi.13087.
Abdurakhmonov, I. Y., & Abdukarimov, A. (2008). Application of association mapping to understanding the genetic diversity of plant germplasm resources. International Journal of Plant Genomics. https://doi.org/10.1155/2008/574927
Alexander, D. H., Shringarpure, S. S., Novembre, J., & Lange, K. (2015). Admixture 1.3 software manual.
Allen, T. T. (2019). Software overview and methods review: Minitab. In T.T. Allen (Ed.), Introduction to engineering statistics and lean six sigma (pp. 575–600). London: Springer.
Asimit, J., & Zeggini, E. (2010). Rare variant association analysis methods for complex traits. Annual Review of Genetics, 44, 293–308. https://doi.org/10.1146/annurev-genet-102209-163421
Bhandari, A., Sandhu, N., Bartholome, J., Cao-Hamadoun, T.-V., Ahmadi, N., Kumari, N., & Kumar, A. (2020). Genome-wide association study for yield and yield related traits under reproductive stage drought in a diverse indica-aus rice panel. Rice, 13(1), 1–22. https://doi.org/10.1186/s12284-020-00406-3
Bradbury, P. J., Zhang, Z., Kroon, D. E., Casstevens, T. M., Ramdoss, Y., & Buckler, E. S. (2007). TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics, 23(19), 2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Browning, B. L., & Browning, S. R. (2016). Genotype imputation with millions of reference samples. The American Journal of Human Genetics, 98(1), 116–126. https://doi.org/10.1016/j.ajhg.2015.11.020
FAOSTAT. (2019). Food and agricultural organisation. http://www.fao.org/faostat/en/#data/QC. Accessed 17 May, 2021, 2021
Gonzalez, D., Bowen, A. J., Carroll, T. S., & Conlan, R. S. (2007). The transcription corepressor LEUNIG interacts with the histone deacetylase HDA19 and mediator components MED14 (SWP) and CDK8 (HEN3) to repress transcription. Molecular and Cellular Biology, 27(15), 5306–5315. https://doi.org/10.1128/MCB.01912-06
https://www.indiastat.com/data/agriculture/rice/data-year/all-years
Korte, A., & Farlow, A. (2013). The advantages and limitations of trait analysis with GWAS: A review. Plant Methods, 9(1), 1–9. https://doi.org/10.1186/1746-4811-9-29
Kushwaha, U. K. S., Mangal, V., Bairwa, A. K., Adhikari, S., Ahmed, T., Bhat, P., Yadav, A., Dhaka, N., Prajapati, D. R., & Gaur, A. (2017). Association mapping, principles and techniques. J Biol Environ Eng, 2(1), 1–9.
Li, G., Zhang, H., Li, J., Zhang, Z., & Li, Z. (2021). Genetic control of panicle architecture in rice. The Crop Journal. https://doi.org/10.1016/j.cj.2021.02.004
Lin, Y. L., Wu, D. H., Wu, C. C., & Huang, Y. F. (2021). Explore the genetics of weedy traits using rice 3K database. Botanical Studies, 62(1), 1–16. https://doi.org/10.1186/s40529-020-00309-y
Ling, S., Chen, C., Wang, Y., Sun, X., Lu, Z., Ouyang, Y., & Yao, J. (2015). The mature anther-preferentially expressed genes are associated with pollen fertility, pollen germination and anther dehiscence in rice. BMC Genomics, 16(1), 1–17. https://doi.org/10.1186/s12864-015-1305-y
Ray, D. K., Mueller, N. D., West, P. C., & Foley, J. A. (2013). Yield Trends Are Insufficient to Double Global Crop Production by 2050. PLoS ONE. https://doi.org/10.1371/journal.pone.0066428.
Ren, M., Huang, M., Qiu, H., Chun, Y., Li, L., Kumar, A., Fang, J., Zhao, J., He, H., & Li, X. (2021). Genome-wide association study of the genetic basis of effective tiller number in rice. Rice, 14(1), 1–13. https://doi.org/10.1186/s12284-021-00495-8
RGP, K. (2014). The 3000 rice genomes project. Gigascience, 3(1), 7. https://doi.org/10.1186/2047-217X-3-7
Sakai, H., Lee, S. S., Tanaka, T., Numa, H., Kim, J., Kawahara, Y., Wakimoto, H., Yang, C. C., Iwamoto, M., & Abe, T. (2013). Rice annotation project database (RAP-DB): an integrative and interactive database for rice genomics. Plant and Cell Physiology, 54(2), e6–e6. https://doi.org/10.1093/pcp/pcs183
Sang, J., Zou, D., Wang, Z., Wang, F., Zhang, Y., Xia, L., Li, Z., Ma, L., Li, M., & Xu, B. (2020). IC4R-2.0: rice genome reannotation using massive RNA-seq data. Genomics, Proteomics and Bioinformatics, 18(2), 161–172. https://doi.org/10.1016/j.gpb.2018.12.011
Subedi, S. R., Sandhu, N., Singh, V. K., Sinha, P., Kumar, S., Singh, S., Ghimire, S. K., Pandey, M., Yadaw, R. B., & Varshney, R. K. (2019). Genome-wide association study reveals significant genomic regions for improving yield, adaptability of rice under dry direct seeded cultivation condition. BMC Genomics, 20(1), 1–20. https://doi.org/10.1186/s12864-019-5840-9
Tian, C., Gregersen, P. K., & Seldin, M. F. (2008). Accounting for ancestry: Population substructure and genome-wide association studies. Human Molecular Genetics, 17(R2), R143–R150. https://doi.org/10.1093/hmg/ddn268
Turner, S. D. (2014). qqman: An R package for visualizing GWAS results using QQ and manhattan plots. Biorxiv. https://doi.org/10.1101/005165
Wang, J., & Zhang, Z. (2021). GAPIT Version 3: Boosting power and accuracy for genomic association and prediction. Genomics, Proteomics and Bioinformatics. https://doi.org/10.1016/j.gpb.2021.08.005
Zhao, K., Tung, C. W., Eizenga, G. C., Wright, M. H., Ali, M. L., Price, A. H., Norton, G. J., Islam, M. R., Reynolds, A., & Mezey, J. (2011). Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nature Communications, 2(1), 1–10. https://doi.org/10.1038/ncomms1467
Acknowledgements
Authors are thankful to IRRI-South Asia Breeding Hub, Hyderabad for providing the valuable genetic materials. Authors also thank ICAR-NASF for the funding support.
Funding
Funding was provided by Indian Council of Agricultural Research (GTR 8030).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sundaramoorthy, M., Ramasamy, S.P., Rajagopalan, V.R. et al. Pilot scale genome wide association mapping identified novel loci for grain yield traits in rice. Plant Physiol. Rep. 27, 11–21 (2022). https://doi.org/10.1007/s40502-021-00641-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40502-021-00641-w